Abstract
The identification of convergent phenotypes in different models of psychiatric illness highlights robust phenotypes that are more likely to be implicated in disease pathophysiology. Here, we utilize human iPSCs harboring distinct mutations in DISC1 that have been found in families with major mental illness. One mutation was engineered to mimic the consequences on DISC1 protein of a balanced translocation linked to mental illness in a Scottish pedigree; the other mutation was identified in an American pedigree with a high incidence of mental illness. Directed differentiation of these iPSCs using NGN2 expression shows rapid conversion to a homogenous population of mature excitatory neurons. Both DISC1 mutations result in reduced DISC1 protein expression, and show subtle effects on certain presynaptic proteins. In addition, RNA sequencing and qPCR showed decreased expression of UNC5D, DPP10, PCDHA6, and ZNF506 in neurons with both DISC1 mutations. Longitudinal analysis of neurite outgrowth revealed decreased neurite outgrowth in neurons with each DISC1 mutation, which was mimicked by UNC5D knockdown and rescued by transient upregulation of endogenous UNC5D. This study shows a narrow range of convergent phenotypes of two mutations found in families with major mental illness, and implicates dysregulated netrin signaling in DISC1 biology.
Similar content being viewed by others
Introduction
One approach to the study of major mental illness is to study disease-associated mutations in cell and animal model systems. However, such mutations often result in relatively subtle alterations in neuronal structure, function, and connectivity. The study of multiple mutations linked to major mental illness in parallel enables the identification of shared phenotypes across disease-associated mutations. This allows for discernment of those phenotypes that are unique to a specific disease-associated mutation from those that may be common to the disease process. Identification of convergent phenotypes will therefore further elucidate disease pathophysiology and hopefully facilitate development of interventions that may alter the course of the disease.
DISC1 has been implicated in major mental illness by rare mutations that have been linked to neuropsychiatric disease, including schizophrenia, bipolar disorder, and autism spectrum disorder1,2,3,4. Controversy exists surrounding the relevance of DISC1 mutation to psychiatric disease given the lack of DISC1 association in large GWAS studies, and the limited number of pedigrees with DISC1 disruption and mental illness (two described to date with more than one carrier). DISC1 was first identified in a Scottish pedigree in which a t(1;11) translocation interrupting DISC1 co-segregates with major mental illness2. One concern with this pedigree is that the translocation does not perfectly segregate with the disease, and other aspects of the genetic background may be contributing to risk. Subsequent studies of this family have shown an association of the balanced translocation with a reduction in cortical thickness, a phenotype which was shared with subjects with schizophrenia2,5,6. The second pedigree is from a smaller American family that harbors a 4 bp deletion at the C-terminus of the gene3. Studies of DISC1 to date implicate the gene as a rare variant that predisposes to disease in select individuals, which GWAS studies are not adept in identifying7. This principle is demonstrated in Alzheimer disease by the strong pathophysiological insights gained from rare mutations in APP, PSEN1, and PSEN2 despite the absence of linkage of these genes to disease in GWAS analyses8. Given these examples and the extensive data linking DISC1 variants to defects in neurodevelopmental processes, we believe that the study of rare disease-associated DISC1 mutations provides an opportunity to elucidate disease-relevant phenotypes important to the proper development of the human brain. In spite of the controversy over DISC1 genetic findings, there is consensus that DISC1 has an important role in neurodevelopment1.
Two different DISC1 mutations have been studied in human induced pluripotent stem cell (iPSC)-derived neurons. One, developed by our group, is modeled after the balanced t(1;11) translocation described above that interrupts the coding sequence of DISC1 early in exon 2,9,10. We used TALENs to create isogenic iPSC lines with a frameshift mutation in exon 8 of DISC1, which induced a nearby premature stop codon, thereby closely recapitulating the effects of DISC1 interruption at the site of the balanced translocation11. Another mutation is a 4 bp deletion at the end of DISC1 exon 12, which was identified in a proband with schizophrenia in an American pedigree3,12. In the Song and Ming labs, iPSCs were generated from family members of this pedigree and were additionally genetically modified to either introduce or correct the DISC1 mutation of interest12. Both our group and the Song/Ming groups employed an embryoid aggregate-based differentiation protocol in our published studies, with some differences in the specific methods across studies. While the embryoid aggregate-based protocol provides a highly valuable technique for studying neurodevelopment, this differentiation method results in a heterogeneous pool of neural cells that follows a protracted human developmental timeline13. This can complicate analyses requiring a homogenous cell population of mature neurons, which may not develop in these cultures for several months.
The advancement of lineage reprogramming has yielded differentiation protocols with accelerated timescales and, depending on the protocol, relatively homogenous cell fates. These methods employ forced expression of critical regulators of cell fate, driving differentiation toward-specific cell types. In comparison to their embryoid-aggregate-based counterparts, transdifferentiation protocols often result in cells that mature more quickly than aggregate-derived cells, and that overcome the protracted development and differentiation of aggregate-based methods. While protocols that faithfully recapitulate development offer insights into disease-associated neurodevelopmental defects, lineage reprogramming provides an opportunity to examine a homogenous population of mature neurons. Increased culture homogeneity also reduces line-to-line differentiation variability, which may reveal otherwise-obscured, subtle, or cell-type-specific effects of disease mutations. Lineage reprogramming thus allows an investigator to bypass neurodevelopmental phenotypes to study the effects of a given mutation in a relatively mature neuron.
Multiple protocols exist for generating glutamatergic neurons from other cell types14,15,16,17,18,19,20,21. Prior studies have used expression of ASCL1, POU3F2, and MYTL to generate mature neurons from cells containing autism and psychiatric disease risk variants22,23. Another method utilizes Neurogenin-2 (NGN2) expression, which directly converts iPSCs to cerebral cortical layer 2/3 excitatory neurons14, and has been used to study disease-associated mutations in models of autism and schizophrenia24,25,26, tuberous sclerosis27, epileptic encephalopathy28, and neurodegenerative diseases29,30.
We previously showed that DISC1 exon 8 disruption alters WNT activity and neural progenitor and neuronal cell fate based on gene and protein expression11, without global effects of altered pre- or post-synaptic gene or protein expression. The study of the DISC1 4 bp deletion found that DISC1 mutation led to altered expression of presynaptic proteins, thereby reducing neuronal activity12. We sought to directly compare these DISC1 mutation models using the same differentiation protocol, in order to determine the convergent phenotypes that exist with both mutations and to better understand altered biology across these models. Given the synaptic phenotype found with DISC1 4 bp deletion, we aimed to compare phenotypes of DISC1 mutation in mature, excitatory neurons using NGN2 transduction. This protocol both enhances consistency due to the homogenous cell population generated14, and aids in the elimination of confounding effects of the developmental phenotype observed with DISC1 exon 8 mutation, which could potentially mask a later synaptic phenotype through alteration of cell fates.
We therefore utilized iPSCs with DISC1 exon 8 mutation (with isogenic wild-type lines) and with the DISC1 exon 12 4 bp deletion (with wild-type lines derived from family members, kindly provided by Drs. Song and Ming) to generate mature glutamatergic neurons using NGN2 transduction and compare neuronal characteristics. In neurons, neither DISC1 mutation resulted in altered global expression of synaptic proteins, but did show subtle defects in a subset of presynaptic proteins. Using RNAseq to evaluate the transcriptome, we found convergence of both DISC1 mutations on significantly decreased expression of just 4 genes after controlling for multiple comparisons, including UNC5D, a netrin receptor. Analysis of neurite outgrowth showed a persistent decrease in neurite length with both exon 8 and exon 12 DISC1 mutations. This phenotype was induced in wild-type neurons by UNC5D knockdown and rescued in DISC1 mutant neurons by UNC5D upregulation. Thus, two independent DISC1 mutations linked to major mental illness result in impaired neurite outgrowth via decreased expression of UNC5D, implicating this pathway in the pathogenesis of psychiatric disorders.
