Abstract
The ecophysiology of complete ammonia-oxidizing bacteria (CMX) of the genus Nitrospira and their widespread occurrence in groundwater suggests that CMX bacteria have a competitive advantage over ammonia-oxidizing bacteria (AOB) and archaea (AOA) in these environments. However, the specific contribution of their activity to nitrification processes has remained unclear. We aimed to disentangle the contribution of CMX, AOA and AOB to nitrification and to identify the environmental drivers of their niche differentiation at different levels of ammonium and oxygen in oligotrophic carbonate rock aquifers. CMX ammonia monooxygenase sub-unit A (amoA) genes accounted on average for 16 to 75% of the total groundwater amoA genes detected. Nitrification rates were positively correlated to CMX clade A associated phylotypes and AOB affiliated with Nitrosomonas ureae. Short-term incubations amended with the nitrification inhibitors allylthiourea and chlorate suggested that AOB contributed a large fraction to overall ammonia oxidation, while metaproteomics analysis confirmed an active role of CMX in both ammonia and nitrite oxidation. Ecophysiological niche differentiation of CMX clades A and B, AOB and AOA was linked to their requirements for ammonium, oxygen tolerance, and metabolic versatility. Our results demonstrate that despite numerical predominance of CMX, the first step of nitrification in oligotrophic groundwater appears to be primarily governed by AOB. Higher growth yields at lower ammonia turnover rates and energy derived from nitrite oxidation most likely enable CMX to maintain consistently high populations.

Similar content being viewed by others
Introduction
Nitrification is a globally relevant process of the nitrogen cycle mediated by a polyphyletic group of microorganisms [1]. The discovery of complete ammonia oxidizers (comammox bacteria, CMX) challenged the paradigm of a strict division of labor in the process of nitrification [2]. While ammonia oxidation is carried out by ammonia-oxidizing bacteria (AOB) and archaea (AOA) [3] and nitrite oxidation by nitrite-oxidizing bacteria (NOB) [2], CMX of the genus Nitrospira can perform complete oxidation of ammonia (NH3) to nitrate (NO3-) in one cell [4, 5]. CMX have been frequently reported from engineered environments such as drinking water or wastewater treatment facilities [6,7,8] and from natural environments like soils [9, 10], surface freshwater [11, 12] and groundwater [13, 14]. However, our knowledge about the role and relevance of CMX for nitrification activity in natural environments is still scarce.
In most environments, CMX will compete with canonical nitrifiers for either ammonia (NH3) or nitrite (NO2-) or both, raising the question of the actual contribution of CMX to nitrification processes and of the factors that control their co-occurrence with AOB, AOA, and NOB. Recent investigations consider affinity to NH3, growth rates, growth yields, and metabolic versatility as key factors that determine the niche differentiation and competition between CMX, AOB, and AOA [2, 15]. Since CMX exhibit higher affinity to NH3 and oxygen than terrestrial AOB [15, 16] along with higher growth yields compared to AOB and AOA [2], CMX might have a competitive advantage over AOB in environments that are limited in NH3 and ammonium (NH4+) such as oligotrophic groundwater. Particular lineages of AOB and AOA exhibit adaptation to NH4+ limited conditions [17,18,19,20] and have previously been reported from groundwater environments [21,22,23]. As such, oligotrophic groundwater represents an excellent microbial habitat to study the co-occurrence of CMX, AOB and AOA and their contribution to nitrification activity.
Based on the phylogeny of the ammonia monooxygenase (Amo), the key enzyme of ammonia oxidation, CMX Nitrospira are divided into two clades, A and B [2, 4]. Metagenomic studies uncovering the phylogeny and metabolic features of CMX proposed an ecophysiological niche differentiation between the two clades in natural habitats [4, 21, 24,25,26]. Those findings are linked to genomic variability in nitrogen uptake and alternative energy metabolism [25, 26], and several studies suggest a more prominent role of clade A in nitrification. So far, CMX enrichments and isolates comprise only clade A representatives [27]. 13C stable isotope probing confirmed growth of CMX clade A with NH4+ during incubations of wetland coast sediments [28], and CMX clade A were highly abundant in salt marsh sediments showing strong potential for nitrification [29]. However, the ecophysiological niche differentiation between the clades has rarely been studied in the context of their co-occurrence and competition with canonical nitrifiers.
Previous studies from karstic carbonate rock aquifers of the Hainich Critical Zone Exploratory in central Germany pointed to strong links between in situ nitrification and chemolithoautotrophic carbon fixation [30, 31]. In addition, genome resolved metagenomics revealed an important role of CMX Nitrospira in the nitrogen and carbon cycle in this oligotrophic habitat [21]. The aquifers are characterized by a high spatial heterogeneity of NH4+ and NO3- concentrations across an oxygen gradient, linked to differences in groundwater flow regime and distance to groundwater recharge areas [28,29,30]. These settings are ideal for the investigation of the ecophysiological niche differentiation of CMX clade A and B, AOB and AOA.
We combined 15N-based nitrification rate measurements with selective nitrification inhibitors and genome-resolved metagenomics and metaproteomics to assess the contributions of CMX Nitrospira, AOB and AOA to overall groundwater nitrification in these aquifers and to follow the composition of the groundwater nitrifier populations across the hydrochemical gradients. We hypothesized that (i) CMX act as key ammonia oxidizers in these oligotrophic groundwater habitats and account for a large fraction of overall nitrification activity. We further hypothesized that (ii) distribution patterns of ammonia oxidizers across gradients of NH4+ and oxygen availability are linked to their ecophysiological niche differentiation, indicated by differences in encoded metabolic potentials.
Materials and methods
Study site characteristics and sampling of groundwater
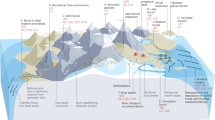
Groundwater was obtained from karstic limestone aquifers at the Hainich Critical Zone Exploratory (CZE) in Thuringia (Germany) within the framework of the Collaborative Research Center (CRC) AquaDiva [32]. Further details about location and geological setting are reported elsewhere [32, 33]. Sampling wells H14, H32, H42, H43, H52 and H53 at varying depths provide access to oxygen-deficient (less than 15 µM DO) groundwater from an upper mudstone-dominated low-flow aquifer (HTU) and wells H13, H31, H41 and H51 (mean values 154 to 672 µM DO) provide access to oxic groundwater from a lower fast-flow limestone-dominated karstified aquifer (HTL) [32, 34] (Fig. 1A, B). The main recharge zone for groundwater is located at the hilltop of the transect near wells H13 and H14. Groundwater collected from regular monthly sampling between 2018 and 2021 was obtained using submersible motor pumps (Grundfos, Denmark). Hydrochemical parameters (pH, T, TOC, DOC) were determined as described in [33, 34]. NH4+ (sum of NH3 and NH4+), NO2- and NO3- concentrations were measured with colorimetric methods [35, 36] from groundwater filtered through 0.2 µm-pore size sterile polyvinylidine fluoride filters. Concentrations of free NH3 were calculated based on pH and NH4+ concentrations (pKa NH4+/NH3 = 9.73 at 10 °C). Groundwater for molecular analysis was collected in sterile 10 L fluorinated polyethylene containers and filtered through 0.2 µm polycarbonate filters (Nuclepore, Whatman). Additional groundwater samples for metaproteomic analysis were collected from wells H14, H41, H43 and H52 in January 2019 and filtered through 0.2 µm pore size filters (Merck Millipore, Germany). All filters were immediately frozen on dry ice and stored at −80 °C until extraction.
