Abstract
Prochlorococcus and Synechococcus are the most abundant photosynthesizing organisms in the oceans. Gene content variation among picocyanobacterial populations in separate ocean basins often mirrors the selective pressures imposed by the region’s distinct biogeochemistry. By pairing genomic datasets with trace metal concentrations from across the global ocean, we show that the genomic capacity for siderophore-mediated iron uptake is widespread in Synechococcus and low-light adapted Prochlorococcus populations from deep chlorophyll maximum layers of iron-depleted regions of the oligotrophic Pacific and S. Atlantic oceans: Prochlorococcus siderophore consumers were absent in the N. Atlantic ocean (higher new iron flux) but constituted up to half of all Prochlorococcus genomes from metagenomes in the N. Pacific (lower new iron flux). Picocyanobacterial siderophore consumers, like many other bacteria with this trait, also lack siderophore biosynthesis genes indicating that they scavenge exogenous siderophores from seawater. Statistical modeling suggests that the capacity for siderophore uptake is endemic to remote ocean regions where atmospheric iron fluxes are the smallest, especially at deep chlorophyll maximum and primary nitrite maximum layers. We argue that abundant siderophore consumers at these two common oceanographic features could be a symptom of wider community iron stress, consistent with prior hypotheses. Our results provide a clear example of iron as a selective force driving the evolution of marine picocyanobacteria.
Similar content being viewed by others
Introduction
Prochlorococcus and its sister lineage Synechococcus are some of the smallest known photosynthetic organisms and are among the most numerically abundant life forms on the planet. These marine picocyanobacteria are unicellular, free-living, geographically widespread, and highly abundant in the oligotrophic subtropical/tropical ocean, often comprising half of the total chlorophyll [1]. Prochlorococcus and Synechococcus account for approximately 25% of global marine net primary productivity [2], making the picocyanobacteria key drivers of marine biogeochemical cycles [3]. Light gradients drive the vertical distribution of Prochlorococcus with low-light (LL) adapted clades occupying the deeper parts of the euphotic zone, and high-light (HL) adapted clades near the surface. This broad light-driven ecological and evolutionary division is further partitioned into mostly coherent genomic clusters (clades), each with distinct ecological and physiological properties [4]. Synechococcus clades partition along temperature gradients in the sea, but lack clear association with light and depth [5]. At the finest scale of diversity, picocyanobacterial clades further separate into distinct sympatric subpopulations, which share the majority of their genes but also contain segments of unique genetic material [6]. These variable genomic regions encode functionally adaptive traits that tune each population’s physiology to its local environment.
Iron (Fe) is a crucial micronutrient for marine phytoplankton due to its central role as an enzyme cofactor in cellular processes, including respiration and photosynthesis. As a result, the concentrations and chemical forms of Fe influence global carbon cycle dynamics [7]. Dissolved Fe (dFe) is scarce in much of the ocean and is mostly (>99%) complexed with organic chelating ligands that solubilize and stabilize the ions in solution [8]. Concentrations of these ligands quickly increase in response to Fe fertilization events, which implies that they are actively produced by members of the microbial community [9]. It is challenging to determine the chemical structure of these ligands [10], so their abundance and stability coefficients (a measure of how “strongly” the ligand binds Fe) are typically inferred electrochemically [11]. Many electrochemical studies operationally partition the ligand pool into weak (L2) and strong (L1) binding classes. The weak L2 class includes humic-like substances, exopolysaccharides, and undefined colloids. The strong L1 class includes siderophores, small Fe-binding molecules that microbes produce during periods of Fe starvation, but it is not clear how much of the L1 class are genuine siderophores. Under some conditions, siderophores appear to account for much of the strong L1 ligand fraction in seawater [12, 13].
There are two primary mechanisms by which marine microbes extract dFe bound to the organic ligands in the ocean. First, dFe can dissociate from organic ligands in the extracellular environment (via kinetic control, photodegradation, or cell surface reductases) and is imported across the outer membrane as an unbound, inorganic ion [14]. Second, whole Fe-ligand complexes can be directly translocated across cell membranes (Supplementary Fig. S1) [15]. Direct uptake pathways are prevalent in fast-growing copiotrophic marine bacteria with large genomes but absent in free-living marine bacteria with streamlined genomes such as Prochlorococcus, Synechococcus, and SAR11 [16, 17]. In these organisms, selection favors the minimization of genome size and metabolic complexity over the versatility of maintaining multiple direct Fe uptake pathways [16]. A decade ago, it was believed that marine picocyanobacteria fulfilled their Fe requirements only via the dissociation mechanism while relying upon a single inner membrane Fe(III) ATP binding cassette transporter [18, 19]. Prior work also showed that Prochlorococcus and marine Synechococcus isolate genomes lacked the genes necessary for siderophore biosynthesis and uptake [19]. This image changed when putative siderophore uptake gene clusters were identified from a handful of genomes from Prochlorococcus surface clades HLII and HLIV collected from remote, low-Fe regions of the global ocean [17, 20]. This exciting finding suggested that some Prochlorococcus populations had adapted to Fe scarcity by supplementing the uptake of dissociated dFe ions with the direct uptake of siderophore-bound Fe.
Siderophore uptake genes from cells belonging to HL adapted Prochlorococcus clades have been shown to be most abundant in surface waters that have low modeled Fe concentrations [21, 22]. However, the extent of siderophore uptake potential in cells belonging to LL-adapted clades, which are uniquely adapted to waters deep in the euphotic zone, is unknown. This is important because interactions between Fe and light limitation or Fe-light co-limitation may increase Fe demand for Prochlorococcus clades inhabiting deeper waters and the deep chlorophyll maximum layer (DCM) [23, 24]. Also of interest is understanding what environmental features, in addition to Fe, are associated with picocyanobacterial siderophore consumers. Here, we survey an extensive data set of picocyanobacterial uncultivated single-cell and cultivated isolate genomes to obtain a general picture of the phylogenetic and biogeographic structure of picocyanobacterial siderophore traits. Next, we combine biogeochemical and metagenomic time-series datasets from the Hawai’i Ocean Time-series (HOT) station ALOHA and the Bermuda Atlantic Time-series (BATS) station BATS [25] to more deeply understand seasonal dynamics and the depth distributions of Prochlorococcus siderophore traits in a community context. Finally, we quantify the Prochlorococcus and Synechococcus siderophore traits in 645 metagenomic samples from Tara Oceans [26] and GEOTRACES [25] and connect the distribution of the trait to thousands of trace metal and other biogeochemical measurements from across the world’s oceans. Using these rich data, we identify environmental features associated with picocyanobacterial siderophore uptake and reveal that siderophore uptake is predominantly a feature of low-light adapted LLI clade Prochlorococcus residing near DCM layers from the lower limits of the euphotic zone. These findings reveal new regions of phytoplankton Fe stress in the global ocean and underscore light and Fe availability as key features shaping the evolution of Prochlorococcus and Synechococcus.
Materials and methods
Data sources
We used a collection of over 700 Prochlorococcus and Synechococcus single-cell and isolate genomes collected from across the global ocean [27, 28], 195 surface and DCM metagenomes from Tara Oceans project [29], 480 metagenomes acquired from GEOTRACES cruises [25], and 133 metagenomes from the HOT and BATS time-series [25]. Trace metal and other chemical concentrations are from the GEOTRACES Intermediate Data Product IDP2017 version 2 (accessed January 2019) [30]. Samples were from sections GA02 [31, 32], GA03, GA10 [33], and GP13. Biogeochemical data from the Tara Oceans project was obtained from https://doi.pangaea.de/10.1594/PANGAEA.875579. Modeled climatological dFe from the MIT Darwin model v0.1_llc90 was obtained from the Simons Collaborative Marine Atlas Project (CMAP) [34]. Modeled dFe was averaged over 12 months at a 0.5 degree grid in 5 m depth bins from the upper 250 meters. Metagenome sequence data and associated metagenomes with environmental variables were quality controlled as described earlier [35, 36]. See supplementary material for details.
