Abstract
Background
Post-colonoscopy colorectal cancers (PCCRCs) pose challenges in clinical practice. PCCRCs occur due to a combination of procedural and biological causes. In a nested case–control study, we compared clinical and molecular features of PCCRCs and detected CRCs (DCRCs).
Methods
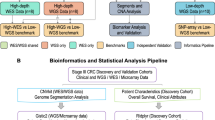
Whole-genome chromosomal copy number changes and mutation status of genes commonly affected in CRC were examined by low-coverage WGS and targeted sequencing, respectively. MSI and CIMP status was also determined.
Results
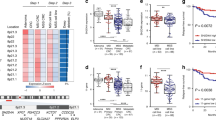
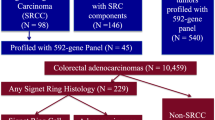
In total, 122 PCCRCs and 98 DCRCs with high-quality DNA were examined. PCCRCs were more often located proximally (P < 0.001), non-polypoid appearing (P = 0.004), early stage (P = 0.009) and poorly differentiated (P = 0.006). PCCRCs showed significantly less 18q loss (FDR < 0.2), compared to DCRCs. No significant differences in mutations were observed. PCCRCs were more commonly CIMP high (P = 0.014) and MSI (P = 0.029). After correction for tumour location, only less 18q loss remained significant (P = 0.005).
Conclusion
Molecular features associated with the sessile serrated lesions (SSLs) and non-polypoid colorectal neoplasms (CRNs) are more commonly seen in PCCRCs than in DCRCs. These together with the clinical features observed support the hypothesis that SSLs and non-polypoid CRNs are contributors to the development of PCCRCs. The future focus should be directed at improving the detection and endoscopic removal of these non-polypoid CRN and SSLs.
Clinical trial registration
NTR3093 in the Dutch trial register (www.trialregister.nl).
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 24 print issues and online access
$259.00 per year
only $10.79 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
The sequencing data used in this study are available in the European Genomes and phenomes Archive (ega-archive.org), with provisory number EGAS00001004686.
References
Singh S, Singh PP, Murad MH, Singh H, Samadder NJ. Prevalence, risk factors, and outcomes of interval colorectal cancers: a systematic review and meta-analysis. Am J Gastroenterol. 2014;109:1375–89. https://doi.org/10.1038/ajg.2014.171
Rex DK. Preventing colorectal cancer and cancer mortality with colonoscopy: what we know and what we don’t know. Endoscopy. 2010;42:320–3. https://doi.org/10.1055/s-0029-1244067
le Clercq CM, Sanduleanu S. Interval colorectal cancers: what and why. Curr Gastroenterol Rep. 2014;16:375. https://doi.org/10.1007/s11894-014-0375-3
Pabby A, Schoen RE, Weissfeld JL, Burt R, Kikendall JW, Lance P, et al. Analysis of colorectal cancer occurrence during surveillance colonoscopy in the dietary Polyp Prevention Trial. Gastrointest Endosc. 2005;61:385–91.
le Clercq CM, Bouwens MW, Rondagh EJ, Bakker CM, Keulen ET, de Ridder RJ, et al. Postcolonoscopy colorectal cancers are preventable: a population-based study. Gut. 2014;63:957–63. https://doi.org/10.1136/gutjnl-2013-304880
Robertson DJ, Lieberman DA, Winawer SJ, Ahnen DJ, Baron JA, Schatzkin A, et al. Colorectal cancers soon after colonoscopy: a pooled multicohort analysis. Gut. 2014;63:949–56. https://doi.org/10.1136/gutjnl-2012-303796
Rondagh EJ, Bouwens MW, Riedl RG, Winkens B, de Ridder R, Kaltenbach T, et al. Endoscopic appearance of proximal colorectal neoplasms and potential implications for colonoscopy in cancer prevention. Gastrointest Endosc. 2012;75:1218–25. https://doi.org/10.1016/j.gie.2012.02.010
Kudo S, Lambert R, Allen JI, Fujii H, Fujii T, Kashida H, et al. Nonpolypoid neoplastic lesions of the colorectal mucosa. Gastrointest Endosc. 2008;68:S3–47. https://doi.org/10.1016/j.gie.2008.07.052.