A Schematic cross-section of the Hainich groundwater monitoring transect showing two superimposed aquifer assemblages accessible by 10 wells with various depths. B Overview of the concentrations of dissolved oxygen, ammonium, and nitrate from oxic (blue) and hypoxic/ anoxic (green) groundwater wells (n = 24 per well). Single dots represent one measurement, the crossbar indicates the mean of all values, and the error bars show the variation from the mean. Outliers are beyond the error bars. Lowercase letter code indicates significant differences based on Dunn’s multiple comparison test (p < 0.05). C Nitrification rates assessed by 15N based labeling of groundwater from five days incubation close to in situ conditions (n ≥ 3 per well; n.d. = not detected; n.a. = not available). D Gene abundances of amoA per L and E fractions of amoA genes in groundwater from different groups of ammonia-oxidizing prokaryotes in 10 groundwater wells (n ≥ 3).
15N-isotope labeling and rate measurements
Groundwater was collected in sterile 0.5 L glass bottles and filled from the bottom using sterile tubing. Bottles were overfilled with three volumes exchanges before they were capped with silicone septa without headspace to minimize alterations of in situ oxygen concentrations. At each sampling campaign, groundwater was collected in triplicate bottles for each well along with one control and kept at 4 °C until further processing within 2 h. Prior to the 15N labelling, 10 ml of each sampling bottle was removed for analysis of inorganic nitrogen concentrations and pH, and replaced with N2. Bottles containing anoxic groundwater were amended with 1% sterile oxygen (4 ml 100% oxygen) to stimulate nitrification under oxygen-deficient conditions. Groundwater of control bottles was sterile filtered through 0.2 µm filters (Supor, Pall Corporation, USA). Sterile filtered 15N-labeled ammonium sulfate solution ((15NH4)2SO4) (98 at%, Cambridge Isotope Laboratories, Tewksbury) as the substrate source for nitrifiers was added to all 0.5 L bottles to a final concentration of 50 µM, at a natural background of 0.5 to 55 µM NH4+. Nitrification rate measurements were conducted according to Overholt et al. [21] and detailed descriptions are provided in Supplementary Methods.
Incubation experiments in the presence of nitrification inhibitors
The ammonia and nitrite oxidation activity of CMX, and the nitrite oxidation activity of NOB were selectively inhibited by 1 mM potassium chlorate (KClO3) [7], and 100 µM allylthiourea (ATU) was used to inhibit the ammonia oxidation activity of CMX and AOB [7, 37]. Nitrification rate measurements with 15N tracer were carried out over five days as described above. Groundwater from well H41 was collected in February 2020 and amended with 50 µM NH4+ as (15NH4)2SO4 and one of the nitrification inhibitors along with untreated controls.
Groundwater for mesocosm experiments was collected in March and May 2021 and was filled in sterile 2 L glass bottles without headspace to assess potential growth of ammonia oxidizers with NH4+ in the presence of nitrification inhibitors at the same concentrations as before. Groundwater from oxic well H41 (10 µM NH4+ in situ) was adjusted to 10 and 50 µM NH4+, and groundwater from hypoxic well H53 (36 µM NH4+ in situ) was used without additional supplement. All samples were incubated for 15 to 25 days at 15 °C without agitation in the dark. Subsamples for nitrogen compound analysis were taken at the onset and every two to three days until the end of the incubation to monitor changes of the nitrogen chemistry (Supplementary Methods). Incubation of well H41 with an initially 10 µM NH4+ was adjusted again to 15 µM NH4+ after 15 days. After incubation, the groundwater was filtered on 0.2 µm polycarbonate filters and stored at −80 °C for later extraction of nucleic acids.
Nucleic acid extraction, cDNA-synthesis and quantitative PCR
Genomic DNA and total RNA of microbial biomass from the 0.2 µm filters were extracted using the DNeasy PowerSoil kit (Qiagen, Germany) and the RNeasy PowerWater kit (Qiagen, Germany), respectively, according to the manufacturer’s specifications. RNA was further processed with the TURBO DNA-free kit (Thermo Fischer Scientific, Germany) to remove remnant DNA in the samples before cDNA synthesis [38]. Reverse transcription was performed using the GoScript Reverse Transcription System (Promega, USA). Gene and transcript abundances of amoA from CMX Nitrospira, AOB, AOA and of nxrB from Nitrospira-like NOB were determined using a CFX96 qPCR cycler (Bio-Rad, USA). Primer pairs for quantification were comamoAF/ comamoAR for CMX amoA [11], AmoA-1F/ AmoA-2R for AOB amoA [39], Arch-amoAF/ Arch-amoAR for AOA amoA [40] and nxrB169f/ nxrB638r for Nitrospira-like nxrB [41]. More details are provided in the Supplementary Methods. For testing of potential correlations between CMX amoA and Nitrospira nxrB genes, we assumed on average one copy of amoA and 1.5 copies of nxrB per cell. Calculation of the correction factor 1.5 was based on the analysis of putative canonical Nitrospira-affiliated MAGs from the Hainich groundwater metagenome, which harbored 1 to 2 nxrB gene copies per genome [21].
Amplicon sequencing of amoA and 16 S rRNA genes and sequence analysis
Amplicon libraries of CMX, AOB and AOA amoA genes were created using the NEBNext Ultra DNA Library Prep Kit for Illumina (New England Biolabs, USA) according to the manufacturer’s guideline. The same primers as for qPCR were used for CMX and AOB amoA, while primers Arch-amoA-104F/ Arch-amoA-616Rmod [42, 43] were chosen for sequencing of AOA amoA genes due to shorter fragment length (Supplementary Methods). Sequencing was conducted on an Illumina MiSeq using v3 chemistry. After trimming primer sequences, raw reads were first quality filtered with DADA2 1.22.0 [44] and further processed with Mothur v.1.46.1 [45] (Supplemental Methods). OTUs were assigned at 95% sequence identity for CMX, 95% for AOB [46] and 96% for AOA [42] (see Dataset S3).
Amplicon libraries of the bacterial 16 S rRNA gene were constructed as described elsewhere [47] to assess the relative abundance of nitrite oxidizers present in the groundwater, including non-Nitrospira NOB. Raw sequence reads were processed using DADA2 1.22.0. Taxonomy was assessed using the SILVA reference data base release 138.1 [48]. Sequence analysis was performed using the R package phyloseq [49].
Inferring metabolic potential from metagenome assembled genomes of CMX Nitrospira and other nitrifiers
To assess metabolic capabilities of the two CMX clades, genomic features of metagenome assembled genomes (MAGs) were investigated. Groundwater sampling for metagenomic sequencing and generation of MAGs were described in detail in [21, 50]. Here, Hainich groundwater MAGs, which were taxonomically classified as Nitrospira on genus level and 243 Nitrospira reference genomes downloaded from NCBI (July 2021) were aligned and screened for both amoA and nxrB genes with HMMER in Anvi’o v7.1 [21] using custom AmoA and NxrB databases containing sequences from the FunGene repository [51].
