Abstract
Autism spectrum disorder (ASD) is a complex neurodevelopmental disorder caused by genetic or environmental perturbations during early development. Diagnoses are dependent on the identification of behavioral abnormalities that likely emerge well after the disorder is established, leaving critical developmental windows uncharacterized. This is further complicated by the incredible clinical and genetic heterogeneity of the disorder that is not captured in most mammalian models. In recent years, advancements in stem cell technology have created the opportunity to model ASD in a human context through the use of pluripotent stem cells (hPSCs), which can be used to generate 2D cellular models as well as 3D unguided- and region-specific neural organoids. These models produce profoundly intricate systems, capable of modeling the developing brain spatiotemporally to reproduce key developmental milestones throughout early development. When complemented with multi-omics, genome editing, and electrophysiology analysis, they can be used as a powerful tool to profile the neurobiological mechanisms underlying this complex disorder. In this review, we will explore the recent advancements in hPSC-based modeling, discuss present and future applications of the model to ASD research, and finally consider the limitations and future directions within the field to make this system more robust and broadly applicable.
Similar content being viewed by others
ASD overview and 2D/3D modeling
Autism spectrum disorder (ASD) is a highly prevalent neurodevelopmental disorder (NDD) that impacts as many as 1/44 of children in the United States [1]. Clinical presentations of ASD vary widely among individuals but must include repetitive, restricted behaviors and social deficits [2]. To add to this complexity, comorbidities often include epilepsy and seizure disorders (up to 30%), intellectual disability (>30%), ADHD, gastrointestinal disorders (up to 70%), anxiety, and depression [3,4,5,6,7,8,9]. Risk factors for ASD can occur prenatally, perinatally, and postnatally and include genetic disruptions and environmental insults, of which there is likely a combinatorial or synergistic effect.
Twin studies were one of the first indicators of the genetic component to ASD, and while heritability estimates can range from 45 to 90%, it is more broadly thought to be 70–80% [10, 11]. More recently, cohort-based sequencing studies have provided a genetic framework to studying ASD and have identified several hundred implicated genes. Genetic disruptions include inherited rare variants and less common de novo mutations that exist as single nucleotide polymorphisms (SNPs), copy number variants (CNVs), and chromosomal abnormalities [12,13,14,15,16,17,18,19] Despite the immense progress in identifying ASD-risk genes, the encoded proteins and resulting pathobiology remains elusive. Scientists have turned to genetic modeling to better understand the molecular, cellular, and functional (circuit-based) consequences to disruption in these ASD-risk genes [20].
Genetic models of ASD have most commonly included mouse and human cell lines, which provide biologically and clinically relevant opportunities for study [21, 22]. These models are not without their caveats, however, as the development of a mouse brain excludes human-specific processes such as brain gyrification, the protracted development and formation of particular neural cell types, and human-specific gene transcriptional programs. In addition, mouse behavioral assays are often not translatable to the complex clinical presentations of those with ASD (reviewed in refs. [23, 24]). Conversely, human cell lines produce reliable, replicable environments for testing simple pathways, but are reductive and lack the ability to mimic complex developmental brain processes. Further, they can lack specific cell types and structures that play a crucial role in development, such as brain vascularization.
Induced pluripotent stem cells (iPSCs), which can be generated from human blood or skin-derived fibroblasts, have transformed the use of human cellular models [25]. iPSCs retain the unique genetic background from the individual, which is important given that idiopathic ASD represents roughly 80–85% of the ASD population [26]. Through directed differentiation, iPSCs can produce any germ layer cell types to model complex and inaccessible tissue such as the developing brain, allowing for a putatively unlimited supply of patient-specific tissues to study disease processes or drug screening [27] (reviewed in ref. [28], summarized in Table 1).
An essential process during brain development is the genesis and differentiation of neural stem cells (neurogenesis), which can be captured using hPSCs. Neurogenesis describes the emergence of various differentiated brain cell types from neural stem cells and neural progenitor cells (NPCs). Both precursor cell types are important for the formation of the neurons and glia that populate the cerebral cortex, and can be classified based on their mitotic state, location, and polarity (apical or basal). The polarity of an NPC or neural stem cell reflects the positioning of essential proteins and organelles such as the Golgi apparatus and can influence the cell fate and diversity of daughter cells. Disruptions in cell polarity have been associated with a number of NDDs including Fragile X, SCZ, and ASD [29,30,31,32].
NPCs differ from neural stem cells in that their pluripotent fate is more restricted; they have limited proliferation and are capable of producing most neural and glial cell types in the CNS. Given the frequent presentations of macrocephaly amongst ASD individuals, it is possible that excessive neural growth is an underlying factor that may contribute towards ASD pathogenesis, which has been examined using hPSC-derived NPCs [33,34,35,36].
NPC proliferation has been characterized using patient-derived hPSCs and can even be used to stratify subpopulations amongst patients with ASD. In a recent study, hPSCs derived from an ASD cohort that had either idiopathic ASD or a 16p11.2 microdeletion were used to generate NPCs to examine proliferating pathways. The team found that hPSCs derived from macrocephalic individuals with either the 16p11.2 deletion or idiopathic ASD showed increased proliferation and DNA synthesis and proliferation, whereas the remaining probands displayed the opposite trend [37]. The lines were categorized as hyperproliferative and hypoproliferative NPCs and were then treated with basic fibroblast growth factor (bFGF), a mitogen response element that can prime cells for cortical progenitor proliferation. Interestingly, NPCs from the hyperproliferative group displayed a blunted response to bFGF, whereas the hypoproliferative group showed an increase in DNA synthesis sensitivity and response to stimulation. This work highlights the complexities of disease modeling and how patient-derived NPCs can be used to identify subpopulations amongst heterogeneous clinical datasets.
hPSC-derived NPCs enable researchers to examine complex biological processes relating to proliferation and neuronal differentiation. Directed differentiation can produce robust cultures that can be subjected to high-throughput screening, drug testing and phenotyping [38]. Terminal differentiation can be achieved through the addition of various compounds and transcription factors to broaden the window of development that is examined [39,40,41,42], an example being the Ngn2 system that produces glutamatergic-like excitatory neurons. Two major caveats of this system are the reductive and overly simplistic 2D nature of the cultures, and their short lifespans. Unlike hPSCs, NPCs can only be passaged a discreet number of times, limiting the scalability of the model.
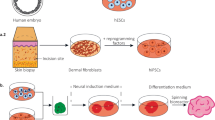
Although hPSC-derived monolayer cultures have deepened our understanding of CNS development, function, and pathology, 2D spatial-organizational constraints limit their ability to model three-dimensional (3D) tissue architecture with complex cell-cell and cell-extracellular matrix (ECM) interactions [43, 44]. Advances in stem cell technology over the past decade have led to the emergence of 3D self-organizing brain organoid models that recapitulate key cellular, structural, and circuital features of human development and disease (summarized in Table 2) [45]. These models were first pioneered with the use of hPSCs by Dr. Yoshiki Sasai’s group with the generation of cortical tissues and 3D optic cup structures in the early 2000s, which has since broadened towards established protocols that exploit either intrinsic or extrinsic signaling pathways to coax the differentiation of hPSCs toward cellular lineages reminiscent of whole or region-specific brain development, respectively [46] (Fig. 1).
Unguided approaches rely on the spontaneous differentiation of hPSCs to ECM-embedded heterogeneous cerebral tissue [47]. The resultant unguided neural organoids (UNOs), formerly known as cerebral organoids [48], exhibit discrete regionalization reminiscent of in vivo human whole-brain development, such as markers of forebrain, midbrain, hindbrain, dorsal cortex, prefrontal cortex, hippocampal, occipital lobe, ventral forebrain, choroid plexus, meningeal, and retinal identity [47, 49]. Furthermore, they demonstrate cellular and structural features unique to human cortical progenitor zone organization, such as layers resemblant of ventricular and subventricular zones [49]. Epigenomic and single-cell transcriptomic analyses of UNO tissue have revealed a remarkable similarity to the early developing fetal cortex [50, 51]. However, undirected hPSC differentiation inherently results in stochastic organoid cellular composition that can hinders batch reproducibility [49]. Alternatively, guided organoid approaches incorporate exogenous signaling factors to direct hPSC differentiation towards region-specific lineages, such as those present in the cerebral cortex [52,53,54,55], forebrain [56,57,58,59], medial ganglionic eminence [54], midbrain [56, 60], thalamus [61], striatum [62], pituitary [63], hypothalamus [56, 64, 65], choroid plexus [66], cerebellum [67], brainstem [68], and spinal cord [69, 70]. These organoids generally display less batch-to-batch heterogeneity than their undirected counterparts and therefore may be more conducive to quantitative analyses [46]. Guided organoids have even been combined to generate assembloids comprised of different brain regions, which provide incredible promise to study pathophysiology within affected circuits.
Brain structure, assembloids, and circuitry
Broad structural and circuit abnormalities have been identified in multiple brain regions of individuals with ASD. In addition to generalized macrocephaly, ASD brains can have structural abnormalities within areas of higher order cognitive processing such as the cerebellum, frontal lobe, and limbic system [71, 72], and even manifest in enlargement of the ventricular cavities where newborn neurons originate [73, 74]. Due to a lack of standardized clinical imaging and EEG recordings, it is impossible to know how pervasive these brain abnormalities are within ASD populations alone, but recent population studies have estimated a frequency of 30–50% [75]. These structural changes are often subtle and variable across individuals with ASD, suggesting that dysregulated circuitry between affected regions and altered molecular pathways may be the underlying cause to this presentation [76,77,78].
Unguided neural organoids (UNOs) have been used to model both microcephaly and macrocephaly in disease contexts [49, 56]. A primary example includes studies of PTEN variants in ASD populations that are comorbid for macrocephaly [79]. Loss of PTEN function was investigated in UNOs by use of isogenic hPSC homozygous mutant lines; concordant with the loss of function mutations found in NDD populations with macrocephaly, the UNOs displayed an increase in size across development, in addition to aberrant tissue folding identified through light sheet microscopy [79]. More recently, PTEN gene-dosage sensitivity was assessed by comparing the isogenic KO to a lentiviral overexpression hPSC line, to model the 10q23.31 microduplication associated with patients with autosomal dominant primary microcephaly. Here it was found that UNO size was inversely proportional to PTEN expression, and could be rescued by an AKT inhibitor that acts on a known PTEN pathway [80]. This demonstrates the use of UNOs to model whole-brain structural abnormalities in ASD populations, and how they can be mined for pharmacological rescue.
Due to the developmental nature of the disorder, as well as the multiple brain regions affected, ASD may arise from miswiring amongst neural circuits during fetal development, with an enrichment in the developing cortex. Advances in guided neural organoids (GNOs, or brain-region-specific organoids) have enabled investigation into how different areas of the brain interact in a disease model. When merged, the resulting assembloids provide the necessary environment for cell-cell interactions and complex developmental processes including integration into circuitry. Assembloids can be examined to assess gross structural abnormalities, the migration of neuronal subpopulations, as well as inter-assembloid circuitry. Assembloids have included the combination of cortical (dorsal) and ventral forebrain organoids [58], cortical-thalamus [61], cortical-striatum [62], cortical-subpial spheroids [81], and even tri-part assembloids consisting of cortical-spinal fused to skeletal muscle assembloids [70]. These new model systems allow for the de novo generation of synthetic circuits in the lab, which have been shown to generate spontaneous neural oscillations comparable to that of the developing human brain [82].
Migration and circuit-based disruptions have been described in multiple ASD models, and in patients are often identified through MRI of gross structural abnormalities in the brain or inferred from EEG recordings of epileptic or paroxysmal activity [75, 83, 84] (reviewed in refs. [85,86,87]). EEG abnormalities include an increased frequency of focal spikes, or localized activity to a particular area of the brain [84, 88, 89]. Several wavelength frequency abnormalities have been characterized within ASD cohorts, including an increase in low-frequency (delta and theta) and high frequency (beta and gamma) wavelengths which is contrasted by a reduction in mid-range alpha frequencies, producing a characteristic U-shaped electrophysiological profile, in which the extremities of the power spectrum are enhanced in ASD populations and the mid-range values are reduced. Organoids generate many of the neural stem cell populations and mature cell types in the brain, and are capable of producing many of the EEG wavelengths mentioned above, in addition to increased firing rate, burst frequency, synchronicity, and population spiking across several months of development [45]. These qualities make organoids a promising model to examine functional aspects of ASD in a developing model. Assembloid systems are likely capable of complex neural activity and ossciations [90], and importantly can be used to probe for the innervation and migration of specific cell types in order to assess cellular circuitry between distinct brain regions.
