Abstract
Background
Children with low birth weight (LBW) have a higher risk of developing endocrine-metabolic disorders later in life. Deregulation of specific microRNAs (miRNAs) could underscore the programming of adult pathologies. We analyzed the miRNA expression pattern in both umbilical cord serum samples from LBW and appropriate-for-gestational-age (AGA) newborns and maternal serum samples in the 3rd trimester of gestation, and delineated the relationships with fetal growth, body composition, and markers of metabolic risk.
Methods
Serum samples of 12 selected mother–newborn pairs, including 6 LBW and 6 AGA newborns, were used for assessing miRNA profile by RNA-sequencing. The miRNAs with differential expression were validated in a larger cohort [49 maternal samples and 49 umbilical cord samples (24 LBW, 25 AGA)] by RT-qPCR. Anthropometric, endocrine-metabolic markers and body composition (by DXA) in infants were determined longitudinally over 12 months.
Results
LBW newborns presented reduced circulating concentrations of miR-191-3p (P = 0.015). miR-191-3p levels reliably differentiated LBW from AGA individuals (ROC AUC = 0.76) and were positively associated with anthropometric and body composition measures at birth and weight Z-score at 12 months (P < 0.05).
Conclusions
miR-191-3p was reliably different in LBW individuals, and could be a new player in the epigenetic mechanisms linking LBW and future endocrine-metabolic adverse outcomes.
Impact
-
Children with low birth weight (LBW) have a higher risk of developing endocrine-metabolic disorders. Deregulation of specific microRNAs (miRNAs) could underscore the programming of those pathologies.
-
miR-191-3p is downregulated in serum of LBW newborns, and its concentrations associate positively with neonatal anthropometric measures, with lean mass and bone accretion at age 15 days and with weight Z-score at age 12 months.
-
miR-191-3p was reliably different in individuals with LBW, and could be a new player in the epigenetic mechanisms connecting LBW and future endocrine-metabolic adverse outcomes.
Similar content being viewed by others
Introduction
Children with low birth weight (LBW), in particular those who undergo rapid postnatal catch-up growth, have a higher chance of developing endocrine-metabolic disorders later in life.1 A “mismatch” between decreased prenatal weight gain and increased postnatal weight gain favors fat excess in subcutaneous adipose tissue resulting in ectopic lipid accretion, particularly in the viscera and liver, which in turn could foster the development of insulin resistance and low-grade inflammation status.2 The mechanisms underlying this sequence of events remain poorly understood, but epigenetic modulation of gene expression could contribute to the programming of adult pathologies.3,4,5
microRNAs (miRNAs), which represent a layer of epigenetic control, are short non-coding molecules that are able to promote post-transcriptional modifications in gene expression by binding the 3-UTR (untranslated region) of its target mRNAs.6 miRNAs are expressed in a broad variety of tissues, including cord blood and placenta,7,8 and can also be packed into micro-vesicles and be actively secreted into circulation.9 miRNAs participate in the onset and development of a variety of disorders and have emerged as diagnostic biomarkers in several diseases.10 Concerning the LBW condition, recent studies have identified altered circulating levels of different miRNAs in mothers delivering LBW infants,11,12 together with differentially expressed miRNAs in placentas13,14,15,16 and umbilical cord of LBW newborns.17 Indeed, we have previously shown that specific umbilical cord miRNAs may serve as new biomarkers for identifying precociously those LBW newborns who will experience catch-up growth and could also take part to the augmented risk for cardiometabolic alterations described in later stages of LBW infants’ life.17 However, to our knowledge, the circulating miRNA profile in serum samples of mothers delivering LBW infants and in umbilical cord serum samples of their newborns have not been simultaneously characterized so far.
In the present work, we analyzed the miRNA expression profile in umbilical cord serum samples from LBW and appropriate-for-gestational-age (AGA) newborns and the corresponding maternal serum samples in the 3rd trimester of gestation, to identify novel candidate biomarkers of the LBW condition as well as to underscore the relationships between the identified miRNAs and fetal growth, body composition and markers of metabolic risk over the first year of life.
Materials and methods
Study population
The study population was comprised of 49 mother–newborn pairs who were enrolled during the customary third-trimester appointment among those pregnant mothers successively visited in the outpatient clinics of Sant Joan de Déu University Hospital and Hospital de Sant Boi – Parc Sanitari Sant Joan de Déu (Barcelona, Spain) (Supplementary Fig. 1). The inclusion criteria were: [1] singleton uncomplicated pregnancies at term (37–42 weeks); [2] birth weight Z-score > −0.5 standard deviation (SD) for sex and gestational age (AGA) or below −1.5 SD for sex and gestational age [LBW; defined as <2.5 Kg independently of gestational age;18] [3] co-availability of blood samples from mothers and newborns (umbilical cord blood), obtained at the third trimester of pregnancy and at delivery, respectively; [4] acquisition of informed written consent. The exclusion criteria were: [1] maternal conditions (infectious, hypertension, pre-eclampsia, gestational diabetes, or pre-existing type 1 or type 2 diabetes), and drug or alcohol abuse, and [2] fetal conditions (malformations or complications at birth).