After sorting good quality MAGs (>90 % completeness, except N. inopinata with 42.25 %) and dereplication, 35 CMX Nitrospira genomes were kept for final analysis. The affiliation of those genomes to CMX clade A and clade B was achieved by alignment of protein sequences of bacterial single copy core genes detected with default ‘Bacteria_71’ hmm profile using MUSCLE [52] in Anvi’o and subsequent construction of a maximum likelihood phylogenetic tree using FastTree with 1000 bootstrap iterations. The tree was then processed with iTOL for visualization [53]. The genomic repertoire of representative groundwater nitrifier MAGs was inferred by KEGG orthology (KO) previously generated using kofamscan [21] (Supplementary Methods). Relative transcriptional activity of each investigated MAG was extracted from Overholt et al. [21], who determined the transcriptomic coverages for each open reading frame from each MAG from previously generated metatranscriptomic libraries of the groundwater [30].
Metaproteomic analysis
Proteins were extracted from filters using a phenol-chloroform based protocol as previously described [54]. Proteins were subjected to SDS polyacrylamide gel electrophoresis, subsequently in-gel tryptic cleavage and LC-MS/MS analysis was performed as previously described [55] (Supplemental Methods). Translated amino acid sequences from all genes of the MAGs obtained from the metagenomic datasets [21, 50] were used as a reference database for protein identification. Taxonomic and functional annotations of identified peptides were transferred from the metagenomic dataset [21, 50]. Mass spectrometry proteomics data have been deposited into the ProteomeXchange Consortium via the PRIDE [56] partner repository.
Statistical analysis
All statistical analyses were conducted using R 4.12 (R Core Team, 2020) [57], including packages FSA [58], ComplexHeatmap [59] and vegan [60]. Analysis details are explained in the Supplementary Methods. To analyze communities of CMX, AOB and AOA together, the absolute amoA read abundances of OTUs >2% of all reads for each ammonia oxidizer community were determined as followed: The percentage of each OTU was calculated from the relative abundance obtained by amoA amplicon sequencing and was multiplied by the total amoA counts from corresponding samples measured with qPCR.
Results
Nitrification activity and distribution of nitrifiers across the different groundwater wells
Nitrification rates in groundwater varied over two orders of magnitude ranging from 0.54 to 136 nmol N L−1 d−1 across the wells (Fig. 1C). We observed the highest rates at oxic well H41 with 93 ± 24 nmol N L−1 d−1 (mean ± standard deviation (s.d.)) and high activity at hypoxic well H52 with 58 ± 4.0 nmol N L−1 d−1. Rates were positively correlated with NH4+ concentrations in the groundwater (rs = 0.67, p < 0.05), which were on average below 40 µM and significantly higher in the hypoxic wells (Fig. 1B). NO3- concentrations ranged from 1.7 to 38 µM in hypoxic and from 80 to 384 µM in oxic groundwater and showed no correlation with the rates across the transect.
CMX amoA genes accounted for 16 ± 7.2 to 75 ± 12.3% (mean ± s.d.) of the total groundwater amoA genes detected (mean values per well, see Dataset S2). Proportions of CMX amoA and AOB amoA genes increased substantially to maxima of 95% and 34% at wells with higher NH4+ concentrations (H41, H51, H52, H53), and gene abundances were positively correlated to NH4+ (rs = 0.48, p < 0.01; rs = 0.44, p < 0.01), respectively (Figs. 1E, 2D). In contrast, AOA dominated the ammonia-oxidizing community by up to 90% in wells with lower NH4+ concentrations located closer to the main recharge area (H13 and H14). Oxic well H41 and anoxic well H53 harbored the highest absolute amoA gene and transcript abundances of CMX and AOB (Fig. 1D, Fig. S1A). Nitrification rates were strongly correlated with AOB amoA gene abundances (rs = 0.87, p < 0.001) (Fig. 2D), and AOB also had the highest amoA transcript/ gene ratios across all wells (Fig. S1B).
A Redundancy analysis (RDA) demonstrates association between distribution of ammonia-oxidizing community (relative amoA read abundances of CMX, AOB and AOA OTUs which account for <2% sequence reads of each community were normalized to amoA gene abundances to generate absolute amoA read abundances) and significant hydrochemical parameters fitting (p < 0.05) with nitrifier community distribution across eight groundwater wells (n ≥ 3). Dotted ellipses show the confidence of clusters assuming a multivariant normal distribution. B Community composition of CMX Nitrospira showing dominant OTUs (< 2% of sequence reads) affiliated to clade A and clade B across eight groundwater wells (n ≥ 3 per well). Each bar displays the mean absolute amoA read abundance of OTUs. C Maximum likelihood phylogenetic tree of deduced AmoA amino acid sequences of CMX Nitrospira depicting evolutionary relationship between reference genomes, and sequences of OTUs and MAGs from Hainich groundwater. The tree was constructed using the JTT substitution model with gamma distribution and 1000 bootstrap iterations. Bootstrap support values greater than 75% are denoted with black dots. An AmoA sequence from Nitrosomonas oligotropha served as outgroup indicated by the arrow. D Pairwise correlations between total amoA gene abundances of CMX, AOB and AOA, absolute amoA read abundances of OTUs and groundwater hydrochemical parameters (TIC = total inorganic carbon, TOC = total organic carbon, DOC = dissolved organic carbon, ORP = redox potential, EC = electrical conductivity) using Spearman’s rank correlation. Hierarchical clustering is based on Euclidean distance. Color code of correlation coefficient rs displays strength of association using following significance levels: *p < 0.05, **p < 0.01, ***p < 0.001.
Across all wells, abundances of CMX amoA genes and Nitrospira nxrB genes encoding nitrite oxidoreductase were positively correlated to each other (rs = 0.72, p < 0.001) (Fig. S1D). Assuming one amoA and 1.5 nxrB copies per genome (see Dataset S4), CMX accounted for 18 to 44% and 48 to 70% of the Nitrospira population at oxic well H41 and hypoxic well H52, respectively (Fig. S1C), which exhibited the highest observed nitrification rates (Fig. 1C). Furthermore, CMX and Nitrosomonadaceae had the highest relative abundance of peptides at wells H41 and H52 (Fig. S2). Out of the Nitrosomonadaceae affiliated peptides, 88.3% were assigned to Nitrosomonas.
The ammonia oxidizer community composition showed formation of clusters depending on local groundwater chemistry and the location in the transect (Fig. 2A). The CMX community was dominated by three OTUs affiliated to clade A and six OTUs affiliated to clade B, of which clade A OTUs were closely related to Ca. Nitrospira nitrificans (Fig. 2C). These clade A OTUs increased in abundance at wells H41, H51, H52 and H53 and accounted for up to 50% of the CMX community (Fig. 2B). CMX clade A affiliated OTUs were also positively correlated to the nitrification rates (rs = 0.72–0.88, p < 0.05), while only two low abundant clade B affiliated OTUs correlated with the rates (Fig. 2D). The AOB community was mainly dominated by AOB OTU1 affiliated to Nitrosomonas ureae, which was the only AOB phylotype showing a strong correlation with nitrification rates (rs = 0.88, p < 0.01). The most dominant AOA OTU1 related to Nitrosoarchaeum koreense accounted for an average of 70% of all AOA across the groundwater sites (Fig. S3). Clustering based on the correlation with hydrochemical parameters suggested similar ecological preferences of (i) AOA and some clade B affiliated CMX OTUs, and (ii) Nitrosomonas ureae affiliated AOB OTU1 and CMX clade A affiliated OTUs (Fig. 2D). Among potential nitrite oxidizers, Nitrospira sp. were consistently present in all the samples, while 16 S rRNA gene sequences affiliated with Nitrospinaceae, Ca. Nitrotoga, and Nitrobacter occurred only occasionally, and relative abundances of these groups were at least one order of magnitude lower than those of Nitrospira (Fig. S4).