Materials and methods
Induced neuron differentiation
Induced neurons were generated as described14, with minor modifications described below. iPSCs were plated in mTeSR1 media at a density of 95K cells/cm2 on Matrigel-coated plates for viral transduction. Viral plasmids were obtained from Addgene (plasmids #19780, 52047, 30130). Lentiviruses were obtained from Alstem with ultrahigh titers and used at the following concentrations: pTet-O-NGN2-puro: 0.1 µl/50K cells; Tet-O-FUW-eGFP: 0.05 µl/50K cells; Fudelta GW-rtTA: 0.11 µl/50K cells. Transduced cells were dissociated with Accutase and plated onto Matrigel-coated plates at 50,000 cells/cm2 in mTeSR1 (day 0). On day 1, media were changed to KSR media with doxycycline (2 µg/ml, Sigma). Doxycyline was maintained in the media for the remainder of the differentiation. On day 2, media were changed to 1:1 KSR: N2B media with puromycin (5 µg/ml, Gibco). Puromycin was maintained in the media throughout the differentiation. On day 3, media were changed to N2B media + 1:100 B27 supplement (Life Technologies), and puromycin (10 µg/ml). From day 4 on, cells were cultured in NBM media + 1:50 B27 + BDNF, GDNF, CNTF (10 ng/ml, Peprotech).
Embryoid aggregate differentiation
Neuronal differentiation via embryoid aggregate method was performed as previously described31. Briefly, iPSC colonies were removed from MEFs and cultured as embryoid aggregates in suspension for 4 days in iPSC media (without FGF2), followed by 2 days in N2 neural induction media. Day 7 aggregates were plated onto Matrigel-coated 6-well plates and maintained in N2 neural induction media, forming neuroepithelial structures. At day 17, neural rosettes were enzymatically isolated using STEMDiff Neural Rosette Selection Reagent (Stemcell Technologies) and cultured in suspension for 7 days in N2/B27 neural induction media containing cAMP (1 µM, Sigma) and IGF-1 (10 ng/ml, Peprotech). Day 24 neural aggregates were dissociated using Accutase in the presence of 10 µM ROCK inhibitor and plated for final differentiation in neural differentiation media containing cAMP (10 µM, Sigma), IGF-1, BDNF, and GDNF (10 ng/ml, Peprotech).
Induced neuron protocol
-
KSR media: Knockout DMEM, 15% KOSR, 1× MEM-NEAA, 55 µM beta-mercaptoethanol, 1× GlutaMAX (Life Technologies).
-
N2B media: DMEM/F12, 1× GlutaMAX (Life Technologies), 1× N2 supplement B (Stemcell Technologies), 0.3% dextrose (d-(+)-glucose, Sigma).
-
NBM media: Neurobasal medium, 0.5× MEM-NEAA, 1× GlutaMAX (Life Technologies), 0.3% dextrose (d-(+)-glucose, Sigma).
Embryoid aggregate protocol
-
MEF media: DMEM, 10% FBS, 100 U/ml Penicillin-Streptomycin, 2 mM l-glutamine (Life Technologies).
-
iPSC media: DMEM/F12, 20% KOSR, 1× MEM-NEAA, 1× Penicillin-Streptomycin-Glutamine, 55 µM beta- mercaptoethanol (Life Technologies).
-
N2 neural induction media: DMEM/F12, 1× N2 supplement, 1× MEM-NEAA (Life Technologies), 2 µg/ml heparin (Sigma).
-
N2/B27 neural induction media: DMEM/F12, 1× N2 supplement, 1× B27 supplement, 1× MEM-NEAA (Life Technologies), 2 µg/ml heparin (Sigma).
-
Neural differentiation media: Neurobasal medium, 1× N2 supplement, 1× B27 supplement, 1× MEM-NEAA (Life Technologies), 2 µg/ml heparin (Sigma).
RNA sequencing
Double-stranded cDNA was synthesized using the SuperScript III reverse transcriptase protocol with random hexamers on 1 ng mRNA from each sample analyzed. Sequencing libraries were generated by processing the double-stranded cDNA product through the Illumina Nextera tagmentation library protocol. Multiplexed libraries were sequenced on an Illumina NextSeq 500 to a depth of 433 million paired-end reads (75 bases per read) total. RNAseq reads were quality trimmed, then quantified using the Kallisto pseudoalignment quantification program (58) (v0.42.4) running 100 bootstraps against a kallisto index generated from ChGR38 with a k-mer length of 31. Kallisto quantified samples are compared using Sleuth (59) (v0.28.1) in R Studio (v3.4.0 of R; v1.0.143 of R Studio). The ComBat algorithm was used to remove variance induced by differentiation round32.
Single-cell qRT-PCR
For single cell experiments, the NGN2 protocol was modified to include dual SMAD inhibition and WNT inhibition during a critical window of differentiation33. Day 4-induced neurons were co-cultured with mouse astrocytes. WTex8 lines A & B and MUTex8 lines A & D were used for this single-cell analysis. Day 28 cultures were single-cell sorted for GFP expression (marking transduced neurons) into lysis buffer (10 mM Tris, pH 8.0, 0.1 mM EDTA with 0.5% NP40 [Thermo Scientific PI-28324] and 100 U/ml SUPERase•In™ [Ambion AM2696]). Sorted cells in 96-well plates were flash-frozen on dry ice and kept at −80 °C until further processing. Single-cell gene expression profiling was performed using the Fluidigm Biomark dynamic array according to the manufacturer’s protocol. Quantitative RT-PCR was performed using TaqMan Gene Expression Assays (Applied Biosystems) on the Biomark 96.96 Dynamic Array system (Fluidigm).
qRT-PCR
RNA was extracted using the Pure Link RNA Mini Kit (Life Technologies) and reverse transcribed using SuperScript II (Life Technologies). cDNA was used for qPCR with Fast SYBR Green Master Mix (Life Technologies) on a ViiA 7 System (Life Technologies). Samples were run using at least 3 technical replicates (n in figures represent biological replicates only). Data were normalized to GAPDH expression using the ΔΔCT method as previously described34. Primers are listed in Table 1.
Immunocytochemistry and microscopy
Cells were fixed with 4% paraformaldehyde (Sigma), followed by membrane permeabilization and blocking with 0.2% Triton X-100 (Sigma) and 2% donkey serum (Jackson Immunoresearch) in PBS for 1 h at room temperature. Cells were then incubated with primary antibodies overnight at 4 °C, secondary antibodies for 1 h at room temperature, and 1:1000 DAPI (Life Technologies) for 10 min, with multiple washes between each step. Antibodies used are listed in Table 2. Cells were stained for F-actin as indicated, using 670 Fluorescent Phalloidin (Cytoskeleon, Inc., PHDN1) following protocol from manufacturer. Samples were imaged using Zeiss LSM710 or LSM880 confocal microscopes and Zen black software. Zen black and FIJI were used to pseudo-color images and add scale bars.
Western blots
Lysates were prepared in a buffer containing 1% NP40, 10 mM EDTA, 150 mM NaCl, 50 mM Tris, cOmplete Protease Inhibitors and phosSTOP (Roche). BCA protein assays were performed on all samples to normalize for protein content (Pierce). Equal protein amounts were loaded onto 4–12% Bis-Tris NuPAGE gels (Life Technologies) and transferred to nitrocellulose membranes. Blots were either incubated with HRP-conjugated secondary antibodies and developed using ECL substrate (Pierce) or incubated with fluorophore-conjugated secondary antibodies and imaged on the Odyssey system (LI-COR).