Due to the nature of these tools and an inability to examine ASD pathology at a cellular resolution, the causative cell populations remain unknown. One potential cell population that may drive these global abnormalities are GABAergic interneurons, which are known to regulate the power of upper and lower- frequencies in the developing brain [91]. It is possible that disruptions in the connectivity of these and other cell types in the fetal brain are what produce the epileptiform changes, which can occur through local miswiring or the failure of a cell population to migrate to its intended destination. It should be noted that these processes arise in early fetal development and occur well before the postnatal time point of clinical assessments such as MRI and EEG in ASD populations.
Neuronal migration is an essential process in the developing brain, where excitatory cells emerge from the ventricular zone to create laminar structures in a well-defined, spatiotemporal manner. Recently, it has been found that a subpopulation of inhibitory neurons is also born from cortical progenitor cells, a phenomenon that appears to be human-specific [92, 93]. The remaining inhibitory neurons follow later in development to emerge from proliferative zones in the ventral telencephalon to migrate into the cortex [94,95,96]. This migratory process is well characterized in the human brain and known to be disrupted in NDDs such as ASD, Tourette Syndrome, and epilepsy [97]. Interneuron migration was recently investigated in human 3D organoid models of Timothy syndrome (TS), a severe neurodevelopmental disorder caused by mutations in the calcium channel, LTCC. Using patient-derived forebrain assembloids composed of cortical (dorsal) and subpallium (ventral) organoids, the researchers were able to identify disruptions in GABAergic interneuron migration originating from the ventral organoid; specifically, their saltatory movements were more frequent but less efficient, moving a lesser distance than control lines. Abnormal calcium signaling was thought to underlie the migration defects, and targeted pharmacological activation of the mutated calcium channel was found to rescue the migration phenotype. Importantly, this abnormality was found exclusively in assembloid-derived ventral organoids, and not in ventral organoids alone, demonstrating the utility of this system in modeling complex circuitry.
Circuitry-based disruptions have also been described more broadly in copy number variant (CNV) models of ASD (15q11.3, 15q13.3, 22q11.2, 22q13.3, 1q21.1) [98,99,100,101,102], which frequently include epileptic comorbidities. These functional deficits in ASD have been explained as an imbalance in the ratio of excitatory: inhibitory cells and have been explored in a cohort of NDD patients with the known ASD-related CNV, 22q13.3. Importantly, this deletion encompasses a lead ASD-risk gene, SHANK3, which is highly expressed in human striatal tissue and has been implicated in corticostriatal circuitry disruptions in ASD individuals [62, 103]. Using an assembloid model of cortical organoids fused to striatal organoids, the group examined the axonal innervation from glutamatergic excitatory neurons into the striatum, which functionally connected to medium spiny neurons in the striatum organoid, similar to a developing brain [62]. Patient-derived assembloids were sliced or dissociated for single-cell patch clamp and calcium signaling, respectively, and both assays showed a hyperexcitable phenotype. Interestingly, this change was not present in individual striatal organoids, demonstrating the importance of assembloid modeling to capture complex circuit-based abnormalities in ASD models.
Migration defects can result in a failure for cellular integration and may contribute to downstream disruptions in cell signaling and activity that is present in ASD models. Disruptions in cell circuitry can be probed on a functional level using tools such as single-cell patch electrophysiology, multielectrode array (MEA) recordings of large neuronal populations, and [two-photon] calcium imaging to record abnormalities in firing patterns. These techniques have been used in the aforementioned migration studies as a means of complimenting the findings, as well as in epileptic studies [90, 104,105,106] to profile synaptic activity in a more sophisticated manner to include recordings of neural oscillations in both single organoids and assembloid systems [82, 90, 107, 108]. Using MEA recordings paired with traditional fMRI and calcium imaging, researchers have identified epileptiform changes that are unique to the brains of those with ASD and epilepsy [109,110,111], and have even used the models to explore unconventional pharmaceutical rescue of these abnormalities [90]. This method has also been used to show dosage-dependent responses of brain organoids to convulsant and antiepileptic compounds in seizure liability drug screens [108], demonstrating the utility of this system as a putative translational medicine tool. It is with these tools that researchers and clinicians can begin to understand circuitry pathogenesis in ASD to better provide targeted therapeutics (Fig. 2). hPSC modeling has truly revolutionized the field of ASD research and has enabled scientists to examine the pathophysiology from the top (phenotyping broad structural and growth components) to the integrated circuitry between different cell types, all the way down to the core mechanistic components to the disorder (Table 2).
Schematic representation of human pluripotent stem cell (hPSC) modeling of autism spectrum disorder (ASD). The top panel describes modeling avenues as genetic, clinical, or environmental, which can be assessed on the molecular, biological, and functional level (bottom left panel). Therapeutic outcomes from each of these assessments are described in the bottom right panel.
Despite the versatility of recording neural activity in 3D models, they are not without caveats. Techniques such as MEA and electrophysiology are difficult to scale up, and often record superficial neuronal populations on the direct surface of the organoid. Furthermore, a clear caveat comes from the assembloid system itself, as it includes the merging of regionalized systems that are created independently (and artificially) rather than together as with a truly developing brain. Despite being able to generate innervation and achieve neuronal migration in a biologically relevant manner, the order in which these processes occur does not represent that of the fetal brain and would require a more sophisticated approach (concurrently guiding merged organoids/co-cultures) to better represent the complex development of neural circuitry in a fetal brain. It is possible that the artificial timing is what hinders production of rare neural cell types and more complex circuitry [112]. There is also considerable variability in the cell types produced that later participate in organoid circuitry; select long-term organoid cultures have been shown to produce cortical progenitor-derived interneurons [82] however this is not the case for all protocols [113]. The absence of these cell types may result from a failure to reach maturation, or a lack of guidance factors provided by neighboring cell types or directed in the neural medium [114]. Current organoid protocols differ in terms of media components, extracellular matrix use, embedding, shaking, and even so far as nomenclature itself. The latter of which has recently been addressed in a joint call for a standardized naming system within the larger organoid community [48], and will likely expand to include more universal protocols in years to come.
Modeling environmental insults in ASD using brain organoids
Prenatal environmental insults, comprising either acquired (e.g., infection, substance use, heavy metal exposure, etc.) or inherent (e.g., vitamin deficiencies, stress, diabetes, etc.) pregnancy and birth complications, have been increasingly linked to NDDs including ASD [115]. However, evidence and specificity of these associations is mostly observational. Organoid systems offer unrestricted temporal access to early human neurodevelopmental milestones to examine these epidemiological associations in vitro through perturbation studies (summarized in Table 3) [115].
Since 2016, 3D hPSC models have been used to study the effect of infectious agents on early brain development. Maternal infection during pregnancy has been associated with an increased risk of ASD in offspring [116], and strong pathogen-host specificity has previously hindered the potential of traditional animal models to reliably recapitulate human transplacental and intrauterine infection [115]. The advantage of brain organoids to model environmental insults first became apparent in 2016, when the cellular basis of Zika virus (ZIKV)-associated microencephaly was investigated in human UNOs [117,118,119]. ZIKVBR-infected organoids exhibited a reduced growth rate and average growth area compared to mock-infected controls [119], which allowed investigators to provide supporting evidence of a causal link between the 2015 Brazilian ZIKV outbreak and increased incidences of congenital brain malformations in the surrounding population. More recently, organoid studies have provided critical insight into the virulence and putative cellular tropisms of SARS-CoV-2 infection in the developing brain [120,121,122,123], as well as potential therapeutic strategies [124]. Other groups have used UNOs or GNOs to explore the consequences of ToRCH infections (e.g., toxoplasmosis [125], cytomegalovirus [126], herpes simplex virus [127], human immunodeficiency virus [128]) on early neurodevelopment [122].
Growing epidemiological evidence implicates substance use and in utero chemical exposure with an increased risk for ASD [129]. To date, various groups have modeled the developmental effect of early exposure to chemical substances using brain organoids, including alcohol [130], nicotine [131], cocaine [132], heavy metals [133], valproic acid [134], and diesel particulate matter [135]. For instance, UNOs continuously exposed to ethanol from day 10–30 of differentiation exhibited increased apoptosis, impaired neurogenesis, and attenuated neurite outgrowth [136]. Furthermore, 2-month UNOs exposed to alcohol levels comparable to binge drinking displayed apoptosis in a cell-type-specific manner, increased metabolic stress, and altered gene expression in key pathways implicated in various neurological diseases [130]. Scalable organoid-based toxicological screens have also shown promise in identifying and assessing the cellular basis of species-specific neurotoxicity [137].
Parental factors and pregnancy complications have also been associated with ASD diagnosis [138]. Organoids allow researchers to investigate the influence of discrete environmental stressors in a controlled environment and precise genetic background, such as maternal stress (e.g., induced by glucocorticoid hormones [139]) and birth complications (e.g., hypoxia [140]). However, despite their clear advantages, organoid modeling of environmental programming is constrained by their inherent limitations. Groups should thoroughly consider the biological implications of missing cell diversity and circuitry during project design and interpretation. For example, 3D cellular models lack intrinsic maternal protective barriers (e.g., placenta, blood-brain separation, xenobiotic inactivation, etc.) that play an essential role in preserving neurodevelopment against environmental insults and may themselves be influenced during environmental programming [115]. Likewise, current environmental perturbation studies using a direct application of a given insult (e.g., toxin) to organoids is not physiologically representative and likely causes exacerbated effects. However, future studies could improve this using organoid transplanted into rodent models where physiological concentrations and drug metabolic processing may be better obtained.
Profiling developmental trajectories across time
Two of the major barriers to ASD therapeutics are a lack of available biomarkers and a poor understanding of disease trajectory. Prior to the use of human-derived hPSC modeling, scientists were limited to postmortem brain tissue to identify neural biomarkers for ASD, which are subject to degradation and often depict less relevant developmental timepoints. Less invasive imaging techniques can be used at more pertinent timepoints, and even across a developmental continuum, however their low resolution fails to capture relevant biological pathways and the repeated measures across time are often between different individuals and underpowered to provide a conclusive understanding of the brain in a disease context. This has been addressed in recent longitudinal imaging studies [141, 142], but require more patient representation to capture the spectrum of the disorder. Capturing the disease trajectory is further complicated by the time at which ASD diagnoses occur; behavioral abnormalities are likely present well after the disorder and many important neurodevelopmental processes are established [2]. For example, deficits in neuron migration or differentiation may be identified at a later time point, but the causative cell population or biological pathways will remain unidentified using basic clinical assessments. Similarly, many critical synaptic pruning events that are disrupted in ASD and identified through MRI are undetectable by adulthood [143], indicating a critical developmental window that must be more thoroughly examined. The identification of vulnerable windows in development can better guide when populations should be assessed for biomarkers. Should biomarkers be identified during a pre-symptomatic period, at-risk children could be better supported, which is critical when considering the increased risk factor for neuropsychiatric illnesses for individuals with ASD diagnosis later in adulthood [144]. 3D organoids faithfully produce cell types in a spatiotemporal manner similar to that of a developing brain, and so disruptions in any of these processes can be assessed spatially, functionally, and through multi-omic approaches (Fig. 2).
Bulk RNA sequencing can be used to elucidate affected pathways in both pre and postnatal development to identify mechanisms underlying alterations in developmental trajectories [111, 145, 146]. In 1- to 3-month-old cortical GNOs, bulk RNA transcriptomic signatures demonstrated that 1-month-old organoids most closely resembled early mid-fetal (13–16 gestational weeks) through late mid-fetal (19–2 gestational weeks) periods, whereas 3-month-old organoids capture profiles of late mid-fetal (19–24 gestational weeks) through neonatal-early infancy (up to 6 months postnatal) developmental periods. The ability of this model to mirror developmental windows across time lends itself to studying early developmental processes, particularly prenatal neurodevelopment that were previously inaccessible using human samples. Downstream patient clinical information can then be combined with these models to guide core developmental questions surrounding neurodevelopment.
