Abstract
Background
Prematurity is a severe pathophysiological condition, however, little is known about the gestational age-dependent development of the neonatal metabolome.
Methods
Using an untargeted liquid chromatography-tandem mass spectrometry metabolomics protocol, we measured over 9000 metabolites in 298 neonatal residual heel prick dried blood spots retrieved from the Danish Neonatal Screening Biobank. By combining multiple state-of-the-art metabolome mining tools, we retrieved chemical structural information at a broad level for over 5000 (60%) metabolites and assessed their relation to gestational age.
Results
A total of 1459 (~16%) metabolites were significantly correlated with gestational age (false discovery rate-adjusted P < 0.05), whereas 83 metabolites explained on average 48% of the variance in gestational age. Using a custom algorithm based on hypergeometric testing, we identified compound classes (617 metabolites) overrepresented with metabolites correlating with gestational age (P < 0.05). Metabolites significantly related to gestational age included bile acids, carnitines, polyamines, amino acid-derived compounds, nucleotides, phosphatidylcholines and dipeptides, as well as treatment-related metabolites, such as antibiotics and caffeine.
Conclusions
Our findings elucidate the gestational age-dependent development of the neonatal blood metabolome and suggest that the application of metabolomics tools has great potential to reveal novel biochemical underpinnings of disease and improve our understanding of complex pathophysiological mechanisms underlying prematurity-associated disorders.
Impact
-
A large variation in the neonatal dried blood spot metabolome from residual heel pricks stored at the Danish Neonatal Screening Biobank can be explained by gestational age.
-
While previous studies have assessed the relation of selected metabolic markers to gestational age, this study assesses metabolome-wide changes related to prematurity. Using a combination of recently developed metabolome mining tools, we assess the relation of over 9000 metabolic features to gestational age.
-
The ability to assess metabolome-wide changes related to prematurity in neonates could pave the way to finding novel biochemical underpinnings of health complications related to preterm birth.
Similar content being viewed by others
Introduction
Prematurity is a complex and challenging pathophysiological condition associated with increased morbidity and mortality.1,2 Numerous early- and late-onset disorders are associated with preterm birth, including psychiatric disorders.3 The risk of complications is inversely correlated to the gestational age at birth but with large variations within age groups. It is likely that the individual degree of metabolic prematurity more accurately represents the complication risk. Metabolomic analyses could allow for an individual assessment of multiorgan function and maturity, and may thus contribute to improved understanding of pathophysiological mechanisms behind the development of prematurity-associated disorders. However, currently, there is limited knowledge on the metabolomic profile of preterm neonates4,5,6 and on how to assess metabolic maturity.
Growing evidence suggests a strong link between early-life microbiota and disease,7 as well as short- and as long-term complications associated with preterm birth, such as necrotizing enterocolitis, diabetes, cardiovascular disease, neurodevelopmental disorders and neuropsychiatric disorders have been related to the underdevelopment of the gut microbiota, gastrointestinal tract and immune system.3,8,9,10,11,12,13 Although the exact timing of the establishment of the intestinal microbiome in human life remains unknown,14 it is generally agreed upon that the gut microbial colonization starts at birth at the latest and undergoes shifts in composition and structure as the host matures over time.15,16,17,18,19
Recent studies have shown that marker metabolites of microbial metabolism are readily detectable in human blood and that the human blood metabolome may predict gut bacterial α-diversity.20 However, gut microbiome-derived metabolites that become available to the preterm infant and may impact gut maturation and overall host metabolism and health remain unknown.
Monitoring gut microbial health and the metabolic degree of prematurity during newborn screening could offer a powerful tool for the early detection and possible early intervention in prematurity-associated disease progression through probiotics, diet or microbial transplants. Dried blood spots (DBS) routinely collected for newborn screening are minimally invasive and metabolomics approaches enable the simultaneous measurement of thousands of metabolites, thus offering unique insights into metabolic underpinnings of complex pathophysiological conditions.21,22
Here, we hypothesize that metabolic gestational age-dependent degree of prematurity, which impacts overall host metabolism and health, may be monitored during newborn screening. Using DBS retrieved from the Danish Neonatal Screening Biobank in combination with recently developed computational metabolomics tools, including mass spectral molecular networking, unsupervised substructure discovery and in silico structure annotation,23,24,25,26 we assess the gestational age-dependent development of the blood metabolome of very preterm to term neonates.