LC–MS/MS proteomics
Day 21 iNs were lysed in urea lysis buffer (8 M urea, 100 mM NaHPO4, pH 8.5), including 5 µl (100× stock) HALT protease and phosphatase inhibitor cocktail (Thermo Fisher), and used for label-free LC–MS/MS proteomics through the Emory School of Medicine Proteomics Core, as described35. The ComBat algorithm was used to remove variance induced by differentiation round32.
Neurite outgrowth assay
Day 4 iNs were dissociated and plated on Matrigel coated 96-well clear plates (Greiner Bio-One, 655090) at ~15,000 cells/cm2. Plates were cultured and imaged in an Essen live cell analysis IncuCyte system for up to 120 h (DIV 9). Neurite outgrowth analysis was performed using IncuCyte ZOOM software.
UNC5D knockdown and overexpression
UNC5D knockdown: Day 4 iNs were dissociated and plated in Matrigel coated 96-well µclear plates (Greiner Bio-One, 655090) at 78,125 cells/cm2. Approximately 12 h later, iNs were transduced with either vehicle, empty lentivirus, or 4 lentiviruses encoding UNC5D shRNA (Sigma: TRCN0000419223, TRCN0000061318, TRCN0000438007, TRCN0000061322). Twelve hours after infection, lentivirus was removed and fresh D4 media were added to all wells. Cells were cultured in the IncuCyte for 120 h before fixation with 4% paraformaldehyde (Sigma) or lysing for RNA analysis.
UNC5D activation: UNC5D activation studies using CRISPR/Cas were designed based upon concepts presented in two studies36,37. A dCas9-VPR plasmid (Addgene #6379838) was co-transfected with red fluorescent protein (RFP) and gRNAs targeted to the promoter of UNC5D: #1 CGAGCGGGGGGGGGAGCTGC, #2 GAGGCCGCTCCACGTGCCCC, #3 GGCGGGGCCCGAAGCGCCCC. NGN2-transduced iPSCs were plated onto Matrigel-coated plates in mTeSR1 (day 0). On D2 of iN differentiation, cells were transfected with RFP, gRNAs, and dCas9-VPR following the Lipofectamine 2000 protocol (Thermo Fisher Scientific). On D4, iNs were dissociated and plated on Matrigel coated 96-well µclear plates (Greiner Bio-One, 655090) at ~75,000 cells/cm2. Plates were cultured and imaged in the Essen live cell analysis IncuCyte system for up until day 12. Due to low transfection efficiency, neurite outgrowth analysis was performed using IncuCyte ZOOM software on RFP positive neurites only.
Data collection and statistics
Data were analyzed using GraphPad PRISM 7 software. Values are expressed as means ± SEM. Statistical significance was tested as indicated in figure legends.
Results
DISC1 mutation does not significantly alter cell fate or electrophysiological activity in NGN2 induced neurons
Glutamatergic neurons were generated using viral NGN2 transduction14. Immunostaining of day 21 cells confirmed that NGN2 expression induced rapid neuronal fate conversion and resulted in induced neurons (iNs) expressing excitatory synaptic markers VGLUT1 and VGLUT2, the presynaptic marker SYP, neuronal cytoskeletal proteins MAP2, TUJ1, and TAU (MAPT), the mature neuronal marker NEUN, and cortical upper-layer neuronal transcriptional proteins CUX2 and BRN2 (Fig. 1a). Isogenic wild type and DISC1 exon 8 mutant lines (WTex8 and MUTex8) and familial wild type and DISC1 exon 12 4 bp deletion lines (WTex12 and MUTex12) were all similarly differentiated (Fig. 1a). Two independent lines of each genotype were used, with nomenclature corresponding to prior publications shown in Fig. 1b11,12. Comparison of gene expression in iNs versus embryoid aggregate-differentiated neurons revealed expected decreases in expression of neural progenitor markers and increased expression of mature neuronal and excitatory synaptic markers (Supp Fig. 1). RNA sequencing was performed on day 21 cultures; RNA expression of the genes corresponding to immunostaining markers in Fig. 1a is shown in Fig. 1c. There were no significant differences in expression of these neuronal markers across the 4 genotypes studied. WTex8 and MUTex8 lines additionally were used for single-cell multiplex qRT-PCR with the Fluidigm platform, analyzing expression of 187 genes relevant to neuronal differentiation and psychiatric disease (Supp. Table 1). At a single cell level, expression of the majorinterrupts the coding sequenceity of these genes was very consistent across individual cells, reinforcing the efficiency and consistency of the NGN2 transduction protocol (Fig. 1d, e). There were subtle changes in gene expression, with the most divergent genes shown in the bottom panel of Fig. 1d. Overall, immunostaining, RNA sequencing of pooled cultures, and single-cell Fluidigm analyses revealed consistent and comparable expression of mature excitatory neuronal markers in iNs regardless of DISC1 genotype. NGN2 transduced cultures demonstrated spontaneous action potentials by DIV 8 (Supp Fig. 2a, b). Spontaneous activity increased over differentiation time, but was not significantly different across the 4 genotypes studied (Supp Fig. 2c). These data show that two distinct DISC1 mutations do not dramatically alter cell fate or activity in NGN2-transduced mature excitatory neurons.
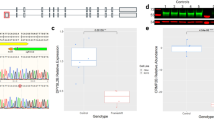
a iPSCs were infected with NGN2 and cultured as iNs to day 21, and immunostained for neuronal markers as shown. Scale bars = 50 μm. b Table summarizing lines used, including genotype as referred to here, lines as named in prior papers11,12, DISC1 genotype, and type of control. c RNA sequencing was performed on day 21 iNs; TPM is shown for the same neuronal markers shown in a. d Single-cell multiplex qRT-PCR (Fluidigm) of 184 cells each from WTex8 and MUTex8 lines for 187 genes. Y-axis shows background-CT, with higher numbers representing higher expression. Genes are arranged left to right and top to bottom from most similar expression across WTex8 and MUTex8 to most divergent expression. e Heatmap of single-cell data in D along with housekeeping genes, grouped by line and genotype. Columns single cells, rows single genes
DISC1 mutations similarly decrease DISC1 protein levels but do not globally dysregulate presynaptic protein expression
The two DISC1 mutations described each induce frameshift mutations at different locations in the coding sequence of the DISC1 gene. However, the proposed mechanisms of these mutations differ. The exon 8 mutation has been shown to decrease DISC1 protein expression in iPSC-derived human neurons via non-sense-medicated decay of the frameshift mutation RNA11. Based on overexpression studies in HEK293 cells, the exon 12 mutation has been proposed to generate a mutant protein that aggregates with wild-type protein, depleting soluble DISC112. Western blot of iN lysates showed similar decreases in protein levels to approximately half wild-type levels with both exon 8 and exon 12 mutation (Fig. 2a, b). This suggests that, regardless of mechanism, both mutations result in haploinsufficiency with a decrease in wild-type DISC1 protein expression. Prior studies in embryoid aggregate-derived neurons showed either no change in presynaptic protein expression (MUTex8)11, or increased expression of presynaptic proteins (MUTex12)12. Using the NGN2 induction protocol in parallel, neither exon 8 nor exon 12 DISC1 mutation significantly altered SYP or SYN1 expression (Fig. 2c, d).