Bogie RMM, Veldman MHJ, Snijders LARS, Winkens B, Kaltenbach T, Masclee AAM, et al. Endoscopic subtypes of colorectal laterally spreading tumors (LSTs) and risk of submucosal invasion: a meta-analysis. Endoscopy. 2018;50:263–82.
Oka S, Tanaka S, Saito Y, Iishi H, Kudo SE, Ikematsu H, et al. Local recurrence after endoscopic resection for large colorectal neoplasia: a multicenter prospective study in Japan. Am J Gastroenterol. 2015;110:697–707. https://doi.org/10.1038/ajg.2015.96.
Hazewinkel Y, Lopez-Ceron M, East JE, Rastogi A, Pellise M, Nakajima T, et al. Endoscopic features of sessile serrated adenomas: validation by international experts using high-resolution white-light endoscopy and narrow-band imaging. Gastrointest Endosc. 2013;77:916–24. https://doi.org/10.1016/j.gie.2012.12.018
O’Brien MJ, Zhao Q, Yang S. Colorectal serrated pathway cancers and precursors. Histopathology. 2015;66:49–65.
Cisyk AL, Singh H, McManus KJ. Establishing a biological profile for interval colorectal cancers. Dig Dis Sci. 2014;59:2390–402. https://doi.org/10.1007/s10620-014-3210-7.
Sawhney MS, Farrar WD, Gudiseva S, Nelson DB, Lederle FA, Rector TS, et al. Microsatellite instability in interval colon cancers. Gastroenterology. 2006;131:1700–5. https://doi.org/10.1053/j.gastro.2006.10.022
Arain MA, Sawhney M, Sheikh S, Anway R, Thyagarajan B, Bond JH, et al. CIMP status of interval colon cancers: another piece to the puzzle. Am J Gastroenterol. 2010;105:1189–95. https://doi.org/10.1038/ajg.2009.699
Nishihara R, Wu K, Lochhead P, Morikawa T, Liao X, Qian ZR, et al. Long-term colorectal-cancer incidence and mortality after lower endoscopy. N. Engl J Med. 2013;369:1095–105. https://doi.org/10.1056/NEJMoa1301969
Soong TR, Nayor J, Stachler MD, Perencevich M, Jajoo K, Saltzman JR, et al. Clinicopathologic and genetic characteristics of interval colorectal carcinomas favor origin from missed or incompletely excised precursors. Modern Pathol. 2018; e-pub ahead of print 21 November, 2018. https://doi.org/10.1038/s41379-018-0176-6
Cisyk AL, Nugent Z, Wightman RH, Singh H, McManus KJ. Characterizing microsatellite instability and chromosome instability in interval colorectal cancers. Neoplasia. 2018;20:943–50. https://doi.org/10.1016/j.neo.2018.07.007.
Statline CBS. Bevolkingsontwikkeling; kevendgeborenen, overledenen en migratie per regio. 2011; http://statline.cbs.nl/statweb/.
Rutter MD, Beintaris I, Valori R, Chiu HM, Corley DA, Cuatrecasas M, et al. World endoscopy organization consensus statements on post-colonoscopy and post-imaging colorectal cancer. Gastroenterology. 2018;155:909–25 e903. https://doi.org/10.1053/j.gastro.2018.05.038.
Carvalho B, Postma C, Mongera S, Hopmans E, Diskin S, van de Wiel MA, et al. Multiple putative oncogenes at the chromosome 20q amplicon contribute to colorectal adenoma to carcinoma progression. Gut. 2009;58:79–89. https://doi.org/10.1136/gut.2007.143065
Scheinin I, Sie D, Bengtsson H, Van De Wiel MA, Olshen AB, Van Thuijl HF, et al. DNA copy number analysis of fresh and formalin-fixed specimens by shallow whole-genome sequencing with identification and exclusion of problematic regions in the genome assembly. Genome Res. 2014;24:2022–32.