Metabolic capabilities of CMX Nitrospira clades and AOB
We analyzed nine CMX affiliated MAGs from the Hainich groundwater [21], which were phylogenomically affiliated to two CMX clade A MAGs and seven clade B MAGs (Fig. 3A). Both clade A MAGs (H51-bin291-1 and H41-bin216-1) were closely related to Ca. Nitrospira nitrificans on the genome level and showed 99% amino acid sequence identity to the AmoA of CMX OTU5 (Fig. 2C). AmoA sequences of Clade B representative MAGs showed closest relation to AmoA of CMX OTU2, OTU3, OTU8 and OTU9.
A Maximum likelihood tree of bacterial core protein sequences of CMX Nitrospira from nine MAGs originating from Hainich groundwater depicting the evolutionary relationship to reference genomes. The tree was constructed using the JTT substitution model with gamma distribution and 1000 bootstrap iterations. Bootstrap support values greater than 75% are shown as black dots. AmoA sequences from Nitrosomonas oligotropha and Nitrosomonas europaea served as an outgroup indicated by the arrow. B Metabolic capacities of nine groundwater CMX Nitrospira MAGs inferring genomic potential for nitrogen and sulfur energy metabolism, and corresponding transporter types as well as protective systems against radical oxygen species, and genes involved in carbon fixation (rTCA = reductive tricarboxylic acid cycle). Sizes of circles display the transcript coverage of each gene mapped from groundwater metatranscriptome. If more than one gene was present, maximum transcriptional coverage is shown. Filled circles indicate peptides from metaproteomic data matching for displayed genes. rTCA = reverse tricarboxylic acid cycle.
Genes encoding essential components for ammonia oxidation, nitrite oxidation and urea degradation were shared in MAGs representing both CMX clades (Fig. 3B) and most of these genes were highly transcribed in the groundwater. Metaproteomic analysis revealed the presence of peptides related to AmoA, AmoB, AmoC, Hao and NxrA of Nitrospira, demonstrating that nitrification-related proteins of CMX Nitrospira populations were formed in situ. We found different genomic repertoires between CMX clade A and clade B affiliated Nitrospira MAGs containing genes for sulfur and alternative energy metabolism such as oxidation of formate and hydrogen, nitrogen transporter types and cellular regulation of reactive oxygen species. CMX clade B MAGs exclusively harbored genes involved in conversion of bicarbonate to carbon dioxide, oxidation of thiosulfate via the sox pathway, dissimilatory sulfate reduction, as well as genes encoding two superoxide dismutases (SOD1/ SOD2) and a formate transporter (focA). Transporters for cellular import of urea as well as for NO2-/NO3- were present in representative genomes from both CMX clade A and clade B. We found that clade B MAGs encoded Amt-type NH4+ transporters, while clade A MAGs employed Rh-type NH4+ transporters. MAGs from both clades harbored genes for ATP citrate lyase (acl), which were transcriptionally active, indicating an active role in carbon fixation.
Compared to the CMX and AOA affiliated MAGs, transcription of amo genes was more than 100 times higher for AOB affiliated MAG H51-bin202-1 (Fig. S4). The AmoA sequence of this MAG had 93% sequence identity to the AmoA of the dominant, N. ureae affiliated AOB-OTU1 obtained from amplicon sequencing in this study (Fig. S5), suggesting that both approaches probably identified the same population of AOB. This MAG employed Rh-type NH4+ transporters as did representatives of CMX clade A, while AOA affiliated MAGs had Amt-type NH4+ transporters as representatives of CMX clade B (Fig. S4).
Contributions of ammonia-oxidizing groups to ammonia oxidation activity
Short-term incubations in the presence of the nitrite oxidation inhibitor chlorate did not affect the ammonia oxidation rates (74.5 ± 9.4 nmol N L−1 d−1) compared to the rates in controls without inhibitor (75.9 ± 3.7 nmol N L−1 d−1) (Fig. 4A). Chlorate is supposed to inhibit nitrite oxidation of NOB, and both ammonia oxidation and nitrite oxidation by CMX, as nitrite oxidoreductase catalyzes the formation of toxic chlorite from chlorate [7]. In the presence of the ammonia oxidation inhibitor allylthiourea, ammonia oxidation rates were not significantly different from zero. Allylthiourea is supposed to block ammonia oxidation, as it inhibits the ammonia monooxygenases of AOB as well as of CMX but affects those of AOA much less at the 100 µM concentrations used in this study [37, 61].
A Nitrification rates from short-term incubation of oxic groundwater supplemented with 50 µM NH4+ and selective inhibition with chlorate and allylthiourea (n.d. = not detected). Dots depict single rates of each replicate. Crossbar shows the average of three replicates and error bars show the variation between rates. B Fold-change of amoA genes from ammonia oxidizers and nxrB genes from Nitrospira-like nitrite oxidizers after long-term incubation of groundwater. Single points represent one replicate and crossbar shows the average. Error bars display variance between conditions (n = 3). Condition 1: oxic groundwater supplemented with 10 µM NH4+. Condition 2: oxic groundwater supplemented with 50 µM NH4+. Condition 3: hypoxic groundwater without additional NH4+. C Composition of CMX Nitrospira clade A and clade B affiliated OTUs of groundwater and after long-term incubation (n = 3). Bars show the mean relative amoA read abundance of OTUs representing >2%. D Composition of bacterial ammonia-oxidizing community and taxonomic affiliation of OTUs.
Long-term incubations (15–28 days) were performed to further investigate the potential growth and shifts in community composition of CMX, AOB, AOA and Nitrospira-affiliated NOB under the inhibitor treatments, using oxic groundwater from well H41 (condition 1: oxic/ 10 µM NH4+; condition 2: oxic/ 50 µM NH4+) and hypoxic groundwater from H53 (condition 3: hypoxic/ without supplement). Ammonia oxidation was always completely inhibited in the presence of allylthiourea, regardless of groundwater source or initial NH4+ concentrations (Fig. S6). NH4+ was only oxidized in incubations without inhibitor and in the chlorate treatment, while nitrite oxidation was only measured in incubations without inhibitor. Condition 3 without inhibitor showed complete consumption of NH4+ after 13 days, while it took 19 and 25 days under condition 1 and 2, respectively.
In incubations without inhibitor, CMX increased about two-fold (6.5 × 105 and 2.5 × 106 amoA L−1 after 25 and 20 days of incubation) compared to initial groundwater (2.8 × 105 and 1.44 × 106 amoA L−1) under condition 1 and 2, respectively, while we did not detect growth of CMX under condition 3 (Fig. 4B). CMX amoA gene abundance also increased in the allylthiourea treatment under condition 1 but decreased in the chlorate treatments. Nitrospira-affiliated NOB targeted using nxrB genes had similar growth patterns across condition and treatment types as CMX, and CMX accounted for an estimated 14 to 63% of the Nitrospira-affiliated NOB population. AOB amoA abundance increased by one to two orders of magnitude in all untreated and chlorate treatments of the different conditions, while they decreased in the presence of allylthiourea. AOA amoA gene abundance showed only minor variations during all incubation settings. CMX OTU1 affiliated to CMX clade A (Fig. 4C) and AOB-OTU1 related to Nitrosomonas ureae (Fig. 4D) increased in abundance after incubations without inhibitor. These two OTUs were already dominant in the ammonia oxidizer communities of the original groundwater.