Prochlorococcus cell concentrations (qPCR-calibrated) from HOT and BATS [37] were obtained from the Biological and Chemical Oceanography Data Management Office https://www.bco-dmo.org/dataset/3381. L1 organic Fe-binding ligand data are from previous studies at HOT and BATS [12, 38, 39]. Data for BATS was derived from the occupation of BATS (Station 12) during the U.S. GEOTRACES GA03 cruise [39]. Siderophore concentrations (ferrioxamine E and G) from surface waters of the North Atlantic at stations 41–62 (data not available from the DCM) and the surface and DCM at HOT are from previous studies [12, 40]. In these samples, the DCM was defined as the depth range of maximum chlorophyll fluorescence at HOT (100–125 m) and BATS (90–135 m). The surface was defined as all depths shallower than the DCM.
Measurements of Fe-binding ligands and siderophores
We analyzed organic Fe-binding ligand data from BATS and HOT using competitive ligand exchange adsorptive cathodic stripping voltammetry (CLE-ACSV) as described previously [12, 38, 41]. We measured ferrioxamine E and G siderophore concentrations using liquid chromatography (LC) coupled to inductively coupled plasma mass spectrometry (ICP-MS) after pre-concentration via solid-phase extraction [12, 13, 42]. DCM samples were from a depth range of 70–200 meters, and surface samples were within a depth range of 3–45 meters.
Comparative genomics
The steps for producing the genome phylogeny in Fig. 1 are detailed in a prior publication [36]. All Prochlorococcus (n = 663) and Synechococcus (n = 96) genomes were annotated against eggNOG 4.5.1 [43] using eggNOG-Mapper v1.0.3–3-g3e22728 [44]. Siderophore transport gene clusters were identified preliminarily using best hits to COGs and PFAMs for the outer membrane TonB dependent receptor (Pfam ID: PF00593, COG ID: COG4771). This gene was used as an anchor gene for examining adjacent gene neighborhoods (Supplementary Fig. S1). Genomes were considered to be authentic siderophore consumers if at least five of the following families were detected adjacent to the outer membrane receptor or if the contig was interrupted: ATP binding cassette (ABC) solute binding protein (Pfam ID: PF01497, COG ID: COG0614), ABC permease (Pfam ID: PF01032, COG ID: COG0609), ABC ATPase (Pfam ID: PF00005, COG ID: COG1120), TonB protein (Pfam ID: PF03544, COG ID: COG0810), and ExbB (Pfam ID: PF01618, COG ID: COG0811), and ExbD (Pfam ID: PF02472). All Prochlorococcus genomes were surveyed for siderophore biosynthesis potential using antiSMASH v5.0 [45], which searches for known nonribosomal peptide synthetases and polyketide synthase siderophore biosynthesis pathways. The true proportion of genomes with the siderophore transport cluster was estimated by accounting for genome incompleteness in the single-cell genomes as described earlier [35] and in the supplementary material.
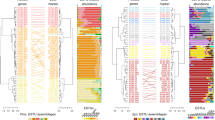
A Sampling locations of picocyanobacteria isolate and single-cell genomes used for this study. Point size is proportional to the total number of isolated genomes from each location, and point color shows whether a siderophore consumer genome was isolated at that location. Map color displays the climatological annual mean total dissolved Fe concentration (nmol kg−1) integrated over the upper 55 meters of the water column. Oceanographic stations ALOHA and BATS are denoted for reference. B Siderophore uptake potential across picocyanobacterial clades. The phylogenetic tree was built with 96 Synechococcus genomes and 605 Prochlorococcus genomes and is rooted at Synechococcus sp. WH5701; nodes show bootstrap values (250 bootstraps). Triangles depict monophyletic clades, and the triangle area is proportional to the number of siderophore consumer genomes in each clade.
Metagenomic read classification and count processing
Metagenome reads were mapped to the MARMICRODB marine microbial database using Kaiju v1.6.0 [46] as described before [36]. Reads mapping best to Prochlorococcus and Synechococcus TonB dependent outer membrane receptors or single-copy, core gene families from Prochlorococcus and Synechococcus were retained for further analysis. The TonB dependent receptor was selected as an indicator for the presence of the entire siderophore uptake cluster because it is the longest gene in the cluster and because its sequence composition is conserved and distinct in Prochlorococcus and Synechococcus (Supplementary Fig. S2). There was no clear similarity cutoff between metagenomic reads and the receptor sequence that differentiated Prochlorococcus or Synechococcus clades, and we did not attempt to resolve individual clades by the TonB receptor (Supplementary Fig. S2). Receptor and core gene counts were length-normalized to the median length of the corresponding gene, then divided by the length-normalized median abundance of all Prochlorococcus or Synechococcus core genes. Samples where the median core gene coverage was less than 100 were excluded. See supplementary material for details.
Statistics and data analysis
Random forest regression was performed using the normalized Prochlorococcus TonB dependent receptor abundance as the response variable and a collection of 45 potential explanatory variables with the R package Ranger v0.12.1 [47]. Hyperparameters were tuned and the model trained using nested ten-fold cross-validation, reserving 20% of the data for estimating final model performance. Signal extraction using principal component analysis (PCA) was performed on scaled and centered predictors [48] in cross-validation. All principal components (PCs) cumulatively explaining 99% of the variance in at least one training partition were retained (27 PCs total). Signal extraction was performed to ensure that model predictors were statistically independent and to circumvent the significant inter-variable correlations in the environmental data. PC importance rankings were determined in each training partition using the Boruta heuristic [49], resulting in all predictor PCs consistently performing better than random. This step buffers against overfitting the model while assessing variable importance relative to baseline importance (i.e., noise). Boruta rankings were aggregated over the training partitions, resulting in a total of 20 PCs that, in combination, were better predictors of siderophore abundance than randomly generated data. Cumulatively, these PCs explained 97% of the variance in the original data.
After training and testing the final random forest, the original predictor variables were related to PCs using PCA loadings by taking the square of the eigenvector matrix to get the variance explained (R2) by each variable for each PC. All variables with an R2 ≥ 0.1 with at least one of the 20 informative PCs were retained (n = 26). These variables were then ordered by their rank contribution to predicting the distribution of Prochlorococcus siderophore consumers. The rank contribution was estimated as the sum of the variance explained by each variable across all 20 informative PCs, with each PC weighted by the relative importance obtained from the feature importance algorithm. Pearson’s correlation was used to measure the strength of the linear relationship between siderophore relative abundance and each predictor variable.
Beta regression was used to model the relationship between Prochlorococcus siderophore transporter relative abundance, nitrite, and DCM depth, while allowing the effect of nitrite to vary as a function of DCM depth. This model was selected because light-limited phytoplankton generate nitrite [50] and deeper DCMs are, presumably, more light-limited than shallower DCMs [51]. Continuous covariates were transformed to categorical covariates by binning into three roughly equally sized groups: 3.3e–4 < Lo NO2 ≤ 4.0e–2; 4.0e-2 < Md NO2 ≤ 7.3e-2; 7.3e-2 < Hi NO2 ≤ 1.4. The emmeans v1.6.0 R package was used to estimate the magnitude and statistical significance of marginal means for each covariate in the model. The seasonal effects in time-series metagenomes were estimated using Generalized Additive Mixed Models and Linear Mixed-Effect Models with the mgcv v1.8–26 and nlme v3.1–148 libraries in R as described in the supplementary material.