Wanders LK, Cordes M, Voorham Q, Sie D, de Vries SD, d’haens G, et al. IBD-associated dysplastic lesions show more chromosomal instability than sporadic adenomas. Inflamm bowel Dis. 2019. https://doi.org/10.1093/ibd/izz171.
Voorham QJ, Carvalho B, Spiertz AJ, van Grieken NC, Mongera S, Rondagh EJ, et al. Chromosome 5q loss in colorectal flat adenomas. Clin Cancer Res. 2012;18:4560–9. https://doi.org/10.1158/1078-0432.CCR-11-2385
Weisenberger DJ, Siegmund KD, Campan M, Young J, Long TI, Faasse MA, et al. CpG island methylator phenotype underlies sporadic microsatellite instability and is tightly associated with BRAF mutation in colorectal cancer. Nat Genet. 2006;38:787–93.
Derks S, Postma C, Moerkerk P, van den Bosch SM, Carvalho B, Hermsen MA, et al. Promoter methylation precedes chromosomal alterations in colorectal cancer development. Anal Cell Pathol. 2006;28:247–57.
Herman JG, Graff JR, Myöhänen S, Nelkin BD, Baylin SB, Methylation-specific PCR. a novel PCR assay for methylation status of CpG islands. Proc Natl Acad Sci USA. 1996;93:9821–6.
Van De Wiel MA, Smeets SJ, Brakenhoff RH, Ylstra B. CGHMultiArray: exact P-values for multi-array comparative genomic hybridization data. Bioinformatics. 2005;21:3193–4.
van Buuren S, Groothuis-Oudshoorn K. MICE: multivariate imputation by chained equations in R. J Stat Softw. 2011;45:1–67.
R: A language and environment for statistical computing. Vienna, Austria: R Foundation for Statistical Computing; 2015.
Dorenkamp S, Mesters I, Schepers J, Vos R, van den Akker M, Teijink J, et al. Disease combinations associated with physical activity identified: The SMILE Cohort Study. Biomed Res Int. 2016;2016:9053578. https://doi.org/10.1155/2016/9053578.
Finch H. Comparison of distance measures in cluster analysis with dichotomous data. J Data Sci. 2005;3:85–100.
Warnes GR, Bolker B, Bonebakker L, Gentleman R, Huber W, Liaw A, et al. gplots: Various R Programming Tools for Plotting Data [computer program]. Version 3.1.1. 2016. https://github.com/talgalili/gplots.
Cancer Genome Atlas N. Comprehensive molecular characterization of human colon and rectal cancer. Nature. 2012;487:330–7. https://doi.org/10.1038/nature11252
Stoffel EM, Erichsen R, Froslev T, Pedersen L, Vyberg M, Koeppe E, et al. Clinical and molecular characteristics of post-colonoscopy colorectal cancer: a population-based study. Gastroenterology. 2016. https://doi.org/10.1053/j.gastro.2016.07.010.
Cisyk AL, Penner-Goeke S, Lichtensztejn Z, Nugent Z, Wightman RH, Singh H, et al. Characterizing the prevalence of chromosome instability in interval colorectal cancer. Neoplasia. 2015;17:306–16. https://doi.org/10.1016/j.neo.2015.02.001.