Discussion
We present evidence that despite numerical predominance of CMX bacteria, they contribute only a small fraction to ammonia oxidation in the studied oligotrophic groundwater. Instead, ammonia oxidation in this system appeared to be largely driven by AOB. Ranging from 0.54 up to 136 nmol N L−1 d−1, the measured nitrification rates (ammonia oxidation + nitrite oxidation) matched those from oligotrophic lakes and open marine environments including oxygen minimum zones [62,63,64,65], but were lower than in rivers, and estuaries [66, 67] (Table S1). The observed nitrification rates did not reflect the spatial heterogeneity of groundwater NO3- concentrations, suggesting that surface-derived inputs mask the effects of in situ NO3 formation on groundwater NO3- loads in these carbonate-rock aquifers.
The oligotrophic conditions in the groundwater appeared to select for ammonia oxidizers with the highest reported NH3 affinities of their respective group. As the three groups differ in their NH3 affinity, with 0.049 to 0.083 µM for Ca. Nitrospira inopinata [15] (corresponding to 11 to 19 µM NH4+ at pH 7.3), 0.3 to 4 µM for the Nitrosomonas oligotropha lineage [15, 20] (corresponding to 55 to 829 µM NH4+ at pH 7.3), and <0.01 µM for AOA closely related to Nitrosoarchaeum koreensis [15] (corresponding to <2.3 µM NH4+ at pH 7.3), fluctuating resource availability at overall limiting conditions may have enabled their coexistence. Previous metagenomics analysis work at this study site identified CMX as key nitrifiers and demonstrated that measured groundwater carbon fixation rates could fully be accounted for by carbon fixation associated to nitrification, assuming nitrification and carbon fixation kinetics of CMX Nitrospira [21]. Similarly, in our study, CMX accounted for up to 80% of the ammonia oxidizer community at the groundwater well showing the highest nitrification activity, supporting our original assumption that CMX were the most competitive ammonia-oxidizing group. Our inhibitor-based approach strongly suggested that ammonia oxidation appeared to be largely driven by AOB, even though CMX outnumbered AOB by a factor of 10 in the samples used for rate measurements.
Treatment with 1 mM chlorate in nitrification assays was expected to inhibit the nitrite oxidation activity of canonical nitrite oxidizers as well as both steps of nitrification in CMX, since reduction of chlorate to toxic chlorite from the activity of nitrite oxidoreductase would eventually also inhibit ammonia oxidation in the same cell [7]. Ammonia oxidation was not affected in the chlorate treatment, but was completely inhibited in the treatment with allylthiourea, which specifically targets the ammonia monooxygenase and acts as a reversible inhibitor preventing the oxidation of NH4+ [61]. At a concentration of below 160 µM, it selectively inhibits the activity of AOB and CMX but not of AOA [37]. Consequently, this inhibitor approach strongly suggested that AOB were primarily responsible for the highest observed ammonia oxidation activity at well H41. A strong bias of our results by potential residual activity of CMX in the presence of chlorate appears unlikely. Although Nitrospira is capable of detoxifying hazardous chlorite by chlorite dismutases [68], the chlorate concentrations used here obviously resulted in growth inhibition and cell death, which is in line with previous observations in wetland sediment mesocosms [28]. Similarly, it appears unlikely that our results were caused by excessive growth of AOB, since nitrification rates were linear across these five-days incubations. A negligible contribution of AOA to ammonia oxidation was indicated by no detectable activity in the presence of allylthiourea.
We cannot completely rule out that activity by AOB was enhanced in the chlorate treatment due to absence of competition with CMX. This is one of the pitfalls associated with inhibition experiments and could have led to an overestimation of the contribution of AOB in our case. In fact, increased activity and growth of AOB in soil mesocosms where AOA were inhibited was recently demonstrated by Zhao et al. [69]. Our transcriptional and metaproteomics-based findings indicate a very active role of AOB in ammonia oxidation, supporting a substantial contribution of this group to ammonia oxidation rates. Ammonium concentrations at well H41, which had the highest nitrification activity, were around 10 µM NH4+ in situ and 50 µM were used for rate measurements, corresponding to 0.04 or 0.22 µM free NH3, respectively. The NH3 affinities and per cell ammonia oxidation rates differ between AOB and CMX [15]. It remains unclear to what extend competition for NH4+ would restrict their ammonia oxidation rates under the given ammonia-limited conditions, which presumably affect AOB more than CMX, and if this would result in increased activity of AOB when CMX were suppressed due to inhibition. According to Vilardi et al. [70], CMX and AOB can fill independent niches in the nitrifying community, which would reduce potential ammonia oxidation competition between both groups. More detailed knowledge of the exact ammonia oxidation kinetics of the groundwater AOB and CMX representatives is needed to further disentangle the mechanisms of their coexistence.
Despite the presumably small contribution of CMX to ammonia oxidation rates, we obtained strong evidence of their active role in overall nitrification. CMX expressed genes and formed peptides involved in both ammonia and nitrite oxidation in situ, which was confirmed by growth of CMX in the presence of NH4+ in long-term incubations. However, it remains unclear if the major energy source to sustain cell metabolism originated from complete ammonia oxidation or rather from nitrite oxidation. In fact, a recent subsurface study detected transcriptional activity in association with nitrite oxidation but not ammonia oxidation for a CMX-related MAG [71], while CMX Nitrospira in water from rapid sand filters did not prefer to oxidize external nitrite alone [7]. As we found NO2- transporters in representative groundwater MAGs from both CMX clades, the cells may also be capable of taking up the NO2- formed by AOB, supporting their primary role as nitrite oxidizers along with Nitrosomonas ureae and Nitrosospira sp. affiliated AOB as key ammonia oxidizers.
AOB with faster ammonia oxidation rates than CMX [3, 15] did not outcompete CMX for NH4+ but simply processed a larger fraction of the available NH4+, thereby driving the ammonia oxidation rates and providing the resulting NO2- to both CMX and canonical NOB. Overall, the genus Nitrospira clearly dominated the NOB community, as NOB affiliated with Nitrospinaceae, Ca. Nitrotoga, and Nitrobacter occurred only occasionally and were one or two orders of magnitude less abundant than Nitrospira. The question remains unclear, why CMX which accounted for more than 50% of the Nitrospira population were apparently favored as nitrite oxidizers over canonical nitrite-oxidizing Nitrospira in the groundwater. Physiological studies of Ca. Nitrospira inopinata demonstrated a much lower affinity to external NO2- compared to canonical Nitrospira [15], a trait which is, however, not shared among all CMX Nitrospira [72]. A recent microcosm-based study of drinking water biofilter media suggested that CMX outcompete canonical NOB in natural nitrifier communities from nitrogen-limited environments over time [70]. However, co-occurrence of CMX and canonical Nitrospira appears to be common across a broad range of terrestrial and aquatic habitats [27], and their respective roles in nitrification might be highly variable.