Twenty-five newborns were born AGA (N = 11 girls and N = 14 boys) and 24 were born with LBW (N = 10 girls and N = 14 boys). The postnatal follow-up to age 15 days was completed in 20 out of 25 AGA and in 22 out of 24 LBW newborns. Over the subsequent year of prospective follow-up, about 50% of the initial population dropped out from the study, and thus, the assessments could only be completed in N = 14 AGA and N = 15 LBW infants at age 4 months and in N = 11 AGA and N = 12 LBW infants at age 1 year (Supplementary Fig. 1).
Assessments
Maternal age at delivery, smoking habits, parity, pre-gestational weight, weight at delivery, and maternal body mass index (BMI) calculated as weight (in kg)/the square of height (in meters), were obtained from the maternal clinical history. Gestational age was estimated based on the last menses and validated by first-trimester ultrasound.
Infants were weighed and measured immediately after delivery, and again at age 4 and 12 months, and BMI was derived at each time point. Sex-adjusted and gestational age-adjusted Z-scores (standard deviation scores) for weight, length, and BMI were calculated using regional normative data.19
A maternal venous sample was obtained during the third trimester of pregnancy, between week 28 and delivery. Umbilical cord blood samples were collected promptly after childbirth, and processed before the separation of the placenta.20 At age 4 and 12 months, venous samples were obtained during the morning in the fasting state. The serum fraction was separated by centrifugation (at 3000 r.p.m. for 10 min) and stored at −80 °C until analysis.
Serum glucose was determined by the glucose oxidase method. Insulin-like growth factor I (IGF-I) was measured by immuno-chemiluminiscence (DPC IMMULITE 2500, Siemens, Germany); the detection limit was 25 ng/mL; the coefficients of variability (CVs) were <10%. Insulin resistance index (homeostasis model assessment of insulin resistance; HOMA-IR) was assessed as [fasting insulin (mU/L)] × [fasting glucose (mmol/L)]/22.5.
Body composition was evaluated at age 15 days, 4 months, and 1 year by dual X-ray absorptiometry (DXA) with a Lunar Prodigy coupled to Lunar software (Lunar Corp, Madison, WI), adjusted for infants.20 Fat and lean mass CVs were <3%.
Small RNA-sequencing
The serum samples of 12 mother–newborn pairs, including six AGA and six LBW newborns (discovery cohort), were selected for miRNA profiling (Supplementary Fig. 1). Those newborns fulfilled specifically the following birth weight Z-score criterium: −0.5 ≤SD ≤ + 1.0 (AGA) and SD ≤ −2.0 (LBW), in order to select the most severe cases and avoid overlapping. The sample was stratified to represent the entire cohort regarding gestational age, BMI, and newborn sex. The studied subgroups were also comparable concerning maternal (pre-pregnancy BMI, age, and gestational weight gain) and delivery (newborn sex and gestational age) characteristics.
miRNA profiling was performed by RNA-sequencing (Making Genetics S.L.; Villava, Navarra, Spain; www.making-genetics.eu). Total RNA containing the small RNA fraction was obtained from 250 µL of serum by the use of miRNeasy Serum/Plasma kit (Qiagen, Hilden, Germany) and small cDNA libraries were generated using Small RNA TruSeq kit (Illumina, San Diego, CA) following the instructions of the manufacturer. Indexed libraries were pooled in an equimolar ratio and sequenced on an Illumina NextSeq 550 machine with NextSeq 500/550 High Output Kit v2.5 (75 cycles). From the RNA-seq data, libraries’ miRNA expression was extracted using the featureCounts function (Bioconductor R package Rsubread),21 after correct alignment on GRCh38 using the STAR22 of adapter-trimmed and filtered reads as stated by the manufacturer.
As previously described,23 data were filtered by genes that had > one reads per million mapped reads in at least two libraries; intra- and inter-sample normalization were done using the trimmed mean of M-values (TMM) method implemented in the edgeR Bioconductor package.24 After preprocessing, linear model fitting and differential miRNA expression analysis were performed using the eBayes moderated t-statistic by limma package, and individual miRNAs P-values were derived.25 For each miRNA the log2 fold change was calculated from the readcount. The results are expressed as the average expression in AGA samples versus the average expression in LBW samples (both maternal and umbilical cord). Benjamini–Hochberg procedure was used to adjust the raw P-values, and a FDR (false discovery rate) cut-off of 0.05 was used as a statistically significant threshold in the analysis.