Lengauer C, Kinzler KW, Vogelstein B. Genetic instabilities in human cancers. Nature. 1998;396:643–9. https://doi.org/10.1038/25292
Simons CC, Hughes LA, Smits KM, Khalid-de Bakker CA, de Bruine AP, Carvalho B, et al. A novel classification of colorectal tumors based on microsatellite instability, the CpG island methylator phenotype and chromosomal instability: implications for prognosis. Ann Oncol. 2013;24:2048–56. https://doi.org/10.1093/annonc/mdt076
JEIJ, Medema JP, Dekker E. Colorectal neoplasia pathways: state of the art. Gastrointest Endosc Clin North Am. 2015;25:169–82. https://doi.org/10.1016/j.giec.2014.11.004
Fearon ER, Carethers JM. Molecular subtyping of colorectal cancer: time to explore both intertumoral and intratumoral heterogeneity to evaluate patient outcome. Gastroenterology. 2015;148:10–13. https://doi.org/10.1053/j.gastro.2014.11.024
Shaukat A, Arain M, Thaygarajan B, Bond JH, Sawhney M. Is BRAF mutation associated with interval colorectal cancers? Dig Dis Sci. 2010;55:2352–6. https://doi.org/10.1007/s10620-010-1182-9
Hoffmeister M, Blaker H, Jansen L, Alwers E, Amitay EL, Carr PR, et al. Colonoscopy and reduction of colorectal cancer risk by molecular tumor subtypes: a population-based case-control study. Am J Gastroenterol. 2020;115:2007–16. https://doi.org/10.14309/ajg.0000000000000819.
Okamoto K, Kitamura S, Kimura T, Nakagawa T, Sogabe M, Miyamoto H, et al. Clinicopathological characteristics of serrated polyps as precursors to colorectal cancer: current status and management. J Gastroenterol Hepatol. 2017;32:358–67. https://doi.org/10.1111/jgh.13482
Pohl H, Srivastava A, Bensen SP, Anderson P, Rothstein RI, Gordon SR, et al. Incomplete polyp resection during colonoscopy-results of the complete adenoma resection (CARE) study. Gastroenterology. 2013;144:74–80 e71. https://doi.org/10.1053/j.gastro.2012.09.043
Bettington M, Walker N, Rosty C, Brown I, Clouston A, McKeone D, et al. Clinicopathological and molecular features of sessile serrated adenomas with dysplasia or carcinoma. Gut. 2017;66:97–106. https://doi.org/10.1136/gutjnl-2015-310456
Leggett B, Whitehall V. Role of the serrated pathway in colorectal cancer pathogenesis. Gastroenterology. 2010;138:2088–2100. https://doi.org/10.1053/j.gastro.2009.12.066
Voorham QJ, Rondagh EJ, Knol DL, van Engeland M, Carvalho B, Meijer GA, et al. Tracking the molecular features of nonpolypoid colorectal neoplasms: a systematic review and meta-analysis. Am J Gastroenterol. 2013;108:1042–56. https://doi.org/10.1038/ajg.2013.126
Voorham QJ, Carvalho B, Spiertz AJ, Claes B, Mongera S, van Grieken NC, et al. Comprehensive mutation analysis in colorectal flat adenomas. PLoS ONE. 2012;7:e41963. https://doi.org/10.1371/journal.pone.0041963
Konda K, Konishi K, Yamochi T, Ito YM, Nozawa H, Tojo M, et al. Distinct molecular features of different macroscopic subtypes of colorectal neoplasms. PLoS ONE. 2014;9:e103822. https://doi.org/10.1371/journal.pone.0103822
Sanduleanu S, Rondagh EJ, Masclee AA. Development of expertise in the detection and classification of non-polypoid colorectal neoplasia: experience-based data at an academic GI unit. Gastrointest Endosc Clin North Am. 2010;20:449–60. https://doi.org/10.1016/j.giec.2010.03.006
Jover R, Zapater P, Polania E, Bujanda L, Lanas A, Hermo JA, et al. Modifiable endoscopic factors that influence the adenoma detection rate in colorectal cancer screening colonoscopies. Gastrointest Endosc. 2013;77:381–9 e381. https://doi.org/10.1016/j.gie.2012.09.027.
le Clercq CM, Mooi RJ, Winkens B, Salden BN, Bakker CM, van Nunen AB, et al. Temporal trends and variability of colonoscopy performance in a gastroenterology practice. Endoscopy. 2016;48:248–55. https://doi.org/10.1055/s-0041-111117
Waldmann E, Gessl I, Sallinger D, Jeschek P, Britto-Arias M, Heinze G, et al. Trends in quality of screening colonoscopy in Austria. Endoscopy. 2016;48:1102–9. https://doi.org/10.1055/s-0042-113185.
