Abstract
Background
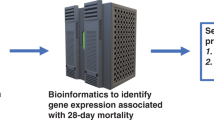
Prognostic biomarker research neonatal sepsis is lacking. We assessed the utility of a validated pediatric prognostic tool called PERSEVERE II that uses decision tree methodology to predict mortality at discharge in neonates who experienced sepsis.
Methods
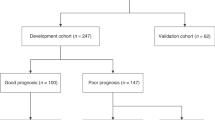
Prospective study in a dual-center cohort of neonates with sepsis admitted between June 2020 and December 2021. Biomarker analysis was done on serum samples obtained at the time of evaluation for the event.
Results
In a cohort of 59 neonates with a mortality rate of 15.3%, PERSEVERE II was 67% sensitive and 59% specific for mortality, p 0.27. Amongst PERSEVERE II biomarkers, IL-8 showed good prognostic performance for mortality prediction with a cutoff of 300 pg/mL (sensitivity 100%, specificity 65%, negative predictive value 100%, AUC 0.87, p 0.0003). We derived a new decision tree that is neonate specific (nPERSEVERE) with improved performance compared to IL-8 (sensitivity 100%, specificity 86%, negative predictive value 100%, AUC 0.95, p < 0.0001).
Conclusions
IL-8 and nPERSEVERE demonstrated good prognostic performance in a small cohort of neonates with sepsis. Moving toward precision medicine in sepsis, our study proposes an important tool for clinical trial prognostic enrichment that needs to be validated in larger studies.
Impact
-
Prognostic and predictive biomarker research is lacking in the newborn intensive care unit.
-
Biomarkers can be used at the time of evaluation for neonatal sepsis (blood culture acquisition) to identify neonates with high baseline mortality risk.
-
Stratification is an important step toward precision medicine in neonatal sepsis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 14 print issues and online access
$259.00 per year
only $18.50 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Datasets generated from this work are available upon reasonable request to the corresponding author.
References
Stoll, B. J. et al. Neonatal outcomes of extremely preterm infants from the NICHD Neonatal Research Network. Pediatrics 126, 443–456 (2010).
Camacho-Gonzalez, A., Spearman, P. W. & Stoll, B. J. Neonatal infectious diseases: evaluation of neonatal sepsis. Pediatr. Clin. North Am. 60, 367–389 (2013).
Carr, R., Modi, N. & Dore, C. G-CSF and GM-CSF for treating or preventing neonatal infections. Cochrane Database Syst. Rev. 2003, CD003066 (2003).
Ohlsson, A. & Lacy, J. B. Intravenous immunoglobulin for suspected or proven infection in neonates. Cochrane Database Syst. Rev. 1, CD001239 (2020).
Brown, J. V. E., Meader, N., Wright, K., Cleminson, J. & McGuire, W. Assessment of C-reactive protein diagnostic test accuracy for late-onset infection in newborn infants: a systematic review and meta-analysis. JAMA Pediatr. 174, 260–268 (2020).
Ye, Q., Du, L. Z., Shao, W. X. & Shang, S. Q. Utility of cytokines to predict neonatal sepsis. Pediatr. Res. 81, 616–621 (2017).
Zhou, M., Cheng, S., Yu, J. & Lu, Q. Interleukin-8 for diagnosis of neonatal sepsis: a meta-analysis. PLoS One 10, e0127170 (2015).
Sherwin, C. et al. Utility of interleukin-12 and interleukin-10 in comparison with other cytokines and acute-phase reactants in the diagnosis of neonatal sepsis. Am. J. Perinatol. 25, 629–636 (2008).
Wong, H. R. et al. Prospective clinical testing and experimental validation of the Pediatric Sepsis Biomarker Risk Model. Sci. Transl. Med. 11, eaax9000 (2019).
Wong, H. R. et al. The pediatric sepsis biomarker risk model. Crit. Care 16, R174 (2012).
Wong, H. R. et al. Testing the prognostic accuracy of the updated pediatric sepsis biomarker risk model. PLoS One 9, e86242 (2014).