Pathway enrichment analyses
Target Scan (v7.2; targetscan.org)26 was used for predicting biological targets of the miRNAs with differential expression between the AGA and LBW subgroups. In mammals, predictions are classified according to the predicted efficacy of targeting, determined using cumulative weighted context++ scores of the sites.25 The cumulative weighted context++ score cut-off was set up at −0.9 with the objective of minimizing false-positive associations between miRNA and their targets, in addition, to yield a feasible amount of targets to be considered in the analysis of enrichment.27 Once target genes were identified, Enrichr tools, interrogating KEEG, BIOPLANET, and WIKIPATHWAY databases, were used to predict biological pathways and biological processes.28
miRNA validation by RT-qPCR analysis
The five most differentially expressed miRNAs in the 12 mother–newborn samples in which miRNA profiling were validated in the whole study cohort [validation cohort: 49 maternal serum samples and 49 umbilical cord serum samples (25 AGA; 24 LBW)] by RT-qPCR. Total RNA containing the small RNA fraction was obtained from 250 µL of serum by the use of the miRNeasy serum/plasma advanced kit (Qiagen) and the RNA spike-in kit for miRCURY LNA™ universal RT (Qiagen-Exiqon). Total RNA was reversed-transcribed by the use of a miRCURY LNA™ universal RT cDNA synthesis kit (Qiagen-Exiqon). miRNA expression was assessed by RT-qPCR using miRCURY LNA™ SYBR Green PCR kit and specific miRCURY LNA™ miRNA PCR assays (Qiagen-Exiqon) using a LightCycler 480 real-time PCR System (Roche Diagnostics, Pleasanton, CA). The standard cycling conditions were as follows: 2-min polymerase activation at 95 °C, 45 cycles of 10 s at 95 °C for denaturation and 60 s at 56 °C for annealing and extension. Housekeeping control miRNAs were selected for normalization according to our RNA-seq profiling (Supplementary Table 1) and to published data.29,30,31 Relative miRNA expression levels were calculated using the comparative threshold 2−ΔΔCT method.
Statistical analysis and ethics Statement
Data were analyzed using the SPSS statistics version 22.0 (SPSS software, IBM, Chicago, IL). Results are shown as mean ± interquartile range (CI 95%). Variables with normal distribution were compared with a two-tailed Student’s t-test. When necessary, logarithmic transformation was used to achieve the normal distribution of continuous variables. Chi-square test was used to compare qualitative variables. The area under the ROC curve (AUC) was used to assess the diagnostic value for distinguishing between AGA and LBW subjects. The associations between miRNA expression and clinical and body composition parameters were assessed by Pearson correlation; followed by multivariate regression analysis adjusted for confounding variables such as sex of the newborn and maternal smoking. A P-value < 0.05 was considered statistically significant. All data provided in this work have been deposited in an online repository (BioProject ID: PRJNA747266) and are acceccible at: https://www.ncbi.nlm.nih.gov/bioproject/PRJNA747266.
The study was approved by the Institutional Review Board of the University of Barcelona, Sant Joan de Déu Hospital; mothers signed an informed consent at recruitment.
Results
Maternal and offspring characteristics
Table 1 depicts the anthropometric variables in the mothers and in the newborns by subgroups according to birth weight.
No differences were observed between mothers delivering LBW newborns as compared to those delivering AGA newborns. At birth, LBW newborns were smaller, less adipose, had less lean mass, less bone mineral density (BMD), and less bone mineral content (BMC), as expected.20 At age 4 months, LBW infants remained smaller and had less fat mass, and less lean mass, BMD, and BMC. At age 12 months, LBW infants normalized their adiposity and BMC, but remained lighter and had less lean mass and BMC.
miRNA profiling
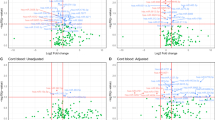
RNA-sequencing detected 20 differentially expressed miRNAs in maternal serum samples, and 87 differentially expressed miRNAs in umbilical cord serum samples. Among all detected miRNAs, one (miR-9-5p) was shared by the mothers and newborns (Fig. 1a). Among the 20 differentially expressed miRNAs in maternal samples, 11 were downregulated and 9 were upregulated in the samples from mothers delivering LBW infants. Among the 87 differentially expressed miRNAs found in umbilical cord serum samples, 9 were downregulated and 78 were upregulated in LBW newborns (Fig. 1b, c).
a Venn diagram depicting the miRNAs differentially expressed in serum samples from mothers delivering appropriate-for-gestational-age (AGA) and low birth weight (LBW) newborns, and in umbilical cord samples from those AGA and LBW newborns. b, c Hierarchical clustering heatmap depicting differentially expressed miRNAs identified in maternal serum (b) and in umbilical cord samples (c). miRNAs with a P-value < 0.05 and a Log2 fold change > ±2 in expression levels are shown.
Predicted miRNA pathways and target genes
TargetScan was used to predict the hypothetical targets for the 20 miRNAs detected in maternal samples and the 87 detected in newborn samples found to be differentially expressed. After applying a cumulative weighted context score cut-off level of −0.9 and removing duplicates (since two or more miRNAs could be shared by more than one target gene) the predicted target genes were 35 in the mothers and 391 in the newborns (Supplementary Table 2). Enrichment analysis hinted that the putative miRNA targets were mostly involved in insulin and inflammation pathways (Table 2).