Methods
Study cohort
Residual extracts from the Danish Newborn Screening programme comprising all preterm (28–36 weeks) and randomly selected term to late-term births (37–42 weeks) were collected during the period from March to June 2016 and stored at −20 °C until analysis. All original DBS samples were collected 48–72 h after birth. Repeat analyses (i.e., DBS collected after 72 h after birth) were not included. The samples were grouped per gestational age in weeks at birth, resulting in a cohort of a total of 298 newborns (133 girls) with gestational ages from 28 to 42 weeks (n = 8–68 each) (Table 1). The study was conducted in accordance with the Declaration of Helsinki and the protocol complies with the Danish Ethical Committee law by not being a health research project (Section 2,1), but a method development study not requiring an ethical approval.27
Metabolomic profiling
All samples (including blank and pooled quality control samples) were submitted to untargeted metabolomic profiling using liquid chromatography-tandem mass spectrometry (LC-MS/MS) at Statens Serum Institut, Copenhagen, Denmark between July 6, 2016 and July 14, 2016. Raw data files were preprocessed using MZmine (version 2.40.1).28 A detailed description of LC-MS/MS as well as preprocessing parameters can be found in the Supplementary Methods.
Statistical analyses
Overall variation in the metabolome related to gestational age was assessed using a principal coordinates analysis (PCoA) plot with the Bray–Curtis dissimilarity. A permutational multivariate analysis of variance (PERMANOVA)29 model was fitted to the Bray–Curtis distance matrix to assess the variation in the metabolome explained by gestational age. To estimate gestational age from the neonatal metabolome, we used a tenfold cross-validation (CV) implementation of the least absolute shrinkage and selection operator (LASSO) method, including comparing its performance to a Ridge regression, such as that described in Wilmanski et al.20 Metabolite richness and α-diversity was assessed using the mean number of metabolites and Shannon index, respectively, measured per sample and stratified by categories of prematurity and gestational age. Subsequently, a Kruskal–Wallis test was used to compare mean metabolite richness and α-diversity per prematurity category, whereas correlation between metabolite richness and gestational age was evaluated using Kendall’s τ.30
Univariate correlation at the individual metabolite level was assessed using Kendall’s τ and P values were adjusted for multiple hypothesis testing using the false discovery rate (FDR) method.31 To equalize the statistical power for the univariate correlation analysis, we randomly subsampled 10 individuals per week of gestational age, with exception of week 30, for which we only had eight samples available. Within this subsample, mass spectral features for which FDR-adjusted P < 0.05 were found. A molecular network was created, and molecular families enriched for significant features were detected (hypergeometric test, see Supplementary Methods). To demonstrate that our conclusions are not sensitive to the particular subsample, we repeated the univariate analysis across 1000 subsamples and explored the spectral relationships across the entire sample set. Specifically, for each feature we assessed whether it appeared more often than would be expected, considering a feature significant if its frequency of appearance had a probability below 5% of occurring by chance (≥130 times based upon 11% features being chosen, on average, from each subsample, for more details, see Supplementary Methods). To check whether the same relationships between correlation and spectral similarity are recovered when considering all 1000 subsamples, we identified features whose spectral neighbourhood (cosine similarity ≥ 0.7) was enriched for features identified as significant across the subsamples as described above (hypergeometric test, for more details, see Supplementary Methods). Additional information is provided in the Supplementary Methods. All statistical analyses were performed in R 3.6.132 or Python 3.7.33 All Jupyter notebooks used for statistical analysis are publicly available at: https://github.com/madeleineernst/Prematurity_SupplementaryMaterial.
Metabolite identification
Aggregated MZmine preprocessed MS/MS fragmentation spectra were submitted to feature-based mass spectral molecular networking through the Global Natural Products Social Molecular Networking Platform (GNPS)24,34,35 and searched against all GNPS spectral libraries. To further enhance chemical structural information within the network, substructure information was incorporated using the GNPS MS2LDA workflow (https://ccms-ucsd.github.io/GNPSDocumentation/ms2lda/).23 Information from in silico structure annotations from Network Annotation Propagation25 and Dereplicator36 were incorporated using the GNPS MolNetEnhancer workflow (https://ccms-ucsd.github.io/GNPSDocumentation/molnetenhancer/)26 with chemical class annotations retrieved from the ClassyFire chemical ontology.37 A detailed description of all workflow parameters can be found in the Supplementary Methods. Mass spectral molecular network data, data from MS2LDA unsupervised substructure discovery, and in silico structure annotation are available upon request.
Results
A total of 9010 metabolites (mass spectral features with unique MS/MS fragmentation patterns) were measured. Using a combination of metabolome mining tools, including mass spectral molecular networking (GNPS), unsupervised substructure discovery (MS2LDA) and in silico annotation through the MolNetEnhancer workflow,26 putative chemical structural information at the chemical class level, corresponding to a level 3 metabolite identification according to the Metabolomics Standard Initiative’s reporting standards38 could be retrieved for over 60% (5687) of the detected metabolites (Supplementary Fig. 1).
PCoA and permutational analysis of variance demonstrated that 2.6% of the variation in the metabolomics data could be explained by gestational age (PERMANOVA, P < 0.05, Adonis R2 = 0.026) with strongest separation observed along PCo3 (Fig. 1a).