Bulk RNA sequencing was also used in patient-derived forebrain assembloids from individuals with Timothy Syndrome, a monogenic form of ASD. Using ventral tissue at multiple timepoints, the team identified alterations in GABAergic signaling at early differentiation stages where they had previously identified disruptions in interneuron migration [57]. They then used weighted gene co-expression network analysis (WGCNA) after gene set enrichment analysis to generate modules of highly correlated genes within the datasets. From these modules, they concluded that GABAergic signaling disruption was likely due to dysregulation of calcium signaling, which was rescued pharmacologically. Taken together, this transcriptomics approach identified early windows in development that are impaired in models of TS syndrome and further distilled the disruptions down to core mechanistic pathways amendable to pharmacological rescue.
Similarly, RNA sequencing in patient-derived brain organoids was used in the common 22q11.2 CNV to profile developmental processes across 100 days of development. This CNV presents highly variable clinical presentations, ranging from cardiac impairments to general developmental delays [147]. Multiple timepoints were used to capture disruptions in biologically relevant pathways such as pattern specification, NPC proliferation, membrane potential regulation, and glial differentiation [148]. The authors were able to identify biologically relevant windows sensitive to neuronal excitability, which they corroborated with functional assays such as single-cell electrophysiology and calcium imaging. Despite the high variability in clinical presentations, these cellular phenotypes remained consistent across multiple patient-derived lines and were even recapitulated with the heterozygous KO of a single gene, DGCR8. This demonstrates the versatility of using organoids to explore potential therapeutic avenues and driver genes within CNVs.
Bulk transcriptomics can identify unbiased biological pathways and biomarkers that may otherwise be missed with conventional phenotyping methods such as immunofluorescence or single gene expression. One notable disadvantage to the technique is the homogenization of highly heterogeneous tissue to capture the average global gene expression; in doing so, subtle intracellular signatures among heterogeneous populations are overlooked. Single-cell RNA sequencing (scRNA Seq) is an alternative transcriptomic approach to capture cell-type identity and individual transcriptomic profiles and trajectories over time. Downstream analyses have been aided with the release of publicly available databases, which include hPSCs, ESCs, embryoid bodies, and neural organoids at multiple developmental timepoints [149, 150]. Information from these databases can be combined with spatial anatomical tools such as the Allen Brain Atlas to provide reliable cell-type identification and pseudotemporal gene expression alignment for identification of developmental trajectories. More recently, scRNA Seq has been combined with lineage tracing inducible CRISPR technology, termed iTracer, to identify cell lineage dynamics and clonality across UNO development [151]. This technique introduces a barcoding library to identify cell types from an hPSC pool, which is retained in daughter cells throughout division and differentiation, and when paired with an inducible CRISPR scar can track lineage dynamics during a discrete window of time. This can putatively be used to identify small windows of changes to cellular fate during UNO development and can be complemented with techniques such as 4D light sheet microscopy to track migration of daughter cells and newly generated neurons. This platform identified lineage disruptions in a neurodevelopmental dysplasia KO model [151], which was paired with 4D spatial sequencing to show disruptions in brain regionalization consistent with these lineage disruptions. Using sophisticated lineage tracing in a heterogeneous human model enables us to ask questions about population-specific dynamics throughout space and time—formerly something that was restricted to animal models.
Understanding transcriptomics on a single-cell level compliments the diversity of cell types that organoids can produce and has helped establish vulnerable cell-type populations to ASD [152, 153]. The exact localization of these cell types in an ASD model has yet to be fully established but can be aided with the use of spatial sequencing to determine the cytoarchitectural microenvironment within individuals with ASD. A recent study examined the adult human cortex using 10x Genomics’ barcoding platform, Visium, to generate spatial maps of gene expression within the human dorsolateral prefrontal cortex. When this dataset was integrated with other NDD datasets, including those of ASD patients, there was a profound layer-specific enrichment of known ASD genes, highlighting the need to understand ASD genetics both spatially and functionally [154]. Defining cellular transcriptomes with spatial resolution is especially relevant when using 3D models that establish brain laminar structure and distinct cytoarchitecture. To this end, spatial transcriptomics lends an unbiased perspective on cell-type-specific abnormalities through generation of spatial gene maps that, when paired with imaging techniques such as MRI, could help delineate structural abnormalities and underlying circuit defects not identified through scRNA Seq alone. Cell population microenvironments can even be probed for activity-dependent pathways to help establish the affected circuits and their regionalization. Spatial transcriptomics have been used in organoids to establish neural lineage dynamics with spatial resolution (iTracer), neurodevelopmental patterning factors [155], and can be paired with fluorescent tagging to isolate or identify specific regionalization within heterogeneous organoid or diseased tissue [156].
The use of transcriptomic profiling provides powerful information about cell identity, lineage, and localization. Analysis pipelines enable the user to infer cell trajectory, intercellular communication [157], and can even be used to predict drug response [158, 159]. Recent developments in electrophysiology can also allow a glimpse into the synaptic activity of a given cell via Patch-Seq, a modified version of whole-cell patch-clamp electrophysiology that enables transcriptomic capturing as well as morphological rendering of a given cell. This three-in-one platform provides comprehensive information about the functionality of a cell as well as its operative biological pathways [160] and could have powerful implications in understanding ASD pathophysiology. It is a low-throughput alternative, however, and should be used selectivity within cell populations that are known to be disrupted in the disorder.
Another tool to capture functionality across time includes multielectrode arrays, which are capable of recording neural populations in 2D and intact 3D cultures across development. Importantly, these recordings are done in an unbiased manner to capture population-wide recordings and synchronous activity of diverse neural populations [107, 108]. MEA recordings in cortical organoids have been shown to correlate with that of human preterm neonatal EEG signatures [82], and can therefore provide a glimpse into the network activity of ASD populations during critical developmental windows. Of note, these oscillations can even be captured in assembloid systems [90], offering the ability to capture inter-organoid circuitry, generalized EEG patterns between both organoids, as well as focal signatures to a particular brain region [after stimulating the other]. Knowledge of how particular brain regions are affected functionally can help clinicians decide which pharmaceutical approaches may be most beneficial to their patients [161,162,163]. They can provide screening opportunities for clinicians to modify pharmaceutical compounds in a controlled environment to target ASD-specific pathways [82, 164,165,166], and can even identify causative driver genes that can be targeted by AAV- or ASO- based gene therapy [166,167,168,169,170,171,172,173]. Following refinement and rigorous testing, patient-derived neural organoids can be used to enhance and personalize cell therapies, gene therapy, and drug discovery, thereby accelerating their transition from the benchtop to the clinic (Fig. 2).
Limitations to human modeling and future directions
2D and 3D human models have made enormous progress in the past decade; with the emergence of stem cell reprogramming, patient-derived skin and blood samples are now capable of producing hPSCs that can later go on to mimic general and brain-region-specific processes. These models have great potential for clinical applications and to understand the mechanisms of ASD pathology. They are not without their caveats, however, which include limitations to growth, tissue maturity, and an absence of vascularization and external input from the peripheral nervous system (summarized in Table 1).
Brain organoids have undergone extensive transcriptomic profiling to show the presence of many different brain-region cell types that emerge in a spatial-temporal manner [44]. Multiple studies have revealed the persistence of a stem cell niche alongside these mature cell types, which supports the use of brain organoids to model fetal development [111]. The presence of this niche is unique to organoids and makes late-stage developmental modeling difficult to achieve. In addition, long-term cultures are further hindered by a lack of vascularization and nutrient flow to the inner organoid core [174], which is compounded by the absence of the blood-brain barrier and its inclusion of immune cells such as microglia. This is especially a limitation to modeling autism, as microglia are a proposed vulnerable cell type within ASD and are thought to contribute towards its immunopathology [175, 176]. Luckily this caveat has been addressed with the introduction of blood vessel organoids that provide vascularization networks at the cellular level, which in turn increase NPC populations, and introduce microglia into the environment [177, 178]. This is incredibly important given the prenatal time point where neurovascularization occurs, the human-specific expression pattern of vascular cells, and its influence on brain structure and development [179].
Microglia populations have also been incorporated into growing organoids through direct co-culture or merging of NPCs and primitive macrophages, which are capable of synaptic pruning and phagocytic activity once mature. These models can be used to investigate the effects of the immune environment on brain pathology [180, 181]. The addition of microglia would provide critical developmental cues to all cell types in the organoids, while supplying a cellular substrate to understand how neuroinflammatory processes occur in NDDs. For example, over-pruning of synapses is one type of deleterious function of abnormal microglia that have yet to be modeled in organoids and would allow complex modeling in 3D.
Despite the enormous progress in modeling specific brain regions through guided differentiation, an element of the CNS that has been underrepresented in human ASD research is the eye. Multiple NDDs are associated with vision disorders, and there has been tremendous advancement of retinal organoid protocols. Retina morphogenesis is a highly regulated process both temporally and spatially, and much like the developing brain requires a stem cell niche that is present in early development [182,183,184]. Individuals who are blind are at least ten times as likely to have ASD, and clinical studies have shown comorbid vision impairment within ASD populations, although the underlying pathogenesis between these two conditions remains unexplained [185, 186]. Retinal organoids are capable of producing retinal pigment epithelia and functional photoreceptors, and their application to ASD modeling would provide novel insights into how retinal development may be impaired and later give rise to visual impairment and dysfunction. Further, the emergence of retinal-cortical assembloids [187], provides the necessary tool to study eye-brain connections in NDDs to understand how dysfunctional sensory input and function may arise.
In recent years, growing evidence suggests the importance of exploring ASD models beyond the CNS due to the high proportion of sensory dysfunctions reported amongst individuals with ASD (as high as 90% [188]). Mouse and fly studies highlight the role of the somatosensory nervous system in ASD sensory and behavioral deficits [189,190,191,192]. Building from the knowledge gained by 2D cultures, dorsal root ganglion-like organoids [193] and neuromuscular organoids [194, 195] have emerged and offer a promising opportunity to investigate the role of the PNS in ASD pathophysiology, such as altered sensory functioning. This approach would integrate external input into what has traditionally been a CNS-exclusive model, providing a more complete understanding of ASD pathogenesis.
An exciting model to examine developmental biology more comprehensively, and with theoretical inclusion of all the systems noted above, include synthetic embryos, or embryoids. These novel systems produce gastrulating embryo-like structures that are capable of undergoing organogenesis [196, 197]. While prototypes have been formed in mouse ESCs, and can only reach 8 days of development, optimizations in a human background may achieve month-long growth periods that would enable scientists a novel glimpse into human fetal development (reviewed in ref. [198]). An important consideration, however, would be the inclusion of stimuli to mimic true gastrulation both within and external to the womb. Notwithstanding, these exciting advancements also give rise to several important ethical considerations. While there have been some preliminary discussions concerning hPSC-based ethics in research [199,200,201,202], these discussions must be formalized, informed by science, and made jointly between experts in the field, policymakers, and activists in order to develop appropriate universal standards.
While each of the models discussed in this review provide novel insight into neural development and circuitry, they remain undoubtedly limited by the natural heterogeneity within both the model itself and the clinical pathophysiology of ASD. It is more likely that these models will provide a starting point for understanding ASD pathogenesis that, when coupled with a multitude of animal and clinical modeling, may ultimately result in a therapeutic breakthrough [203]. Importantly, we also acknowledge the essential role that members of the ASD community play in conducting thoughtful and meaningful research. Self-advocates have expressed their need for improved social support systems, and we hope that the incorporation of those personally affected into decision making will bolster the research done at the bench, and ultimately provide a more comprehensive and compassionate approach to addressing ASD therapeutics and clinical outcomes [204,205,206,207,208]. Given the accelerated pace of brain organoid research over the last few years, this human and patient-specific model system will undoubtedly play a critical role in helping to develop future therapies.