Results and discussion
Siderophore uptake potential in picocyanobacterial genomes
We searched a collection of over 700 marine Prochlorococcus and Synechococcus genomes (including cultivated isolate genomes and uncultivated single-cell genomes) for siderophore biosynthesis and siderophore uptake gene clusters. These genomes are from over 40 distinct geographic locations across the world’s oceans (Fig. 1A, Supplementary Fig. S3), including the well-studied oceanographic stations ALOHA and BATS, and span the breadth of Prochlorococcus and Synechococcus phylogenetic diversity (Fig. 1B). Samples were collected throughout the euphotic zone at depths ranging from five meters to over 200 meters depth. In agreement with previous studies, none of the Prochlorococcus and marine Synechococcus genomes contained siderophore biosynthesis gene clusters [19, 52]. However, we identified 47 genomes with siderophore transport gene clusters (Supplementary Fig. S1) - hereafter siderophore consumers - and they were all either found in, or isolated from remote regions spanning the oligotrophic N. and S. Pacific Ocean gyres (Fig. 1A). Synechococcus siderophore consumers were from the North Pacific subtropical gyre (Supplementary Fig. S3) and polar frontal regions between 28 °N and 37 °N [27], where surface dFe concentrations are typically low. The restriction of picocyanobacterial siderophore consumers to the Pacific is analogous to the strong ocean basin segregation of genomic adaptations to nitrogen and phosphorus limitation in Prochlorococcus [53,54,55]. Because the Pacific Ocean encompasses over 70% of Fe-limited ocean regions [56] and siderophore uptake is a form of Fe scavenging [57], together these findings suggest that Fe scarcity is an important selective pressure on picocyanobacterial genome content in the oligotrophic Pacific ocean.
We next examined the distribution of the siderophore uptake trait among clades within the marine picocyanobacterial phylogeny. The trait is distributed unevenly among different clades of Prochlorococcus: 2% and 14% of the genomes from HLII and HLI clades of Prochlorococcus, respectively, contained these clusters, while 31% of those from LLI Prochlorococcus contained them (Fig. 1B, Supplementary Fig. S1). The relative frequency of the siderophore trait increased within Prochlorococcus ecotypes with lower temperature and light level optima: for example, HLI (14%, N = 89) and LLI (31%, N = 84) [4]. We also identified the siderophore transport gene cluster within genomes from Synechococcus clade 5.1 A (37%, N = 42) in subclades II, III, IV, UC-B, and CRD2, with subclades IV and CRD2 accounting for 70% of all Synechococcus siderophore consumers. Although members of Synechococcus clades do not stratify with depth like Prochlorococcus clades, some members of clade 5.1 A display low-light adapted phenotypes [58], and subclades III and I (whose distribution follows clade IV) in particular are optimized for growth at low irradiance [58].
One caveat is that the total number of picocyanobacterial genomes and the clade composition between ocean basins were different, so the observed gene frequencies are likely biased to some extent. Furthermore, many sampling sites were represented by only a handful of genomes, which is too small of a sample size to reliably assess whether a trait is absent in the sampled population. Still, these findings from complete and partial genomes clearly hint towards genuine trends of higher picocyanobacterial siderophore consumer frequency in the pacific ocean and in clades that thrive deep in the euphotic zone.
Previous studies of siderophore uptake genes in Prochlorococcus have been limited to Fe-limited surface waters which are dominated by cells belonging to HL adapted clades [20,21,22]. Our finding that potential siderophore uptake is most prevalent within the LLI clade (Fig. 1B) implies that picocyanobacterial siderophore use is most likely to be associated with low-light conditions. LLI Prochlorococcus is the dominant clade at DCM layers, which form near the base of the euphotic zone and are shaped by a balance between diminished light and enhanced macronutrients (N, P, Si) from deep waters. Furthermore, Fe-limitation or Fe/light co-limitation can be a persistent feature of this layer [23]. Phytoplankton Fe-limitation at the DCM may emerge due to the upregulation of the Fe-rich photosynthetic apparatus under low-light [59], which may increase Fe demand relative to supply [60] and thus low-light Prochlorococcus clades likely require more Fe than high-light clades [24]. Indeed, Prochlorococcus has relatively high photosynthetic Fe requirements under low irradiance, and there is evidence that LLI Prochlorococcus, in particular, is uniquely adapted to the low-Fe and low-light conditions typical of the DCM [24]. In some cases, the high Fe requirements of LLI Prochlorococcus may make them more sensitive to fluctuations in Fe concentration than eukaryotic phytoplankton [61].
Furthermore, low irradiance at the DCM should suppress the photodegradation of siderophores and the release of free dFe [62], which implies that direct siderophore uptake traits may provide higher relative fitness for low-light adapted Prochlorococcus. Thus, competition for inorganic dFe ions at the DCM is intense, and the ability to utilize siderophores may allow Prochlorococcus to reduce relative competition for Fe through niche differentiation.
Spatial and temporal patterns at two long-term ocean study sites
We further explored the Pacific/Atlantic divide and prevalence of the uptake trait in picocyanobacteria genomes by examining metagenomes from the surface, the DCM, and the bottom of the euphotic zone at Stations ALOHA and BATS. We focused on the distributions of Prochlorococcus because Synechococcus genomes did not recruit enough reads at ALOHA for a robust comparison between the two oceans. Indeed, Prochlorococcus is generally two orders of magnitude more abundant than Synechococcus at ALOHA [63]. ALOHA is located in the N. Pacific subtropical gyre [64], while BATS is located in the N. Atlantic subtropical gyre. Both stations are oligotrophic, have comparable net primary productivity and carbon export [65], and are numerically dominated by Prochlorococcus and SAR11 [66]. However, atmospheric iron fluxes are significantly higher at BATS compared with ALOHA [67]. Siderophores, strong Fe-binding ligands (L1), and dFe concentrations have been measured in many studies at these locations [12, 38,39,40]. This wealth of data allowed us to examine relationships between siderophore consumers and Fe-binding ligands, total dFe, and siderophore concentrations on seasonal time scales.
Prochlorococcus siderophore consumers were absent at BATS but constituted up to half of all Prochlorococcus genomes from DCM metagenomes at ALOHA (Fig. 2A), where Prochlrocococus is typically responsible for the majority of productivity, biomass, and chlorophyll concentration [68]. We estimated the clade-integrated proportion of siderophore consumers within the total Prochlorococcus population because we could not identify one sequence similarity threshold for short metagenomic reads that distinguished Prochlorococcus outer membrane siderophore receptors by clade (Supplementary Fig. S2, see methods). The LLI clade dominates the DCM at both ALOHA and BATS [37] and has the highest frequency of single-cell genomes with siderophore transporters. Likewise, the frequency of siderophore consumers in the total Prochlorococcus community at the DCM (i.e., from metagenomes) was strongly positively correlated with the relative abundance of the LLI clade (Fig. 2A). Siderophore consumers also followed a seasonal pattern at the ALOHA DCM, coinciding with peaks in LLI abundance in late summer and early fall (Fig. 2B, Supplementary Fig. S4). This pattern was not present at BATS, where the trait was effectively absent. The annual spring/winter Asian dust flux is the dominant new Fe source to the N. Pacific subtropical gyre [38, 69], and this annual deposition event coincides with the lowest abundance of siderophore consumers at the ALOHA DCM (Fig. 2B). Seasonal variations in Fe supply may contribute to the seasonal succession of siderophore consumers and Prochlorococcus clades [37] at the DCM in the subtropical N. Pacific.
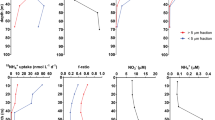
A Prochlorococcus HLII and LLI cell density over 2 years at HOT and BATS as determined by qPCR [37]. Contour plots of cell concentrations are cube root transformed. The deep chlorophyll maximum (DCM) depth range is highlighted in red. fsidero. consumer is the fraction of Prochlorococcus siderophore consumers, and point size is proportional to relative abundance. B Modeled seasonality of of siderophore consumers, LLI, and HLII Prochlorococcus clade abundance at the DCM at stations BATS and ALOHA. Seasonality is represented with a smoother function in a Generalized Additive Model (see methods). Periods with a positive seasonal effect can be interpreted as having higher abundance than the annual mean. Bands are 95% confidence intervals of the smooth function fit. Only terms with a significant seasonal effect smoother (P < 0.05) are shown. C dFe, L1 strong Fe-binding ligands, and summed ferrioxamine E and G siderophore concentrations in the N. Pacific and N. Atlantic subtropical gyres. Points and error bars are marginal population means and 95% confidence intervals from regression. Red asterisk indicates p < 0.05. Full model results are in Supplementary Table S1.
