Abstract
Meromictic Lake Cadagno is a permanently stratified system with a persistent microbial bloom within the oxic-anoxic boundary called the chemocline. The association between oxygenic and anoxygenic photosynthesis within the chemocline has been known for at least two decades. Although anoxygenic purple and green sulfur bacteria have been well studied, reports on oxygenic phytoplankton have remained sparse since their discovery in the 1920s. Nearly a century later, this study presents the first near-complete genome of a photosynthetic microbial eukaryote from the chemocline of Lake Cadagno, provisionally named Chlorella-like MAG. The 18.9 Mbp nuclear genome displays a high GC content (71.5%), and the phylogenetic placement suggests that it is a novel species of the genus Chlorella of Chlorophytes. Functional annotation of the Chlorella-like metagenome-assembled genome predicted 10,732 protein-coding genes, with an approximate 0.6% proportion potentially involved in carbon, sulfur, and nitrogen (C, N, and S) metabolism. In addition to C4 photosynthesis, this study detected genes for heat shock proteins (HSPs) in the Chlorella-like algae, consistent with the other Chlorella species. Altogether, the genomic insights in this study suggest the cooperation of photosynthetic algae with phototrophic sulfur bacteria via C, N, and S metabolism, which may aid their collective persistence in the Lake Cadagno chemocline. Furthermore, this work additionally presents the chloroplast genome of Cryptomonas-like species, which was likely to be presumed as cyanobacteria in previous studies because of the presence of phycobilisomes.
Similar content being viewed by others
Introduction
Meromictic Lake Cadagno is situated at an altitude of 1921 m within the Swiss Alps, with permanent stratification into three zones: mixolimnion (upper oxic), monimolimnion (lower anoxic), and chemocline (oxic-anoxic boundary) [1,2,3,4,5,6]. The chemocline harbors a persistent microbial bloom that coincides with decreased oxygen and light concentrations but increased ammonium, iron, and sulfide concentrations [7,8,9,10,11,12,13]. Within these physicochemical changes of the chemocline, Chl a and turbidity peaks have been used as a proxies for oxygenic phytoplankton (Chl a) and anoxygenic photo- and chemotrophic sulfur bacteria (turbidity) for at least two decades [14,15,16]. Oxygenic photosynthesis specifically facilitates dark aerobic sulfide oxidation [17] by Chromatium okenii, a purple sulfur bacterium, which is also known to contribute most to the biomass of Lake Cadagno chemocline [1, 18]. Oxygenic photosynthesis also fuels in situ oxygen for methane oxidation [19] and may create microaerophilic conditions for iron oxidation [20]. Since in situ oxygen production is proposed to be essential for methane oxidation, iron oxidation, and dark aerobic sulfide oxidation in the chemocline, identifying oxygen-producing phototrophs, including photosynthetic algae and diatoms, remains limited to microscopy [17, 19].
Phycobilin-containing cells and phycocyanin signals are often used as proxies for cyanobacteria in Lake Cadagno (June and October 2013 [21], August [12] and September 2017 [22], and August 2019 [15]) and were hypothesized to create microoxic conditions by facilitating in situ oxygen production [12, 22]. However, information on cyanobacterial species contributing to phycobilin and phycocyanin signals was lacking. While these studies used flow cytometry to identify cyanobacteria, a parallel 16S amplicon gene phylogeny from samples collected in August 2017 revealed that cyanobacteria were rare and that the chloroplasts of Chlorophyta (green algae) and Ochrophyta species (diatoms) were abundant in the chemocline [16]. These contrasting findings on cyanobacteria and chloroplasts warrant the genomic characterization of the microbial communities present in the chemocline of Lake Cadagno, as both can perform oxygenic photosynthesis.
The photosynthetic pigments (chlorophyll a and phycocyanin) of oxygenic phototrophs or phytoplankton peak at the beginning of the chemocline [7, 14, 15], where particulate sulfur, hydrogen sulfide, particulate organic nitrogen, and ammonium concentrations also start to rise [16]. Sulfide is toxic to most eukaryotes, but algae may acquire sulfur from sulfate [23], and some microalgae, such as Chlorella sorokiniana, are also capable of reducing sulfide to sulfate [24]. In addition to sulfur, some reports have highlighted that microbial eukaryotes [25] and cyanobacteria [26] can also metabolize nitrogen. However, their prospective roles in sulfur and nitrogen cycling in Lake Cadagno remain unknown.
The prokaryotic population of Lake Cadagno has been rigorously studied, and the genomes of anoxygenic purple sulfur bacteria (for example, C. okenii and Thiodictyon syntrophicum) that modulate sulfur and nitrogen metabolism have been uncovered [11, 27]. No microbial eukaryotic genome has been described for the Lake Cadagno water column since its initial observation in the 1920s [28]. This study presents the first near-complete genome of the most abundant photosynthetic microbial eukaryote at the beginning of the Lake Cadagno chemocline, focusing on its potential for carbon, sulfur, and nitrogen (C, N, and S) metabolism.
Results and discussion
Phytoplankton, and phototrophic sulfur bacteria peak in the oxic-anoxic boundary called the chemocline of Lake Cadagno, as indicated by photosynthetic pigments (chlorophyll a and phycocyanin) and turbidity (13–15.5 m, Fig. S1) [16]. These microbial peaks distinguish upper-oxic mixolimnion from lower anoxic-monimolimnion, and shotgun DNA sequencing in this study investigated these communities following prior hypotheses of in situ oxygen production by photosynthetic algae [17, 19], and cyanobacteria [22].
Protist community composition and reconstruction of algae and diatoms genomes
From the four chemocline samples, millions of raw reads were quality checked, normalized (29,747,969–80,546,876; Table S1), and assembled into contigs using SPAdes. The total size of assembled contigs ranged between 360–890 Mbp with N50 values between 3,496–8,393 bp and a total number of contigs between 87,929–232,398 (Fig. S2A–D). At 15 m depth, where phytoplankton (Chl a, phycocyanin) were close to the maximum, 7,659 contigs (L50) contributed to 50% of the whole metagenome assembly with a minimum length of 8,393 bp (N50), indicating the contribution of longer contigs (Fig. S2C; 15-w). From each depth of the chemocline, hundreds of Metagenome-Assembled Genomes (MAGs) were obtained based on the coverage and sequence composition using CONCOCT (Fig. 1A) [29]. Most of the resulting MAGs belonged to bacterial lineages, with only a few microbial eukaryotic genomes (4.7 to 13.6% of total MAGs; Fig. 1A, B). This low occurrence of eukaryotic MAGs may be due to their low abundance and may also reflect the challenges associated with eukaryotic genome binning owing to their larger genome size than most prokaryotes observed in the Lake Cadagno chemocline (Fig. 1B, C) [30]. MAGs of phytoplankton, including cyanobacteria and photosynthetic microbial eukaryotes, have been further scrutinized for their involvement in in situ oxygen production via oxygenic photosynthesis [17, 19, 22]. Typical cyanobacteria-specific phycocyanin and phycobilin signals have been reported (years 2013 [21], 2017 [12, 22], and 2019 [15]) for the chemocline of Lake Cadagno. In contrast, this study recovered only one putative cyanobacteria-like MAG with 11.4% completion, based on marker genes detected by BUSCO (Benchmarking Universal Single-Copy Orthologs) (13m_Bin_1, Table S2), which may be due to their previously observed low abundance in the chemocline [16].
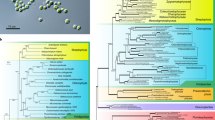
A Bacterial and eukaryotic MAGs obtained through competitive binning via CONCOCT. B Size of MAGs classified by CAT/BAT taxonomy (hits >0.5 bit-scores). C Quality assessment of MAGs using BUSCO (only eukaryotic MAGs with >10% completeness are showed). D Eukaryotic community classification using 18S rRNA gene sequences extracted from the libraries with Phyloflash. The light-lilac, lilac, and dark-lilac color columns on the right y-axis of the plots indicate the mixolimnion-chemocline transition, chemocline, and lower chemocline, respectively.