References
Maenner MJ, Shaw KA, Bakian AV, Bilder DA, Durkin MS, Esler A, et al. Prevalence and characteristics of autism spectrum disorder among children aged 8 years—autism and developmental disabilities monitoring network, 11 sites, United States, 2018. MMWR Surveill Summ. 2021;70:1–16.
Association AP. Diagnostic and statistical manual of mental disorders: DSM-5, 5th edn. American Psychiatry Association; Arlington, 2013.
Mazarati AM, Lewis ML, Pittman QJ. Neurobehavioral comorbidities of epilepsy: role of inflammation. Epilepsia. 2017;58(Suppl 3):48–56.
Bozzi Y, Provenzano G, Casarosa S. Neurobiological bases of autism-epilepsy comorbidity: a focus on excitation/inhibition imbalance. Eur J Neurosci. 2018;47:534–48.
Zeidan J, Fombonne E, Scorah J, Ibrahim A, Durkin MS, Saxena S, et al. Global prevalence of autism: a systematic review update. Autism Res. 2022;15:778–90.
McElhanon BO, McCracken C, Karpen S, Sharp WG. Gastrointestinal symptoms in autism spectrum disorder: a meta-analysis. Pediatrics. 2014;133:872–83.
Holingue C, Newill C, Lee LC, Pasricha PJ, Daniele Fallin M. Gastrointestinal symptoms in autism spectrum disorder: A review of the literature on ascertainment and prevalence. Autism Res. 2018;11:24–36.
Mutluer T, Aslan Genc H, Ozcan Morey A, Yapici Eser H, Ertinmaz B, Can M, et al. Population-based psychiatric comorbidity in children and adolescents with autism spectrum disorder: a meta-analysis. Front Psychiatry. 2022;13:856208.
Joshi G, Faraone SV, Wozniak J, Tarko L, Fried R, Galdo M, et al. Symptom profile of ADHD in youth with high-functioning autism spectrum disorder: a comparative study in psychiatrically referred populations. J Atten Disord. 2017;21:846–55.
Eyring KW, Geschwind DH. Three decades of ASD genetics: building a foundation for neurobiological understanding and treatment. Hum Mol Genet. 2021;30:R236–R244.
Tick B, Bolton P, Happe F, Rutter M, Rijsdijk F. Heritability of autism spectrum disorders: a meta-analysis of twin studies. J Child Psychol Psychiatry. 2016;57:585–95.
Uddin M, Unda BK, Kwan V, Holzapfel NT, White SH, Chalil L, et al. OTUD7A regulates neurodevelopmental phenotypes in the 15q13.3 microdeletion syndrome. Am J Hum Genet. 2018;102:278–95.
Richter M, Murtaza N, Scharrenberg R, White SH, Johanns O, Walker S, et al. Altered TAOK2 activity causes autism-related neurodevelopmental and cognitive abnormalities through RhoA signaling. Mol Psychiatry. 2019;24:1329–50.
Satterstrom FK, Kosmicki JA, Wang J, Breen MS, De Rubeis S, An JY, et al. Large-scale exome sequencing study implicates both developmental and functional changes in the neurobiology of autism. Cell. 2020;180:568–584 e523.
Scharrenberg R, Richter M, Johanns O, Meka DP, Rucker T, Murtaza N, et al. TAOK2 rescues autism-linked developmental deficits in a 16p11.2 microdeletion mouse model. Mol Psychiatry. 2022; https://doi.org/10.1038/s41380-022-01785-3.
Yan QJ, Asafo-Adjei PK, Arnold HM, Brown RE, Bauchwitz RP. A phenotypic and molecular characterization of the fmr1-tm1Cgr fragile X mouse. Genes Brain Behav. 2004;3:337–59.
Zhou X, Feliciano P, Shu C, Wang T, Astrovskaya I, Hall JB, et al. Integrating de novo and inherited variants in 42,607 autism cases identifies mutations in new moderate-risk genes. Nat Genet. 2022;54:1305–19.
Unda BK, Chalil L, Yoon S, Kilpatrick S, Irwin C, Xing S, et al. Impaired OTUD7A-dependent Ankyrin regulation mediates neuronal dysfunction in mouse and human models of the 15q13.3 microdeletion syndrome. Mol Psychiatry. 2023. https://doi.org/10.1038/s41380-022-01937-5.
Chailangkarn T, Trujillo CA, Freitas BC, Hrvoj-Mihic B, Herai RH, Yu DX, et al. A human neurodevelopmental model for Williams syndrome. Nature. 2016;536:338–43.
Deneault E, White SH, Rodrigues DC, Ross PJ, Faheem M, Zaslavsky K, et al. Complete disruption of autism-susceptibility genes by gene editing predominantly reduces functional connectivity of isogenic human neurons. Stem Cell Rep. 2018;11:1211–25.
Roberts JE, Bradshaw J, Will E, Hogan AL, McQuillin S, Hills K. Emergence and rate of autism in fragile X syndrome across the first years of life. Dev Psychopathol. 2020;32:1335–52.
The Dutch-Belgian Fragile X Corsorthium, Bakker CE, Verheij C, Willemsen R, van der Helm R, Oerlemans F, et al. Fmr1 knockout mice: a model to study fragile X mental retardation. Cell. 1994;78:23–33.
Marshall JJ, Mason JO. Mouse vs man: organoid models of brain development & disease. Brain Res. 2019;1724:146427.
Nestler EJ, Hyman SE. Animal models of neuropsychiatric disorders. Nat Neurosci. 2010;13:1161–9.
Takahashi K, Yamanaka S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell. 2006;126:663–76.
de la Torre-Ubieta L, Won H, Stein JL, Geschwind DH. Advancing the understanding of autism disease mechanisms through genetics. Nat Med. 2016;22:345–61.
Zhang W, Ross PJ, Ellis J, Salter MW. Targeting NMDA receptors in neuropsychiatric disorders by drug screening on human neurons derived from pluripotent stem cells. Transl Psychiatry. 2022;12:243.
Dixon TA, Muotri AR. Advancing preclinical models of psychiatric disorders with human brain organoid cultures. Mol Psychiatry. 2022; https://doi.org/10.1038/s41380-022-01708-2.
Yoon KJ, Nguyen HN, Ursini G, Zhang F, Kim NS, Wen Z, et al. Modeling a genetic risk for schizophrenia in iPSCs and mice reveals neural stem cell deficits associated with adherens junctions and polarity. Cell Stem Cell. 2014;15:79–91.
Topol A, English JA, Flaherty E, Rajarajan P, Hartley BJ, Gupta S, et al. Increased abundance of translation machinery in stem cell-derived neural progenitor cells from four schizophrenia patients. Transl Psychiatry. 2015;5:e662.
Sans N, Ezan J, Moreau MM, Montcouquiol M. In: Sala C, Verpelli C, editors. Neuronal and synaptic dysfunction in autism spectrum disorder and intellectual disability. Ch. 13. Academic Press; p. 189–219 2016.
de Anda FC, Rosario AL, Durak O, Tran T, Graff J, Meletis K, et al. Autism spectrum disorder susceptibility gene TAOK2 affects basal dendrite formation in the neocortex. Nat Neurosci. 2012;15:1022–31.
Courchesne E, Carper R, Akshoomoff N. Evidence of brain overgrowth in the first year of life in autism. J Am Med Assoc. 2003;290:337–44.
Dementieva YA, Vance DD, Donnelly SL, Elston LA, Wolpert CM, Ravan SA, et al. Accelerated head growth in early development of individuals with autism. Pediatr Neurol. 2005;32:102–8.
Wang M, Wei PC, Lim CK, Gallina IS, Marshall S, Marchetto MC, et al. Increased neural progenitor proliferation in a hiPSC model of autism induces replication stress-associated genome instability. Cell Stem Cell. 2020;26:221–233 e226.
Marchetto MC, Belinson H, Tian Y, Freitas BC, Fu C, Vadodaria K, et al. Altered proliferation and networks in neural cells derived from idiopathic autistic individuals. Mol Psychiatry. 2017;22:820–35.
Connacher R, Williams M, Prem S, Yeung PL, Matteson P, Mehta M, et al. Autism NPCs from both idiopathic and CNV 16p11.2 deletion patients exhibit dysregulation of proliferation and mitogenic responses. Stem Cell Rep. 2022;17:1380–94.
Readhead B, Hartley BJ, Eastwood BJ, Collier DA, Evans D, Farias R, et al. Expression-based drug screening of neural progenitor cells from individuals with schizophrenia. Nat Commun. 2018;9:4412.
Caiazzo M, Dell’Anno MT, Dvoretskova E, Lazarevic D, Taverna S, Leo D, et al. Direct generation of functional dopaminergic neurons from mouse and human fibroblasts. Nature. 2011;476:224–7.
Chanoumidou K, Hernandez-Rodriguez B, Windener F, Thomas C, Stehling M, Mozafari S, et al. One-step reprogramming of human fibroblasts into oligodendrocyte-like cells by SOX10, OLIG2, and NKX6.2. Stem Cell Rep. 2021;16:771–83.
Pang ZP, Yang N, Vierbuchen T, Ostermeier A, Fuentes DR, Yang TQ, et al. Induction of human neuronal cells by defined transcription factors. Nature. 2011;476:220–3.
Zhang Y, Pak C, Han Y, Ahlenius H, Zhang Z, Chanda S, et al. Rapid single-step induction of functional neurons from human pluripotent stem cells. Neuron. 2013;78:785–98.
Mertens J, Marchetto MC, Bardy C, Gage FH. Evaluating cell reprogramming, differentiation and conversion technologies in neuroscience. Nat Rev Neurosci. 2016;17:424–37.
Amin ND, Pasca SP. Building models of brain disorders with three-dimensional organoids. Neuron. 2018;100:389–405.
Kelley KW, Pasca SP. Human brain organogenesis: toward a cellular understanding of development and disease. Cell. 2022;185:42–61.
Qian X, Song H, Ming GL. Brain organoids: advances, applications and challenges. Development. 2019;146:dev166074.
Lancaster MA, Knoblich JA. Generation of cerebral organoids from human pluripotent stem cells. Nat Protoc. 2014;9:2329–40.
Pasca SP, Arlotta P, Bateup HS, Camp JG, Cappello S, Gage FH, et al. A nomenclature consensus for nervous system organoids and assembloids. Nature. 2022;609:907–10.
Lancaster MA, Renner M, Martin CA, Wenzel D, Bicknell LS, Hurles ME, et al. Cerebral organoids model human brain development and microcephaly. Nature. 2013;501:373–9.
Camp JG, Badsha F, Florio M, Kanton S, Gerber T, Wilsch-Brauninger M, et al. Human cerebral organoids recapitulate gene expression programs of fetal neocortex development. Proc Natl Acad Sci USA. 2015;112:15672–7.
Luo C, Lancaster MA, Castanon R, Nery JR, Knoblich JA, Ecker JR. Cerebral organoids recapitulate epigenomic signatures of the human fetal brain. Cell Rep. 2016;17:3369–84.
Pasca AM, Sloan SA, Clarke LE, Tian Y, Makinson CD, Huber N, et al. Functional cortical neurons and astrocytes from human pluripotent stem cells in 3D culture. Nat Methods. 2015;12:671–8.
Lancaster MA, Corsini NS, Wolfinger S, Gustafson EH, Phillips AW, Burkard TR, et al. Guided self-organization and cortical plate formation in human brain organoids. Nat Biotechnol. 2017;35:659–66.
Xiang Y, Tanaka Y, Patterson B, Kang YJ, Govindaiah G, Roselaar N, et al. Fusion of regionally specified hPSC-derived organoids models human brain development and interneuron migration. Cell Stem Cell. 2017;21:383–398 e387.
Eiraku M, Watanabe K, Matsuo-Takasaki M, Kawada M, Yonemura S, Matsumura M, et al. Self-organized formation of polarized cortical tissues from ESCs and its active manipulation by extrinsic signals. Cell Stem Cell. 2008;3:519–32.
Qian X, Nguyen HN, Song MM, Hadiono C, Ogden SC, Hammack C, et al. Brain-region-specific organoids using mini-bioreactors for modeling ZIKV exposure. Cell. 2016;165:1238–54.