Validation of differentially expressed miRNAs
The miRNAs with differential expression were classified based on their expression fold change, and the top 5 differentially expressed miRNAs from both maternal and umbilical cord samples were chosen for validation by individual RT-qPCR in the whole study population (Table 3). The validation was performed although the rate of differentially expressed genes did not surpass the expected FDR (cut-off of 0.05). miR-9-5p was also selected for validation in umbilical cord serum samples as it was common in mothers and newborns.
The validation of the 11 differentially expressed miRNAs by RT-qPCR confirmed that miR-191-3p was downregulated in LBW newborns compared to AGA newborns (Fig. 2a). The other studied miRNAs in the umbilical cord (miR-355-3p, miR-127-5p, miR-655-3p, miR-5000-3p, and miR-9-5-p) and maternal serum samples (miR-382-5p, miR-1307-3p, miR-375, miR-144-3p, and miR-9-5p) were not significantly different between LBW and AGA newborns. The corresponding results are shown in Supplementary Fig. 2.
a Relative expression of miR-191-3p in umbilical cord samples of appropriate-for-gestational-age (AGA, N = 25) and low birth weight (LBW, N = 24) newborns. Data are mean + interquartile range (CI 95%). P-value is adjusted for newborns sex and maternal smoking. b Receiver operating characteristics (ROC) curve of miR-191-3p for discriminating LBW from AGA newborns. AUC area under the ROC curve. The AUC between 0.75 and 0.9, indicates good discriminatory accuracy.
To assess the predictive value of miR-191-3p for differentiating LBW from AGA newborns, a ROC curve was performed and the AUC was calculated (Fig. 2b). The discriminatory accuracy of circulating miR-191-3p was good with an AUC of 0.76.
Correlations between miRNAs expression and clinical and body composition variables
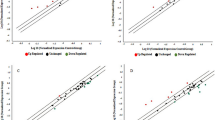
A summary of the correlation coefficient analysis is shown in Fig. 3. The expression of miR-191-3p in umbilical cord positively correlated with birth weight Z-score (R = 0.40; P = 0.03), birth length Z-score (R = 0.34; P = 0.04), and with BMI Z-score at birth (R = 0.38; P = 0.04), with BMC (R = 0.47; P = 0.04) and lean mass at age 15 days (R = 0.42; P = 0.04), and with weight Z-score at age 12 months (R = 0.46; P = 0.02). These correlations were maintained after adjusting for confounding variables such as newborn sex and maternal smoking.
Pearson correlations between circulating miR-191-3p and birth weight Z-score (a), birth length Z-score (b), body mass index (BMI) Z-score (c), bone mineral content (d), and lean mass (e) at age 15 days by dual X-ray absorptiometry, and weight Z-score (f) at age 12 months in appropriate-for-gestational-age (AGA) and in low birth weight (LBW) newborns. White dots represent AGA newborns and black dots correspond to LBW newborns.
In addition, circulating miR-375 was positively related to maternal pre-gestational BMI (R = 0.43; P = 0.006) and with newborns’ fat mass at age 15 days (R = 0.42; P = 0.01). No additional significant correlations between the remaining miRNAs and the study parameters were identified.
Discussion
To our understanding, this is the first study simultaneously profiling miRNAs in pregnant women in the 3rd trimester and in their newborns, and relating those findings to neonatal body composition markers (as judged by DXA). Here, we disclose that miR-191-3p is downregulated in the serum of LBW newborns, and that this miRNA may reliably differentiate LBW from AGA individuals. In addition, circulating miR-191-3p concentrations were found to associate positively with anthropometric measures in the newborn, with lean mass and bone accretion at age 15 days, the latter two markers being known to be reduced in infants with LBW32 and with weight Z-score at age 12 months. Interestingly, circulating levels of miR-375 correlated with maternal pre-gestational BMI and with body fat in the newborns at age 15 days.
Most LBW individuals experience spontaneous catch-up in weight and length starting in the first months of life, which, if excessive, may trigger ectopic fat deposition and ultimately, confer cardiovascular risk.33,34,35 Epigenetic regulation of gene expression by miRNAs may be among the contributors to the variation in the predisposition to follow this track. Along these lines, we have previously identified several miRNAs in umbilical cord tissue associated with LBW children experiencing catch-up growth.17 Even though only three of the differential miRNAs previously described in umbilical cord tissue were also found differentially expressed in the present study in umbilical cord serum (miR-576-5p, miR-873-5p, and miR-222-5p), all miRNAs found differentially expressed in both tissue and serum are involved in similar predicted pathways, including insulin and inflammatory pathways, which are closely related to metabolism regulation. Previous studies have described additional miRNAs as deregulated in LBW newborns;8,13,14,15,16,36,37 however, those studies involved dissimilar types of samples8,13,14,15,16,36 or methods8,13,36 and different participants’ ages.37 Nonetheless, most of the miRNAs detected as differentially expressed in those studies were essentially involved in the same metabolic pathways related to insulin and inflammation.