By analyzing metagenomics reads for 18S rRNA genes, this study observed a mixed population of microbial eukaryotes (Fig. 1D). The relative abundance of eukaryotic community composition in the chemocline was as follows: Alveolata (4.3–13.8%), Cryptomonadales (6.0–16.2%), Obozoa (12.7–27.8%), Rhizaria (8.3–12.1 %), Chloroplastida (10.7–19.5%) and Stramenopiles (5.3–26.2%). Previously, the relative abundance (%) based on 18 S rRNA gene sequences indicated the dominance (>70%) of Cryptophyta with approximately 5–10% of Perkinsozoa (Alveolata) and Stramenopiles [6]. Dinoflagellates (Alveolata), cryptophytes (Cryptomonadales), chlorophytes (Chloroplastida), and diatoms (Stramenopiles) contain chlorophyll a (Chl a) [31], which may contribute to oxygenic photosynthesis in the chemocline. However, genus or species-level investigations are required to know the identity of microorganisms, and eukaryotic MAGs provide further insights.
From the overall eukaryotic populations, BUSCO in the auto-lineage mode identified eukaryotic MAGs with a wide range of gene content completeness (38–94.8%), including Chlorophyta (algae; genus Chlorella, up to 94.8% completeness) and Stramenopile (diatom; genus Nitzschia, up to 62% completeness), when using both competitive (Fig. 1C; Table S2), and non-competitive binning (Fig. S2G; Table S3). Genomes from these photosynthetic clades were expected because chloroplast amplicons of Chlorophyta and Ochrophyta (Stramenopile) were identified in a previous study [16]. Putative MAGs of other eukaryotic algae (Cryptophyceae), including the genus Cryptomonas and Gullardia, were also obtained but had low completion (<10%, Table S10). Overall, these results provide the first assembled genomes of algae and diatoms from the chemocline of Lake Cadagno, which have the potential to contribute to oxygenic photosynthesis.
High-quality eukaryotic genome of a novel species of Chlorella-like microorganism
More genomes of protists are needed to advance our understanding of their biology [32, 33]; however, obtaining well-curated microbial eukaryotic genomes from metagenomes remains challenging owing to their large genome size and complexity [30, 34]. Although there are pioneering studies in other systems [35, 36], there have been no prior reports on eukaryotic genomics from the Lake Cadagno chemocline. In this study, after co-assembling the Chlorophyta-specific reads, a representative Chlorophyta genome of 19.4 Mb (636 contigs; minimum length: 2500 bp, N50: 45 Kbp, Table S4) was obtained by re-assembling and re-binning using SPAdes and CONCOCT respectively. From this representative genome, prospective contaminant contigs (n = 40, Table S5) were removed by referring to GC content and coverage using the Anvi’o interface guided by CAT taxonomy. After removing the contaminants, a final 18.9 Mbp-genome with 596 contigs with a maximum contig length of 181 Kbp was obtained (N50: 46 Kbp, Table S6). The refined Chlorophyta MAG had 93.2% BUSCO completion (n = 1519, Chlorophyta dataset), with 10,732 protein-coding genes and an average genome coverage of 141X (Fig. 2A–C). Not only did the rebinning and refining step result in a decrease in the total number of contigs, but the contribution of long contigs also increased (total contig = 596, of which 19 > 100 Kbp, 119 > 50 Kbp, and 282 > 25 Kbp) compared to the best-quality MAG obtained during primary competitive binning (15-w_36, total contig = 1104, of which 3 > 100 Kbp, 49 > 50 Kbp, and 257 > 25 Kbp). However, the BUSCO completeness score of the refined MAG slightly decreased (by 1.6%) compared with that before refinement MAG (15-w_36, 94.8% complete; Fig. 1C).
A Snail plot of the 18.97 Mbp Chlorophyta genome assembly (Maximum contig length 181 Kbp, shown in red). Orange and pale-orange arcs indicate the N50 and N90 values, respectively, where the longest scaffold is displayed in red. B Eukaryotic genome quality assessment by BUSCO using Chlorophyta dataset (n = 1519). C The number of protein-coding genes predicted in Chlorophyta MAG by EukMetaSanity based on Augustus and MetaEuk gene predictors. D Hexagon-binned plot of GC content and coverage based on mapping with Chlorophyta-specific raw reads obtained from the chemocline samples (13, 15-w, 15, and 15.5 m). E Taxonomic classification of Lake Cadagno MAG contigs calculated using the Diamond BLASTx nr database (updated 24 July 2021).
The taxonomic classification with contig annotation tool (CAT) and the diamond BLASTx results against the nr database indicated that this MAG belonged to Chlorellaceae family (Table S2, Fig. 2E). The phylogenomic analysis also placed it close to other Chlorella species (Fig. 3A). Hence, we provisionally refer to this newly assembled genome as Chlorella-like MAG. It is likely that this Chlorella-like species belongs to an early branching lineage of small microbial eukaryotes such as Nannochloris and Chlorella desiccata, but had a significantly higher GC content when compared to the closest relatives (Fig. 3A–C; 71.50 vs. 40–45). Microscopic images of the closest relatives (species from Chlorellaceae) indicated that the Chlorella-like MAG belonged to the nanophytoplankton community (2–20 µm) [37]. The Chlorella-like assembly exhibited high genome quality (93.2% single copy, 1.1% duplicated, 4.7% missing, and 1.0% fragmented) compared to the mean completeness of Chlorellaceae assemblies available at NCBI (87.9%, Fig. 3D).
A Phylogenomic tree of Chlorella-like MAG with other species from the Chlorellaceae family (NCBI TaxID: 35461). The maximum likelihood phylogeny was estimated from a super-alignment (43,679 aa) of 95 single-copy orthologs, using Chlamydomonas reinhardtii as the outgroup species. Branch length represents the substitutions per site. Values on the nodes indicate bootstrap support. B Pictures of Chlorella species obtained from the culture collection of algae and protozoa (CCAP; https://www.ccap.ac.uk). C Size and GC content comparison. D BUSCO quality assessment using the Chlorophyta_odb10 dataset (number of markers = 1519). The values displayed on the bars represent the number of contigs in the genome assembly.
The predicted proteome from the Chlorella-like MAG was mapped to the OrthoDB v10.1 [38] online database at the Trebouxiophyceae level (Chlorophyta Class) which includes 5 species: Auxenochlorella protothecoides (GCF_000733215.1), C. sorokiniana (GCA_002245835.2), Chlorella variabilis (GCF_000147415.1), Coccomyxa subellipsoidea C-169 (GCF_000258705.1), and Helicosporidium sp. ATCC 50920, (GCA_000690575.1). Approximately 6,000 protein-coding genes from the Chlorella-like MAG had orthologs in at least one of the other five species (Fig. S3). From the chemocline of the meromictic Lake Cadagno, these findings represent the first extensively curated high-quality eukaryotic genome and its predicted proteome.