Birey F, Andersen J, Makinson CD, Islam S, Wei W, Huber N, et al. Assembly of functionally integrated human forebrain spheroids. Nature. 2017;545:54–59.
Bagley JA, Reumann D, Bian S, Levi-Strauss J, Knoblich JA. Fused cerebral organoids model interactions between brain regions. Nat Methods. 2017;14:743–51.
Mariani J, Simonini MV, Palejev D, Tomasini L, Coppola G, Szekely AM, et al. Modeling human cortical development in vitro using induced pluripotent stem cells. Proc Natl Acad Sci USA. 2012;109:12770–5.
Jo J, Xiao Y, Sun AX, Cukuroglu E, Tran HD, Goke J, et al. Midbrain-like organoids from human pluripotent stem cells contain functional dopaminergic and neuromelanin-producing neurons. Cell Stem Cell. 2016;19:248–57.
Xiang Y, Tanaka Y, Cakir B, Patterson B, Kim KY, Sun P, et al. hESC-derived thalamic organoids form reciprocal projections when fused with cortical organoids. Cell Stem Cell. 2019;24:487–497 e487.
Miura Y, Li MY, Birey F, Ikeda K, Revah O, Thete MV, et al. Generation of human striatal organoids and cortico-striatal assembloids from human pluripotent stem cells. Nat Biotechnol. 2020;38:1421–30.
Ozone C, Suga H, Eiraku M, Kadoshima T, Yonemura S, Takata N, et al. Functional anterior pituitary generated in self-organizing culture of human embryonic stem cells. Nat Commun. 2016;7:10351.
Huang WK, Wong SZH, Pather SR, Nguyen PTT, Zhang F, Zhang DY, et al. Generation of hypothalamic arcuate organoids from human induced pluripotent stem cells. Cell Stem Cell. 2021;28:1657–1670 e1610.
Sakaguchi H, Kadoshima T, Soen M, Narii N, Ishida Y, Ohgushi M, et al. Generation of functional hippocampal neurons from self-organizing human embryonic stem cell-derived dorsomedial telencephalic tissue. Nat Commun. 2015;6:8896.
Pellegrini L, Bonfio C, Chadwick J, Begum F, Skehel M, Lancaster MA. Human CNS barrier-forming organoids with cerebrospinal fluid production. Science. 2020;369:eaaz5626.
Muguruma K, Nishiyama A, Kawakami H, Hashimoto K, Sasai Y. Self-organization of polarized cerebellar tissue in 3D culture of human pluripotent stem cells. Cell Rep. 2015;10:537–50.
Eura N, Matsui TK, Luginbuhl J, Matsubayashi M, Nanaura H, Shiota T, et al. Brainstem organoids from human pluripotent stem cells. Front Neurosci. 2020;14:538.
Ogura T, Sakaguchi H, Miyamoto S, Takahashi J. Three-dimensional induction of dorsal, intermediate and ventral spinal cord tissues from human pluripotent stem cells. Development. 2018;145:dev162214.
Andersen J, Revah O, Miura Y, Thom N, Amin ND, Kelley KW, et al. Generation of functional human 3D cortico-motor assembloids. Cell. 2020;183:1913–1929 e1926.
Sparks BF, Friedman SD, Shaw DW, Aylward EH, Echelard D, Artru AA, et al. Brain structural abnormalities in young children with autism spectrum disorder. Neurology. 2002;59:184–92.
Stanfield AC, McIntosh AM, Spencer MD, Philip R, Gaur S, Lawrie SM. Towards a neuroanatomy of autism: a systematic review and meta-analysis of structural magnetic resonance imaging studies. Eur Psychiatry. 2008;23:289–99.
Richards R, Greimel E, Kliemann D, Koerte IK, Schulte-Korne G, Reuter M, et al. Increased hippocampal shape asymmetry and volumetric ventricular asymmetry in autism spectrum disorder. Neuroimage Clin. 2020;26:102207.
Turner AH, Greenspan KS, van Erp TGM. Pallidum and lateral ventricle volume enlargement in autism spectrum disorder. Psychiatry Res Neuroimaging. 2016;252:40–45.
Pan YH, Wu N, Yuan XB. Toward a better understanding of neuronal migration deficits in autism spectrum disorders. Front Cell Dev Biol. 2019;7:205.
D’Mello AM, Stoodley CJ. Cerebro-cerebellar circuits in autism spectrum disorder. Front Neurosci. 2015;9:408.
Di Martino A, Kelly C, Grzadzinski R, Zuo XN, Mennes M, Mairena MA, et al. Aberrant striatal functional connectivity in children with autism. Biol Psychiatry. 2011;69:847–56.
Hull JV, Dokovna LB, Jacokes ZJ, Torgerson CM, Irimia A, Van Horn JD. Resting-state functional connectivity in autism spectrum disorders: a review. Front Psychiatry. 2016;7:205.
Li Y, Muffat J, Omer A, Bosch I, Lancaster MA, Sur M, et al. Induction of expansion and folding in human cerebral organoids. Cell Stem Cell. 2017;20:385–396 e383.
Dhaliwal N, Choi WWY, Muffat J, Li Y. Modeling PTEN overexpression-induced microcephaly in human brain organoids. Mol Brain. 2021;14:131.
Birey F, Li MY, Gordon A, Thete MV, Valencia AM, Revah O, et al. Dissecting the molecular basis of human interneuron migration in forebrain assembloids from Timothy syndrome. Cell Stem Cell. 2022;29:248–264 e247.
Trujillo CA, Gao R, Negraes PD, Gu J, Buchanan J, Preissl S, et al. Complex oscillatory waves emerging from cortical organoids model early human brain network development. Cell Stem Cell. 2019;25:558–569 e557.
Rubenstein JL, Merzenich MM. Model of autism: increased ratio of excitation/inhibition in key neural systems. Genes Brain Behav. 2003;2:255–67.
Tuchman R, Rapin I. Epilepsy in autism. Lancet Neurol. 2002;1:352–8.
Chen JA, Penagarikano O, Belgard TG, Swarup V, Geschwind DH. The emerging picture of autism spectrum disorder: genetics and pathology. Annu Rev Pathol. 2015;10:111–44.
DiCicco-Bloom E, Lord C, Zwaigenbaum L, Courchesne E, Dager SR, Schmitz C, et al. The developmental neurobiology of autism spectrum disorder. J Neurosci. 2006;26:6897–906.
Penagarikano O, Abrahams BS, Herman EI, Winden KD, Gdalyahu A, Dong H, et al. Absence of CNTNAP2 leads to epilepsy, neuronal migration abnormalities, and core autism-related deficits. Cell. 2011;147:235–46.
McVicar KA, Ballaban-Gil K, Rapin I, Moshe SL, Shinnar S. Epileptiform EEG abnormalities in children with language regression. Neurology. 2005;65:129–31.
Milovanovic M, Grujicic R. Electroencephalography in assessment of autism spectrum disorders: a review. Front Psychiatry. 2021;12:686021.
Samarasinghe RA, Miranda OA, Buth JE, Mitchell S, Ferando I, Watanabe M, et al. Identification of neural oscillations and epileptiform changes in human brain organoids. Nat Neurosci. 2021;24:1488–1500.
Tierney AL, Gabard-Durnam L, Vogel-Farley V, Tager-Flusberg H, Nelson CA. Developmental trajectories of resting EEG power: an endophenotype of autism spectrum disorder. PLoS ONE. 2012;7:e39127.
Bandler RC, Vitali I, Delgado RN, Ho MC, Dvoretskova E, Ibarra Molinas JS, et al. Single-cell delineation of lineage and genetic identity in the mouse brain. Nature. 2022;601:404–9.
Delgado RN, Allen DE, Keefe MG, Mancia Leon WR, Ziffra RS, Crouch EE, et al. Individual human cortical progenitors can produce excitatory and inhibitory neurons. Nature. 2022;601:397–403.
Bandler RC, Mayer C. Deciphering inhibitory neuron development: the paths to diversity. Curr Opin Neurobiol. 2023;79:102691.
Gelman D, Griveau A, Dehorter N, Teissier A, Varela C, Pla R, et al. A wide diversity of cortical GABAergic interneurons derives from the embryonic preoptic area. J Neurosci. 2011;31:16570–80.
Wonders CP, Anderson SA. The origin and specification of cortical interneurons. Nat Rev Neurosci. 2006;7:687–96.
Rapanelli M, Frick LR, Pittenger C. The role of interneurons in autism and Tourette syndrome. Trends Neurosci. 2017;40:397–407.
Fejgin K, Nielsen J, Birknow MR, Bastlund JF, Nielsen V, Lauridsen JB, et al. A mouse model that recapitulates cardinal features of the 15q13.3 microdeletion syndrome including schizophrenia- and epilepsy-related alterations. Biol Psychiatry. 2014;76:128–37.
Meechan DW, Maynard TM, Tucker ES, Fernandez A, Karpinski BA, Rothblat LA, et al. Modeling a model: mouse genetics, 22q11.2 Deletion Syndrome, and disorders of cortical circuit development. Prog Neurobiol. 2015;130:1–28.
Sanders SJ, He X, Willsey AJ, Ercan-Sencicek AG, Samocha KE, Cicek AE, et al. Insights into autism spectrum disorder genomic architecture and biology from 71 risk loci. Neuron. 2015;87:1215–33.
Silva AI, Haddon JE, Ahmed Syed Y, Trent S, Lin TE, Patel Y, et al. Cyfip1 haploinsufficient rats show white matter changes, myelin thinning, abnormal oligodendrocytes and behavioural inflexibility. Nat Commun. 2019;10:3455.
Walsh JJ, Christoffel DJ, Heifets BD, Ben-Dor GA, Selimbeyoglu A, Hung LW, et al. 5-HT release in nucleus accumbens rescues social deficits in mouse autism model. Nature. 2018;560:589–94.
Bey AL, Wang X, Yan H, Kim N, Passman RL, Yang Y, et al. Brain region-specific disruption of Shank3 in mice reveals a dissociation for cortical and striatal circuits in autism-related behaviors. Transl Psychiatry. 2018;8:94.
Ammothumkandy A, Ravina K, Wolseley V, Tartt AN, Yu PN, Corona L, et al. Altered adult neurogenesis and gliogenesis in patients with mesial temporal lobe epilepsy. Nat Neurosci. 2022;25:493–503.
DeRosa BA, El Hokayem J, Artimovich E, Garcia-Serje C, Phillips AW, Van Booven D, et al. Convergent pathways in idiopathic autism revealed by time course transcriptomic analysis of patient-derived neurons. Sci Rep. 2018;8:8423.
Russo FB, Freitas BC, Pignatari GC, Fernandes IR, Sebat J, Muotri AR, et al. Modeling the interplay between neurons and astrocytes in autism using human induced pluripotent stem cells. Biol Psychiatry. 2018;83:569–78.
Foliaki ST, Schwarz B, Groveman BR, Walters RO, Ferreira NC, Orru CD, et al. Neuronal excitatory-to-inhibitory balance is altered in cerebral organoid models of genetic neurological diseases. Mol Brain. 2021;14:156.
Yokoi R, Shibata M, Odawara A, Ishibashi Y, Nagafuku N, Matsuda N, et al. Analysis of signal components <500 Hz in brain organoids coupled to microelectrode arrays: a reliable test-bed for preclinical seizure liability assessment of drugs and screening of antiepileptic drugs. Biochem Biophys Rep. 2021;28:101148.
Du Y, Fu Z, Xing Y, Lin D, Pearlson G, Kochunov P, et al. Evidence of shared and distinct functional and structural brain signatures in schizophrenia and autism spectrum disorder. Commun Biol. 2021;4:1073.
Griesi-Oliveira K, Acab A, Gupta AR, Sunaga DY, Chailangkarn T, Nicol X, et al. Modeling non-syndromic autism and the impact of TRPC6 disruption in human neurons. Mol Psychiatry. 2015;20:1350–65.
Urresti J, Zhang P, Moran-Losada P, Yu NK, Negraes PD, Trujillo CA, et al. Cortical organoids model early brain development disrupted by 16p11.2 copy number variants in autism. Mol Psychiatry. 2021;26:7560–80.