Day 21 iN cultures were lysed and used for western blot or proteomics as shown. a Representative DISC1 western blot. b Quantification of DISC1 expression by western blot, normalized to GAPDH. n = 6 for WTex8, 6 for MUTex8, 6 for WTex12, 6 for MUTex12. c Representative SYN1 and SYP western blot. d, e Quantification of SYN1 and SYP expression by western blot, normalized to GAPDH. n = 10 for WTex8, 10 for MUTex8, 10 for WTex12, 10 for MUTex12. Statistics a–d: Two-tailed Student’s t-test, **p < 0.01. f–j Day 21 iN lysates were used for proteomics by LC–MS/MS, n = 4 for each genotype. f Volcano plot of proteomic data of WTex8 vs MUTex8, with significantly upregulated genes colored blue and significantly downregulated genes colored red. g Volcano plot of proteomic data of WTex12 vs MUTex12, with no genes achieving statistical significance. h Differential expression of select proteins involved in neural adhesion, calcium signaling, and the synapse in MUTex8 vs WTex8 lysates. i, j Expression of presynaptic proteins by proteomics in WTex8/MUTex8i and WTex12/MUTex12j. Statistics f–j: Paired t-test with two-stage linear step-up procedure of Benjamini, Krieger, and Yejutieli, with false discovery rate Q = 5%. See also Supp. Table 2
Unbiased analyses of proteomic data comparing all DISC1 mutant lines to all wild-type lines did not result in any changes in protein expression that were significant after correction for multiple comparisons. This was largely due to variability in protein expression, especially in the WTex12 and MUTex12 samples. As these 4 lines are familial but not isogenic, there may be increased variability both within and across genotypes, reducing the power to detect significant differences in protein expression. Separate paired analyses of WTex8/MUTex8 and WTex12/MUTex12 proteomic data confirmed this, as WTex8/MUTex8 analysis revealed multiple significant changes in protein expression (Fig. 2f), whereas WTex12/MUTex12 data showed no significant changes in expression (Fig. 2g). Altered protein expression with DISC1 exon 8 mutation included dysregulation of proteins involved in synaptic and calcium regulation (Fig. 2h), cholesterol and lysosomal biology, chromatin regulation and DNA synthesis, RNA regulation, and the stress response (Supp Fig. 3). Proteomic analyses of both genotypes showed overall unchanged expression of many presynaptic proteins (Fig. 2i, j). Interestingly, however, and in line with the previous study of MUTex12 neurons12, we observed a trend for increased expression of a subset of presynaptic proteins, with SYT1 being the only protein in this category that was significantly increased in MUTex8 lysates (Fig. 2i, j).
RNA sequencing shows altered expression of UNC5D, PCDHA6, DPP10, and ZNF506 with both DISC1 models
As proteomic analyses are limited by peptide detection of only the most abundant proteins, with peptide detection representing ~4000–5000 proteins, we next performed RNA sequencing of day 21 iNs to assess gene expression changes at a whole-transcriptome level. Although there were similar challenges of variability as in proteomic data, analysis of all mutant (MUTex8-ex12) vs all wild-type (WTex8-ex12) RNA sequencing data showed decreased expression of 4 genes after genome-wide correction for multiple comparisons: UNC5D, PCDHA6, DPP10, and ZNF506 (Fig. 3a). Representation of data across all 4 genotypes showed similar decreases in expression with MUTex8 and MUTex12 for each of these genes (Fig. 3b, d–f). As UNC5D encodes a netrin receptor, it was interesting that expression of LRRC4C (a member of the netrin family and a binding partner for netrin G1) also was decreased though did not meet statistical significance after multiple comparison correction (Fig. 3c). Of note, none of these gene products were detected in proteomic analysis.
a–f RNA was collected from day 21 iNs and used for RNA sequencing. n = 4 for WTex8, 4 for MUTex8, 3 for WTex12, 4 for MUTex12. a Volcano plot of RNA sequencing data of all wild type (WTex8 and WTex12) vs all mutant (MUTex8 and MUTex12). Statistics: two-stage linear step-up procedure of Benjamini, Krieger, and Yejutieli, with Q = 5%. Dotted line shows threshold of significance for Q = 5%. b–f RNAseq data for individual genes as shown, by genotype. g–k Day 21 iN culture RNA was collected and used for qRT-PCR for select genes as shown, normalized to GAPDH. n = 12 for WTex8, 12 for MUTex8, 12 for WTex12, 12 for MUTex12, from three independent differentiations. See also Supp. Table 3
RNA sequencing results were validated in separate RNA samples using qRT-PCR on three independent differentiations, which confirmed decreased expression of UNC5D and DPP10 in MUTex8 and MUTex12 samples (Fig. 3g, k). The remaining genes showed significantly decreased expression in only one mutant genotype by qRT-PCR (Fig. 3h–j). Although variability of expression across lines and genotypes limited the discovery of significant gene expression changes, the few identified here were robust enough to be significant at a whole-transcriptome level despite this variability. Furthermore, decreased UNC5D and DPP10 expression was validated in MUTex8 and MUTex12 genotypes in independent samples by qRT-PCR.
Independent DISC1 mutations converge on reduced neurite outgrowth via decreased UNC5D expression
UNC5D is a co-receptor for netrin with the netrin receptor DCC, and modulates DCC signaling. UNC5D is expressed in the subventricular zone (SVZ) and upper layer neurons, and has been found to regulate the transition to radial migration39–43. The interaction of netrins with netrin receptors allow for chemotropic guidance of migrating axons and cells in the developing brain44. DISC1 has independently been implicated in neurite outgrowth via its interacting proteins as well as in models of DISC1 mutation12,45,46,47,48,49,50,51. In order to study the effects of DISC1 mutation on neurite outgrowth in postmitotic neurons, we used the IncuCyte live cell imaging system to morphologically assay differentiating iNs from all genotypes starting at day 4, allowing for automated measurements of neurite characteristics over time. These studies showed that both exon 8 and exon 12 DISC1 mutations resulted in significant decreases in neurite length relative to wild-type controls, which persisted up to 120 h in vitro (DIV 9, Fig. 4a–d).
Day 4 iNs were dissociated and plated for longitudinal IncuCyte imaging and analysis of neurite length. a Example images from the IncuCyte are shown from day 9 cultures (120 h post D4 plating). Top row = brightfield, middle row = overlay of cell body mask in blue and neurite mask in green on brightfield, bottom row = cell body mask in blue and neurite mask in green. Scale bars = 200 μm. b, c Example longitudinal measurements of neurite length are shown from one experiment for WTex8 vs MUTex8b and WTex12 vs MUTex12c. d Neurite length measurements from 2 to 120 h for all 4 genotypes are shown, with WTex8 and MUTex8 data normalized to WTex8t = 2 h and WTex12 and MUTex12 data normalized to WTex12t = 2 h. n = 6 for each group, three independent differentiations. Statistics: Holm-Sidak method with multiple comparison correction, FDR = 5%. *q < 0.05, ***q < 0.001, ***q < 0.0001
In order to investigate the effects of altered UNC5D expression on neurite outgrowth, we manipulated UNC5D expression and measured neurite outgrowth. In order to first visualize whether UNC5D was expressed at the protein level, we performed immunostaining in day 21 iNs. Immunostaining for UNC5D showed localization to neurites, dendritic spines, and growth cones (example images shown in Fig. 5a). Knockdown of UNC5D expression with four independent shRNAs resulted in a decrease in UNC5D expression in iNs (expressed as % expression relative to controls, mean ± SD: shRNA #1–66.5 ± 11.4, shRNA#2–57.5 ± 7.4, shRNA#3: 77.2 ± 10.3, shRNA#4: 77.1 ± 9.3). Viral transduction of day 4 iNs with each of these shRNAs impaired neurite outgrowth in wild-type iNs (Fig. 5b, c), mimicking the effect of DISC1 exon 8 and exon 12 mutations. We then upregulated endogenous UNC5D expression using a catalytically inactive Cas9 fused to C-terminal VP64 acidic transactivation domain36,37 targeted to the UNC5D promoter sequence. Transfection of these plasmids dramatically increased UNC5D expression in HEK293T cells (Supp Fig. 4A). Although, the transfection efficiency of iNs was low, making increased UNC5D expression difficult to assess in pooled cultures, co-transfection of UNC5D-targeted dCas9-VP64 with RFP allowed for neurite length analysis of only RFP-positive, transfected neurites (Supp Fig. 4B). Transient transactivation of UNC5D expression in MUTex8 iNs resulted in increased RFP-positive neurite length relative to control by 84 h after transfection, rescuing the DISC1 mutant phenotype (Fig. 5d, e). These data suggest that two independent DISC1 mutations result in impaired neurite outgrowth in vitro that is mediated via decreased UNC5D expression.