We also found that biogeochemical patterns of dFe and Fe-binding ligands were inversely related to patterns of siderophore consumer abundance at stations ALOHA and BATS (Fig. 2C, Supplementary Table S1). Station ALOHA had the lowest total dFe and the lowest L1 ligand concentrations. The significant enrichment of siderophore consumers at the ALOHA DCM relative to the surface also coincided with a sharp decrease in total L1 concentration, potentially due to biological demand and uptake. However, measurements of dFe and dFe-binding ligand concentrations are positively correlated [70], and this correlation may be, in part, an artifact of the analytical methods used to estimate complexometric titrations parameters in seawater [71]. Thus, the local L1 minimum at the ALOHA DCM cannot be unambiguously attributed to biological uptake. Furthermore, L1 does not represent the concentration of any particular siderophore molecule but is an aggregate metric for the Fe-binding organic matter in the sample. Ferrioxamine siderophores showed no systematic concentration difference between the Atlantic and Pacific nor between the DCM and surface at ALOHA (Fig. 2C), which suggests that ferrioxamine siderophore production may be similar in both oceans. Thus, there is a potential for differences in the chemistry of iron-binding ligands at the DCMs of BATS and ALOHA, but the pattern of Prochlorococcus siderophore consumers most strongly follows differences in total dFe concentrations between the DCM at BATS and ALOHA. These concentration differences reflect higher atmospheric Fe fluxes in the Atlantic [72], as well as differences in dFe flux from below the DCM. For example, there is a weak ferric line below the DCM at ALOHA [38], while the dFe gradient below the DCM at BATS is comparatively strong [73], which implies a higher upwards flux at BATS (Supplementary Fig. S5).
Global patterns of siderophore uptake genes
We next examined the abundance of Prochlorococcus and Synechococcus siderophore consumers in ocean areas not covered by the genomic and time-series metagenomic datasets by quantifying the abundance of picocyanobacterial siderophore consumers in metagenomes from the upper 300 meters of the global tropical and subtropical ocean. In general, results from the global metagenomic data set reinforced our findings from the time-series metagenomes for Prochlorococcus (Fig. 2) and the single-cell and isolate picocyanobacterial genomes (Fig. 1, Supplementary Fig. S3). Collectively, the global metagenomes revealed: (1) a high frequency of Prochlorococcus siderophore consumers in the subtropical and tropical Pacific relative to the N. Atlantic Ocean, and (2) high frequencies of Prochlorococcus siderophore consumers at the DCM - especially oligotrophic DCMs from the Pacific Ocean from depths over 100 meters (Fig. 3A). Synechococcus consumers generally followed a similar trend to Prochlorococcus with highest abundance in the Pacific ocean and at the DCM. However, there was generally low total abundance of total Synechococcus at many sites from the interior of the gyres - e.g., GEOTRACES line GP13 (Fig. 3B). Both Prochlorococcus and Synechococcus siderophore consumers were also particularly abundant in the eastern South Pacific near known oxygen minimum zones. All samples used in our study were collected from fully oxygenated seawater and little variance in our analysis was explained by oxygen concentrations (Fig. 4, Supplementary Fig. S6). However, prokaryotic biomass in oxygen minimum zones has elevated trace metal signatures which may indirectly influence Fe cycling above the oxycline [74].
Fraction of (A) Prochlorococcus genomes and (B) Synechococcus genomes with siderophore uptake gene clusters (fsidero. consumer) at the DCM in the GEOTRACES (circles) and Tara Oceans (triangles) metagenome dataset. Samples where median core gene coverage was less than 100 are denoted by X. In (A) samples from the GP13 section (S. Pacific) are expanded for increased visibility. Point color and size are proportional to abundance.
Environmental variables (heatmap left) are colored by variable type and ordered (heatmap right) by their rank contribution to predicting the distribution of Prochlorococcus siderophore consumers (see methods). Top barplot shows the informative principal components derived from the combined dataset and ranked by their importance (unitless) to the random forest model. The lower barplot shows the variance explained (%) by each of the informative principal components. Heatmap color shows the correlation (R2) between each variable and each informative principal component.
In most Pacific DCM samples, over half of Prochlorococcus cells could potentially use siderophores (assuming the gene cluster is single-copy), which implies that a substantial fraction of Prochlorococcus Fe-demand at the DCM may be fulfilled by siderophores. Unexpectedly, we observed a high abundance of Prochlorococcus siderophore consumers in the S. Atlantic subtropical gyre between 5 °S and 25 °S (Fig. 3A), where the predicted climatological mean of dFe is relatively high (Fig. 1A). Furthermore, Synechococcus consumers were highly abundant in samples south of 40 °S in the S. Atlantic gyre (Fig. 3B), which is at the limit of Prochlorococcus’ latitudinal range [2]. Over 40% of Prochlorococcus genomes at the South Atlantic ocean DCM between 5 °S and 25 °S were putative siderophore consumers, consistent with recent studies suggesting Fe-deficiency and Fe-N co-limitation in this region [75, 76]. Indeed, there is a persistent local minimum in dFe concentrations within the euphotic zone between 5 °S and 25 °S on GEOTRACES transect GA02 [32], and the thermocline waters (approximately 200–400 meters depth) of this region are Fe-deficient relative to macronutrients [77]. Most biogeochemical models predict community N-limitation in the S. Atlantic gyre [78, 79], but there are an increasing number of studies demonstrating Fe-limitation or Fe-macronutrient co-limitation in phytoplankton communities from this region [75, 76]. Like the equatorial Pacific, primary productivity in the S. Atlantic gyre may be primarily sustained by rapid and efficient internal Fe recycling due to low new Fe inputs [80, 81].
The finding of abundant Prochlorococcus and Synechococcus siderophore consumers at the DCM and in the S. Atlantic subtropical gyre led us to ask what specific chemical, biological, and hydrographic mechanisms might influence these distributions. We focused on the distribution of Prochlorococcus siderophore consumers because they were more abundant with a wider geographic distribution than Synechococcus consumers (Fig. 3). We trained a random forest regression for Prochlorococcus siderophore consumer abundance using a set of 27 predictive features derived from PCA on a 45 variable dataset of hydrographic, geochemical, and biological measurements (see methods, supplementary material). We then ranked the original variables by their weighted contribution to explaining the variance in the 20 informative PCs (Fig. 4). The final model had excellent predictive performance in the test dataset (R2 = 0.93, RMSE = 0.00017) indicating that the abundance of Prochlorococcus siderophore consumers can accurately be predicted from GEOTRACES and Tara Oceans biogeochemical data alone. Prochlorococcus siderophore consumers covaried with the biogeochemical parameters that distinguish the subtropical/tropical N. Atlantic ocean, the Mediterranean Sea, and the Red Sea from the rest of the ocean. The N. Atlantic, Mediterranean Sea, and the Red Sea have the highest salinity [82] and receive a large flux of atmospherically deposited material due to their proximity to the Sahara desert and the Arabian peninsula [67]. Correspondingly, salinity and atmospherically deposited trace elements - such as Cu, Al, Zn, Mn, Fe, and Pb [72] - were strongly negatively associated with Prochlorococcus siderophore consumers (Figs. 4, 5, and Supplementary Fig. S6). Siderophore transporters were highly abundant in equatorial Pacific samples dominated by the Prochlorococcus HLIV clade (>60% HLIV), which has been shown previously to have siderophore transport genes [20] and is associated with Fe-limited ocean regions [83]. Overall, our findings reveal that picocyanobacterial siderophore uptake is common in the remote reaches of the subtropical/tropical oligotrophic ocean, where the atmospheric input fluxes of trace elements to the upper ocean are the smallest.
These ten environmental variables (ranked 1–10) have the greatest cumulative predictive contribution to the random forest model (Fig. 4, see methods). Each point is a metagenomic observation colored by ocean basin. Small squares represent samples with missing data that was imputed for each variable (see methods). dFe climatological means are from the MIT Darwin model. The remaining measurements are in situ chemical, biological, or hydrographic measurements from GEOTRACES and Tara Oceans. Boxes show Pearson correlation coefficients (r) and the p value testing whether true r is equal to 0.