Genomes of Chlorella-like and Cryptomonas-like chloroplasts in the chemocline
Previous studies have reported the possible presence of cyanobacteria in the chemocline of Lake Cadagno based on phycocyanin and phycobilin signals [12, 21, 22]; however, a 16S amplicon gene phylogenetic study limited to amplicon data identified chloroplasts [16]. Here, the metagenomic dataset provides evidence for Cryptophyceae (Cryptomonas curvata and Guillard theta) with 9 putative MAGs from the chemocline samples (Table S2). Cryptomonas are known for their phycobiliproteins, two of which (phycocyanobilin and phycoerythrobilin) are present in cyanobacteria [39]. The putative cyanobacterial-like MAG detected in Lake Cadagno was classified as C. curvata using the contig annotation tool (CAT) (Table S2). The phycobiliproteins are located at the thylakoid lumen of Cryptomonas chloroplast [40], and Cryptomonas have also been previously identified in Lake Cadagno [6, 14]. Thus, in addition to identifying the chloroplasts of newly identified Chlorella-like species, the Cryptomonas chloroplasts were also targeted. Using the available chloroplast genomes of Chlorellaceae (Parachlorella kessleri; NC_012978.1) and Cryptophyceae (G. theta; NC_000926.1) from NCBI, BLASTn identified two prospective contigs with a size of at least 100 Kbp, here referred to as Chloroplast A (cpA) and Chloroplast B (cpB) (Fig. 4A, B). Followed by the circularization of cpA and cpB using NOVOPlasty, phylogenetics analysis confirmed that these prospective chloroplasts belonged to the Chlorellaceae and Cryptophyceae (Fig. 5A).
A Phylogenetic tree based on 18 chloroplast marker genes. Bootstrap support values are shown on nodes. B, C Chloroplast coverage and the number of reads (log) were calculated throughout the water column using Bowtie 2 and SAMtools. The light-green, dark-green, light-lilac, lilac, dark-lilac, and light-brown columns on the right y-axis indicate the zones of high-O2 mixolimnion, medium-O2 mixolimnion, mixolimnion-chemocline transition zone, chemocline, lower anoxic chemocline, and monimolimnion, respectively.
Chlorella-like chloroplast (cpA) coverage and the number of reads (log) peaked in the chemocline and persisted in the monimolimnion, a pattern also matching the nuclear genome (Fig. 5B, Fig. 6A–C). Comparing the mean coverage depths of the nuclear and chloroplast genome within the chemocline (48.6 vs 55.6X at 13 m, Fig. 5B, Fig. 6B), each Chlorella-like cell is likely to have a single copy of the chloroplast genome and thus one chloroplast. A single chloroplast was also observed in Chlorella protothecoides using microscopy [41].
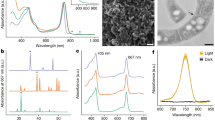
Cryptomonas-like chloroplast (cpB) coverage and read patterns coincided with phycobilisome-containing cells that peaked in the chemocline for sampling season (August 2017) [16] as this study (Fig. 5C). The genes coding for these photosynthetic reaction centers (PSI psa and PSII psb) exist in cpA and cpB chloroplasts (Fig. 4A, B). However, the phycobilisome-specific phycoerythrin protein (cpeB) has only been found in Cryptomonas-like chloroplast, and phycoerythrin has also been identified in Cryptomonas in Lake Cadagno [14]. In previous studies, Cyanobacteria in Lake Cadagno have been identified by targeting phycocyanins in phycobilisomes [12, 21, 22]. In contrast, this study did not recover phycocyanin genes in the Cryptomonas-like chloroplasts, although phycoerythrin is attached to the phycocyanins and is part of the overall phycobilisome structure [42,43,44,45]. Overall, this genomics evidence on MAGs [12, 21, 22] and chloroplast suggests that peaks of phycobilisome-containing cells by flow cytometry [12, 21, 22] and phycocyanin signals [15, 16] in chemocline may have been sourced from Cryptomonas-like cells.
The persistence of Chlorella-like algae and their chloroplasts provides new evidence of ongoing oxygenic photosynthesis in the bottom-monimolimnion. This scenario has also been proposed before for chemocline [19], where a limited amount of light may still be available for oxygenic photosynthesis. The occurrence of Chlorella-like eukaryotic phototrophs in the monimolimnion may also be due to sinking particles in the lower layers of the lake. However, if this scenario is true, this study would expect Cryptomonas-like chloroplasts to sink. Still, their read and coverage patterns decrease in monimolimnion contrary to the Chlorella-like algae and suggest their abundance is restricted to chemocline (Fig. 5B, C). Taken together, the phylogenetic tree, coverage, and read pattern provide evidence that both Chlorella-like and Cryptomonas-like algae may synergistically contribute to oxygenic photosynthesis in the chemocline. However, Chlorella-like eukaryotic algae may also persist in dark monimolimnion, where hydrogen sulfide and ammonia concentrations are at their maximum.
A–C Comparison of the abundance of Chlorella-like eukaryotic algae with purple and green sulfur bacteria based on mapping statistics: A number of reads mapped, B mean depth of coverage and C genome’s relative abundance (%) based on mapping using the total metagenomic reads. D Coverage of protein-coding genes involved in C, N, and S metabolism in the Chlorella-like algae. Protein-coding genes were predicted by EukMetaSanity and mapped to KEGG pathways using eggNOG. A–D The light-green, dark-green, light-lilac, lilac, dark-lilac, and light-brown columns on the right y-axis indicate the zones of high-O2 mixolimnion, medium-O2 mixolimnion, mixolimnion–chemocline transition zone, chemocline, lower anoxic chemocline, and monimolimnion, respectively.
Metabolic potential of Chlorella-like algae with the focus on C, N, and S pathways
The contributions of purple sulfur bacteria (C. okenii, Lamprocystis purpurea, T. syntrophicum, and Thiocystis sp.) and green sulfur bacteria (Chlorobium phaeobacteroides, Chlorobium sp.) for carbon, sulfur, and nitrogen metabolism in the chemocline of Lake Cadagno have been rigorously studied [1, 11, 14, 15, 17, 21, 22, 27, 46, 47]. In contrast, direct evidence of eukaryotic metabolism is yet to be established for the Lake Cadagno chemocline. New Chlorella-like MAG had higher read counts, coverage (48.6X), and relative abundance (3.4%) at the beginning of the chemocline than the purple sulfur bacteria (PSB) and green sulfur bacteria (GSB) (Fig. 6A–C at 13 m). PSB C. okenii and GSB Chlorobium sp. coverage (100 and 61X) and relative abundance (6.3 and 3.5%) were maximum at 15 m whole water sample from the turbidity peak (15-w, Fig. 6B, C). The increased coverage of Chlorella-like green algae, followed by the dominance of Chromatium and Chlorobium is expected as phytoplankton tend to stay above the phototrophic sulfur bacteria, as observed by the peak of phycocyanin, Chl a, and turbidity.
Functional annotations of Chlorella-like MAG using OrthoLoger (Table S7) and eggNOG (Table S8) mappers revealed a repertoire of genes potentially involved in carbon [PATH:ko00710], nitrogen [PATH:ko00910], and sulfur metabolism [PATH:ko00920], with higher coverage in the chemocline and monimolimnion than in the mixolimnion (Fig. 6D; Table S9). The C, N, and S genes for Chlorella-like MAG constituted approximately 0.6% (n = 66) of the total predicted genes (n = 10,732) (Fig. 7A, B). The majority of C, N, and S genes belonged to carbon fixation (50%), followed by sulfur (31%) and nitrogen (18%) metabolism (Fig. 7A, B). Although this study did not detect pathways particularly enriched for Chlorella-like MAG, the C, N, and S metabolism, on the other hand, is consistent with the other Chlorella species (Fig. 7B, C). Yet, the relative abundance comparison indicates that other Chlorella species were not present in the chemocline (Fig. 7D). Further investigation on carbon metabolism (Figs. 6D, 7B; PATH:ko00710) identifies near-complete pathways for the C4 dicarboxylic cycle and crassulacean acid metabolism (CAM) (Fig. S4). These carbon fixation metabolisms are specialized to uptake CO2 in the dark and increase the CO2 availability for photosynthesis [48, 49]. Some genes involved in C4 photosynthesis are shown for Chlorella variabilis; surprisingly, they are not well described for Chlorella species but have been identified in other Chlorophyta genomes [50]. The functional annotations of Chlorella species in this study suggest genes for C4 photosynthesis are consistently present within the Chlorellaceae family (Fig. 7B; PATH:ko00710, Tables S7, S8). Under limited light in the chemocline of Lake Cadagno, such specialized carbon fixation pathways may be used for in situ oxygen production by Chlorella-like photosynthetic algae, which is also coupled with the dark aerobic sulfide oxidation of C. okenii [17]. The sulfur metabolic genes [PATH:00920: Table S9] in Chlorella-like algae may preferentially acquire sulfate [51] resulting from sulfide oxidation by C. okenii and explain its coupling with phototrophic sulfur bacteria in the chemocline (Fig. 6A–C). Genes involved in nitrogen metabolism [PATH:00910: Table S9] may be used for biomass synthesis, as it has been shown in other microalgae, including C. sorokiniana [52], Chlorella vulgaris, and Chlamydomonas [53,54,55]. Nitrate storage in microbial eukaryotes has been suggested to facilitate survival under anoxic conditions [56], supporting the potential ability of Chlorella-like microorganisms to persist in the anoxic zones of Lake Cadagno.