miR-191 is part of the miR-191/245 cluster located on human chromosome 3 (3p2.31), is widely expressed in human tissues, and encodes for several mature miRNAs, including miR-191-3p.38 Deregulation of miR-191-3p has been widely associated with cancer progression through different pathways, including through enhancement of transcription levels;39 accordingly, miR-191-3p is grouped as being an oncogenic miRNA.40,41 miR-191-3p is also downregulated in patients with hyperlipidemia and in children with overweight/obesity.42,43 Recently, miR-191-3p was found to be under-expressed in mice with cholestatic liver disease and was shown to be regulated by Limb expression 1-like protein (LIX1L), an RNA-binding protein that regulates bile acid metabolism and causes liver injury.44 These data suggest that under-expression of this miRNA might be among the factors that could contribute to the increased metabolic risk in LBW individuals. Moreover, miR-191-3p proved to be a reliable discriminator between LBW and AGA newborns, and thus may become an early biomarker that helps to identify the LBW phenotype.
Gene enriched analysis disclosed that Wnt Family Member 9B (WNT9B) and Inhibitor of Nuclear Factor Kappa B Kinase Regulatory Subunit Gamma (IKBKG) are target genes of miR-191-3p. WNT9B correlated with bone mass accumulation during growth through interaction with the encoding of the low-density lipoprotein receptor-related protein-5 (LRP5), a node of Wnt signaling pathway.45 Thus, down-regulation of mi-191-3p in LBW individuals could, in turn, dysregulate WNT9B and negatively influence bone accretion, accounting at least in part, for the reduced bone mineral content in LBW subjects in the long term.46 Additionally, the Wnt signaling pathway was closely related to the LBW condition,47 obesity, non-alcoholic fatty liver disease (NAFLD), type 2 diabetes, and chronic kidney and cardiovascular disease.48 IKBKG is involved in the activation of the transcription factor nuclear factor-kappa- B (NFκB), which is implicated in inflammatory responses, obesity, and metabolic syndrome.49 Improvement of insulin sensitivity after weight loss attenuates IKBKG expression and NFκB binding activity.50 In addition, IKBKG upregulation may prompt an inflammatory adipocyte population, which in turn, could alter adipocyte differentiation, insulin action, and lipid storage.51 Thus, it is tempting to speculate that mi-191-3p could induce changes in IKBKG expression and the ensuing modifications in the activity of NFκB could be among the mechanisms predisposing LBW individuals to an altered adipogenesis and thus, to metabolic dysfunction.
Although no differences were found in the circulating levels of miR-375 between samples of mothers delivering AGA and LBW newborns, circulating concentrations of this miRNA were found to be associated with maternal pre-gestational BMI. miR-375 is highly expressed in pancreatic beta cells52 and controls beta cell maintenance and insulin secretion.52 miR-375 is downregulated in obese pregnant women, both before and during pregnancy, and is related to an adverse metabolic status in the mother.53 These data suggest that the differential expression of specific miRNAs, including miR-375, can be modified by environmental factors, including lifestyle and nutritional interventions.42 In addition, miR-375 levels were also related to total body fat in both AGA and LBW newborns. Notably, it has been shown that miR-375 is able to promote adipogenesis in animal models via regulation of ERK1/2, which is part of the upstream signaling pathway of peroxisome proliferator-activated receptor-gamma.54 Therefore, it would be interesting to prospectively ascertain whether miR-375 expression follows a different pattern over the first years of life in LBW individuals, and whether this pattern relates to environmental factors, for example, to the type of postnatal feeding. Indeed, insulin-like growth factor I, high-molecular-weight adiponectin, and body composition (by DXA) are among the early markers that diverge in the first year of life between breastfed LBW newborns and formula-fed LBW newborns.32,33
The differentially expressed miRNAs in maternal serum samples, were mainly associated with cell growth regulation and migration in several cancers (miR-382-5p, miR-1307-3p)55,56 and with metabolic disturbances during pregnancy including gestational diabetes (miR-9-5p)57 and pre-eclampsia (miR-144-3p, miR-375).58 Growing evidence has shown that the circulating miRNA profile during pregnancy is influenced by several maternal factors (i.e., maternal nutrition and metabolic perturbations) that have been in turn associated with later impairment in the child’s development and could be used as potential predictors of subsequent children’s health.59
The present work presents several limitations. First, the relatively low number of participants (included in both the discovery and the validation cohort), which may have hindered disclosing statistically significant results. Second, the inclusion of LBW newborns with a birth weight Z-score below −1.5 SD for validation of miRNA profiling, and thus, not strictly small-for-gestational-age.60 Third, the miRNAs selected for the validation did not exceed the FDR cut-off of 0.05. Fourth, the relatively significant rate of drop-outs in the longitudinal follow-up. Further studies in larger cohorts of LBW children with birth weight Z-score strictly SD ≤ −2.0 may be helpful to validate the present findings. Study strengths include the simultaneous assessment of miRNAs profiling in the mothers and newborns, and the longitudinal data availability on body composition and endocrine-metabolic parameters.