In contrast to our findings, CMX Nitrospira were identified as the driver of the two steps of nitrification in groundwater-fed rapid gravity sand filters (RGSF) [7, 73]. Moreover, they made a similar contribution as AOB to ammonia oxidation in wetland soils [28]. At low NH4+ concentrations of 71 µM and 1.1 µmol g−1, CMX also dominated the ammonia oxidizer communities in the RGSF [7] and in coastal wetland sediments [28], respectively. However, our results demonstrate that high abundance of CMX in an environment does not necessarily imply that they also control the complete nitrification process. The competitive advantage of CMX has been associated with a biofilm-associated lifestyle such as in RGSF. However, division of labor in nitrification may remain the preferred strategy in the planktonic communities of groundwater environments at low NH4+ concentrations.
Distribution patterns of ammonia oxidizers across the groundwater wells suggested different niche preferences for AOB and CMX clade A compared to AOA and some clade B affiliated CMX. Both AOA and CMX clade B dominated in groundwater closer to the major recharge area with lower NH4+ availability. AOA can be the most abundant ammonia oxidizers in soils [74] and several studies reported CMX clade B in soil habitats [9, 75]. Consequently, transfer from soils to shallow groundwater could result in the observed high fractions of AOA and CMX clade B in the ammonia oxidizer communities of the respective wells [47]. In turn, deeper and more distant groundwater favored both AOB and clade A affiliated CMX under slightly increased availability of NH4+ and lower DO concentrations.
The observed ecophysiological niche differentiation of CMX clade A and clade B across different groundwater conditions was not reflected by differences in central nitrogen metabolism of representative MAGs [21, 30] and metaproteomic data from the groundwater. Genes encoding enzymes of ammonia and nitrite oxidation were highly transcribed and respective peptides represented both CMX clades. However, differences in the encoded NH4+ transporters could be linked to different ecological preferences. Rh-type NH4+ transporters in CMX clade A convey a better adaptation to higher NH4+ concentrations than Amt-type transporters. As Rh-type transporters have higher uptake capacity but lower NH3 affinity than Amt-type transporters [25] and are also encoded in AOB [76], CMX clade A might have a competitive advantage against clade B in the groundwater when more NH4+ is available. Along with NH4+ uptake efficiency, also oxygen tolerance appeared to drive the distribution patterns of the two clades, as CMX clade B but not clade A encode two types of superoxide dismutases (SOD1, SOD2) [25, 26]. Finally, given the broader metabolic versatility of clade B including the potential for formate and thiosulfate utilization, CMX clade B representatives might be less strongly tied to energy generation by nitrification compared to the clade A representatives.
In conclusion, our findings point to a strong contribution of AOB to ammonia oxidation activity in oligotrophic groundwater, despite numerical predominance of CMX. Given the limitations of differential inhibitor assays, the exact contribution of CMX to ammonia oxidation in situ remains unclear. Conditions of low NH4+ availability and varying oxygen concentrations appeared to generally support coexistence of CMX, AOB and AOA with representatives closely related to those with the highest NH4+ affinities in their respective groups. Our results suggest that high substrate affinity, high growth yields, and a potentially major role in nitrite oxidation enable CMX Nitrospira to maintain consistently high populations in the groundwater. We further propose niche differentiation of CMX clade A and B with a more important role in nitrification for clade A and a higher metabolic versatility, including the use of alternative electron donors, for clade B.
Data availability
Sequence data generated in this study was submitted to the European Nucleotide Sequence Archive with project number PRJEB57321 for amoA genes under accessions ERS13665998 to ERS13665900, and with project number PRJEB63087 for bacterial 16 S rRNA genes under accessions ERS15585018 to ERS15584991.
The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE [56] partner repository with the dataset identifier PXD039573. Access to this data can be granted upon request.
References
Kuypers MMM, Marchant HK, Kartal B. The microbial nitrogen-cycling network. Nat Rev Microbiol. 2018;16:263–76.
Koch H, van Kessel MAHJ, Lücker S. Complete nitrification: insights into the ecophysiology of comammox Nitrospira. Appl Microbiol Biotechnol. 2018;103:177–89.
Lehtovirta-Morley LE Ammonia oxidation: ecology, physiology, biochemistry and why they must all come together. FEMS Microbiol Lett. 2018;365:fny058.
Daims H, Lebedeva EV, Pjevac P, Han P, Herbold C, Albertsen M, et al. Complete nitrification by nitrospira bacteria. Nature. 2015;528:504–9.
Van Kessel MAHJ, Speth DR, Albertsen M, Nielsen PH, Op Den Camp HJM, Kartal B, et al. Complete nitrification by a single microorganism. Nature. 2015;528:555–9.
Fowler SJ, Palomo A, Dechesne A, Mines PD, Smets BF. Comammox Nitrospira are abundant ammonia oxidizers in diverse groundwater-fed rapid sand filter communities. Environ Microbiol. 2018;20:1002–15.
Gülay A, Fowler SJ, Tatari K, Thamdrup B, Albrechtsen H-J, Al-Soud WA, et al. DNA- and RNA-SIP Reveal Nitrospira spp. as Key Drivers of Nitrification in Groundwater-Fed Biofilters. MBio 2019;10:e01870-19.
Spasov E, Tsuji JM, Hug LA, Doxey AC, Sauder LA, Parker WJ, et al. High functional diversity among Nitrospira populations that dominate rotating biological contactor microbial communities in a municipal wastewater treatment plant. ISME J. 2020;14:1857–72.
Li C, Hu HW, Chen QL, Chen D, He JZ. Niche differentiation of clade a comammox Nitrospira and canonical ammonia oxidizers in selected forest soils. Soil Biol Biochem. 2020;149:107925.
Osburn ED, Barrett JE. Abundance and functional importance of complete ammonia-oxidizing bacteria (comammox) versus canonical nitrifiers in temperate forest soils. Soil Biol Biochem. 2020;145:2015–8.
Zhao Z, Huang G, He S, Zhou N, Wang M, Dang C, et al. Abundance and community composition of comammox bacteria in different ecosystems by a universal primer set. Sci Total Environ. 2019;691:146–55.
Shi Y, Jiang Y, Wang S, Wang X, Zhu G. Biogeographic distribution of comammox bacteria in diverse terrestrial habitats. Sci Total Environ. 2020;717:137257.
Liu S, Wang H, Chen L, Wang J, Zheng M, Liu S, et al. Comammox Nitrospira within the Yangtze River continuum: community, biogeography, and ecological drivers. ISME J. 2020;14:2488–504.
Yu C, Hou L, Zheng Y, Liu M, Yin G, Gao J, et al. Evidence for complete nitrification in enrichment culture of tidal sediments and diversity analysis of clade a comammox Nitrospira in natural environments. Appl Microbiol Biotechnol. 2018;102:9363–77.
Kits DK, Sedlacek CJ, Lebedeva EV, Han P, Bulaev A, Pjevac P, et al. Kinetic analysis of a complete nitrifier reveals an oligotrophic lifestyle. Nature. 2017;549:269–72.
Roots P, Wang Y, Rosenthal AF, Griffin JS, Sabba F, Petrovich M, et al. Comammox Nitrospira are the dominant ammonia oxidizers in a mainstream low dissolved oxygen nitrification reactor. Water Res. 2019;157:396–405.
Martens-Habbena W, Berube PM, Urakawa H, De La Torre JR, Stahl DA. Ammonia oxidation kinetics determine niche separation of nitrifying Archaea and Bacteria. Nature. 2009;461:976–9.
Prosser JI, Nicol GW. Archaeal and bacterial ammonia-oxidisers in soil: the quest for niche specialisation and differentiation. Trends Microbiol. 2012;20:523–31.