A the total number of protein-coding genes predicted in the present study (number of genes from the official annotation available in NCBI are shown when available); B Relative composition of C, N, and S genes. Values on the barplot represent the number of C, N, and S genes; C Percentage of C, N, and S genes among the total predicted genes; D Relative abundance of Chlorella genomes in the chemocline of Lake Cadagno. E The number and types of heat shock proteins identified in each genome. Only genes belonging to the Chlorophyta and Viridiplantae taxonomy were used.
Additionally, the Chlorella-like microbial genome contains 17 genes belonging to heat shock protein (HSP) families consistent with other Chlorella species (Fig. 7E). Compared to other HSPs in the chemocline, HSP70 has a relatively higher number of genes (Fig. 7E). The HSPs are known for participating in environmental stress response [57], including extreme temperature [58] and exposure to redox metals [57, 59], as shown for Chlorella [60] and other algae [57, 58, 61]. Such environmental stress might frequently occur in Lake Cadagno microbial bloom, owing to the internal oscillations of the chemocline bringing fluctuations in the temperature, light, sulfur, ammonia, and trace metal (iron and manganese) concentrations [9, 16, 62]. Thus, the abundance of heat shock proteins may aid Lake Cadagno Chlorella in thriving in the chemocline while maintaining cooperation with phototrophic sulfur bacteria via C, N, and S metabolism.
Conclusion
Microbial eukaryotes are essential members of the Lake Cadagno chemocline because of autotrophy and their interactions with phototrophic sulfur bacteria via C, N, and S metabolism. This work presented the first near-complete genome (including nuclear and chloroplast assemblies) of a novel green algae species related to Chlorellaceae, providing genomic and phylogenomic evidence for this overlooked microbial eukaryote in Lake Cadagno. Based on this genomic work, primers can be designed to monitor the seasonal abundance of Chlorella-like microorganisms and morphologically characterize them using microscopy. Notably, Chlorella-like species thrive in almost no light and persist in anoxia; thus, its carbon fixation metabolism and genes for heat shock proteins may warrant its potential for biotechnological applications. Additionally, the chloroplast genomes of Chlorella-like and Cryptomonas-like species indicated that both microorganisms are capable of in situ oxygenic photosynthesis, a process that has been previously proposed in the anoxic waters of Lake Cadagno [19]. However, the similarities between Cryptomonas and cyanobacteria containing phycobilisomes suggest caution when reporting cyanobacteria in Lake Cadagno using flow cytometry [12, 21, 22].
Materials and methods
Sample collection and DNA extraction for shotgun DNA metagenomics
Samples were collected from Lake Cadagno, situated at an altitude of 1921 m above sea level (46.5504 °N, 8.7119 °E) in the Swiss Alps. The sampling strategy has been described in a previous study [16]. Briefly, 20 L of water was collected between two subsequent days, 28–29 August 2017, from the stratified zones of Lake Cadagno. The upper-oxic mixolimnion (5, 9, and 11 m) was sampled on the 28th of August (day 1), the chemocline (13, 15, and 15.5 m), and the monimolimnion (17 m) was sampled on day 2 (29th of August). The collected water samples were pre-filtered using a 55 µm mesh to remove zooplankton [14] and subsequently passed through a filtration setup equipped with 0.22 µm-filters (cat. #GPWP14250 142 mm Express Plus filter, Millipore, Darmstadt, Germany). After filtration, the 0.22 µm-filters were flash-frozen at −196 °C and stored at −80 °C until DNA extraction, performed in October 2018, as explained previously [16]. An additional whole water sample without a mesh (15-w) was collected from the turbidity peak.
DNA sequencing and metagenome assembly
PCR-free libraries were prepared with a read length and insert size of 250 bp and sequenced using HiSeq 4000 (Illumina, San Diego, CA, USA) at the University of Michigan Advanced Genomics sequencing core facility. A step-by-step guide on microbial eukaryote genome hunting is available at GitHub (https://github.com/JSSaini/Pipeline_For_Lake_Cadagno_Eukaryotic_Metagenomics; Fig. 8). In summary, raw reads were trimmed, quality checked, and normalized using BBDuk and BBNorm in BBTools (v38.00) [63]. These normalized reads were assembled using SPAdes (v3.15.0) at default kmer lengths of 21, 33, and 55 using 16 CPUs and 250GB of memory [64]. Metagenomics mode (--meta) was used for all samples except 15-w, for which the assembly procedure was successful (i.e., analysis run to completion) without using the “--meta” flag. The contig names of each assembly were simplified using Anvi’o (anvi-script-reformat-fasta, v7), and contigs less than 1,000 nucleotides (-l 1,000) were removed [65].
Raw reads were mapped to the metagenome assemblies (including nuclear and chloroplast) using Bowtie 2 (v2.4.2) [66] and SAMtools (v1.12) [67]. The resulting binary alignment map (BAM) was sorted and indexed using Anvi’o (anvi-init-bam, v7) [65]. Contigs were then binned into MAGs using CONCOCT (clustering contigs with coverage and composition, v1.1.0 [29]) with default parameters using two strategies: by providing coverage information from (a) the individual library and (b) from all libraries, referred to as non-competitive and competitive binning, respectively.
Obtaining representative Chlorophyta MAG from the chemocline
Chlorophyta MAGs with >90% BUSCO (v5.2.0) completeness were detected in all four chemocline samples (13, 15, 15-w, and 15.5 m) using competitive and non-competitive binning. To check for similarity, these MAGs were compared based on average nucleotide identity (Fig. S5) using dRep (v2.2.3) [68]. Chlorophyta-specific reads were extracted by mapping raw reads using Bowtie 2 (v2.4.2) and SAMtools (v1.12) to the concatenated Chlorophyta MAGs from the non-competitive binning to obtain the maximum number of Chlorophyta reads. Next, the Chlorophyta-specific raw reads (R1 and R2) were extracted from the resulting BAM files using bamtofastq (v1.1.0). SPAdes (v3.15.0) [64] coassembled Chlorophyta-specific raw reads at a default kmer length of 21, 33, and 55 nucleotides using two strategies. In the first strategy, SPAdes coassembled Chlorophyta-specific raw reads from all four samples (13, 15, 15-w, and 15.5 m). And in the second strategy, Chlorophyta-specific raw reads from only two samples were coassembled (13 and 15.5 m). The latter assembly was prioritized for rebinning because of the higher N50 value.
The contig names of each assembly were simplified using Anvi’o (anvi-script-reformat-fasta, v7), and contigs less than 2500 nucleotides (-l 2,500) were removed [65]. The Chlorophyta-specific assembly was rebinned using CONCOCT [29] (v1.1.0), providing coverage information (BAM) from all four chemocline samples.