In conclusion, in the current study, miR-191-3p was downregulated in the serum of LBW newborns and associated with anthropometric measures and body composition variables at birth and with weight Z-score at age 12 months. In addition, it may have an effect on the expression of target genes involved in bone accretion, adipocyte differentiation, and insulin action. miR-191-3p was reliably different in LBW individuals, and thus could be a new player in the epigenetic mechanism linking LBW and future endocrine-metabolic adverse outcomes. Follow-up of LBW individuals beyond the first year of life will disclose further the potential contribution of miR-191-3p deregulation to these risks.
Data availability
Data generated during the study have been archived in a public repository (BioProject ID: PRJNA747266) and are acceccible at: https://www.ncbi.nlm.nih.gov/bioproject/PRJNA747266.
References
Barker, D. J. et al. Type 2 (non-insulin dependent) diabetes mellitus, hypertension and hyperlipidaemia (syndrome X): relation to reduced fetal growth. Diabetologia 36, 62–67 (1993).
de Zegher, F., Malpique, R., Garcia-Beltran, C. & Ibáñez, L. Towards a simple marker of hepato-visceral adiposity and insulin resistance: the Z-score change from weight-at-birth to BMI-in-childhood. Pediatr. Obes. 14, e12533 (2019).
Lukaszewski, M. A., Eberlé, D., Vieau, D. & Breton, C. Nutritional manipulations in the perinatal period program adipose tissue in offspring. Am. J. Physiol. Endocrinol. Metab. 305, E1195–E1207 (2013).
Rodríguez-Rodríguez, P. et al. Implication of oxidative stress in fetal programming of cardiovascular disease. Front. Physiol. 9, 602 (2018).
Einstein, F. et al. Cytosine methylation dysregulation in neonates following intrauterine growth restriction. PLoS ONE 5, e8887 (2010).
Baek, D. et al. The impact of microRNAs on protein output. Nature 455, 64–71 (2008).
Tang, Q. et al. miR-141 contributes to fetal growth restriction by regulating PLAG1 expression. PLoS ONE 8, e58737 (2013).
Wang, D., Na, Q., Song, W. W. & Song, G. Y. Altered expression of miR-518b and miR-519a in the placenta is associated with low fetal birth weight. Am. J. Perinatol. 31, 729–734 (2014).
Tsochandaridis, M., Nasca, L., Toga, C. & Levy-Mozziconacci, A. Circulating microRNAs as clinical biomarkers in the predictions of pregnancy complications. Biomed. Res. Int. 2015, 294954 (2015).
Deiuliis, J. A. MicroRNAs as regulators of metabolic disease: pathophysiologic significance and emerging role as biomarkers and therapeutics. Int. J. Obes. 40, 88–101 (2016).
Rodosthenous, R. S. et al. Second trimester extracellular microRNAs in maternal blood and fetal growth: an exploratory study. Epigenetics 12, 804–810 (2017).
Kim, S. H. et al. Maternal plasma miRNAs as potential biomarkers for detecting risk of small-for-gestational-age births. EBioMedicine 62, 103145 (2020).
Maccani, M. A., Padbury, J. F. & Marsit, C. J. miR-16 and miR-21 expression in the placenta is associated with fetal growth. PLoS ONE 6, e21210 (2011).
Östling, H., Kruse, R., Helenius, G. & Lodefalk, M. Placental expression of microRNAs in infants born small for gestational age. Placenta 81, 46–53 (2019).
Kennedy, E. M. et al. Placental microRNA expression associates with birthweight through control of adipokines: results from two independent cohorts. Epigenetics 16, 770–782 (2021).
Roxenlund, F., Kruse, R., Östling, H. & Lodefalk, M. Differential microRNA expression in placentas of small-for-gestational age neonates with and without exposure to poor maternal gestational weight gain. J. Perinat. Med. 49, 632–635 (2021).
Mas-Parés, B. et al. Umbilical cord miRNAs in small-for-gestational-age children and association with catch-up growth: a pilot study. J. Clin. Endocrinol. Metab. 104, 5285–5298 (2019).
Nakano, Y. Adult-onset diseases in low birth weight infants: association with adipose tissue maldevelopment. J. Atheroscler. Thromb. 27, 397–405 (2020).
Ferrández-Longas, A. et al. Longitudinal study of normal Spanish children from birth to adulthood: anthropometric, pubertal, radiological and intellectual data. Pediatr. Endocrinol. Rev. 2, 425–642 (2005).
Díaz, M. et al. Placental and cord blood methylation of genes involved in energy homeostasis: association with fetal growth and neonatal body composition. Diabetes 66, 779–784 (2017).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general-purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Díaz, M., Bassols, J., López-Bermejo, A., de Zegher, F. & Ibáñez, L. Low circulating levels of miR-451a in girls with polycystic ovary syndrome: different effects of randomized treatments. J. Clin. Endocrinol. Metab. 105, dgz204 (2020).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Agarwal, V., Bell, G. W., Nam, J. W. & Bartel, D. P. Predicting effective microRNA target sites in mammalian mRNAs. Elife 4, e05005 (2015).