Herber J, Klotz F, Frommeyer B, Weis S, Straile D, Kolar A, et al. A single Thaumarchaeon drives nitrification in deep oligotrophic lake constance. Environ Microbiol. 2020;22:212–28.
Sedlacek CJ, McGowan B, Suwa Y, Sayavedra-Soto L, Laanbroek HJ, Stein LY, et al. A physiological and genomic comparison of nitrosomonas cluster 6a and 7 Ammonia-oxidizing bacteria. Micro Ecol. 2019;78:985–94.
Overholt WA, Trumbore S, Xu X, Bornemann TLV, Probst AJ, Krüger M, et al. Carbon fixation rates in groundwater similar to those in oligotrophic marine systems. Nat Geosci. 2022;15:561–7.
Opitz S, Küsel K, Spott O, Totsche KU, Herrmann M. Oxygen availability and distance to surface environments determine community composition and abundance of ammonia-oxidizing prokaroytes in two superimposed pristine limestone aquifers in the Hainich region, Germany. FEMS Microbiol Ecol. 2014;90:39–53.
Van Der Wielen PWJJ, Voost S, Van Der Kooij D. Ammonia-oxidizing bacteria and archaea in groundwater treatment and drinking water distribution systems. Appl Environ Microbiol. 2009;75:4687–95.
Palomo A, Dechesne A, Cordero OX, Smets BF. Evolutionary ecology of natural comammox nitrospira populations. mSystems. 2022;7:e0113921.
Palomo A, Pedersen AG, Fowler SJ, Dechesne A, Sicheritz-Pontén T, Smets BF. Comparative genomics sheds light on niche differentiation and the evolutionary history of comammox Nitrospira. ISME J. 2018;12:1779–93.
Poghosyan L, Koch H, Lavy A, Frank J, van Kessel MAHJ, Jetten MSM, et al. Metagenomic recovery of two distinct comammox Nitrospira from the terrestrial subsurface. Environ Microbiol. 2019;21:3627–37.
Palomo A, Dechesne A, Pedersen AG, Smets BF. Genomic profiling of Nitrospira species reveals ecological success of comammox Nitrospira. Microbiome. 2022;10:204.
Sun D, Tang X, Li J, Liu M, Hou L, Yin G, et al. Chlorate as a comammox Nitrospira specific inhibitor reveals nitrification and N2O production activity in coastal wetland. Soil Biol Biochem. 2022;173:108782.
Wang DQ, Zhou CH, Nie M, Gu JD, Quan ZX. Abundance and niche specificity of different types of complete ammonia oxidizers (comammox) in salt marshes covered by different plants. Sci Total Environ. 2021;768:144993.
Wegner CE, Gaspar M, Geesink P, Herrmann M, Marz M, Küsel K. Biogeochemical regimes in shallow aquifers reflect the metabolic coupling of the elements nitrogen, sulfur, and carbon. Appl Environ Microbiol. 2019;85:e02346–18.
Herrmann M, Rusznyák A, Akob DM, Schulze I, Opitz S, Totsche KU, et al. Large Fractions of CO 2 -fixing microorganisms in Pristine Limestone Aquifers Appear to be involved in the oxidation of reduced sulfur and nitrogen compounds. Appl Environ Microbiol. 2015;81:2384–94.
Küsel K, Totsche KU, Trumbore SE, Lehmann R, Steinhäuser C, Herrmann M. How deep can surface signals be traced in the Critical Zone? merging biodiversity with biogeochemistry research in a central German Muschelkalk landscape. Front Earth Sci. 2016;4:32.
Kohlhepp B, Lehmann R, Seeber P, Küsel K, Trumbore SE, Totsche KU. Aquifer configuration and geostructural links control the groundwater quality in thin-bedded carbonate-siliciclastic alternations of the Hainich CZE, central Germany. Hydrol Earth Syst Sci. 2017;21:6091–116.
Lehmann R, Totsche KU. Multi-directional flow dynamics shape groundwater quality in sloping bedrock strata. J Hydrol. 2020;580:124291.
Scheiner D. A modified version of the sodium salicylate method for analysis of wastewater nitrates. Water Res. 1974;8:835–40.
Seelmeyer G. Deutsche Einheitsverfahren zur Wasseruntersuchung. Herausgegeben im Auftrage der Fachgruppe Wasserchemie in der Gesellschaft Deutscher Chemiker E. V. Bearbeitet von L. W. Haase, H. Stooff, G. Gad, W. Wesly u.a. Verlag Chemie, Weinheim/Bergstraße 1954, 180. Mater Corros. 1954;5:357.
Shen T, Stieglmeier M, Dai J, Urich T, Schleper C. Responses of the terrestrial ammonia-oxidizing archaeon Ca. Nitrososphaera viennensis and the ammonia-oxidizing bacterium Nitrosospira multiformis to nitrification inhibitors. FEMS Microbiol Lett. 2013;344:121–9.
Herrmann M, Hädrich A, Küsel K. Predominance of thaumarchaeal ammonia oxidizer abundance and transcriptional activity in an acidic fen. Environ Microbiol. 2012;14:3013–25.
Rotthauwe JH, Witzel KP, Liesack W. The ammonia monooxygenase structural gene amoA as a functional marker: molecular fine-scale analysis of natural ammonia-oxidizing populations. Appl Environ Microbiol. 1997;63:4704–12.
Francis CA, Roberts KJ, Beman JM, Santoro AE, Oakley BB. Ubiquity and diversity of ammonia-oxidizing archaea in water columns and sediments of the ocean. Proc Natl Acad Sci. 2005;102:14683–8.
Pester M, Maixner F, Berry D, Rattei T, Koch H, Lücker S, et al. NxrB encoding the beta subunit of nitrite oxidoreductase as functional and phylogenetic marker for nitrite-oxidizing Nitrospira. Environ Microbiol. 2014;16:3055–71.
Alves RJE, Minh BQ, Urich T, Von Haeseler A, Schleper C. Unifying the global phylogeny and environmental distribution of ammonia-oxidising archaea based on amoA genes. Nat Commun. 2018;9:1517.
Tourna M, Freitag TE, Nicol GW, Prosser JI. Growth, activity and temperature responses of ammonia-oxidizing archaea and bacteria in soil microcosms. Environ Microbiol. 2008;10:1357–64.
Callahan BJ, McMurdie PJ, Rosen MJ, Han AW, Johnson AJA, Holmes SP. DADA2: high-resolution sample inference from Illumina amplicon data. Nat Methods. 2016;13:581–3.
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB, et al. Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol. 2009;75:7537–41.
Purkhold U, Wagner M, Timmermann G, Pommerening-Röser A, Koops HP. 16S rRNA and amoA-based phylogeny of 12 novel betaproteobacterial ammonia-oxidizing isolates: Extension of the dataset and proposal of a new lineage within the nitrosomonads. Int J Syst Evol Microbiol. 2003;53:1485–94.
Krüger M, Potthast K, Michalzik B, Tischer A, Küsel K, Deckner FFK, et al. Drought and rewetting events enhance nitrate leaching and seepage-mediated translocation of microbes from beech forest soils. Soil Biol Biochem. 2021;154:2020.08.03.234047.
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P, et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 2013;41:590–6.
McMurdie PJ, Holmes S. Phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS One. 2013;8:e61217.
Chaudhari NM, Overholt WA, Figueroa-Gonzalez PA, Taubert M, Bornemann TLV, Probst AJ, et al. The economical lifestyle of CPR bacteria in groundwater allows little preference for environmental drivers. Environ Microbiome. 2021;16:24.