a Day 9 iNs were immunostained for UNC5D, MAP2, and F-actin as shown, revealing UNC5D expression in neurites, dendritic spines, and growth cones. Scale bar = 20 μm. b–c UNC5D shRNAs were transduced into day 4 iNs. WTex8 day 4 iNs were dissociated and plated for IncuCyte neurite length analysis with media alone (vehicle), viral transduction vehicle (empty), or else transduced with empty virus or 4 shRNAs targeting different sequences within UNC5D. Longitudinal neurite length analysis is shown. d, e MUTex8 day 4 iNs were dissociated and transfected with either dCas9-VP64 + RFP (dCas9) or dCas9-VP64 + UNC5D sgRNA #2 + RFP (dCas9 + UNC5D sgRNA). RFP-positive neurite length was analyzed longitudinally with IncuCyte. e Neurite length analysis at 192 h after plating (DIV 9) shows significantly increased neurite length with transient transactivation of endogenous UNC5D expression with gRNA #2. n = 25 for each group. Statistics: one-way ANOVA, *p < 0.05, **p < 0.01, ***p < 0.001, ***p < 0.0001
Discussion
The advent of “induced neuron” differentiation methods, which accelerate differentiation by overexpressing master regulators of cell fate, allows for detailed study of fate-restricted homogenous populations of mature neurons. In the present study, we use one such method to study the effects of two distinct DISC1 mutations in mature excitatory neurons. NGN2 transduction results in homogenous and mature neuronal cultures, by pooled and single cell gene expression analyses. We find that independent DISC1 mutations decrease levels of detergent-soluble DISC1 protein, consistent with haploinsufficiency. In this cellular model, these mutations do not dramatically dysregulate presynaptic protein expression or spontaneous neural electrophysiological activity. Both DISC1 mutations result in significant decreases in UNC5D and DPP10 expression by unbiased RNA sequencing, confirmed by qRT-PCR. Examination of neurite outgrowth in culture revealed an early and persistent decrease in neurite length with DISC1 exon 8 and exon 12 mutations, which was phenocopied by UNC5D knockdown and rescued by UNC5D upregulation.
These neurite outgrowth findings contrast the prior study of MUTex12 neurons, wherein transient increases in neurite length were observed with DISC1 mutation12. The protein expression findings also differ from the prior study, which found dramatic upregulation of the presynaptic protein apparatus and decreased spontaneous activity in iPSC-derived neurons12. These differences may be a result of the distinct differentiation protocols used—the prior study utilized an embryoid-aggregate method, which recapitulated neurodevelopment, whereas this study bypasses early neurodevelopment via rapid induction of NGN2. Both differentiation protocols have benefits for understanding the neurobiology of DISC1 mutations; however, in order to study convergent effects across two separate mutations, utilizing a protocol that generated a very homogeneous population of neurons ensured less variability across iPSC lines. By using one differentiation method vs another, different results in phenotypes may be uncovered due to the specific population of neurons being studied.
The proteome and transcriptome analyses described here were limited by variability across iPSC lines. This was especially true with the WTex12/MUTex12 lines, which are intra-familial but not isogenic. Conversely, the isogenic WTex8/MUTex8 lines demonstrated increased power to detect significant differences in gene and protein expression, perhaps due to reduction of other genomic variability or else to the relative potency of the mutations. This demonstrates the importance of using isogenic controls in order to strengthen analyses of disease-associated phenotypes. However, identification of gene expression changes and impaired neurite outgrowth even with non-isogenic genomic variability in WTex12/MUTex12 lines demonstrates the robustness of these phenotypes.
While proteomic analysis did not show dramatic dysregulation of synaptic proteins, analysis of WT/MUTex8 lines did reveal altered expression of proteins implicated in neuronal physiology, including the neuronal migration protein CALM152, glutamate generating enzyme GLS, Ca++-sensing protein NCALD53, cell adhesion protein NCAM (implicated in neurite outgrowth, neuronal migration, and synaptic plasticity54), synaptic vesicle recycling protein PPP3Ca (calcineurin A; implicated in neurodevelopmental disorders55,56), alpha-synuclein (SNCA), and SYT1 (a Ca++ sensor critical for fast neurotransmitter release57). These expression perturbations suggest that there may be a subtle dysregulation of synaptic biology in mature glutamatergic neurons with DISC1 disruption, which was only detectable in comparison of isogenic WT/MUTex8 neurons.
We compared the RNAseq data in the current study to previously published RNAseq analyses in neurons with DISC1 disruption. We found that UNC5D, related gene UNC5C, and LRRC4C were downregulated with DISC1 exon 2 homozygous mutation in day 50 embryoid aggregate-derived neurons; ZNF506 was downregulated in MUTex8 day 50 embryoid aggregate-derived neurons; and LRRC4C and ZNF506 were downregulated in MUTex8 day 17 embryoid aggregate-derived NPCs11. The UNC5D-related genes UNC5B and UNC5C were downregulated in MUTex12 day 28 embryoid aggregate-derived neurons12. Despite differences in differentiation protocol and neural cell fate, overlapping gene expression changes of the current study with previous analyses strengthen the association of the observed gene expression changes with DISC1 mutation.
Both DISC1 mutations studied here resulted in a reproducible decrease in UNC5D expression, which was mechanistically linked to decreased neurite outgrowth in DISC1-disrupted neurons. The UNC5 proteins are netrin co-receptors that have been described to interact with DCC to mediate repulsive cues58. UNC5D is expressed in rodents in multipolar migrating cells of the SVZ and intermediate zone (IZ)39,40, as well as layer four cortical neurons40,43,59,60. Migrating cortical projection neurons initially have a bipolar morphology in the ventricular zone (VZ), then transition to a multipolar morphology in the SVZ/IZ where cells extend and retract processes dynamically61, and eventually extend an axon prior to resuming radial migration, transforming back to a bipolar cell morphology and entering the cortical plate41. We previously had shown that DISC1 mutation alters FOXG1 levels11, a telencephalic transcription factor that represses UNC5D expression39. Dynamic FOXG1 expression results in upregulation of UNC5D (required for initiation of multipolar migration), followed by downregulation of UNC5D (required for multipolar-to-bipolar transition and subsequent cortical plate entry)39. UNC5D has also been shown to interact with Netrin-4 to regulate cell survival and interacts with FLRT2 to delay radial migration into the cortical plate via repulsive cue signaling60,62,63. Decreased expression of UNC5D and consequent impaired neurite outgrowth with disease-associated DISC1 mutations thus suggests a possible perturbation of axonal guidance and dysregulation of chemotropic factor-guided radial migration. Interestingly, the UNC5D locus has been associated with cortical brain volume in the Framingham study64, which could implicate this gene in the decreased cortical thickness seen with DISC1 polymorphisms and mutation6,65,66.