Classification, quality assessment, refinement, and visualization of MAGs
Genomic and taxonomic classification
Raw MAGs were classified using the contig and bin annotation tool (CAT and BAT, v5.2.3) [69]. Gene prediction in CAT/BAT was performed using prodigal [70]. The predicted open reading frames (ORFs) were queried to the NCBI non-redundant protein database (updated 24 July 2021) using DIAMOND (v0.9.14.115) [71,72,73]. To obtain an overview of the eukaryotic community composition, eukaryotic 18S rRNA gene sequences were extracted from metagenomics reads using phyloFlash (v3.4) [74]. The composition of the prokaryotic community based on 16 S amplicon gene sequencing was presented in a previous publication [16].
Quality assessment
MAGs were assessed with BUSCO (v5.2.0) using the “--auto-lineage” mode [75, 76]. BUSCO relies on a collection of single-copy orthologs generated from OrthoDB v10 [77] to identify complete, duplicated, fragmented, and missing single-copy genes. With the “--auto- lineage” mode, BUSCO attempts to identify the most suitable dataset for the assessment and allows the analysis of both prokaryotic and eukaryotic MAGs.
Bin refinement
MAGs were refined by referring to the GC content and coverage using the Anvi’o interface (anvi-refine, Fig. S6) [65]. In addition, the Chlorophyta MAG refinement was guided by CAT taxonomy [69], which aided in removing potential contaminants.
Visualization
Statistics on eukaryotic MAGs (size, contig length, GC content, and coverage) were visualized using BlobToolKit (v2) [78], which uses Diamond BLASTx for taxonomic classification (v0.9.14.115) (nr database, updated 24 July 2021) [71,72,73, 79].
Gene prediction, quantification, and functional annotation of microbial eukaryotes
Protein-coding genes in eukaryotic genomes were predicted using EukMetaSanity (v0.1.0) [80] based on AUGUSTUS (v3.4.0) [81] and MetaEuk (v34c21f2bf34c76f852c0441a29b104e5017f2f6d) [82] gene predictors. The mean depth of gene coverage was calculated using Bowtie 2 (v2.4.2) [66], SAMtools (v1.12), and Anvi’o (v7). The identified protein-coding genes were mapped against the OrthoDB database at the Trebouxiophyceae level (which includes 5 species, v10.1) using the online mapping and charting tool [38, 83], and also mapped at the Chlorophyta level (which includes 17 species, v11) using the OrthoLoger (v3.0.3) software [83]. Additionally, the proteome was mapped to eggNOG (v5.0) orthology resource [84] using eggNOG-mapper (v2.1.9) [85]. KEGG orthology (KO; or K numbers) of the respective genes were mapped to KEGG pathways using the codes provided in the R script at Github under code availability. Optionally, GhostKOALA (v2.2) was used to visualize KEGG pathways based on the KEGG mapper (v5) [86,87,88].
Comparative phylogenomics and quality assessment of Chlorellaceae genomes
We performed a phylogenomic analysis to phylogenetically place the newly discovered Chlorophyta species. The assemblies of related species belonging to Chlorellaceae (NCBI TaxID:35461) were retrieved from NCBI and are listed in the supplementary table (Table S10). Chlamydomonas reinhardtii (GCA_000002595.3) was used as the outgroup. To construct the phylogenomic tree, this study followed a slightly modified version of the snakemake workflow described by Manni et al. 2021 [89], relying on BUSCO [75, 76] to find and extract single-copy orthologs to infer phylogenies. BUSCO (v5.2.2) was run in genome mode (BUSCO_Metaeuk workflow) on each genome assembly using the chlorophyta_odb10 dataset (1,519 markers). The identified single-copy genes that were shared across 100% of the species (with no duplicates across all species) were extracted. For each orthologous group, proteins were aligned using MAFFT (v7.505) [90] and trimmed using trimAl (v1.4 rev15) [91]. The single alignments were concatenated with AMAS (v1.0) [92], and the resulting super-alignment was used to infer a maximum likelihood phylogeny with IQ-TREE (v2.1.2) [93]. The phylogenetic tree was visualized using Dendroscope (v3.7.6) [94] and annotated using ggtree (v3.0.1) [95] in R-studio [96] (v1.4.1106) and Adobe Illustrator (v25.2.1). The results from BUSCO were also used to compare the quality of the newly assembled Chlorella-like MAG with the available Chlorellaceae genomes deposited at NCBI.
Organelle hunting and chloroplast phylogenomics
Chloroplast contigs were identified using BLASTn to obtain the best hits (-max_target_seqs 1) for each query sequence. P. kessleri (NC_012978.1; Chlorella) and G. theta (NC_000926.1; Cryptophyceae) chloroplast genomes were used as queries. The two identified tentative chloroplast genomes (contigs c_000000000152_15mw and c_000000000134-15mm) were termed chloroplast A (cpA) and chloroplast B (cpB), and were used as templates for circularization via NOVOPlasty [97] with a genome size range set to 80,000–200,000 nt and the default kmer length of 33 nucleotides.
Chloroplast phylogenomics was performed using the prospective chloroplast genomes of Lake Cadagno (cpA and cpB) with an additional list of chloroplast genomes from the following three families: (1) Chlorellaceae [Taxonomy ID:35461], (2) Cryptophyceae [Taxonomy ID:3027], and (3) Stramenopile [Taxonomy ID:33634]. Stramenopile chloroplasts were used as the outgroups.
The accession numbers of the chloroplast genomes used in the phylogenomic analysis are provided in supplementary (Table S11). Eighteen marker genes were used to construct the chloroplast phylogenetic tree, including ATP synthase (atpA, atpB, and atpC), large ribosomal subunits (rpl2, rpl5, rpl12, rpl14, rpl19, and rpl23), small ribosomal subunits (rps3, rps8, rps9, and rps19), photosystem I (psaC), and photosystem II (psbA, psbB, psbE, and psbH). These markers were individually aligned using MAFFT (v 7.487) [98], followed by quality assessment and removal of ambiguous sequences using Gblocks (v0.91b) [99]. Phylogenomic tree inferences were made using MEGA (v11.0.10) [100] by selecting the maximum likelihood using the Jones-Taylor-Thornston (JTT) method and bootstrapping (n = 100). The tree was visualized using Dendroscope (v3.7.6) [94] and annotated using ggtree (v3.0.1) [95] in R-studio [96] (v1.4.1106) and Adobe Illustrator (v25.2.1).
Metagenome assembly and abundance statistics
Statistics, including size, GC content, number of contigs, N50 and L50 values of metagenomic assemblies, were calculated using stats.sh (individual) and statswrapper.sh (multiple) scripts in BBMap (v38.96) [101]. The N50 metric is the length of the shortest contig for which half of the genome is assembled on contigs of length N50 or longer, and the L50 value is the minimum number of contigs required to reach 50% of the genome assembly. The mean depth of coverage, number of reads, and relative abundance of purple and green sulfur bacteria and eukaryotic algae MAGs were calculated using Bowtie 2 (v2.4.2) [66], SAMtools (v1.12), Anvi’o (v7), and CoverM (v0.6.1) [102].
Data availability
Raw reads are available at the NCBI under the sequence read archive SUB11916861 and under the accessions SRR21025699, SRR21025700, SRR21025701, and SRR21025702. The GenBank ID for the Chlorella-like MAG is JAOAOU000000000.1. The assembled contigs from raw reads, MAGs, and other data were deposited in Zenodo (https://zenodo.org/record/7505505) [103].
Code availability
A step-by-step guide to the eukaryotic metagenomics pipeline has been made available on GitHub, including the R script used to generate figures (https://github.com/JSSaini/Pipeline_For_Lake_Cadagno_Eukaryotic_Metagenomics).