Of all trace metals in the global dataset, dFe measured from GEOTRACES samples explained most of the weighted variance across the principal component predictors used in the random forest. However, the magnitude of its linear correlation with siderophore consumers (r = −0.10) was smaller than for other metals like Al (r = −0.28) or Cu (r = −0.28) (Fig. 5). Furthermore, dFe concentrations from the Darwin biogeochemical model were more strongly correlated with Prochlorococcus siderophore consumers (r = −0.54) than for dFe or any other trace metal measurement from the GEOTRACES program. Although dFe had the weakest linear correlation with siderophore consumers, the dFe variance was partitioned across multiple, strongly predictive principal components (Fig. 4). This implies that dFe’s predictive power in the random forest was primarily derived from its nonlinear associations and interactions with other environmental variables (e.g., LLI abundance, DCM depth, nitrite, and salinity) and not its linear correlation with siderophore consumers. This finding is compatible with current views of marine Fe biogeochemistry: much of the upper ocean dFe inventory is cycled rapidly and shaped by multiple interacting biotic and abiotic biogeochemical processes [7].
The strong linear correlation between modeled climatological dFe concentrations and Prochlorococcus siderophore consumers is likely because atmospheric dust deposition is the dominant Fe source to the ocean in the MIT Darwin model [84]. This is consistent with the strong negative linear relationship we observe between siderophore consumers and atmospherically deposited elements (e.g., aluminum) in the GEOTRACES data and the strong partitioning of Prochlorococcus siderophore consumers to the oligotrophic Pacific and S. Atlantic. We argue that the global biogeography of Prochlorococcus siderophore consumers is driven by basin-scale geological forcing in the marine Fe cycle [72, 85]. Specifically, it appears that patterns of atmospheric dust deposition set the biogeographical boundary for where the relative fitness benefit of siderophore use exceeds that for reduced genome size and gene loss [86]. On a local scale, the fraction of a Prochlorococcus population that can use siderophores will also reflect a combination of physiological and ecological processes (e.g., light availability), which affects local dynamic biogeochemical processes like regeneration, scavenging, uptake, and colloid formation. In this way, the abundance of Prochlorococcus siderophore consumers reflects a balance of both the “fast” and “slow” biogeochemical processes shaping the marine Fe cycle.
Surprisingly, the random forest regression also showed a strong association between Prochlorococcus siderophore consumers, nitrite concentrations, and DCM depth. We explored this relationship further using linear regression (Fig. 6, Supplementary Table S2). The results were most consistent with a scenario where Prochlorococcus siderophore consumers are abundant at depths with the highest nitrite concentrations, especially where the DCM layer is deeper than 100 meters and to a lesser degree where LLI cells are most abundant. We propose that this pattern is due to phytoplankton Fe and light limitation (or co-limitation) at the DCM and the primary nitrite maximum layer. The origin of the primary nitrite maximum is debated, but it likely forms due to incomplete assimilatory nitrate reduction by Fe- or light-limited phytoplankton [51] in parallel with uncoupled chemoautotrophic nitrification [87]. Due to fast phytoplankton uptake kinetics, nitrite should only accumulate in waters where phytoplankton inorganic nitrogen uptake is limited by Fe and light [87]. Nearly all LLI cells can assimilate nitrite and some can also assimilate nitrate [55], but we found no evidence that nitrate assimilation genes were more or less abundant in LLI siderophore consumers than expected by chance. Overall, the significant positive association of Prochlorococcus siderophore consumers with high nitrite samples - especially from the deepest DCMs - provides indirect support for the hypothesis that the primary nitrite maximum layer in the euphotic subtropical/tropical ocean is associated with Fe and light limitation of photoautotrophic cells.
Each point is a direct metagenomic observation. Subplots show different DCM depth ranges. Points and error bars show estimated marginal population means and 95% confidence intervals from beta regression using nitrite concentration and DCM depth as model covariates. These continuous covariates were transformed into categorical covariates for regression by binning into three equal-sized groups (see methods).
Synthesis
High frequency dynamics in the organic Fe-binding ligand pool of the upper ocean balance the interplay between regeneration and removal fluxes which determines the global inventory of oceanic Fe [88]. Siderophores are increasingly understood to be an important molecular mechanism regenerating Fe from biomass and incorporating atmospherically deposited lithogenic Fe into microbial ecosystems [57]. Many copiotrophic marine bacteria appear to use siderophores, but nearly all genome-streamlined oligotrophic marine bacteria do not - likely to minimize overall metabolic complexity [16, 17]. This raises the question of how minimalist cells like Prochlorococcus fulfill their Fe requirements, especially in the most Fe-limited regions.
Using rich genomic and metagenome data sets, we found that large populations of Prochlorococcus and Synechococcus from remote regions of the global ocean have evolved the ability to scavenge exogenous siderophore-bound Fe, which likely helps them fulfill cellular iron demand in limiting or stressful conditions. The Prochlorococcus siderophore uptake trait is prominent in ocean regions where the atmospheric Fe flux to the surface ocean is lowest (Fig. 7). In these regions, Fe recycling mediated by organic ligands fuels a substantial portion of primary productivity [80]. Here siderophores are likely a critical molecular shuttle between particulate, colloidal, and dissolved phases, ultimately retaining Fe in the upper ocean (Fig. 7). The siderophore uptake trait is most abundant in picocyanobacteria inhabiting remote Fe-depleted regions and in low-light adapted Prochlorococcus, particularly those at deep DCMs near the primary nitrite maximum layer. The vertical distribution of this trait in the water column implies that light availability is also an important control on Prochlorococcus Fe demand. Indeed, the association of siderophore consumers with the highest nitrite concentrations suggests that nitrite accumulation in the euphotic zone may be a consequence of both iron and light limitation.
Conceptual diagram depicting differences in Prochlorococcus siderophore use between the Pacific and Atlantic oceans as exemplified by (A) station ALOHA and (B) station BATS. The total abundance of Prochlorococcus is proportional to the number of points at each depth, while the color represents high-light and low-light adaptation. DCM Deep chlorophyll maximum layer.
There are some caveats to the interpretations of our findings that should be considered. First, we can not unambiguously pinpoint the substrate of the picocyanobacterial transport system, but there are clues as to which molecule is targeted. The siderophore gene cluster includes an outer membrane TonB dependent receptor - a protein family with diverse substrates including siderophores, heme, vitamin B12, and polysaccharides. A prior study with a HLII Prochlorococcus strain found that the outer membrane receptor and substrate-binding protein were strongly upregulated only under Fe-deficient culture conditions [20], which suggests polysaccharides and vitamin B12 are not the primary substrates. TonB dependent receptors can also transport heme, an iron-containing enzyme cofactor, but authentic heme uptake operons usually contain a highly expressed heme oxygenase [89, 90], which is missing from the picocyanobacterial gene clusters. Therefore, we argue that the most parsimonious explanation is that the target substrate is a siderophore or a novel iron-containing metallophore. As a final caveat, we note that we have primarily focused on atmospheric Fe supply due to the strong negative correlations between the distribution of Prochlorococcus siderophore transporters and atmospherically sourced metals like Cu, Al, Mn, and Pb (Fig. 5, Supplementary Fig. S6). However, dFe supply from below the euphotic zone is also an important new Fe source [61, 91], and rapid internal recycling of Fe in the euphotic zone may prolong seasonal phytoplankton productivity and fuel macronutrient consumption [80, 81]. These potential dFe supply processes and other aspects of the Fe budget (e.g., particulate Fe lability, variable mesopelagic regeneration efficiencies [92], and mechanisms of dust bioavailability [93]) were not explicitly considered in this work but are likely relevant to the global distribution of picocyanobacteria siderophore consumers.