The netrin family gene LRRC4C, which was decreased in MUTex8 and MUTex12 by RNAseq (but did not meet threshold for whole-transcriptome significance) and in MUTex12 by qRT-PCR, has been associated with neurodevelopmental disorders in genetic studies67. LRRC4C (also referred to as NGL-1) is a netrin-G1 receptor that promotes axonal outgrowth68, and the NGL family is involved in excitatory synapse formation69. LRRC4C is also phosphorylated by autism spectrum disorder vulnerability gene CDKL5, which modulates the interaction of LRRC4C with PSD-9570. Furthermore, Netrin-G1 (NTNG1) disruptions or polymorphisms have been associated with atypical Rett syndrome, intellectual disability, autism spectrum disorder, and schizophrenia71,72,73,74. Decreased expression of both LRRC4C and UNC5D further strengthens the association of DISC1 disruption with perturbation of cell adhesion and chemotropic guidance molecule-mediated axonal guidance and cell migration. These previously described roles for netrin signaling do not preclude additional roles for UNC5D and LRRC4C, but provide clues to their function in the mature brain.
The other genes identified by RNAseq to be altered with DISC1 exon 8 and exon 12 mutation are DPP10, PCDHA6, and ZNF506. DPP10 modulates the activity of Kv4.2 potassium channels, and has been implicated in mental illness and neurodegenerative disease models75–77 Decreased DPP10 expression suggests a possible synaptic phenotype not identified in the current study, which could be further investigated with detailed electrophysiological analyses. PCDHA6 is a member of the alpha protocadherins, a family of Ca++-dependent cell adhesion molecules78 that have been linked to bipolar disorder and schizophrenia79. ZNF506 is a zinc-finger protein without known function.
DISC1 has been implicated in neuronal migration and neurite outgrowth in several studies (reviewed here1). DISC1 knockdown reduces migration of cultured mouse NPCs80, in vivo rodent cortical neurons81,82,83,84 and interneurons48. DISC1 polymorphisms have also been linked to impaired migration85, and DISC1 phosphorylation has been liked to a switch from NPC proliferation to migration86. In rodents, DISC1 polymorphisms or mutation have been linked to decreased neurite length in vivo47,87 and in primary neuronal cultures88. DISC1 has been linked to neurite outgrowth in immortalized rodent cell cultures, with proposed mechanisms of interactions with NDEL1, FEZ1, DBZ, or dysbindin46,50,81,91,92,93. Study of the DISC1 exon 8 mutation in a three-dimensional cerebral organoid system revealed a marked WNT-dependent morphologic alteration94, which could reflect an alteration in migration related to the decrease in UNC5D expression observed here. Indeed, UNC5D RNA levels were found to be reduced in DISC1 ex8 organoids (see Supp Fig. 5).
Although, the causal link between DISC1 mutation and major mental illness remains controversial, the relationship of DISC1 to neurodevelopment has been well-established1. The current study shows that multiple mutations previously linked to mental illness result in shared gene expression changes and decreased neurite outgrowth related to decreased UNC5D expression. Further study of multiple neuropsychiatric disease-associated mutations in parallel will allow for identification of convergent phenotypes and elucidate shared mechanisms of neurodevelopmental disorder pathophysiology.
References
Brandon, N. J. & Sawa, A. Linking neurodevelopmental and synaptic theories of mental illness through DISC1. Nat. Rev. Neurosci. 12, 707–722 (2011).
Millar, J. K. et al. Disruption of two novel genes by a translocation co-segregating with schizophrenia. Hum. Mol. Genet. 9, 1415–1423 (2000).
Sachs, N. A. et al. A frameshift mutation in disrupted in Schizophrenia 1 in an American family with schizophrenia and schizoaffective disorder. Mol. Psychiatry 10, 758–764 (2005).
Chubb, J. E., Bradshaw, N. J., Soares, D. C., Porteous, D. J. & Millar, J. K. The DISC locus in psychiatric illness. Mol. Psychiatry 13, 36–64 (2008).
Sullivan, P. F. Questions about DISC1 as a genetic risk factor for schizophrenia. Mol. Psychiatry 18, 1050–1052 (2013).
Doyle, O. M. et al. The cortical thickness phenotype of individuals with DISC1 translocation resembles schizophrenia. J. Clin. Invest. 125, 3714–3722 (2015).
Porteous, D. J. et al. DISC1 as a genetic risk factor for schizophrenia and related major mental illness: response to Sullivan. Mol. Psychiatry 19, 141–143 (2014).
Bertram, L., Lill, C. M. & Tanzi, R. E. The genetics of Alzheimer disease: back to the future. Neuron 68, 270–281 (2010).
Blackwood, D. H. et al. Schizophrenia and affective disorders--cosegregation with a translocation at chromosome 1q42 that directly disrupts brain-expressed genes: clinical and P300 findings in a family. Am. J. Hum. Genet 69, 428–433 (2001).
St Clair, D. et al. Association within a family of a balanced autosomal translocation with major mental illness. Lancet 336, 13–16 (1990).
Srikanth, P. et al. Genomic DISC1 disruption in hiPSCs alters Wnt signaling and neural cell fate. Cell Rep. 12, 1414–1429 (2015).
Wen, Z. et al. Synaptic dysregulation in a human iPS cell model of mental disorders. Nature 515, 414–418 (2014).
Shi, Y., Kirwan, P., Smith, J., Robinson, H. P. C. & Livesey, F. J. Human cerebral cortex development from pluripotent stem cells to functional excitatory synapses. Nat. Neurosci. 15, 477–486 (2012).
Zhang, Y. et al. Rapid single-step induction of functional neurons from human pluripotent stem cells. Neuron 78, 785–798 (2013).
Thoma, E. C. et al. Ectopic expression of neurogenin 2 alone is sufficient to induce differentiation of embryonic stem cells into mature neurons. PLoS ONE 7, e38651 (2012).
Chanda, S. et al. Generation of induced neuronal cells by the single reprogramming factor ASCL1. Stem Cell Rep. 3, 282–296 (2014).
Ambasudhan, R. et al. Direct reprogramming of adult human fibroblasts to functional neurons under defined conditions. Cell Stem Cell 9, 113–118 (2011).
Heinrich, C. et al. Directing astroglia from the cerebral cortex into subtype specific functional neurons. PLoS Biol. 8, e1000373 (2010).
Yang, Y. et al. Direct conversion of adipocyte progenitors into functional neurons. Cell Reprogram. 15, 484–489 (2013).
Ho, S.-M., Hartley, B. J., Tcw, J., Beaumont, M., Stafford, K. & Slesinger, P. A. et al. Rapid Ngn2-induction of excitatory neurons from hiPSC-derived neural progenitor cells. Methods 101, 113–124 (2016).
Zhao, P., Zhu, T., Lu, X., Zhu, J. & Li, L. Neurogenin 2 enhances the generation of patient-specific induced neuronal cells. Brain Res. 1615, 51–60 (2015).
Chanda, S., Marro, S., Wernig, M. & Südhof, T. C. Neurons generated by direct conversion of fibroblasts reproduce synaptic phenotype caused by autism-associated neuroligin-3 mutation. Proc. Natl Acad. Sci. USA 110, 16622–16627 (2013).
Yoshimizu, T. et al. Functional implications of a psychiatric risk variant within CACNA1C in induced human neurons. Mol. Psychiatry 20, 162–169 (2015).