References
Philippi M, Kitzinger K, Berg JS, Tschitschko B, Kidane AT, Littmann S, et al. Purple sulfur bacteria fix N2 via molybdenum-nitrogenase in a low molybdenum Proterozoic ocean analogue. Nat Commun. 2021;12:4774.
Xiong Y, Guilbaud R, Peacock CL, Cox RP, Canfield DE, Krom MD, et al. Phosphorus cycling in Lake Cadagno, Switzerland: a low sulfate euxinic ocean analogue. Geochim Cosmochim Acta. 2019;251:116–35.
Canfield DE, Farquhar J, Zerkle AL. High isotope fractionations during sulfate reduction in a low-sulfate euxinic ocean analog. Geology. 2010;38:415–8.
Dahl TW, Anbar AD, Gordon GW, Rosing MT, Frei R, Canfield DE. The behavior of molybdenum and its isotopes across the chemocline and in the sediments of sulfidic Lake Cadagno, Switzerland. Geochim Cosmochim Acta. 2010;74:144–63.
Wirth SB, Gilli A, Niemann H, Dahl TW, Ravasi D, Sax N, et al. Combining sedimentological, trace metal (Mn, Mo) and molecular evidence for reconstructing past water-column redox conditions: The example of meromictic Lake Cadagno (Swiss Alps). Geochim Cosmochim Acta. 2013;120:220–38.
Tonolla M, Storelli N, Danza F, Ravasi D, Peduzzi S, Posth NR, et al. Lake Cadagno: Microbial Life in Crenogenic. In: Gulati R, Zadereev E, Degermendzhi A, editors. Ecology of Meromictic Lakes. Springer, Cham; 2017. Ecological Studies, vol 228, pp 155–86.
Gregersen LH, Habicht KS, Peduzzi S, Tonolla M, Canfield DE, Miller M, et al. Dominance of a clonal green sulfur bacterial population in a stratified lake. FEMS Microbiol Ecol. 2009;70:30–41.
Habicht KS, Miller M, Cox RP, Frigaard N-U, Tonolla M, Peduzzi S, et al. Comparative proteomics and activity of a green sulfur bacterium through the water column of Lake Cadagno, Switzerland. Environ Microbiol. 2011;13:203–15.
Ellwood MJ, Hassler C, Moisset S, Pascal L, Danza F, Peduzzi S, et al. Iron isotope transformations in the meromictic Lake Cadagno. Geochim Cosmochim Acta. 2019;255:205–21.
Decristophoris P, Peduzzi S, Ruggeri-Bernardi N, Hahn D, Tonolla M. Fine scale analysis of shifts in bacterial community structure in the chemocline of meromictic Lake Cadagno, Switzerland. J Limnol. 2009;68:16–24.
Luedin S, Liechti N, Cox RP, Danza F, Frigaard N-U, Posth NR, et al. Draft genome sequence of Chromatium okenii isolated from the stratified alpine Lake Cadagno. Sci Rep. 2019;9:1936.
Danza F, Ravasi D, Storelli N, Roman S, Lüdin S, Bueche M, et al. Bacterial diversity in the water column of meromictic Lake Cadagno and evidence for seasonal dynamics. PLoS ONE. 2018;13:e0209743.
Berg J. The microbial impact on Fe & S cycling at oxic-anoxic interfaces: a single-cell view. 2016. Doctoral dissertation, Universität Bremen, Bremen.
Camacho A, Erez J, Chicote A, Florín M, Squires MM, Lehmann C, et al. Microbial microstratification, inorganic carbon photoassimilation and dark carbon fixation at the chemocline of the meromictic Lake Cadagno (Switzerland) and its relevance to the food web. Aquat Sci. 2001;63:91–106.
Di Nezio F, Beney C, Roman S, Danza F, Buetti-Dinh A, Tonolla M, et al. Anoxygenic photo- and chemo-synthesis of phototrophic sulfur bacteria from an alpine meromictic lake. FEMS Microbiol Ecol. 2021;97:fiab010.
Saini JS, Hassler C, Cable R, Fourquez M, Danza F, Roman S, et al. Bacterial, phytoplankton, and viral distributions and their biogeochemical contexts in meromictic Lake Cadagno offer insights into the Proterozoic Ocean microbial loop. MBio. 2022;13:e0005222.
Berg JS, Pjevac P, Sommer T, Buckner CRT, Philippi M, Hach PF, et al. Dark aerobic sulfide oxidation by anoxygenic phototrophs in anoxic waters. Environ Microbiol. 2019;21:1611–26.
Bosshard PP, Stettler R, Bachofen R. Seasonal and spatial community dynamics in the meromictic Lake Cadagno. Arch Microbiol. 2000;174:168–74.
Milucka J, Kirf M, Lu L, Krupke A, Lam P, Littmann S, et al. Methane oxidation coupled to oxygenic photosynthesis in anoxic waters. ISME J. 2015;9:1991–2002.
Berg JS, Michellod D, Pjevac P, Martinez-Perez C, Buckner CRT, Hach PF, et al. Intensive cryptic microbial iron cycling in the low iron water column of the meromictic Lake Cadagno. Environ Microbiol. 2016;18:5288–302.
Posth NR, Bristow LA, Cox RP, Habicht KS, Danza F, Tonolla M, et al. Carbon isotope fractionation by anoxygenic phototrophic bacteria in euxinic Lake Cadagno. Geobiology. 2017;15:798–816.
Luedin SM, Storelli N, Danza F, Roman S, Wittwer M, Pothier JF, et al. Mixotrophic growth under micro-oxic conditions in the purple sulfur bacterium ‘Thiodictyon syntrophicum’. Front Microbiol. 2019;10:384.
Giordano M, Prioretti L. Sulphur and algae: metabolism, ecology and evolution. In: Borowitzka MA, Beardall J, Raven JA, editors. The physiology of microalgae. Cham: Springer International Publishing; 2016. 185–209.
Meier L, Stará D, Bartacek J, Jeison D. Removal of H2S by a continuous microalgae-based photosynthetic biogas upgrading process. Process Saf Environ Prot. 2018;119:65–68.
Terrado R, Monier A, Edgar R, Lovejoy C. Diversity of nitrogen assimilation pathways among microbial photosynthetic eukaryotes. J Phycol. 2015;51:490–506.
Stal LJ. Nitrogen fixation in cyanobacteria. In: eLS. Chichester, UK: John Wiley & Sons, Ltd; 2015. 1–9
Luedin SM, Pothier JF, Danza F, Storelli N, Frigaard N-U, Wittwer M, et al. Complete genome sequence of ‘Thiodictyon syntrophicum’ sp. nov. strain Cad16T, a photolithoautotrophic purple sulfur bacterium isolated from the alpine meromictic Lake Cadagno. Stand Genom Sci. 2018;13:14.
Bachmann H. Das Phytoplankton der Pioraseen nebst einigen Beiträgen zur Kenntnis des Phytoplanktons schweizerischer Alpenseen. Z Hydrol. 1928;4:50–103.
Alneberg J, Bjarnason BS, de Bruijn I, Schirmer M, Quick J, Ijaz UZ, et al. Binning metagenomic contigs by coverage and composition. Nat Methods. 2014;11:1144–6.
Keeling PJ. Combining morphology, behaviour and genomics to understand the evolution and ecology of microbial eukaryotes. Philos Trans R Soc Lond B Biol Sci. 2019;374:20190085.
Sheath RG, Wehr JD. Introduction to the freshwater algae. In: Freshwater algae of North America; 2015. 1–11.
Sibbald SJ, Archibald JM. More protist genomes needed. Nat Ecol Evol. 2017;1:145.
Massana R, López-Escardó D. Metagenome assembled genomes are for eukaryotes too. Cell Genomics. 2022;2:100130.
Chen L-X, Anantharaman K, Shaiber A, Eren AM, Banfield JF. Accurate and complete genomes from metagenomes. Genome Res. 2020;30:315–33.