Fish JA, Chai B, Wang Q, Sun Y, Brown CT, Tiedje JM, et al. FunGene: the functional gene pipeline and repository. Front Microbiol. 2013;4:291.
Edgar RC. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004;32:1792–7.
Letunic I, Bork P. Interactive tree of life (iTOL) v5: an online tool for phylogenetic tree display and annotation. Nucleic Acids Res. 2021;49:W293–W296.
Taubert M, Overholt WA, Heinze BM, Matanfack GA, Houhou R, Jehmlich N, et al. Bolstering fitness via CO2 fixation and organic carbon uptake: mixotrophs in modern groundwater. ISME J. 2021;16:1153–62.
Taubert M, Stöckel S, Geesink P, Girnus S, Jehmlich N, von Bergen M, et al. Tracking active groundwater microbes with D2O labelling to understand their ecosystem function. Environ Microbiol. 2018;20:369–84.
Perez-Riverol Y, Csordas A, Bai J, Bernal-Llinares M, Hewapathirana S, Kundu DJ, et al. The PRIDE database and related tools and resources in 2019: improving support for quantification data. Nucleic Acids Res. 2019;47:D442–D450.
Team RC. A Language and Environment for Statistical Computing. R Found Stat Comput. 2020. R Foundation for Statistical Computing, Vienna, Austria. 2: https://www.R--project.org.
Ogle D, Wheeler P, Dinno A. FSA: Fisheries Stock Analysis. R package version 0.8.31.2020.
Gu Z, Eils R, Schlesner M. Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics. 2016;32:2847–9.
Oksanen J, Kindt R, Legendre P, O’Hara B, Simpson GL, Solymos PM, et al. The vegan package version 2.5-7. Community Ecol Packag 2008;190.
Bédard C, Knowles R. Physiology, biochemistry, and specific inhibitors of CH4, NH4+, and CO oxidation by methanotrophs and nitrifiers. Microbiol Rev. 1989;53:68–84.
Small GE, Bullerjahn GS, Sterner RW, Beall BFN, Brovold S, Finlay JC, et al. Rates and controls of nitrification in a large oligotrophic lake. Limnol Oceanogr. 2013;58:276–86.
Newell SE, Babbin AR, Jayakumar A, Ward BB Ammonia oxidation rates and nitrification in the Arabian Sea. Global Biogeochem Cycles 2011;25:GB4016.
Beman JM, Chow CE, King AL, Feng Y, Fuhrman JA, Andersson A, et al. Global declines in oceanic nitrification rates as a consequence of ocean acidification. Proc Natl Acad Sci USA. 2011;108:208–13.
Bristow LA, Dalsgaard T, Tiano L, Mills DB, Bertagnolli AD, Wright JJ, et al. Ammonium and nitrite oxidation at nanomolar oxygen concentrations in oxygen minimum zone waters. Proc Natl Acad Sci. 2016;113:10601–6.
Gribsholt B, Boschker HTS, Struyf E, Andersson M, Tramper A, De Brabandere L, et al. Nitrogen processing in a tidal freshwater marsh: a whole-ecosystem 15N labeling study. Limnol Oceanogr. 2005;50:1945–59.
Miranda J, Balachandran KK, Ramesh R, Wafar M. Nitrification in Kochi backwaters. Estuar Coast Shelf Sci. 2008;78:291–300.
Kostan J, Sjöblom B, Maixner F, Mlynek G, Furtmüller PG, Obinger C, et al. Structural and functional characterisation of the chlorite dismutase from the nitrite-oxidizing bacterium ‘ Candidatus Nitrospira defluvii’: identification of a catalytically important amino acid residue. J Struct Biol. 2010;172:331–42.
Zhao J, Bello MO, Meng Y, Prosser JI, Gubry-Rangin C. Selective inhibition of ammonia oxidising archaea by simvastatin stimulates growth of ammonia oxidising bacteria. Soil Biol Biochem. 2020;141:107673.
Vilardi KJ, Cotto I, Sevillano M, Dai Z, Anderson CL, Pinto A. Comammox Nitrospira bacteria outnumber canonical nitrifiers irrespective of electron donor mode and availability in biofiltration systems. FEMS Microbiol Ecol. 2022;98:fiac032.
Mosley OE, Gios E, Close M, Weaver L, Daughney C, Handley KM. Nitrogen cycling and microbial cooperation in the terrestrial subsurface. ISME J. 2022;16:2561–73.
Sakoula D, Koch H, Frank J, Jetten MS, van Kessel MA, Lücker S. Enrichment and physiological characterization of a novel comammox Nitrospira indicates ammonium inhibition of complete nitrification. ISME J. 2020;15:1010–24.
Tatari K, Gülay A, Thamdrup B, Albrechtsen HJ, Smets BF. Challenges in using allylthiourea and chlorate as specific nitrification inhibitors. Chemosphere. 2017;182:301–5.
Cardarelli EL, Bargar JR, Francis CA. Diverse Thaumarchaeota dominate subsurface ammonia-oxidizing communities in semi-arid floodplains in the Western United States. Micro Ecol. 2020;80:778–92.
Zhu G, Wang X, Wang S, Yu L, Armanbek G, Yu J, et al. Towards a more labor-saving way in microbial ammonium oxidation: a review on complete ammonia oxidization (comammox). Sci Total Environ. 2022;829:154590. Elsevier
Weidinger K, Neuhäuser B, Gilch S, Ludewig U, Meyer O, Schmidt I. Functional and physiological evidence for a Rhesus-type ammonia transporter in Nitrosomonas europaea. FEMS Microbiol Lett. 2007;273:260–7.
Acknowledgements
We thank Heiko Minkmar, Falko Gutmann, Jens Wurlitzer, Stefan Krümmling, Swantantar Kumar, Patricia Geesink, Lena Carstens and Linda Gorniak for support of field and laboratory work and Gundula Rudolph for ion chromatography measurements. We acknowledge Kai Uwe Totsche and Robert Lehmann for field site management and provision of physicochemical data from groundwater.
Funding
This study was part of the Collaborative Research Centre 1076 AquaDiva (CRC AquaDiva) of the Friedrich Schiller University Jena, funded by the Deutsche Forschungsgemeinschaft under project number 218627073. Climate chambers to conduct experiments under controlled temperature conditions and the infrastructure for high-throughput sequencing were financially supported by the Thüringer Ministerium für Wirtschaft, Wissenschaft und Digitale Gesellschaft (TMWWDG; project B 715-09075 and project 2016 FGI 0024 “BIODIV”). Martin Taubert gratefully acknowledges funding by the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation) under Germany´s Excellence Strategy – EXC 2051 – Project-ID 390713860. Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
MH and MK designed the study. MK conducted all molecular work, incubation experiments and data analysis. BT and LB performed IRMS measurements and analysis. NC and WO performed metagenomic data analysis. MT, NJ and MvB conducted metaproteomic data analysis. MK and MH wrote the manuscript with contributions of all authors.
Corresponding author
Ethics declarations
Competing interests
The author’s declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Krüger, M., Chaudhari, N., Thamdrup, B. et al. Differential contribution of nitrifying prokaryotes to groundwater nitrification. ISME J 17, 1601–1611 (2023). https://doi.org/10.1038/s41396-023-01471-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-023-01471-4