Kanton S, Pasca SP. Human assembloids. Development 2022;149:dev201120.
Gordon A, Yoon SJ, Tran SS, Makinson CD, Park JY, Andersen J, et al. Long-term maturation of human cortical organoids matches key early postnatal transitions. Nat Neurosci. 2021;24:331–42.
Mayhew CN, Singhania R. A review of protocols for brain organoids and applications for disease modeling. STAR Protoc. 2022;4:101860.
Sarieva K, Mayer S. The effects of environmental adversities on human neocortical neurogenesis modeled in brain organoids. Front Mol Biosci. 2021;8:686410.
Jiang HY, Xu LL, Shao L, Xia RM, Yu ZH, Ling ZX, et al. Maternal infection during pregnancy and risk of autism spectrum disorders: a systematic review and meta-analysis. Brain Behav Immun. 2016;58:165–72.
Dang J, Tiwari SK, Lichinchi G, Qin Y, Patil VS, Eroshkin AM, et al. Zika virus depletes neural progenitors in human cerebral organoids through activation of the innate immune receptor TLR3. Cell Stem Cell. 2016;19:258–65.
Garcez PP, Loiola EC, Madeiro da Costa R, Higa LM, Trindade P, Delvecchio R, et al. Zika virus impairs growth in human neurospheres and brain organoids. Science. 2016;352:816–8.
Cugola FR, Fernandes IR, Russo FB, Freitas BC, Dias JL, Guimaraes KP, et al. The Brazilian Zika virus strain causes birth defects in experimental models. Nature. 2016;534:267–71.
Andrews MG, Mukhtar T, Eze UC, Simoneau CR, Perez Y, Mostajo-Radji MA, et al. Tropism of SARS-CoV-2 for developing human cortical astrocytes. Natl Acad. Sci. U.S.A. 2022. https://doi.org/10.1073/pnas.2122236119.
Jacob F, Pather SR, Huang WK, Zhang F, Wong SZH, Zhou H, et al. Human pluripotent stem cell-derived neural cells and brain organoids reveal SARS-CoV-2 neurotropism predominates in choroid plexus epithelium. Cell Stem Cell. 2020;27:937–950 e939.
Ramani A, Muller L, Ostermann PN, Gabriel E, Abida-Islam P, Muller-Schiffmann A, et al. SARS-CoV-2 targets neurons of 3D human brain organoids. EMBO J. 2020;39:e106230.
Zhang BZ, Chu H, Han S, Shuai H, Deng J, Hu YF, et al. SARS-CoV-2 infects human neural progenitor cells and brain organoids. Cell Res. 2020;30:928–31.
Mesci P, de Souza JS, Martin-Sancho L, Macia A, Saleh A, Yin X, et al. SARS-CoV-2 infects human brain organoids causing cell death and loss of synapses that can be rescued by treatment with Sofosbuvir. PLoS Biol. 2022;20:e3001845.
Seo HH, Han HW, Lee SE, Hong SH, Cho SH, Kim SC, et al. Modelling Toxoplasma gondii infection in human cerebral organoids. Emerg Microbes Infect. 2020;9:1943–54.
Sun G, Chiuppesi F, Chen X, Wang C, Tian E, Nguyen J, et al. Modeling human cytomegalovirus-induced microcephaly in human iPSC-derived brain organoids. Cell Rep. Med. 2020;1:100002.
D’Aiuto L, Bloom DC, Naciri JN, Smith A, Edwards TG, McClain L, et al. Modeling herpes simplex virus 1 infections in human central nervous system neuronal cells using two- and three-dimensional cultures derived from induced pluripotent stem cells. J Virol. 2019;93:e00111-19.
Dos Reis RS, Sant S, Keeney H, Wagner MCE, Ayyavoo V. Modeling HIV-1 neuropathogenesis using three-dimensional human brain organoids (hBORGs) with HIV-1 infected microglia. Sci Rep. 2020;10:15209.
Bolte S, Girdler S, Marschik PB. The contribution of environmental exposure to the etiology of autism spectrum disorder. Cell Mol Life Sci. 2019;76:1275–97.
Arzua T, Yan Y, Jiang C, Logan S, Allison RL, Wells C, et al. Modeling alcohol-induced neurotoxicity using human induced pluripotent stem cell-derived three-dimensional cerebral organoids. Transl Psychiatry. 2020;10:347.
Wang Y, Wang L, Zhu Y, Qin J. Human brain organoid-on-a-chip to model prenatal nicotine exposure. Lab Chip. 2018;18:851–60.
Lee CT, Chen J, Kindberg AA, Bendriem RM, Spivak CE, Williams MP, et al. CYP3A5 mediates effects of cocaine on human neocorticogenesis: studies using an in vitro 3D self-organized hPSC model with a single cortex-like unit. Neuropsychopharmacology. 2017;42:774–84.
Yin F, Zhu Y, Wang Y, Qin J. Engineering brain organoids to probe impaired neurogenesis induced by cadmium. ACS Biomater Sci Eng. 2018;4:1908–15.
Cui K, Wang Y, Zhu Y, Tao T, Yin F, Guo Y, et al. Neurodevelopmental impairment induced by prenatal valproic acid exposure shown with the human cortical organoid-on-a-chip model. Microsyst Nanoeng. 2020;6:49.
Bilinovich SM, Uhl KL, Lewis K, Soehnlen X, Williams M, Vogt D, et al. Integrated RNA sequencing reveals epigenetic impacts of diesel particulate matter exposure in human cerebral organoids. Dev Neurosci. 2020;42:195–207.
Zhu Y, Wang L, Yin F, Yu Y, Wang Y, Shepard MJ, et al. Probing impaired neurogenesis in human brain organoids exposed to alcohol. Integr Biol. 2017;9:968–78.
Renner H, Becker KJ, Kagermeier TE, Grabos M, Eliat F, Gunther P, et al. Cell-type-specific high throughput toxicity testing in human midbrain organoids. Front Mol Neurosci. 2021;14:715054.
Kim JY, Son MJ, Son CY, Radua J, Eisenhut M, Gressier F, et al. Environmental risk factors and biomarkers for autism spectrum disorder: an umbrella review of the evidence. Lancet Psychiatry. 2019;6:590–600.
Cruceanu C, Dony L, Krontira AC, Fischer DS, Roeh S, Di Giaimo R, et al. Cell-type-specific impact of glucocorticoid receptor activation on the developing brain: a cerebral organoid study. Am J Psychiatry. 2022;179:375–87.
Pasca AM, Park JY, Shin HW, Qi Q, Revah O, Krasnoff R, et al. Human 3D cellular model of hypoxic brain injury of prematurity. Nat Med. 2019;25:784–91.
Lin HY, Perry A, Cocchi L, Roberts JA, Tseng WI, Breakspear M, et al. Development of frontoparietal connectivity predicts longitudinal symptom changes in young people with autism spectrum disorder. Transl Psychiatry. 2019;9:86.
Oldehinkel M, Mennes M, Marquand A, Charman T, Tillmann J, Ecker C, et al. Altered connectivity between cerebellum, visual, and sensory-motor networks in autism spectrum disorder: results from the EU-AIMS Longitudinal European Autism Project. Biol Psychiatry Cogn Neurosci Neuroimaging. 2019;4:260–70.
Redcay E, Courchesne E. When is the brain enlarged in autism? A meta-analysis of all brain size reports. Biol Psychiatry. 2005;58:1–9.
Jadav N, Bal VH. Associations between co-occurring conditions and age of autism diagnosis: implications for mental health training and adult autism research. Autism Res. 2022; https://doi.org/10.1002/aur.2808.
Chau KK, Zhang P, Urresti J, Amar M, Pramod AB, Chen J, et al. Full-length isoform transcriptome of the developing human brain provides further insights into autism. Cell Rep. 2021;36:109631.
Kang HJ, Kawasawa YI, Cheng F, Zhu Y, Xu X, Li M, et al. Spatio-temporal transcriptome of the human brain. Nature. 2011;478:483–9.
McDonald-McGinn DM, Sullivan KE, Marino B, Philip N, Swillen A, Vorstman JA, et al. 22q11.2 deletion syndrome. Nat Rev Dis Prim. 2015;1:15071.
Khan TA, Revah O, Gordon A, Yoon SJ, Krawisz AK, Goold C, et al. Neuronal defects in a human cellular model of 22q11.2 deletion syndrome. Nat Med. 2020;26:1888–98.
Fleck JS, Sanchis-Calleja F, He Z, Santel M, Boyle MJ, Camp JG, et al. Resolving organoid brain region identities by mapping single-cell genomic data to reference atlases. Cell Stem Cell. 2021;28:1148–1159 e1148.
Kanton S, Boyle MJ, He Z, Santel M, Weigert A, Sanchis-Calleja F, et al. Organoid single-cell genomic atlas uncovers human-specific features of brain development. Nature. 2019;574:418–22.
He Z, Maynard A, Jain A, Gerber T, Petri R, Lin HC, et al. Lineage recording in human cerebral organoids. Nat Methods. 2022;19:90–99.
Guan J, Lin Y, Ji G. Cell type-specific gene network-based analysis depicts the heterogeneity of autism spectrum disorder. Front Cell Neurosci. 2020;14:59.
Velmeshev D, Schirmer L, Jung D, Haeussler M, Perez Y, Mayer S, et al. Single-cell genomics identifies cell type-specific molecular changes in autism. Science. 2019;364:685–9.
Maynard KR, Collado-Torres L, Weber LM, Uytingco C, Barry BK, Williams SR, et al. Transcriptome-scale spatial gene expression in the human dorsolateral prefrontal cortex. Nat Neurosci. 2021;24:425–36.
Legnini I, Emmenegger L, Zappulo A, Wurmus R, Martinez AO, Jara CC, et al. Spatio-temporal, optogenetic control of gene expression in organoids. [Preprint]. 2021. Available from: https://doi.org/10.1101/2021.09.26.461850.
Genshaft AS, Ziegler CGK, Tzouanas CN, Mead BE, Jaeger AM, Navia AW, et al. Live cell tagging tracking and isolation for spatial transcriptomics using photoactivatable cell dyes. Nat Commun. 2021;12:4995.
Jin S, Guerrero-Juarez CF, Zhang L, Chang I, Ramos R, Kuan CH, et al. Inference and analysis of cell-cell communication using CellChat. Nat Commun. 2021;12:1088.
Hu Y, Sui X, Song F, Li Y, Li K, Chen Z, et al. Lung cancer organoids analyzed on microwell arrays predict drug responses of patients within a week. Nat Commun. 2021;12:2581.
Raghavan S, Winter PS, Navia AW, Williams HL, DenAdel A, Lowder KE, et al. Microenvironment drives cell state, plasticity, and drug response in pancreatic cancer. Cell. 2021;184:6119–6137 e6126.
Cadwell CR, Scala F, Li S, Livrizzi G, Shen S, Sandberg R, et al. Multimodal profiling of single-cell morphology, electrophysiology, and gene expression using Patch-seq. Nat Protoc. 2017;12:2531–53.
Izpisua Belmonte JC, Callaway EM, Caddick SJ, Churchland P, Feng G, Homanics GE, et al. Brains, genes, and primates. Neuron. 2015;86:617–31.
Jennings CG, Landman R, Zhou Y, Sharma J, Hyman J, Movshon JA, et al. Opportunities and challenges in modeling human brain disorders in transgenic primates. Nat Neurosci. 2016;19:1123–30.
Kaiser T, Feng G. Modeling psychiatric disorders for developing effective treatments. Nat Med. 2015;21:979–88.
Ilieva M, Aldana BI, Vinten KT, Hohmann S, Woofenden TW, Lukjanska R, et al. Proteomic phenotype of cerebral organoids derived from autism spectrum disorder patients reveal disrupted energy metabolism, cellular components, and biological processes. Mol Psychiatry. 2022;27:3749–59.
Meng Q, Zhang W, Wang X, Jiao C, Xu S, Liu C, et al. Human forebrain organoids reveal connections between valproic acid exposure and autism risk. Transl Psychiatry. 2022;12:130.
Rabeling A, Goolam M. Cerebral organoids as an in vitro model to study autism spectrum disorders. Gene Ther. 2022; https://doi.org/10.1038/s41434-022-00356-z.