Riffo-Campos, Á. L., Riquelme, I. & Brebi-Mieville, P. Tools for sequence-based miRNA target prediction: what to choose? Int. J. Mol. Sci. 17, 1987 (2016).
Kuleshov, M. V. et al. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res. 44, W90–W97 (2016).
Song, J. et al. Identification of suitable reference genes for qPCR analysis of serum microRNA in gastric cancer patients. Dig. Dis. Sci. 57, 897–904 (2012).
Hu, Z. et al. Serum microRNA profiling and breast cancer risk: the use of miR-484/191 as endogenous controls. Carcinogenesis 33, 828–834 (2012).
Wang, X. et al. Evaluation of the performance of serum miRNAs as normalizers in microRNA studies focused on cardiovascular disease. J. Thorac. Dis. 10, 2599–2607 (2018).
de Zegher, F. et al. Body composition and circulating high-molecular-weight adiponectin and IGF-I in infants born small for gestational age: breast- versus formula-feeding. Diabetes 61, 1969–1973 (2012).
de Zegher, F. et al. Breast-feeding vs formula-feeding for infants born small-for-gestational-age: divergent effects on fat mass and on circulating IGF-I and high-molecular-weight adiponectin in late infancy. J. Clin. Endocrinol. Metab. 98, 1242–1247 (2013).
Fabricius-Bjerre, S. et al. Impact of birth weight and early infant weight gain on insulin resistance and associated cardiovascular risk factors in adolescence. PLoS ONE 6, e20595 (2011).
Wang, S. F. et al. Birth weight and risk of coronary heart disease in adults: a meta-analysis of prospective cohort studies. J. Dev. Orig. Health Dis. 5, 408–419 (2014).
Rodil-Garcia, P., Arellanes-Licea, E. D. C., Montoya-Contreras, A. & Salazar-Olivo, L. A. Analysis of microRNA expression in newborns with differential birth weight using newborn screening cards. Int. J. Mol. Sci. 18, 2552 (2017).
Marzano, F. et al. Pilot study on circulating miRNA signature in children with obesity born small for gestational age and appropriate for gestational age. Pediatr. Obes. 13, 803–811 (2018).
Lagos-Quintana, M., Rauhut, R., Meyer, J., Borkhardt, A. & Tuschl, T. New microRNAs from mouse and human. RNA 9, 175–179 (2003).
Nagpal, N. & Kulshreshtha, R. miR-191: an emerging player in disease biology. Front. Genet. 5, 99 (2014).
Vittoria, M. A. et al. A genome-wide microRNA screen identifies regulators of tetraploid cell proliferation. Mol. Biol. Cell 29, 1682–1692 (2018).
Di Leva, G. et al. Estrogen mediated-activation of miR-191/425 cluster modulates tumorigenicity of breast cancer cells depending on estrogen receptor status. PLoS Genet. 9, e1003311 (2013).
Iacomino, G. et al. Circulating microRNAs are associated with early childhood obesity: results of the I. Family Study. Genes Nutr. 14, 2 (2019).
Xu, J. et al. Several circulating miRNAs related to hyperlipidemia and atherosclerotic cardiovascular diseases. Lipids Health Dis. 18, 104 (2019).
Li, J. et al. Limb expression 1-like (LIX1L) protein promotes cholestatic liver injury by regulating bile acid metabolism. J. Hepatol. 75, 400-413 (2021).
Mo, X. B. et al. Gene-based association analysis identified novel genes associated with bone mineral density. PLoS ONE 10, e0121811 (2015).
Laitinen, J. et al. Body size from birth to adulthood and bone mineral content and density at 31 years of age: results from the northern Finland 1966 birth cohort study. Osteoporos Int. 16, 1417–1424 (2005).
Wang, Y. et al. Differentially expressed circular RNAs in maternal and neonatal umbilical cord plasma from SGA compared with AGA. J. Cell Biochem. 121, 713–722 (2020).
Aamir, K., Khan, H. U., Sethi, G., Hossain, M. A. & Arya, A. Wnt signaling mediates TLR pathway and promote unrestrained adipogenesis and metaflammation: therapeutic targets for obesity and type 2 diabetes. Pharmacol. Res. 152, 104602 (2020).
Baker, R. G., Hayden, M. S. & Ghosh, S. NF-κB, inflammation, and metabolic disease. Cell Metab. 13, 11–22 (2011).
de Mello, V. D. et al. Downregulation of genes involved in NFkappaB activation in peripheral blood mononuclear cells after weight loss is associated with the improvement of insulin sensitivity in individuals with the metabolic syndrome: the GENOBIN study. Diabetologia 51, 2060–2067 (2008).