Yi, F. et al. Autism-associated SHANK3 haploinsufficiency causes Ih channelopathy in human neurons. Science 352, aaf2669 (2016).
Pak, C. et al. Human neuropsychiatric disease modeling using conditional deletion reveals synaptic transmission defects caused by heterozygous mutations in NRXN1. Cell Stem Cell 17, 316–328 (2015).
Lee, I. S., et al. Characterization of molecular and cellular phenotypes associated with a heterozygous CNTNAP2 deletion using patient-derived hiPSC neural cells. NPJ Schizophr. 1, (2015) https://doi.org/10.1038/npjschz.2015.19.
Ebrahimi-Fakhari, D. et al. Impaired mitochondrial dynamics and mitophagy in neuronal models of tuberous sclerosis complex. Cell Rep. 17, 1053–1070 (2016).
Patzke, C. et al. Analysis of conditional heterozygous STXBP1 mutations in human neurons. J. Clin. Invest. 125, 3560–3571 (2015).
Aneichyk, T. et al. Dissecting the causal mechanism of X-linked dystonia-parkinsonism by integrating genome and transcriptome assembly. Cell 172, 897–909.e21 (2018).
Muratore, C. R. et al. Cell-type dependent Alzheimer’s disease phenotypes: probing the biology of selective neuronal vulnerability. Stem Cell Rep. 9, 1868–1884 (2017).
Muratore, C. R., Srikanth, P., Callahan, D. G. & Young-Pearse, T. L. Comparison and optimization of hiPSC forebrain cortical differentiation protocols. PLoS ONE 9, e105807 (2014).
Johnson, W. E., Li, C. & Rabinovic, A. Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics 8, 118–127 (2007).
Nehme, R. et al. Combining NGN2 programming with developmental patterning generates human excitatory neurons with NMDAR-mediated synaptic transmission. Cell Rep., 23, 2509–2523 (2018).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods 25, 402–408 (2001).
Seyfried, N. T. et al. A multi-network approach identifies protein-specific co-expression in asymptomatic and symptomatic Alzheimer’s disease. Cell Syst. 4, 60–72.e4 (2017).
Maeder, M. L. et al. CRISPR RNA-guided activation of endogenous human genes. Nat. Methods 10, 977–979 (2013).
Perez-Pinera, P. et al. RNA-guided gene activation by CRISPR-Cas9-based transcription factors. Nat. Methods 10, 973–976 (2013).
Chavez, A. et al. Highly efficient Cas9-mediated transcriptional programming. Nat. Methods 12, 326–328 (2015).
Miyoshi, G. & Fishell, G. Dynamic FoxG1 expression coordinates the integration of multipolar pyramidal neuron precursors into the cortical plate. Neuron 74, 1045–1058 (2012).
Sasaki, S., Tabata, H., Tachikawa, K. & Nakajima, K. The cortical subventricular zone-specific molecule Svet1 is part of the nuclear RNA coded by the putative Netrin receptor gene Unc5d and is expressed in multipolar migrating cells. Mol. Cell Neurosci. 38, 474–483 (2008).
Cooper, J. A. Molecules and mechanisms that regulate multipolar migration in the intermediate zone. Front. Cell Neurosci. 8, 386 (2014).
Molyneaux, B. J., Arlotta, P., Menezes, J. R. L. & Macklis, J. D. Neuronal subtype specification in the cerebral cortex. Nat. Rev. Neurosci. 8, 427–437 (2007).
Zhong, Y. et al. Identification of the genes that are expressed in the upper layers of the neocortex. Cereb. Cortex 14, 1144–1152 (2004).
Lai Wing Sun, K., Correia, J. P. & Kennedy, T. E. Netrins: versatile extracellular cues with diverse functions. Development 138, 2153–2169 (2011).
Camargo, L. M. et al. Disrupted in Schizophrenia 1 Interactome: evidence for the close connectivity of risk genes and a potential synaptic basis for schizophrenia. Mol. Psychiatry 12, 74–86 (2007).
Ozeki, Y. et al. Disrupted-in-Schizophrenia-1 (DISC-1): mutant truncation prevents binding to NudE-like (NUDEL) and inhibits neurite outgrowth. Proc. Natl Acad. Sci. USA 100, 289–294 (2003).
Lepagnol-Bestel, A. M., Kvajo, M., Karayiorgou, M., Simonneau, M. & Gogos, J. A. A Disc1 mutation differentially affects neurites and spines in hippocampal and cortical neurons. Mol. Cell Neurosci. 54, 84–92 (2013).
Steinecke, A., Gampe, C., Valkova, C., Kaether, C. & Bolz, J. Disrupted-in-Schizophrenia 1 (DISC1) is necessary for the correct migration of cortical interneurons. J. Neurosci. 32, 738–745 (2012).
Kvajo, M. et al. Altered axonal targeting and short-term plasticity in the hippocampus of Disc1 mutant mice. Proc. Natl Acad. Sci. USA 108, E1349–E1358 (2011).
Miyoshi, K. et al. Disrupted-In-Schizophrenia 1, a candidate gene for schizophrenia, participates in neurite outgrowth. Mol. Psychiatry 8, 685–694 (2003).
Kang, E. et al. Interaction between FEZ1 and DISC1 in regulation of neuronal development and risk for schizophrenia. Neuron 72, 559–571 (2011).
Kobayashi, H. et al. Calm1 signaling pathway is essential for the migration of mouse precerebellar neurons. Development 142, 375–384 (2015).
Riessland, M. et al. Neurocalcin delta suppression protects against spinal muscular atrophy in humans and across species by restoring impaired endocytosis. Am. J. Hum. Genet. 100, 297–315 (2017).
Weledji, E. P. & Assob, J. C. The ubiquitous neural cell adhesion molecule (N-CAM). Ann. Med Surg. 3, 77–81 (2014).
Myers, C. T. et al. De novo mutations in PPP3CA cause severe neurodevelopmental disease with seizures. Am. J. Hum. Genet. 101, 516–524 (2017).
Mizuguchi, T. et al. Loss-of-function and gain-of-function mutations in PPP3CA cause two distinct disorders.Hum. Mol. Genet. https://doi.org/10.1093/hmg/ddy052 (2018).
Südhof, T. C. Neurotransmitter release: the last millisecond in the life of a synaptic vesicle. Neuron 80, 675–690 (2013).
Hong, K. et al. A ligand-gated association between cytoplasmic domains of UNC5 and DCC family receptors converts netrin-induced growth cone attraction to repulsion. Cell 97, 927–941 (1999).
Tarabykin, V., Stoykova, A., Usman, N. & Gruss, P. Cortical upper layer neurons derive from the subventricular zone as indicated by Svet1 gene expression. Development 128, 1983–1993 (2001).
Takemoto, M. et al. Laminar and areal expression of unc5d and its role in cortical cell survival. Cereb. Cortex 21, 1925–1934 (2011).
Tabata, H. & Nakajima, K. Multipolar migration: the third mode of radial neuronal migration in the developing cerebral cortex. J. Neurosci. 23, 9996–10001 (2003).
Seiradake, E. et al. FLRT structure: balancing repulsion and cell adhesion in cortical and vascular development. Neuron 84, 370–385 (2014).
Yamagishi, S. et al. FLRT2 and FLRT3 act as repulsive guidance cues for Unc5-positive neurons. EMBO J. 30, 2920–2933 (2011).
Sarnowski, C. et al. Whole genome sequence analyses of brain imaging measures in the Framingham Study. Neurology 90, e188–e196 (2018).
Brauns, S. et al. DISC1 is associated with cortical thickness and neural efficiency. Neuroimage 57, 1591–1600 (2011).