Our results imply that the presence of siderophore-consuming Prochlorococcus and Synechococcus could be a valuable biomarker for diagnosing ecosystem Fe deficiency. Indeed, our study highlights two areas of potential iron deficiency that warrant further investigation - the S. Atlantic subtropical gyre and the DCM of the oligotrophic Pacific and S. Atlantic oceans. These areas have historically been considered nitrogen- and light-limited ecosystems, but our results suggest an essential role for Fe. Most of the potentially Fe-deficient regions revealed by our analysis are remote, and sampling Fe at these locations requires specialized knowledge of shipboard trace metal incubations, sampling, and analysis. Because DNA sampling does not require specialized trace metal clean conditions, omics-based approaches based on robust markers of nutrient deficiency could be one complementary tool for increasing the breadth of trace metal surveys in the ocean. Here we have leveraged cross-scale biology of the well-described and ubiquitous marine picocyanobacterium, Prochlorococcus, to reveal vast regions of community iron deficiency in the global ocean using a siderophore transport system as a biomarker. These findings high-light the intricate connection between the ecology and evolution of Prochlorococcus and the Fe cycle of the surface ocean.
Data availability
GEOTRACES and HOT/BATS Metagenome sequencing reads are available from the NCBI Sequence Read Archive under studies SRP110813 and SRP109831 and BioProjects PRJNA385854 and PRJNA385855. Associated sample collection data can be found at https://doi.org/10.1038/sdata.2018.176. Tara Oceans Primary Metagenome sequencing reads are available from the European Nucleotide Archive under project PRJEB402. Associated Tara Oceans chemical and environmental data are available from the Pangaea repository at https://doi.org/10.1594/PANGAEA.875576. Chemical data from the GEOTRACES Intermediate Data Project IDP17 v2 are available at https://www.bodc.ac.uk/geotraces/data/idp2017/. Picocyanobacterial single-cell genomes and other sequence data are available from GenBank 16 S/ITS records MG666579-MG668595, MH074888-MH077527, MH319718-MH319767, and MH327275-MH327492; NCBI Sequence Read Archive study SRP141175; and Genbank assemblies QBVH00000000-QCVZ00000000. Associated sample collection data can be found at https://doi.org/10.1038/sdata.2018.154. Output from the MIT Darwin model and qPCR measurements of Prochlorococcus abundance at HOT/BATS is available from the Simons Collaborative Marine Atlas Project https://simonscmap.com/. The MARMICRODB reference database and instructions for use are available from https://doi.org/10.5281/zenodo.3520509. All computer code for reproducing the results from this specific study is available from https://github.com/slhogle/cyano-sidero-ocean.
References
Partensky F, Garczarek L. Prochlorococcus: advantages and limits of minimalism. Ann Rev Mar Sci. 2010;2:305–31.
Flombaum P, Gallegos JL, Gordillo RA, Rincón J, Zabala LL, Jiao N, et al. Present and future global distributions of the marine cyanobacteria Prochlorococcus and Synechococcus. Proc Natl Acad Sci. 2013;110:9824–9.
Follows MJ, Dutkiewicz S, Grant S, Chisholm SW. Emergent biogeography of microbial communities in a model ocean. Science. 2007;315:1843–6.
Biller SJ, Berube PM, Lindell D, Chisholm SW. Prochlorococcus: the structure and function of collective diversity. Nat Rev Microbiol. 2014;13:13–27.
Farrant GK, Doré H, Cornejo-Castillo FM, Partensky F, Ratin M, Ostrowski M, et al. Delineating ecologically significant taxonomic units from global patterns of marine picocyanobacteria. Proc Natl Acad Sci. 2016;113:E3365–74.
Kashtan N, Roggensack SE, Rodrigue S, Thompson JW, Biller SJ, Coe A, et al. Single-cell genomics reveals hundreds of coexisting subpopulations in wild Prochlorococcus. Science. 2014;344:416–20.
Tagliabue A, Bowie AR, Boyd PW, Buck KN, Johnson KS, Saito MA. The integral role of iron in ocean biogeochemistry. Nature. 2017;543:51–9.
Gledhill M, van den Berg CMG. Determination of complexation of iron(III) with natural organic complexing ligands in seawater using cathodic stripping voltammetry. Mar Chem. 1994;47:41–54.
Rue EL, Bruland KW. The role of organic complexation on ambient iron chemistry in the equatorial Pacific Ocean and the response of a mesoscale iron addition experiment. Limnol Oceanogr 1997;42:901–10.
Hassler CS, van den Berg CMG, Boyd PW. Toward a regional classification to provide a more inclusive examination of the ocean biogeochemistry of iron-binding ligands. Front Mar Sci. 2017;4:1–19.
Gledhill M, Buck KN. The organic complexation of iron in the marine environment: a review. Front Microbiol. 2012;3:1–17.
Bundy RM, Boiteau RM, McLean C, Turk-Kubo KA, McIlvin MR, Saito MA, et al. Distinct siderophores contribute to Iron cycling in the mesopelagic at station ALOHA. Front. 2018;5:1–15.
Boiteau RM, Mende DR, Hawco NJ, McIlvin MR, Fitzsimmons JN, Saito MA, et al. Siderophore-based microbial adaptations to iron scarcity across the eastern Pacific Ocean. Proc Natl Acad Sci. 2016;113:14237–42.
Shaked Y, Lis H. Disassembling iron availability to phytoplankton. Front Microbiol. 2012;3:123.
Morrissey J, Bowler C. Iron utilization in marine cyanobacteria and eukaryotic algae. Front Microbiol. 2012;3:43.
Hogle SL, Cameron Thrash J, Dupont CL, Barbeau KA. Trace metal acquisition by marine heterotrophic bacterioplankton with contrasting trophic strategies. Appl Environ Microbiol. 2016;82:1613–24.
Hopkinson B, Barbeau K. Iron transporters in marine prokaryotic genomes and metagenomes. Environ Microbiol. 2012;14:114–28.
Webb EA, Moffett JW, Waterbury JB. Iron stress in open-ocean Cyanobacteria (Synechococcus, Trichodesmium, and Crocosphaera spp.): Identification of the IdiA protein. Appl Environ Microbiol. 2001;67:5444–52.
Hopkinson BM, Morel FMM. The role of siderophores in iron acquisition by photosynthetic marine microorganisms. Biometals. 2009;22:659–69.
Malmstrom RR, Rodrigue S, Huang KH, Kelly L, Kern SE, Thompson A, et al. Ecology of uncultured Prochlorococcus clades revealed through single-cell genomics and biogeographic analysis. ISME J. 2013;7:184–98.
Ustick LJ, Larkin AA, Garcia CA, Garcia NS, Brock ML, Lee JA, et al. Metagenomic analysis reveals global-scale patterns of ocean nutrient limitation. Science. 2021;372:287–91.
Garcia CA, Hagstrom GI, Larkin AA, Ustick LJ, Levin SA, Lomas MW, et al. Linking regional shifts in microbial genome adaptation with surface ocean biogeochemistry. Philos Trans R Soc Lond B Biol Sci. 2020;375:20190254.
Hogle SL, Dupont CL, Hopkinson BM, King AL, Buck KN, Roe KL, et al. Pervasive iron limitation at subsurface chlorophyll maxima of the California Current. Proc Natl Acad Sci. 2018;115:13300–5.
Hawco NJ, Fu F, Yang N, Hutchins DA, John SG. Independent iron and light limitation in a low-light-adapted Prochlorococcus from the deep chlorophyll maximum. ISME J. 2020;15:359–62.
Biller SJ, Berube PM, Dooley K, Williams M, Satinsky BM, Hackl T, et al. Marine microbial metagenomes sampled across space and time. Sci Data. 2018;5:180176.
Pesant S, Not F, Picheral M, Kandels-Lewis S, Le Bescot N, Gorsky G, et al. Open science resources for the discovery and analysis of Tara Oceans data. Sci Data. 2015;2:150023.
Berube PM, Biller SJ, Hackl T, Hogle SL, Satinsky BM, Becker JW, et al. Single cell genomes of Prochlorococcus, Synechococcus, and sympatric microbes from diverse marine environments. Sci Data. 2018;5:180154.
Biller SJ, Berube PM, Berta-Thompson JW, Kelly L, Roggensack SE, Awad L, et al. Genomes of diverse isolates of the marine cyanobacterium Prochlorococcus. Sci Data. 2014;1:140034.
Karsenti E, Acinas SG, Bork P, Bowler C, De Vargas C, Raes J, et al. A holistic approach to marine eco-systems biology. PLoS Biol. 2011;9:e1001177.