Delmont TO, Eren AM, Vineis JH, Post AF. Genome reconstructions indicate the partitioning of ecological functions inside a phytoplankton bloom in the Amundsen Sea, Antarctica. Front Microbiol. 2015;6:1090.
Delmont TO, Gaia M, Hinsinger DD, Frémont P, Vanni C, Fernandez-Guerra A, et al. Functional repertoire convergence of distantly related eukaryotic plankton lineages abundant in the sunlit ocean. Cell Genomics. 2022;2:100123.
Leblanc K, Quéguiner B, Diaz F, Cornet V, Michel-Rodriguez M, Durrieu de Madron X, et al. Nanoplanktonic diatoms are globally overlooked but play a role in spring blooms and carbon export. Nat Commun. 2018;9:953.
Zdobnov EM, Kuznetsov D, Tegenfeldt F, Manni M, Berkeley M, Kriventseva EV. OrthoDB in 2020: evolutionary and functional annotations of orthologs. Nucleic Acids Res. 2021;49:D389–93.
Glazer AN, Wedemayer GJ. Cryptomonad biliproteins - an evolutionary perspective. Photosynth Res. 1995;46:93–105.
Reith M, Douglas S. Localization of beta-phycoerythrin to the thylakoid lumen of Cryptomonas phi does not involve a signal peptide. Plant Mol Biol. 1990;15:585–92.
Angelova A, Park S-H, Kyndt J, Fitzsimmons K, Brown JK. Sonication-based isolation and enrichment of Chlorella protothecoides chloroplasts for Illumina genome sequencing. J Appl Phycol. 2014;26:209–18.
Dumay J, Morançais M, Munier M, Le Guillard C, Fleurence J. Chapter Eleven - Phycoerythrins: Valuable proteinic pigments in Red Seaweeds. In: Bourgougnon N, editor. Advances in botanical research. Vol 71, Nantes: Academic Press; 2014. pp 321–43.
Beutler M. Spectral fluorescence of chlorophyll and phycobilins as an in situ tool of phytoplankton analysis: Models, Algorithms and Instruments. 2003. Doctoral dissertation, Christian-Albrechts-Universität, Kiel.
Alam T, Najam L, Al-Harrasi A, et al. Extraction of natural pigments from marine algae. J Agric Mar Sci. 2018;23:81–91.
Haverkamp THA, Others. Shades of red and green: the colorful diversity and ecology of picocyanobacteria in the Baltic Sea. Netherlands Institute of Ecology (NIOO)-Royal Netherlands Academy of Arts; 2008.
Storelli N, Peduzzi S, Saad MM, Frigaard N-U, Perret X, Tonolla M. CO2 assimilation in the chemocline of Lake Cadagno is dominated by a few types of phototrophic purple sulfur bacteria. FEMS Microbiol Ecol. 2013;84:421–32.
Halm H, Musat N, Lam P, Langlois R, Musat F, Peduzzi S, et al. Co-occurrence of denitrification and nitrogen fixation in a meromictic lake, Lake Cadagno (Switzerland). Environ Microbiol. 2009;11:1945–58.
Hultine KR, Cushman JC, Williams DG. New perspectives on crassulacean acid metabolism biology. J Exp Bot. 2019;70:6489–93.
Kellogg EA. C4 photosynthesis. Curr Biol. 2013;23:R594–9.
Chi S, Wu S, Yu J, Wang X, Tang X, Liu T. Phylogeny of C4-photosynthesis enzymes based on algal transcriptomic and genomic data supports an archaeal/proteobacterial origin and multiple duplication for most C4-related genes. PLoS One. 2014;9:e110154.
Giordano M, Norici A, Ratti S, Raven JA. Role of sulfur for algae: acquisition, metabolism, ecology and evolution. In: Hell R, Dahl C, Knaff D, Leustek T, editors. Sulfur metabolism in phototrophic organisms. Dordrecht: Springer Netherlands; 2008. 397–415.
Kim S, Lee Y, Hwang S-J. Removal of nitrogen and phosphorus by Chlorella sorokiniana cultured heterotrophically in ammonia and nitrate. Int Biodeterior Biodegrad. 2013;85:511–6.
Pozzobon V, Cui N, Moreaud A, Michiels E, Levasseur W. Nitrate and nitrite as mixed source of nitrogen for Chlorella vulgaris: Growth, nitrogen uptake and pigment contents. Bioresour Technol. 2021;330:124995.
Scherholz ML, Curtis WR. Achieving pH control in microalgal cultures through fed-batch addition of stoichiometrically-balanced growth media. BMC Biotechnol. 2013;13:39.
Sanz-Luque E, Chamizo-Ampudia A, Llamas A, Galvan A, Fernandez E. Understanding nitrate assimilation and its regulation in microalgae. Front Plant Sci. 2015;6:899.
Kamp A, Høgslund S, Risgaard-Petersen N, Stief P. Nitrate storage and dissimilatory nitrate reduction by eukaryotic microbes. Front Microbiol. 2015;6:1492.
Abassi S, Wang H, Ponmani T, Ki J-S. Small heat shock protein genes of the green algae Closterium ehrenbergii: Cloning and differential expression under heat and heavy metal stresses. Environ Toxicol. 2019;34:1013–24.
Kobayashi Y, Harada N, Nishimura Y, Saito T, Nakamura M, Fujiwara T, et al. Algae sense exact temperatures: small heat shock proteins are expressed at the survival threshold temperature in Cyanidioschyzon merolae and Chlamydomonas reinhardtii. Genome Biol Evol. 2014;6:2731–40.
Sathasivam R, Ki J-S. Heat shock protein genes in the green alga Tetraselmis suecica and their role against redox and non-redox active metals. Eur J Protistol. 2019;69:37–51.
Chankova S, Mitrovska Z, Miteva D, Oleskina YP, Yurina NP. Heat shock protein HSP70B as a marker for genotype resistance to environmental stress in Chlorella species from contrasting habitats. Gene. 2013;516:184–9.
Uji T, Gondaira Y, Fukuda S, Mizuta H, Saga N. Characterization and expression profiles of small heat shock proteins in the marine red alga Pyropia yezoensis. Cell Stress Chaperones. 2019;24:223–33.
Sepúlveda Steiner O, Bouffard D, Wüest A. Persistence of bioconvection‐induced mixed layers in a stratified lake. Limnol Oceanogr. 2021;66:1531–47.
Bushnell B. BBTools software package. 2014;578:579. https://sourceforge.net/projects/bbmap/.
Bankevich A, Nurk S, Antipov D, Gurevich AA, Dvorkin M, Kulikov AS, et al. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J Comput Biol. 2012;19:455–77.
Eren AM, Esen ÖC, Quince C, Vineis JH, Morrison HG, Sogin ML, et al. Anvi’o: an advanced analysis and visualization platform for ‘omics data. PeerJ. 2015;3:e1319.
Langmead B, Salzberg SL. Fast gapped-read alignment with Bowtie 2. Nat Methods. 2012;9:357–9.
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, et al. The sequence alignment/map format and SAMtools. Bioinformatics. 2009;25:2078–9.
Olm MR, Brown CT, Brooks B, Banfield JF. dRep: a tool for fast and accurate genomic comparisons that enables improved genome recovery from metagenomes through de-replication. ISME J. 2017;11:2864–8.
von Meijenfeldt FAB, Arkhipova K, Cambuy DD, Coutinho FH, Dutilh BE. Robust taxonomic classification of uncharted microbial sequences and bins with CAT and BAT. Genome Biol. 2019;20:217.
Hyatt D, Chen G-L, Locascio PF, Land ML, Larimer FW, Hauser LJ. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinform. 2010;11:119.
Buchfink B, Reuter K, Drost H-G. Sensitive protein alignments at tree-of-life scale using DIAMOND. Nat Methods. 2021;18:366–8.