Carless, M. A. et al. Impact of DISC1 variation on neuroanatomical and neurocognitive phenotypes. Mol. Psychiatry 16, 1096–104– 1063 (2011).
Maussion, G. et al. Implication of LRRC4C and DPP6 in neurodevelopmental disorders. Am. J. Med. Genet. A 173, 395–406 (2017).
Lin, J. C., Ho, W.-H., Gurney, A. & Rosenthal, A. The netrin-G1 ligand NGL-1 promotes the outgrowth of thalamocortical axons. Nat. Neurosci. 6, 1270–1276 (2003).
Kim, S. et al. NGL family PSD-95-interacting adhesion molecules regulate excitatory synapse formation. Nat. Neurosci. 9, 1294–1301 (2006).
Ricciardi, S. et al. CDKL5 ensures excitatory synapse stability by reinforcing NGL-1-PSD95 interaction in the postsynaptic compartment and is impaired in patient iPSC-derived neurons. Nat. Cell Biol. 14, 911–923 (2012).
van Kuilenburg, A. B. P. et al. Analysis of severely affected patients with dihydropyrimidine dehydrogenase deficiency reveals large intragenic rearrangements of DPYD and a de novo interstitial deletion del(1)(p13.3p21.3). Hum. Genet. 125, 581–590 (2009).
Borg, I. et al. Disruption of Netrin G1 by a balanced chromosome translocation in a girl with Rett syndrome. Eur. J. Hum. Genet. 13, 921–927 (2005).
Ohtsuki, T. et al. Association of polymorphisms in the haplotype block spanning the alternatively spliced exons of the NTNG1 gene at 1p13.3 with schizophrenia in Japanese populations. Neurosci. Lett. 435, 194–197 (2008).
O’Roak, B. J. et al. Sporadic autism exomes reveal a highly interconnected protein network of de novo mutations. Nature 485, 246–250 (2012).
Chen, T., Gai, W.-P. & Abbott, C. A. Dipeptidyl peptidase 10 (DPP10(789)): a voltage gated potassium channel associated protein is abnormally expressed in Alzheimer’s and other neurodegenerative diseases. Biomed. Res. Int. 2014, 209398 (2014).
Zagha, E. et al. DPP10 modulates Kv4-mediated A-type potassium channels. J. Biol. Chem. 280, 18853–18861 (2005).
Lin, M. et al. Heat shock alters the expression of schizophrenia and autism candidate genes in an induced pluripotent stem cell model of the human telencephalon. PLoS ONE 9, e94968 (2014).
Hamada, S. & Yagi, T. The cadherin-related neuronal receptor family: a novel diversified cadherin family at the synapse. Neurosci. Res. 41, 207–215 (2001).
Pedrosa, E. et al. Analysis of protocadherin alpha gene enhancer polymorphism in bipolar disorder and schizophrenia. Schizophr. Res. 102, 210–219 (2008).
Wu, Q. et al. DISC1 regulates the proliferation and migration of mouse neural stem/progenitor cells through Pax5, Sox2, Dll1 and Neurog2. Front. Cell Neurosci. 11, 261 (2017).
Kamiya, A. et al. A schizophrenia-associated mutation of DISC1 perturbs cerebral cortex development. Nat. Cell Biol. 7, 1167–1178 (2005).
Young-Pearse, T. L., Suth, S., Luth, E. S., Sawa, A. & Selkoe, D. J. Biochemical and functional interaction of disrupted-in-schizophrenia 1 and amyloid precursor protein regulates neuronal migration during mammalian cortical development. J. Neurosci. 30, 10431–10440 (2010).
Tomita, K., Kubo, K.-I., Ishii, K. & Nakajima, K. Disrupted-in-Schizophrenia-1 (Disc1) is necessary for migration of the pyramidal neurons during mouse hippocampal development. Hum. Mol. Genet. 20, 2834–2845 (2011).
Kubo, K.-I. et al. Migration defects by DISC1 knockdown in C57BL/6, 129X1/SvJ, and ICR strains via in utero gene transfer and virus-mediated RNAi. Biochem. Biophys. Res. Commun. 400, 631–637 (2010).
Singh, K. K. et al. Common DISC1 polymorphisms disrupt Wnt/GSK3β signaling and brain development. Neuron 72, 545–558 (2011).
Ishizuka, K., et al. DISC1-dependent switch from progenitor proliferation to migration in the developing cortex. 473: 92–96 (2011).
Lee, F. H. F. et al. Disc1 point mutations in mice affect development of the cerebral cortex. J. Neurosci. 31, 3197–3206 (2011).
Zhang, Q. et al. Electrical stimulation using conductive polymer polypyrrole counters reduced neurite outgrowth of primary prefrontal cortical neurons from NRG1-KO and DISC1-LI Mice. Sci. Rep. 7, 42525 (2017).
Hattori, T. et al. DISC1 regulates cell-cell adhesion, cell-matrix adhesion and neurite outgrowth. Mol. Psychiatry 15, 778–798–809 (2010).
Kamiya, A. et al. DISC1-NDEL1/NUDEL protein interaction, an essential component for neurite outgrowth, is modulated by genetic variations of DISC1. Hum. Mol. Genet. 15, 3313–3323 (2006).
Hattori, T. et al. A novel DISC1-interacting partner DISC1-binding zinc-finger protein: implication in the modulation of DISC1-dependent neurite outgrowth. Mol. Psychiatry 12, 398–407 (2007).
Lee, S.-A. et al. Disrupted-in-schizophrenia 1 (DISC1) regulates dysbindin function by enhancing its stability. J. Biol. Chem. 290, 7087–7096 (2015).
Muratore, C. R. et al. The familial Alzheimer’s disease APPV717I mutation alters APP processing and Tau expression in iPSC-derived neurons. Hum. Mol. Genet. 23, 3523–3536 (2014).
Srikanth, P. et al. Shared effects of DISC1 disruption and elevated WNT signaling in human cerebral organoids. Transl. Psychiatry 8, 77 (2018).
Acknowledgements
We thank Guo-Li Ming and Hongjun Song for generously providing the WTex12 and MUTex12 lines used in this study. We thank Dennis Selkoe, Matt LaVoie, and all members of the Young-Pearse lab for continual advice throughout this study, and Dana Callahan for technical assistance. We acknowledge technical assistance from Duc M. Duong (Emory Integrated Proteomics Core) and support from the Emory Neuroscience NINDS Core Facilities (P30NS055077). This work was supported by 01MH101148 and R01MH101148-S1, and the Brain and Behavior Research Foundation (TYP).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Srikanth, P., Lagomarsino, V.N., Pearse, R.V. et al. Convergence of independent DISC1 mutations on impaired neurite growth via decreased UNC5D expression. Transl Psychiatry 8, 245 (2018). https://doi.org/10.1038/s41398-018-0281-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41398-018-0281-9
This article is cited by
-
Modeling disrupted synapse formation in wolfram syndrome using hESCs-derived neural cells and cerebral organoids identifies Riluzole as a therapeutic molecule
Molecular Psychiatry (2023)
-
Predictive network analysis identifies JMJD6 and other potential key drivers in Alzheimer’s disease
Communications Biology (2023)
-
Metabolic contributions to neuronal deficits caused by genomic disruption of schizophrenia risk gene SETD1A
Schizophrenia (2022)
-
APP and DYRK1A regulate axonal and synaptic vesicle protein networks and mediate Alzheimer’s pathology in trisomy 21 neurons
Molecular Psychiatry (2022)
-
The plasma peptides of Alzheimer’s disease
Clinical Proteomics (2021)