Schlitzer R, Anderson RF, Dodas EM, Lohan M, Geibert W, Tagliabue A, et al. The GEOTRACES intermediate data product 2017. Chem Geol. 2018;493:210–23.
Salt LA, van Heuven SMAC, Claus ME, Jones EM, de Baar HJW. Rapid acidification of mode and intermediate waters in the southwestern Atlantic Ocean. Biogeosciences. 2015;12:1387–401.
Rijkenberg MJA, Middag R, Laan P, Gerringa LJA, van Aken HM, Schoemann V, et al. The distribution of dissolved iron in the West Atlantic Ocean. PLoS One. 2014;9:e101323.
Wyatt NJ, Milne A, Woodward EMS, Rees AP, Browning TJ, Bouman HA, et al. Biogeochemical cycling of dissolved zinc along the GEOTRACES South Atlantic transect GA10 at 40°S: Dissolved zinc in the Atlantic at 40o S. Glob Biogeochem Cycles. 2014;28:44–56.
Ashkezari MD, Hagen NR, Denholtz M, Neang A, Burns TC, Morales RL, et al. Simons collaborative marine atlas project (Simons CMAP): an open‐source portal to share, visualize, and analyze ocean data. Limnol Oceanogr Methods. 2021;19:488–96.
Acker M, Hogle SL, Berube PM, Hackl T, Stepanauskas R, Chisholm SW, et al. Phosphonate production by marine microbes: exploring new sources and potential function. Proc Natl Acad Sci. 2022;119:e2113386119.
Becker JW, Hogle SL, Rosendo K, Chisholm SW. Co-culture and biogeography of Prochlorococcus and SAR11. ISME J. 2019;13:1506–19.
Malmstrom RR, Coe A, Kettler GC, Martiny AC, Frias-Lopez J, Zinser ER, et al. Temporal dynamics of Prochlorococcus ecotypes in the Atlantic and Pacific oceans. ISME J. 2010;4:1252–64.
Fitzsimmons JN, Hayes CT, Al-Subiai SN, Zhang R, Morton PL, Weisend RE, et al. Daily to decadal variability of size-fractionated iron and iron-binding ligands at the Hawaii Ocean Time-series Station ALOHA. Geochim Cosmochim Acta. 2015;171:303–24.
Buck KN, Sohst B, Sedwick PN. The organic complexation of dissolved iron along the U.S. GEOTRACES (GA03) North Atlantic Section. Deep Sea Res Part 2. 2015;116:152–65.
Mawji E, Gledhill M, Milton JA, Tarran GA, Ussher S, Thompson A, et al. Hydroxamate siderophores: occurrence and importance in the Atlantic Ocean. Environ Sci Technol. 2008;42:8675–80.
Buck KN, Bruland KW. The physicochemical speciation of dissolved iron in the Bering Sea, Alaska. Limnol Oceanogr. 2007;52:1800–8.
Boiteau RM, Fitzsimmons JN, Repeta DJ, Boyle EA. Detection of iron ligands in seawater and marine cyanobacteria cultures by high-performance liquid chromatography-inductively coupled plasma-mass spectrometry. Anal Chem. 2013;85:4357–62.
Huerta-Cepas J, Szklarczyk D, Forslund K, Cook H, Heller D, Walter MC, et al. eggNOG 4.5: a hierarchical orthology framework with improved functional annotations for eukaryotic, prokaryotic and viral sequences. Nucleic Acids Res. 2016;44:D286–93.
Huerta-Cepas J, Forslund K, Coelho LP, Szklarczyk D, Jensen LJ, von Mering C, et al. Fast genome-wide functional annotation through orthology assignment by eggNOG-mapper. Mol Biol Evol. 2017;34:2115–22.
Blin K, Shaw S, Steinke K, Villebro R, Ziemert N, Lee SY, et al. antiSMASH 5.0: updates to the secondary metabolite genome mining pipeline. Nucleic Acids Res. 2019;47:W81–W87.
Menzel P, Ng KL, Krogh A. Fast and sensitive taxonomic classification for metagenomics with Kaiju. Nat Commun. 2016;7:11257.
Wright M, Ziegler A. ranger: A fast implementation of random forests for high dimensional data in C++ and R. J Stat Softw, Artic. 2017;77:1–17.
Jolliffe IT. Principal Component Analysis. 1986. Springer-Verlag.
Kursa M, Rudnicki W. Feature selection with the Boruta package. J Stat Softw, Artic. 2010;36:1–13.
Milligan AJ, Harrison PJ. Effects of non-steady-state iron limitation on nitrogen assimilatory enzymes in the marine diatom Thalassiosira weissflogii (Bacillariophyceae). J Phycol. 2000;36:78–86.
Lomas MW, Lipschultz F. Forming the primary nitrite maximum: Nitrifiers or phytoplankton? Limnol Oceanogr. 2006;51:2453–67.
Ahlgren NA, Belisle BS, Lee MD. Genomic mosaicism underlies the adaptation of marine Synechococcus ecotypes to distinct oceanic iron niches. Environ Microbiol. 2019;22:1801–15.
Martiny AC, Coleman ML, Chisholm SW. Phosphate acquisition genes in Prochlorococcus ecotypes: evidence for genome-wide adaptation. Proc Natl Acad Sci. 2006;103:12552–7.
Coleman ML, Chisholm SW. Ecosystem-specific selection pressures revealed through comparative population genomics. Proc Natl Acad Sci. 2010;107:18634–9.
Berube PM, Rasmussen A, Braakman R, Stepanauskas R, Chisholm SW. Emergence of trait variability through the lens of nitrogen assimilation in Prochlorococcus. eLife. 2019;8:e41043.
Olgun N, Duggen S, Croot PL, Delmelle P, Dietze H, Schacht U, et al. Surface ocean iron fertilization: The role of airborne volcanic ash from subduction zone and hot spot volcanoes and related iron fluxes into the Pacific Ocean. Global Biogeochem Cycles. 2011;25:GB4001.
Manck LE, Park J, Tully BJ, Poire AM, Bundy RM, Dupont CL, et al. Petrobactin, a siderophore produced by Alteromonas, mediates community iron acquisition in the global ocean. ISME J. 2021;16:358–69.
Mackey KRM, Post AF, McIlvin MR, Saito MA. Physiological and proteomic characterization of light adaptations in marine Synechococcus. Environ Microbiol. 2017;19:2348–65.
Raven JA. Predictions of Mn and Fe use efficiencies of phototrophic growth as a function of light availability for growth and of C assimilation pathway. N. Phytol. 1990;116:1–18.
Sunda WG, Huntsman SA. Interrelated influence of iron, light and cell size on marine phytoplankton growth. Nature. 1997;390:389–92.
Hawco NJ, Barone B, Church MJ, Babcock-Adams L, Repeta DJ, Wear EK, et al. Iron depletion in the deep chlorophyll maximum: Mesoscale eddies as natural iron fertilization experiments. Global Biogeochem Cycles. 2021;35:e2021GB007112.
Barbeau K, Rue EL, Bruland KW, Butler A. Photochemical cycling of iron in the surface ocean mediated by microbial iron(III)-binding ligands. Nature. 2001;413:409–13.
van den Engh GJ, Doggett JK, Thompson AW, Doblin MA, Gimpel CNG, Karl DM. Dynamics of Prochlorococcus and Synechococcus at Station ALOHA revealed through flow cytometry and high-resolution vertical sampling. Front Mar Sci. 2017;4:1–14.
Karl DM, Church MJ. Microbial oceanography and the Hawaii Ocean Time-series programme. Nat Rev Microbiol. 2014;12:699–713.
Neuer S, Davenport R, Freudenthal T, Wefer G, Llinás O, Rueda M-J, et al. Differences in the biological carbon pump at three subtropical ocean sites. Geophys Res Lett 2002;29:32–1.
Giovannoni SJ, Vergin KL. Seasonality in ocean microbial communities. Science. 2012;335:671–6.