Wong, H. R. et al. A multibiomarker-based outcome risk stratification model for adult septic shock. Crit. Care Med. 42, 781–789 (2014).
Jacobs, L. et al. The Pediatric Sepsis Biomarker Risk Model (PERSEVERE) biomarkers predict clinical deterioration and mortality in immunocompromised children evaluated for infection. Sci. Rep. 9, 424 (2019).
Harris, P. A. et al. Research electronic data capture (REDCap)–a metadata-driven methodology and workflow process for providing translational research informatics support. J. Biomed. Inf. 42, 377–381 (2009).
Wynn, J. L. & Wong, H. R. Pathophysiology and treatment of septic shock in neonates. Clin. Perinatol. 37, 439–479 (2010).
Jetton, J. G. et al. Incidence and outcomes of neonatal acute kidney injury (AWAKEN): a multicentre, multinational, observational cohort study. Lancet Child Adolesc. Health 1, 184–194 (2017).
Wynn, J. L. et al. The influence of developmental age on the early transcriptomic response of children with septic shock. Mol. Med. 17, 1146–1156 (2011).
Hack, C. E. et al. Interleukin-8 in sepsis: relation to shock and inflammatory mediators. Infect. Immun. 60, 2835–2842 (1992).
Anderson, B. J. et al. Plasma sTNFR1 and IL8 for prognostic enrichment in sepsis trials: a prospective cohort study. Crit. Care 23, 400 (2019).
Bao, Q., Lv, R. & Lei, M. IL-33 attenuates mortality by promoting IFN-gamma production in sepsis. Inflamm. Res. 67, 531–538 (2018).
Vanden Berghe, T. et al. Simultaneous targeting of IL-1 and IL-18 is required for protection against inflammatory and septic shock. Am. J. Respir. Crit. Care Med. 189, 282–291 (2014).
Lertwattanachai, T. et al. Clinical outcomes of empirical high-dose meropenem in critically ill patients with sepsis and septic shock: a randomized controlled trial. J. Intensive Care 8, 26 (2020).
Singer, M. et al. The Third International Consensus Definitions for Sepsis and Septic Shock (Sepsis-3). JAMA 315, 801–810 (2016).
Goldstein, B., Giroir, B. & Randolph, A., International Consensus Conference on Pediatric Sepsis. International Pediatric Sepsis Consensus Conference: definitions for sepsis and organ dysfunction in pediatrics. Pediatr. Crit. Care Med. 6, 2–8 (2005).
Acknowledgements
Near the end of this work, Dr Hector R. Wong passed away unexpectedly. He was a good mentor, friend, husband, father, and physician scientist. He will be forever remembered by those who had the privilege to work with him. We also appreciate Dr Basillia Zingarelli for reviewing the final drafts of the manuscript and Andrew M. Smith for his input on the analysis. Some of the figures were generated using www.BioRender.com.
Funding
This presented work is supported by Dr Wong’s R35GM126943 grant from the NIH.
Author information
Authors and Affiliations
Contributions
F.N.A., M.N.A., and H.R.W. conceptualized and designed the study, performed the statistical analyses, and reviewed the data generated. P.L. conducted the Luminex assays to obtain serum biomarker levels. F.N.A drafted the initial manuscript, and F.N.A and M.N.A. reviewed and revised the manuscript. All authors approved the work submitted in this manuscript and are accountable for all aspects of this work.
Corresponding author
Ethics declarations
Competing interests
Cincinnati Children’s Research foundation and Hector Wong hold U.S. patents for the PERSEVERE biomarkers. Cincinnati Children’s Research foundation and F.N.A. have filed for a U.S. patent for nPERSEVERE.
Ethics approval and consent to participate
This study was reviewed by the institutional review board at Cincinnati Children’s Hospital Medical Center and the University of Cincinnati and consent was waived.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Al Gharaibeh, F.N., Lahni, P., Alder, M.N. et al. Biomarkers estimating baseline mortality risk for neonatal sepsis: nPERSEVERE: neonate-specific sepsis biomarker risk model. Pediatr Res 94, 1451–1456 (2023). https://doi.org/10.1038/s41390-022-02414-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41390-022-02414-z