Dixon TA, Muotri AR. Advancing preclinical models of psychiatric disorders with human brain organoid cultures. Mol Psychiatry. 2023;28:83–95.
Chen GT, Geschwind DH. Challenges and opportunities for precision medicine in neurodevelopmental disorders. Adv Drug Deliv Rev. 2022;191:114564.
Elamin M, Dumarchey A, Stoddard C, Robinson TM, Cowie C, Gorka D, et al. The role of UBE3A in the autism and epilepsy-related Dup15q syndrome using patient-derived, CRISPR-corrected neurons. Stem Cell Rep. 2023. https://doi.org/10.1016/j.stemcr.2023.02.002.
Foust KD, Nurre E, Montgomery CL, Hernandez A, Chan CM, Kaspar BK. Intravascular AAV9 preferentially targets neonatal neurons and adult astrocytes. Nat Biotechnol. 2009;27:59–65.
Meijboom KE, Abdallah A, Fordham NP, Nagase H, Rodriguez T, Kraus C, et al. CRISPR/Cas9-mediated excision of ALS/FTD-causing hexanucleotide repeat expansion in C9ORF72 rescues major disease mechanisms in vivo and in vitro. Nat Commun. 2022;13:6286.
Bowles KR, Silva MC, Whitney K, Bertucci T, Berlind JE, Lai JD, et al. ELAVL4, splicing, and glutamatergic dysfunction precede neuron loss in MAPT mutation cerebral organoids. Cell. 2021;184:4547–4563 e4517.
Lange J, Zhou H, McTague A. Cerebral organoids and antisense oligonucleotide therapeutics: challenges and opportunities. Front Mol Neurosci. 2022;15:941528.
Giandomenico SL, Mierau SB, Gibbons GM, Wenger LMD, Masullo L, Sit T, et al. Cerebral organoids at the air-liquid interface generate diverse nerve tracts with functional output. Nat Neurosci. 2019;22:669–79.
Krakowiak P, Goines PE, Tancredi DJ, Ashwood P, Hansen RL, Hertz-Picciotto I, et al. Neonatal cytokine profiles associated with autism spectrum disorder. Biol Psychiatry. 2017;81:442–51.
Xu ZX, Kim GH, Tan JW, Riso AE, Sun Y, Xu EY, et al. Elevated protein synthesis in microglia causes autism-like synaptic and behavioral aberrations. Nat Commun. 2020;11:1797.
Matsui TK, Tsuru Y, Hasegawa K, Kuwako KI. Vascularization of human brain organoids. Stem Cells. 2021;39:1017–24.
Sun XY, Ju XC, Li Y, Zeng PM, Wu J, Zhou YY, et al. Generation of vascularized brain organoids to study neurovascular interactions. eLife. 2022;11:e76707.
Chico TJA, Kugler EC. Cerebrovascular development: mechanisms and experimental approaches. Cell Mol Life Sci. 2021;78:4377–98.
Muffat J, Li Y, Omer A, Durbin A, Bosch I, Bakiasi G, et al. Human induced pluripotent stem cell-derived glial cells and neural progenitors display divergent responses to Zika and dengue infections. Proc Natl Acad Sci USA. 2018;115:7117–22.
Xu R, Boreland AJ, Li X, Erickson C, Jin M, Atkins C, et al. Developing human pluripotent stem cell-based cerebral organoids with a controllable microglia ratio for modeling brain development and pathology. Stem Cell Rep. 2021;16:1923–37.
Adler R, Canto-Soler MV. Molecular mechanisms of optic vesicle development: complexities, ambiguities and controversies. Dev Biol. 2007;305:1–13.
Volkner M, Zschatzsch M, Rostovskaya M, Overall RW, Busskamp V, Anastassiadis K, et al. Retinal organoids from pluripotent stem cells efficiently recapitulate retinogenesis. Stem Cell Rep. 2016;6:525–38.
Eiraku M, Takata N, Ishibashi H, Kawada M, Sakakura E, Okuda S, et al. Self-organizing optic-cup morphogenesis in three-dimensional culture. Nature. 2011;472:51–56.
Baron-Cohen S, Scott FJ, Allison C, Williams J, Bolton P, Matthews FE, et al. Prevalence of autism-spectrum conditions: UK school-based population study. Br J Psychiatry. 2009;194:500–9.
Dakin S, Frith U. Vagaries of visual perception in autism. Neuron. 2005;48:497–507.
Fligor CM, Lavekar SS, Harkin J, Shields PK, VanderWall KB, Huang KC, et al. Extension of retinofugal projections in an assembled model of human pluripotent stem cell-derived organoids. Stem Cell Rep. 2021;16:2228–41.
Leekam SR, Nieto C, Libby SJ, Wing L, Gould J. Describing the sensory abnormalities of children and adults with autism. J Autism Dev Disord. 2007;37:894–910.
Mammen MA, Moore GA, Scaramella LV, Reiss D, Ganiban JM, Shaw DS, et al. Infant avoidance during a tactile task predicts autism spectrum behaviors in toddlerhood. Infant Ment Health J. 2015;36:575–87.
Orefice LL, Mosko JR, Morency DT, Wells MF, Tasnim A, Mozeika SM, et al. Targeting peripheral somatosensory neurons to improve tactile-related phenotypes in ASD models. Cell. 2019;178:867–886 e824.
Orefice LL, Zimmerman AL, Chirila AM, Sleboda SJ, Head JP, Ginty DD. Peripheral mechanosensory neuron dysfunction underlies tactile and behavioral deficits in mouse models of ASDs. Cell. 2016;166:299–313.
Wiggins LD, Robins DL, Bakeman R, Adamson LB. Brief report: sensory abnormalities as distinguishing symptoms of autism spectrum disorders in young children. J Autism Dev Disord. 2009;39:1087–91.
Mazzara PG, Muggeo S, Luoni M, Massimino L, Zaghi M, Valverde PT, et al. Frataxin gene editing rescues Friedreich’s ataxia pathology in dorsal root ganglia organoid-derived sensory neurons. Nat Commun. 2020;11:4178.
Faustino Martins JM, Fischer C, Urzi A, Vidal R, Kunz S, Ruffault PL, et al. Self-organizing 3D human trunk neuromuscular organoids. Cell Stem Cell. 2020;27:498.
Pereira JD, DuBreuil DM, Devlin AC, Held A, Sapir Y, Berezovski E, et al. Human sensorimotor organoids derived from healthy and amyotrophic lateral sclerosis stem cells form neuromuscular junctions. Nat Commun. 2021;12:4744.
Amadei G, Handford CE, Qiu C, De Jonghe J, Greenfeld H, Tran M, et al. Embryo model completes gastrulation to neurulation and organogenesis. Nature. 2022;610:143–53.
Tarazi S, Aguilera-Castrejon A, Joubran C, Ghanem N, Ashouokhi S, Roncato F, et al. Post-gastrulation synthetic embryos generated ex utero from mouse naive ESCs. Cell. 2022;185:3290–3306 e3225.
Amin ND, Pasca SP. Mouse embryo models built from stem cells take shape in a dish. Nature. 2022;610:39–40.
Farahany NA, Greely HT, Hyman S, Koch C, Grady C, Pasca SP, et al. The ethics of experimenting with human brain tissue. Nature. 2018;556:429–32.
George RP, Tollefsen C. Embryo: a defense of human life. J Clin Investig. 2008;118:2673–2673.
Jeziorski J, Brandt R, Evans JH, Campana W, Kalichman M, Thompson E, et al. Brain organoids, consciousness, ethics and moral status. Semin Cell Dev Biol. 2022; https://doi.org/10.1016/j.semcdb.2022.03.020.
Lavazza A, Massimini M. Cerebral organoids: ethical issues and consciousness assessment. J Med Ethics. 2018;44:606–10.
Revah O, Gore F, Kelley KW, Andersen J, Sakai N, Chen X, et al. Maturation and circuit integration of transplanted human cortical organoids. Nature. 2022;610:319–26.
Kapp SK, editor. Autistic community and the neurodiversity movement: stories from the frontline. Singapore: Palgrave Macmillan; 2020.
Cameron L, Murphy J. Obtaining consent to participate in research: the issues involved in including people with a range of learning and communication disabilities. Br J Learn Disabil. 2007;35:113–20.
Cascio MA, Weiss JA, Racine E. Making autism research inclusive by attending to intersectionality: a review of the research ethics literature. Rev J Autism Developmental Disord. 2021;8:22–36.
Cascio MA, Weiss JA, Racine E, Autism Research Ethics Task Force. Person-oriented ethics for autism research: Creating best practices through engagement with autism and autistic communities. Autism. 2020;24:1676–90.
Nicolaidis C, Raymaker D, Kapp SK, Baggs A, Ashkenazy E, McDonald K, et al. The AASPIRE practice-based guidelines for the inclusion of autistic adults in research as co-researchers and study participants. Autism. 2019;23:2007–19.
Meganathan K, Prakasam R, Baldridge D, Gontarz P, Zhang B, Urano F, et al. Altered neuronal physiology, development, and function associated with a common chromosome 15 duplication involving CHRNA7. BMC Biol. 2021;19:147.
Mihailovich M, Germain P-L, Shyti R, Pozzi D, Noberini R, Liu Y, et al. 7q11.23 CNV alters protein synthesis and REST-mediated neuronal intrinsic excitability. [Preprint]. 2022. Available from: https://doi.org/10.1101/2022.10.10.511483.
Iefremova V, Manikakis G, Krefft O, Jabali A, Weynans K, Wilkens R, et al. An organoid-based model of cortical development identifies non-cell-autonomous defects in Wnt signaling contributing to Miller-Dieker syndrome. Cell Rep. 2017;19:50–59.
Cavallo F, Troglio F, Faga G, Fancelli D, Shyti R, Trattaro S, et al. High-throughput screening identifies histone deacetylase inhibitors that modulate GTF2I expression in 7q11.23 microduplication autism spectrum disorder patient-derived cortical neurons. Mol Autism. 2020;11:88.
Ross PJ, Zhang WB, Mok RSF, Zaslavsky K, Deneault E, D’Abate L, et al. Synaptic dysfunction in human neurons with autism-associated deletions in PTCHD1-AS. Biol Psychiatry. 2020;87:139–49.
Wegscheid ML, Anastasaki C, Hartigan KA, Cobb OM, Papke JB, Traber JN, et al. Patient-derived iPSC-cerebral organoid modeling of the 17q11.2 microdeletion syndrome establishes CRLF3 as a critical regulator of neurogenesis. Cell Rep. 2021;36:109315.
Bershteyn M, Nowakowski TJ, Pollen AA, Di Lullo E, Nene A, Wynshaw-Boris A, et al. Human iPSC-derived cerebral organoids model cellular features of lissencephaly and reveal prolonged mitosis of outer radial glia. Cell Stem Cell. 2017;20:435–449 e434.
de Jong JO, Llapashtica C, Genestine M, Strauss K, Provenzano F, Sun Y, et al. Cortical overgrowth in a preclinical forebrain organoid model of CNTNAP2-associated autism spectrum disorder. Nat Commun. 2021;12:4087.
Qian X, Su Y, Adam CD, Deutschmann AU, Pather SR, Goldberg EM, et al. Sliced human cortical organoids for modeling distinct cortical layer formation. Cell Stem Cell. 2020;26:766–781 e769.
Raj N, McEachin ZT, Harousseau W, Zhou Y, Zhang F, Merritt-Garza ME, et al. Cell-type-specific profiling of human cellular models of fragile X syndrome reveal PI3K-dependent defects in translation and neurogenesis. Cell Rep. 2021;35:108991.
Trujillo CA, Adams JW, Negraes PD, Carromeu C, Tejwani L, Acab A, et al. Pharmacological reversal of synaptic and network pathology in human MECP2-KO neurons and cortical organoids. EMBO Mol Med. 2021;13:e12523.
Pigoni M, Uzquiano A, Paulsen B, Kedaigle A, Yang SM, Symvoulidis P, et al. Cell-type specific developmental defects in PTEN-mutant cortical organoids converge on abnormal circuit activity. [Preprint]. 2022. Available from: https://doi.org/10.1101/2022.11.15.516664.