Gish W, States DJ. Identification of protein coding regions by database similarity search. Nat Genet. 1993;3:266–72.
NCBI Resource Coordinators. Database resources of the national center for biotechnology information. Nucleic Acids Res. 2018;46:D8–D13.
Gruber-Vodicka HR, Seah BKB, Pruesse E. phyloFlash: rapid small-subunit rRNA profiling and targeted assembly from metagenomes. mSystems. 2020;5:e00920–20.
Seppey M, Manni M, Zdobnov EM. BUSCO: Assessing Genome Assembly and Annotation Completeness. In: Kollmar M, editor. Gene Prediction. Methods in Molecular Biology. Humana: New York; 2019. vol 1962. pp 227–45.
Waterhouse RM, Seppey M, Simão FA, Manni M, Ioannidis P, Klioutchnikov G, et al. BUSCO applications from quality assessments to gene prediction and phylogenomics. Mol Biol Evol. 2018;35:543–8.
Kriventseva EV, Kuznetsov D, Tegenfeldt F, Manni M, Dias R, Simão FA, et al. OrthoDB v10: sampling the diversity of animal, plant, fungal, protist, bacterial and viral genomes for evolutionary and functional annotations of orthologs. Nucleic Acids Res. 2019;47:D807–D811.
Challis R, Richards E, Rajan J, Cochrane G, Blaxter M. BlobToolKit - interactive quality assessment of genome assemblies. G3. 2020;10:1361–74.
Boratyn GM, Schäffer AA, Agarwala R, Altschul SF, Lipman DJ, Madden TL. Domain enhanced lookup time accelerated BLAST. Biol Direct. 2012;7:12.
Neely CJ, Hu SK, Alexander H, Tully BJ. The high-throughput gene prediction of more than 1,700 eukaryote genomes using the software package EukMetaSanity. 2021. https://www.biorxiv.org/content/10.1101/2021.07.25.453296v1.
Stanke M, Steinkamp R, Waack S, Morgenstern B. AUGUSTUS: a web server for gene finding in eukaryotes. Nucleic Acids Res. 2004;32:W309–12.
Levy Karin E, Mirdita M. Söding J. MetaEuk-sensitive, high-throughput gene discovery, and annotation for large-scale eukaryotic metagenomics. Microbiome. 2020;8:48.
Kuznetsov D, Tegenfeldt F, Manni M, Seppey M, Berkeley M, Kriventseva EV, et al. OrthoDB v11: annotation of orthologs in the widest sampling of organismal diversity. Nucleic Acids Res. 2023;51:D445–51.
Huerta-Cepas J, Szklarczyk D, Heller D, Hernández-Plaza A, Forslund SK, Cook H, et al. eggNOG 5.0: a hierarchical, functionally and phylogenetically annotated orthology resource based on 5090 organisms and 2502 viruses. Nucleic Acids Res. 2019;47:D309–14.
Cantalapiedra CP, Hernández-Plaza A, Letunic I, Bork P, Huerta-Cepas J. eggNOG-mapper v2: Functional annotation, orthology assignments, and domain prediction at the metagenomic scale. Mol Biol Evol. 2021;38:5825–9.
Kanehisa M, Sato Y, Kawashima M, Furumichi M, Tanabe M. KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res. 2016;44:D457–62.
Kanehisa M, Sato Y, Morishima K. BlastKOALA and GhostKOALA: KEGG tools for functional characterization of genome and metagenome sequences. J Mol Biol. 2016;428:726–31.
Kanehisa M, Sato Y, Kawashima M. KEGG mapping tools for uncovering hidden features in biological data. Protein Science. 2022;31:47–53.
Manni M, Berkeley MR, Seppey M, Zdobnov EM. BUSCO: assessing genomic data auality and beyond. Curr Protoc. 2021;1:e323.
Kuraku S, Zmasek CM, Nishimura O, Katoh K. aLeaves facilitates on-demand exploration of metazoan gene family trees on MAFFT sequence alignment server with enhanced interactivity. Nucleic Acids Res. 2013;41:W22–8.
Capella-Gutiérrez S, Silla-Martínez JM, Gabaldón T. trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics. 2009;25:1972–3.
Borowiec ML. AMAS: a fast tool for alignment manipulation and computing of summary statistics. PeerJ. 2016;4:e1660.
Minh BQ, Schmidt HA, Chernomor O, Schrempf D, Woodhams MD, von Haeseler A, et al. IQ-TREE 2: New models and efficient methods for phylogenetic inference in the genomic era. Mol Biol Evol. 2020;37:1530–4.
Huson DH, Scornavacca C. Dendroscope 3: an interactive tool for rooted phylogenetic trees and networks. Syst Biol. 2012;61:1061–7.
Yu G. Using ggtree to visualize data on Tree-like structures. Curr Protoc Bioinform. 2020;69:e96.
Team R. RStudio: integrated development for R. PBC, Boston, MA: RStudio; 2020.
Dierckxsens N, Mardulyn P, Smits G. NOVOPlasty: de novo assembly of organelle genomes from whole genome data. Nucleic Acids Res. 2017;45:e18.
Rozewicki J, Li S, Amada KM, Standley DM, Katoh K. MAFFT-DASH: integrated protein sequence and structural alignment. Nucleic Acids Res. 2019;47:W5–W10.
Talavera G, Castresana J. Improvement of phylogenies after removing divergent and ambiguously aligned blocks from protein sequence alignments. Syst Biol. 2007;56:564–77.
Tamura K, Stecher G, Kumar S. MEGA11: molecular evolutionary genetics analysis Version 11. Mol Biol Evol. 2021;38:3022–7.
Bushnell B. BBMap: A fast, accurate, splice-aware aligner. Berkeley, CA, USA: Lawrence Berkeley National Lab. (LBNL); 2014.
Woodcroft BJ. CoverM: Read coverage calculator for metagenomics. Github. Queensland University of Technology, Brisbane. https://github.com/wwood/CoverM.
Saini JS, Manni M, Hassler C, Cable R, Duhaime MB, Zdobnov EM. Metagenomics dataset from chemocline of Lake Cadagno (version 4) [Data set]. Zenodo. 2023. https://doi.org/10.5281/zenodo.7505505.
Acknowledgements
We thank the Swiss National Science Foundation for funding this project (grant no. PP00P2-138955). We are grateful to the Swiss Confederation, and the Ernst and Lucy Schmidheiny Foundation for their financial support to JSS. We acknowledge the Lake Cadagno team and Alpine Biology Centre Foundation (Switzerland) for providing the platform for metagenomic sampling, Duhaime Lab and Advanced Genomics sequencing core at the University of Michigan (USA) for DNA extraction and shotgun metagenomic sequencing of Lake Cadagno. We thank EZ lab members and Silas Kieser for their helpful discussions and constructive feedback in developing this work. The University of Geneva and the University of Michigan provided computing resources for metagenomic analyses. We acknowledge Christof Holliger’s lab for supporting the revision of this work. In the end, thanks to the reviewers and editors for the constructive criticism essential for improving this work.
Funding
Open access funding provided by University of Geneva.
Author information
Authors and Affiliations
Contributions
CH and MD contributed to the conceptualization, with JS conceiving the study. CH, RC, and JS performed the Lake Cadagno sample collection. MD, RC, and JS contributed to the DNA extractions and sequencing of the metagenomics data. JS and MM conducted the bioinformatics analysis with EZ supervising the project. JS drafted the manuscript, with all coauthors contributing to review and editing.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Saini, J.S., Manni, M., Hassler, C. et al. Genomic insights into the coupling of a Chlorella-like microeukaryote and sulfur bacteria in the chemocline of permanently stratified Lake Cadagno. ISME J 17, 903–915 (2023). https://doi.org/10.1038/s41396-023-01396-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-023-01396-y