Yin, Z. et al. Transcriptome analysis of human adipocytes implicates the NOD-like receptor pathway in obesity-induced adipose inflammation. Mol. Cell Endocrinol. 394, 80–87 (2014).
Eliasson, L. The small RNA miR-375 - a pancreatic islet abundant miRNA with multiple roles in endocrine beta cell function. Mol. Cell Endocrinol. 456, 95–101 (2017).
Carreras-Badosa, G. et al. Altered circulating miRNA expression profile in pregestational and gestational obesity. J. Clin. Endocrinol. Metab. 100, E1446–E1456 (2015).
Ling, H. Y. et al. MicroRNA-375 promotes 3T3-L1 adipocyte differentiation through modulation of extracellular signal-regulated kinase signaling. Clin. Exp. Pharmacol. Physiol. 38, 239–246 (2011).
Chen, X., Jiang, C., Sun, R., Yang, D. & Liu, Q. Circular noncoding RNA NR3C1 acts as a miR-382-5p sponge to protect RPE functions via regulating PTEN/AKT/mTOR signaling Pathway. Mol. Ther. 28, 929–945 (2020).
Zheng, Y., Zheng, Y., Lei, W., Xiang, L. & Chen, M. miR-1307-3p overexpression inhibits cell proliferation and promotes cell apoptosis by targeting ISM1 in colon cancer. Mol. Cell Probes 48, 101445 (2019).
Martínez-Ibarra, A. et al. Unhealthy levels of phthalates and bisphenol A in Mexican pregnant women with gestational diabetes and its association to altered expression of miRNAs involved with metabolic disease. Int. J. Mol. Sci. 20, 3343 (2019).
Akgör, U., Ayaz, L. & Çayan, F. Expression levels of maternal plasma microRNAs in preeclamptic pregnancies. J Obstet. Gynaecol. 41, 910–914 (2021).
Franzago, M., Fraticelli, F., Stuppia, L. & Vitacolonna, E. Nutrigenetics, epigenetics and gestational diabetes: consequences in mother and child. Epigenetics 14, 215–235 (2019).
Clayton, P. E. et al. Management of the child born small for gestational age through to adulthood: a consensus statement of the International Societies of Pediatric Endocrinology and the Growth Hormone Research Society. J. Clin. Endocrinol. Metab. 92, 804–810 (2007).
Acknowledgements
C.G.-B. and L.I. are clinical investigators of CIBERDEM (Centro de Investigación Biomédica en Red de Diabetes y Enfermedades Metabólicas Asociadas, Instituto de Salud Carlos III, Madrid, Spain). C.G.-B. is an investigator of the PERIS-AQU program, Generalitat de Catalunya, Spain. G.C.-B. is an Investigator of the Sara Borrell Fund of Scientific Research from Carlos III National Institute of Health (CD19-00172). J.B. is an investigator of the Miguel Servet Fund from Carlos III National Institute of Health (CPII17/00013). A.L.-B. is a Clinical Investigator of the I3 Fund for Scientific Research (Ministry of Science and Innovation, Spain).
Funding
This study was supported by the grant BR201601 from the Research Institute Sant Joan de Déu and by grant PERIS-SLT017/20/000228 from the AQU, Generalitat de Catalunya, Spain.
Author information
Authors and Affiliations
Contributions
C.G.-B. contributed to literature search, the layout of Figures and Tables, collection, analysis and interpretation of data, and wrote the manuscript. G.C.-B. contributed to the collection, analysis, and interpretation of data. J.B. contributed to the design of the study, collection, analysis and interpretation of data, literature search, and wrote the manuscript. R.M. contributed to the design of the study, collection, analysis, and interpretation of data. C.P. contributed collection, analysis, and interpretation of data. F.d.Z. contributed to analysis and interpretation of data, and revision/edition of the manuscript. A.L.-B. contributed to analysis and interpretation of data, and revision/edition of the manuscript. L.I. contributed to the design of the study, literature search, analysis and interpretation of data, wrote the manuscript, and revision/edition of the manuscript. L.I. is the guarantor of this work and, as such, had full access to all the data in the study and takes responsibility for the integrity of the data and the accuracy of the data analysis. Each author listed in the manuscript has seen and approved the submission of this version of the manuscript and takes full reasonability for the contents.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
Written informed consent was acquired from the mothers at recruitment. The study was approved by the Institutional Review Board of the University of Barcelona, Sant Joan de Déu Hospital.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Garcia-Beltran, C., Carreras-Badosa, G., Bassols, J. et al. microRNAs in newborns with low birth weight: relation to birth size and body composition. Pediatr Res 92, 829–837 (2022). https://doi.org/10.1038/s41390-021-01845-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41390-021-01845-4
This article is cited by
-
Epigenetic Responses to Nonchemical Stressors: Potential Molecular Links to Perinatal Health Outcomes
Current Environmental Health Reports (2024)
-
The effect of MicroRNAs variants on idiopathic recurrent pregnancy loss
Journal of Assisted Reproduction and Genetics (2023)