Paulsen B, Velasco S, Kedaigle AJ, Pigoni M, Quadrato G, Deo AJ, et al. Autism genes converge on asynchronous development of shared neuron classes. Nature. 2022;602:268–73.
Papes F, Camargo AP, de Souza JS, Carvalho VMA, Szeto RA, LaMontagne E, et al. Transcription factor 4 loss-of-function is associated with deficits in progenitor proliferation and cortical neuron content. Nat Commun. 2022;13:2387.
Sun AX, Yuan Q, Fukuda M, Yu W, Yan H, Lim GGY, et al. Potassium channel dysfunction in human neuronal models of Angelman syndrome. Science. 2019;366:1486–92.
Sen D, Voulgaropoulos A, Drobna Z, Keung AJ. Human cerebral organoids reveal early spatiotemporal dynamics and pharmacological responses of UBE3A. Stem Cell Rep. 2020;15:845–54.
Villa CE, Cheroni C, Dotter CP, Lopez-Tobon A, Oliveira B, Sacco R, et al. CHD8 haploinsufficiency links autism to transient alterations in excitatory and inhibitory trajectories. Cell Rep. 2022;39:110615.
Wang P, Mokhtari R, Pedrosa E, Kirschenbaum M, Bayrak C, Zheng D, et al. CRISPR/Cas9-mediated heterozygous knockout of the autism gene CHD8 and characterization of its transcriptional networks in cerebral organoids derived from iPS cells. Mol Autism. 2017;8:11.
Mellios N, Feldman DA, Sheridan SD, Ip JPK, Kwok S, Amoah SK, et al. MeCP2-regulated miRNAs control early human neurogenesis through differential effects on ERK and AKT signaling. Mol Psychiatry. 2018;23:1051–65.
Zhang W, Ma L, Yang M, Shao Q, Xu J, Lu Z, et al. Cerebral organoid and mouse models reveal a RAB39b-PI3K-mTOR pathway-dependent dysregulation of cortical development leading to macrocephaly/autism phenotypes. Genes Dev. 2020;34:580–97.
Wenderski W, Wang L, Krokhotin A, Walsh JJ, Li H, Shoji H, et al. Loss of the neural-specific BAF subunit ACTL6B relieves repression of early response genes and causes recessive autism. Proc Natl Acad Sci USA. 2020;117:10055–66.
Malara M, Lutz AK, Incearap B, Bauer HF, Cursano S, Volbracht K, et al. SHANK3 deficiency leads to myelin defects in the central and peripheral nervous system. Cell Mol Life Sci. 2022;79:371.
Wang Y, Chiola S, Yang G, Russell C, Armstrong CJ, Wu Y, et al. Modeling human telencephalic development and autism-associated SHANK3 deficiency using organoids generated from single neural rosettes. Nat Commun. 2022;13:5688.
Mariani J, Coppola G, Zhang P, Abyzov A, Provini L, Tomasini L, et al. FOXG1-dependent dysregulation of GABA/glutamate neuron differentiation in autism spectrum disorders. Cell. 2015;162:375–90.
Notaras M, Lodhi A, Barrio-Alonso E, Foord C, Rodrick T, Jones D, et al. Neurodevelopmental signatures of narcotic and neuropsychiatric risk factors in 3D human-derived forebrain organoids. Mol Psychiatry. 2021;26:7760–83.
Bu Q, Huang Y, Li M, Dai Y, Fang X, Chen K, et al. Acrylamide exposure represses neuronal differentiation, induces cell apoptosis and promotes tau hyperphosphorylation in hESC-derived 3D cerebral organoids. Food Chem Toxicol. 2020;144:111643.
Berdenis van Berlekom A, Kubler R, Hoogeboom JW, Vonk D, Sluijs JA, Pasterkamp RJ, et al. Exposure to the amino acids histidine, lysine, and threonine reduces mTOR activity and affects neurodevelopment in a human cerebral organoid model. Nutrients. 2022;14:2175.
Huang Y, Dai Y, Li M, Guo L, Cao C, Huang Y, et al. Exposure to cadmium induces neuroinflammation and impairs ciliogenesis in hESC-derived 3D cerebral organoids. Sci Total Environ. 2021;797:149043.
Ao Z, Cai H, Havert DJ, Wu Z, Gong Z, Beggs JM, et al. One-stop microfluidic assembly of human brain organoids to model prenatal cannabis exposure. Anal Chem. 2020;92:4630–8.
Brown RM, Rana P, Jaeger HK, O’Dowd JM, Balemba OB, Fortunato EA. Human cytomegalovirus compromises development of cerebral organoids. J Virol. 2019;93:e00957-19.
O’Brien BS, Mokry RL, Schumacher ML, Pulakanti K, Rao S, Terhune SS, et al. Downregulation of neurodevelopmental gene expression in iPSC-derived cerebral organoids upon infection by human cytomegalovirus. iScience. 2022;25:104098.
Sison SL, O’Brien BS, Johnson AJ, Seminary ER, Terhune SS, Ebert AD. Human cytomegalovirus disruption of calcium signaling in neural progenitor cells and organoids. J Virol. 2019;93:e00954-19.
Yang L, Zou J, Zang Z, Wang L, Du Z, Zhang D, et al. Di-(2-ethylhexyl) phthalate exposure impairs cortical development in hESC-derived cerebral organoids. Sci Total Environ. 2023;865:161251.
Qiao H, Guo M, Shang J, Zhao W, Wang Z, Liu N, et al. Herpes simplex virus type 1 infection leads to neurodevelopmental disorder-associated neuropathological changes. PLoS Pathog. 2020;16:e1008899.
Qiao H, Zhao W, Guo M, Zhu L, Chen T, Wang J, et al. Cerebral organoids for modeling of HSV-1-induced-amyloid beta associated neuropathology and phenotypic rescue. Int J Mol Sci. 2022;23:5981.
Qiao H, Chiu Y, Liang X, Xia S, Ayrapetyan M, Liu S, et al. Microglia innate immune response contributes to the antiviral defense and blood-CSF barrier function in human choroid plexus organoids during HSV-1 infection. J Med Virol. 2023;95:e28472.
Zhang X, Lin H, Dong L, Xia Q. Recapitulating influenza virus infection and facilitating antiviral and neuroprotective screening in tractable brain organoids. Theranostics. 2022;12:5317–29.
Zhang B, He Y, Xu Y, Mo F, Mi T, Shen QS, et al. Differential antiviral immunity to Japanese encephalitis virus in developing cortical organoids. Cell Death Dis. 2018;9:719.
Adams Y, Clausen AS, Jensen PO, Lager M, Wilhelmsson P, Henningson AJ, et al. 3D blood-brain barrier-organoids as a model for Lyme neuroborreliosis highlighting genospecies dependent organotropism. iScience. 2023;26:105838.
Harbuzariu A, Pitts S, Cespedes JC, Harp KO, Nti A, Shaw AP, et al. Modelling heme-mediated brain injury associated with cerebral malaria in human brain cortical organoids. Sci Rep. 2019;9:19162.
Yao H, Wu W, Cerf I, Zhao HW, Wang J, Negraes PD, et al. Methadone interrupts neural growth and function in human cortical organoids. Stem Cell Res. 2020;49:102065.
Wu W, Yao H, Dwivedi I, Negraes PD, Zhao HW, Wang J, et al. Methadone suppresses neuronal function and maturation in human cortical organoids. Front Neurosci. 2020;14:593248.
Mokry RL, O’Brien BS, Adelman JW, Rosas S, Schumacher ML, Ebert AD, et al. Nitric oxide attenuates human cytomegalovirus infection yet disrupts neural cell differentiation and tissue organization. J Virol. 2022;96:e0012622.
Cai H, Ao Z, Tian C, Wu Z, Kaurich C, Chen Z, et al. Engineering human spinal microphysiological systems to model opioid-induced tolerance. Bioact Mater. 2023;22:482–90.
Kim J, Lee S, Lee J, Park JC, Kim KH, Ko JM, et al. Neurotoxicity of phenylalanine on human iPSC-derived cerebral organoids. Mol Genet Metab. 2022;136:132–44.
Harbuzariu A, Nti A, Harp KO, Cespedes JC, Driss A, Stiles JK. Neuregulin-1/ErbB4 signaling modulates Plasmodium falciparum HRP2-induced damage to brain cortical organoids. iScience. 2022;25:104407.
Wang L, Sievert D, Clark AE, Lee S, Federman H, Gastfriend BD, et al. A human three-dimensional neural-perivascular ‘assembloid’ promotes astrocytic development and enables modeling of SARS-CoV-2 neuropathology. Nat Med. 2021;27:1600–6.
Andrews MG, Mukhtar T, Eze UC, Simoneau CR, Ross J, Parikshak N, et al. Tropism of SARS-CoV-2 for human cortical astrocytes. Proc Natl Acad Sci USA. 2022;119:e2122236119.
Yi SA, Nam KH, Yun J, Gim D, Joe D, Kim YH, et al. Infection of brain organoids and 2D cortical neurons with SARS-CoV-2 pseudovirus. Viruses. 2020;12:1004.
Samudyata, Oliveira AO, Malwade S, Rufino de Sousa N, Goparaju SK, Gracias J, et al. SARS-CoV-2 promotes microglial synapse elimination in human brain organoids. Mol Psychiatry. 2022;27:3939–50.
McMahon CL, Staples H, Gazi M, Carrion R, Hsieh J. SARS-CoV-2 targets glial cells in human cortical organoids. Stem Cell Rep. 2021;16:1156–64.
Lee JA, Bae DH, Choi WH, Cho CH, Bang YS, Yoo J. Effects of sevoflurane exposure on fetal brain development using cerebral organoids. J Mol Neurosci. 2022;72:2440–50.
Zang Z, Yin H, Du Z, Xie R, Yang L, Cai Y, et al. Valproic acid exposure decreases neurogenic potential of outer radial glia in human brain organoids. Front Mol Neurosci. 2022;15:1023765.
Watanabe M, Buth JE, Vishlaghi N, de la Torre-Ubieta L, Taxidis J, Khakh BS, et al. Self-organized cerebral organoids with human-specific features predict effective drugs to combat zika virus infection. Cell Rep. 2017;21:517–32.
Xu YP, Qiu Y, Zhang B, Chen G, Chen Q, Wang M, et al. Zika virus infection induces RNAi-mediated antiviral immunity in human neural progenitors and brain organoids. Cell Res. 2019;29:265–73.
Yoon KJ, Song G, Qian X, Pan J, Xu D, Rho HS, et al. Zika-virus-encoded NS2A disrupts mammalian cortical neurogenesis by degrading adherens junction proteins. Cell Stem Cell. 2017;21:349–358 e346.
Janssens S, Schotsaert M, Karnik R, Balasubramaniam V, Dejosez M, Meissner A, et al. Zika virus alters DNA methylation of neural genes in an organoid model of the developing human brain. mSystems. 2018;3:e00219-17.
Krenn V, Bosone C, Burkard TR, Spanier J, Kalinke U, Calistri A, et al. Organoid modeling of Zika and herpes simplex virus 1 infections reveals virus-specific responses leading to microcephaly. Cell Stem Cell. 2021;28:1362–1379 e1367.
Acknowledgements
We would like to acknowledge the following agencies as funding support for research in our laboratory on the topics discussed in this review: the Canadian Institute of Health Research (CIHR), the Stem Cell Network (SCN), the National Sciences and Engineering Research Council (NSERC) the Network for European Funding for Neuroscience Research (ERA-NET), and the Donald K. Johnson Eye Institute at the Krembil Research Institute. Figures were created with BioRender.com.
Author information
Authors and Affiliations
Contributions
Article conception was performed by SK and KKS. SK and CI wrote the initial draft and was revised by KKS. The figure design was performed by SK and CI. Final revisions were performed by SK.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kilpatrick, S., Irwin, C. & Singh, K.K. Human pluripotent stem cell (hPSC) and organoid models of autism: opportunities and limitations. Transl Psychiatry 13, 217 (2023). https://doi.org/10.1038/s41398-023-02510-6
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41398-023-02510-6