Pilonis ND, Bugajski M, Wieszczy P, Franczyk R, Didkowska J, Wojciechowska U, et al. Long-term colorectal cancer incidence and mortality after a single negative screening colonoscopy. Ann Intern Med. 2020;173:81–91. https://doi.org/10.7326/M19-2477.
Lam AY, Li Y, Gregory DL, Prinz J, O’Reilly J, Manka M, et al. Association between improved adenoma detection rates and interval colorectal cancer rates after a quality improvement program. Gastrointest Endosc. 2020;92:355–64 e355. https://doi.org/10.1016/j.gie.2020.02.016.
Castaneda D, Popov VB, Verheyen E, Wander P, Gross SA. New technologies improve adenoma detection rate, adenoma miss rate and polyp detection rate: a systematic review and meta-analysis. Gastrointest Endosc. 2018. https://doi.org/10.1016/j.gie.2018.03.022
Bogie RMM, Clercq CMC le, Voorham QJM, Cordes M, Sie D, Broek E vd, et al. OP212 Association of chromosomal instability and microsatellite instability pathways with postcolonoscopy colorectal cancer in a retrospective cohort study. United Eur Gastroenterol J. 2016;4(5_suppl):A1-A156 [Oral presentation 212].
Acknowledgements
We thank the technicians of the pathology laboratory at the Netherlands Cancer Institute in Amsterdam for their help with the DNA isolations. We thank the Tumour Genome Analysis Core at Amsterdam UMC for low-coverage WGS and cancer-related gene panel-targeted sequencing. We thank the Genomics Core Facility at the Netherlands Cancer Institute for cancer-related gene panel-targeted sequencing. We thank the technicians of the pathology laboratory at the Maastricht University Medical Centre for their help with the CIMP analysis. We thank Dr. Silvia Sanduleanu for her contribution to the origin of this study. We acknowledge that a part of this work has been presented at the UEGW 2016 in Vienna [57]. This work was performed within the frame of the COST Action [CA17118], supported by COST (European Cooperation in Science and Technology).
Funding
RB and CLC received an unrestricted educational grant from Pentax Medical BV. BY contributed to the manuscript with the support of the Dutch Cancer Society (Grant No. KWF 2015-7882).
Author information
Authors and Affiliations
Contributions
RB, CC, ME, GM and BC conceived the study concept and design; RB, CC, QV, MC, DS, CR, EB, SV, NG, RR, PS, ES, BY and BC contributed to the acquisition of the data; RB, CC, QV, MC, DS, CR, EB, SV, NG, RV, BW, ME, BY, GM, AM and BC participated in analysis and interpretation of the data; RB, CC, QV, CR, BW, AM and BC drafted the manuscript; all authors did a critical revision of the manuscript for important intellectual content; RB, CR, RV and BW performed the statistical analysis; AM and BC had the study supervision. AM is the guarantor of the article. All authors approved the final version.
Corresponding author
Ethics declarations
Competing interests
AM received funding from the Dutch Cancer Society, from the Dutch Organization for Health Research and Development, from Pentax Europe GmBH. AM has given scientific advice to Kyowa Kirin, Bayer, and Takeda. BC has several patents pending. The remaining authors declare no competing interests.
Ethics approval and consent to participate
The study was approved by the Medical Ethical Committee of the Maastricht University Medical Centre, which waived the need for informed consent. The study is registered as study NTR3093 in the Dutch trial register (www.trialregister.nl). The study was performed in accordance with the Declaration of Helsinki.
Consent to publish
No individual data are presented in this study.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Bogie, R.M.M., le Clercq, C.M.C., Voorham, Q.J.M. et al. Molecular pathways in post-colonoscopy versus detected colorectal cancers: results from a nested case–control study. Br J Cancer 126, 865–873 (2022). https://doi.org/10.1038/s41416-021-01619-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41416-021-01619-z