Jickells T, Moore CM. The importance of atmospheric deposition for ocean productivity. Annu Rev Ecol Evol Syst. 2015;46:481–501.
Rii YM, Karl DM, Church MJ. Temporal and vertical variability in picophytoplankton primary productivity in the North Pacific Subtropical Gyre. Mar Ecol Prog Ser. 2016;562:1–18.
Boyle EA, Bergquist BA, Kayser RA, Mahowald N. Iron, manganese, and lead at Hawaii Ocean Time-series station ALOHA: Temporal variability and an intermediate water hydrothermal plume. Geochim Cosmochim Acta. 2005;69:933–52.
Caprara S, Buck KN, Gerringa LJA, Rijkenberg MJA, Monticelli D. A compilation of iron speciation data for open oceanic waters. Front Mar Sci. 2016;3:1–7.
Gledhill M, Gerringa LJA. The effect of metal concentration on the parameters derived from complexometric titrations of trace elements in seawater: A model study. Front Mar Sci. 2017;4:1–15.
Jickells TD, Baker AR, Chance R. Atmospheric transport of trace elements and nutrients to the oceans. Philos Trans R Soc Lond A. 2016;374:20150286.
Sedwick PN, Church TM, Bowie AR, Marsay CM, Ussher SJ, Achilles KM, et al. Iron in the Sargasso Sea (Bermuda Atlantic Time-series Study region) during summer: Eolian imprint, spatiotemporal variability, and ecological implications. Glob Biogeochem Cycles. 2005;19:GB4006.
Ohnemus DC, Rauschenberg S, Cutter GA, Fitzsimmons JN, Sherrell RM, Twining BS. Elevated trace metal content of prokaryotic communities associated with marine oxygen deficient zones. Limnol Oceanogr. 2016;62:3–25.
Browning TJ, Achterberg EP, Rapp I, Engel A, Bertrand EM, Tagliabue A, et al. Nutrient co-limitation at the boundary of an oceanic gyre. Nature. 2017;551:242–6.
Louropoulou E, Gledhill M, Achterberg EP, Browning TJ, Honey DJ, Schmitz RA, et al. Heme b distributions through the Atlantic Ocean: evidence for ‘anemic’ phytoplankton populations. Sci Rep. 2020;10:4551.
Mark Moore C. Diagnosing oceanic nutrient deficiency. Philos Trans R Soc Lond A. 2016;374:20150290.
Moore CM, Mills MM, Arrigo KR, Berman-Frank I, Bopp L, Boyd PW, et al. Processes and patterns of oceanic nutrient limitation. Nat Geosci. 2013;6:701–10.
Moore JK, Doney SC, Lindsay K. Upper ocean ecosystem dynamics and iron cycling in a global three-dimensional model. Global Biogeochem Cycles 2004;18.
Rafter PA, Sigman DM, Mackey KRM. Recycled iron fuels new production in the eastern equatorial Pacific Ocean. Nat Commun. 2017;8:1100.
Boyd PW, Ellwood MJ, Tagliabue A, Twining BS. Biotic and abiotic retention, recycling and remineralization of metals in the ocean. Nat Geosci. 2017;10:167–73.
Ferreira D, Cessi P, Coxall HK, de Boer A, Dijkstra HA, Drijfhout SS, et al. Atlantic-Pacific asymmetry in deep water formation. Annu Rev Earth Planet Sci. 2018;46:327–52.
Rusch DB, Martiny AC, Dupont CL, Halpern AL, Venter JC. Characterization of Prochlorococcus clades from iron-depleted oceanic regions. Proc Natl Acad Sci. 2010;107:16184–9.
Dutkiewicz S, Ward BA, Monteiro F, Follows MJ. Interconnection of nitrogen fixers and iron in the Pacific Ocean: Theory and numerical simulations. Glob Biogeochem Cycles. 2012;26:GB1012.
Resing JA, Sedwick PN, German CR, Jenkins WJ, Moffett JW, Sohst BM, et al. Basin-scale transport of hydrothermal dissolved metals across the South Pacific Ocean. Nature. 2015;523:200–3.
Sela I, Wolf YI, Koonin EV. Theory of prokaryotic genome evolution. Proc Natl Acad Sci. 2016;113:11399–407.
Zakem EJ, Al-Haj A, Church MJ, van Dijken GL, Dutkiewicz S, Foster SQ, et al. Ecological control of nitrite in the upper ocean. Nat Commun. 2018;9:1206.
Lauderdale JM, Braakman R, Forget G, Dutkiewicz S, Follows MJ. Microbial feedbacks optimize ocean iron availability. Proc Natl Acad Sci. 2020;117:4842–9.
Hogle SL, Barbeau KA, Gledhill M. Heme in the marine environment: from cells to the iron cycle. Metallomics. 2014;6:1107–20.
Hogle SL, Brahamsha B, Barbeau KA. Direct heme uptake by phytoplankton-associated Roseobacter bacteria. mSystems. 2017;2:e00124–16.
Coale KH, Fitzwater SE, Michael Gordon R, Johnson KS, Barber RT. Control of community growth and export production by upwelled iron in the equatorial Pacific Ocean. Nature. 1996;379:621–4.
Bressac M, Guieu C, Ellwood MJ, Tagliabue A, Wagener T, Laurenceau EC, et al. Resupply of mesopelagic dissolved iron controlled by particulate iron composition. Nat Geosci. 2019;12:995–1000.
Basu S, Gledhill M, de Beer D, Prabhu Matondkar SG, Shaked Y. Colonies of marine cyanobacteria Trichodesmium interact with associated bacteria to acquire iron from dust. Commun Biol. 2019;2:284.
Acknowledgements
We thank present and past members of the GEOTRACES consortium for collecting, curating, and sharing trace metal and other biogeochemical data. In particular, we thank the GEOTRACES chief scientists for their support: Andrew Bowie (University of Tasmania), Philip Boyd (University of Tasmania), Edward Boyle (Massachusetts Institute of Technology), Gregory Cutter (Old Dominion University), Loes Gerringa (NIOZ Royal Netherlands Institute for Sea Research), Gideon Henderson (University of Oxford), William Jenkins (Woods Hole Oceanographic Institution), and Micha Rijkenberg (NIOZ Royal Netherlands Institute for Sea Research), as well as Gerhard Herndl (University of Vienna), James Moffett (University of Southern California) and Hein de Baar (NIOZ Royal Netherlands Institute for Sea Research) for their support with GEOTRACES. We thank the HOT and BATS field teams and organizational leaders for their assistance. We also thank the developers, maintainers, and administrators of the Simons Collaborative Marine Atlas Project and the MIT Darwin ecosystem model. This work was supported in part by grants from the National Science Foundation (OCE-1153588, and DBI-0424599 to SWC) and the Simons Foundation (Life Sciences Project Award IDs 337262, 647135 to SWC; SCOPE Award ID 329108 to SWC. This paper is a contribution from the Simons Collaboration on Ocean Processes and Ecology (SCOPE). We kindly thank the senior editor and two anonymous reviewers for feedback that helped us to improve our manuscript.
Funding
Open Access funding provided by University of Turku (UTU) including Turku University Central Hospital.
Author information
Authors and Affiliations
Contributions
SLH: conceptualization, investigation, data curation, software, formal analysis, visualization, writing original draft, writing review, and editing; THackl: investigation, data curation, software, formal analysis, writing review and editing; RMB: investigation, data curation, methodology, writing review and editing; JP: data curation, methodology, BMS: data curation, writing review and editing, THiltunen: resources, investigation, writing review and editing; SJB: data curation, investigation, writing review and editing; PB: data curation, investigation, writing review and editing; SWC: conceptualization, supervision, project administration, resources, funding acquisition, writing review and editing.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hogle, S.L., Hackl, T., Bundy, R.M. et al. Siderophores as an iron source for picocyanobacteria in deep chlorophyll maximum layers of the oligotrophic ocean. ISME J 16, 1636–1646 (2022). https://doi.org/10.1038/s41396-022-01215-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-022-01215-w
This article is cited by
-
Synechococcus nitrogen gene loss in iron-limited ocean regions
ISME Communications (2023)