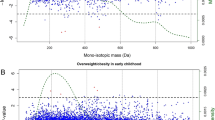
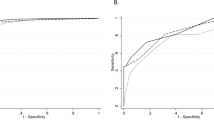
a Principal coordinates analysis using the Bray–Curtis distance metric. Variation of 2.6% in the data is explained by gestational age (PERMANOVA, P < 0.05, Adonis R2 = 0.026). b Metabolome-estimated gestational age versus observed, ultrasound-guided gestational age. Mean R2 across ten cross-validations, Pearson’s correlation coefficient and P value are shown. c Box plots for the number of mass spectral features stratified across different prematurity categories. Significant differences were found across mean number of mass spectral features per prematurity category (very preterm: 28–32 weeks; near term: 33–36 weeks; term: 37–40 weeks; late term: 41–42 weeks) (Kruskal–Wallis, P < 0.05). d Venn diagram illustrating overlapping metabolites significantly associated with gestational age by univariate correlation analysis, hypergeometric testing at the molecular family level and LASSO regression.
The LASSO model suggested that a total of 83 metabolites give a strong estimate of gestational age, explaining an average of 48% of the variance (mean out-of-sample R2 = 0.48, Pearson’s r = 0.73, P = 2.4 × 10−50 for metabolome-estimated gestational age versus observed, ultrasound-guided gestational age) (Fig. 1b and Supplementary Results). Out of these 83 metabolites, four could be matched to GNPS library spectra with a mass spectral similarity score (cosine score) of ≥0.9, including isoleucine–lysine, isoleucine, ophthalmic acid and N1-acetylspermine. Six metabolites were selected in all tenfold CV models and were the most influential in estimating gestational age, whereas 83 metabolites were retained by at least one model. The chemical structural information could be retrieved for one of the six metabolites selected by all tenfold CV models, N1-acetylspermine. No chemical structural information could be retrieved for the remaining five metabolites selected in all tenfold CV models. However, by comparing mass spectral fragmentation spectra to data in the public domain,39 we found that one of the five metabolites was previously found in human sputum samples of patients with cystic fibrosis undergoing antibiotic treatment, faecal samples of children and the surface of a tomato plant. This could suggest that the unknown structure is of (anti)microbial nature. A further unknown metabolite was previously found in diverse human plasma, skin and faecal samples.
Mean metabolite richness was found to vary significantly with different categories of prematurity (Kruskal–Wallis, P < 0.05), with higher numbers of metabolites observed for very preterm versus term neonates (Fig. 1c). Furthermore, mean metabolite richness was found to correlate significantly with gestational age (Kendall’s τ = −0.12, P < 0.05) with more metabolites observed in neonates born at 28 weeks of gestation (Supplementary Fig. 2). Within-sample metabolite diversity (Shannon index), on the other hand, was only found to vary marginally significantly with different categories of prematurity, with higher metabolite diversity observed in near-term and term born children when compared to very-preterm and late-term children (Kruskal–Wallis, P = 0.045) (Supplementary Fig. 3). Within-sample metabolite diversity was not found to vary significantly between preterm (<37 weeks) and term (>37 weeks) born children (Wilcoxon’s rank-sum test, P = 0.81).
In the univariate analyses, 1459 metabolites (~16%) were found to significantly correlate with gestational age (FDR-adjusted P < 0.05, in ≥130 subsamples). Out of these 1459 metabolites, 74 could be chemically structurally annotated through GNPS library matching, manual or in silico annotation propagation throughout the mass spectral molecular network, including amino acid derivatives, carbohydrates (sugars), dipeptides, lipids (bile acids, carnitines and phospholipid catabolites), nucleotides, polyamines (N1-acetylspermine, spermine, spermidine and structural analogues) and xenobiotics (caffeine, acetaminophen, antibiotic-derived metabolites, including penicillamine disulfide, ampicillin and cefuroxime, as well as compounds related to chlorhexidine, a common disinfectant) (Supplementary Data 2). Of the 14 amino acid derivatives significantly correlating with gestational age, eight were found to possibly be related through the histidine, tryptophan or phenylalanine and tyrosine metabolic pathways,40,41,42,43 respectively (Supplementary Data 2 and Fig. 2). Seven amino acids or catabolites were found to correlate positively with gestational age (cis-urocanate/urocanate, glutamine, glutamic acid, histidine, ornithine, serine and valine), whereas seven were found to correlate negatively with gestational age (3-ureidopropionic acid, imidazole propionate, isoleucine, kynurenic acid, methionine, phenylacetylglutamine and quinaldic acid). All peptides and chlorhexidine structural analogues were found to increase with gestational age, whereas antibiotics, caffeine, bile acids (except for cholic and hyocholic acid), carnitines, most nucleotide structural analogues, polyamines and sugars were found to decrease with gestational age. Ophthalmic acid and 4-hydroxynonenal, two potential indicators of oxidative stress,44,45,46 were found to increase and decrease with gestational age, respectively.
Molecules highlighted in red are positively correlated with gestational age, whereas molecules highlighted in blue are negatively correlated with gestational age (Kendall’s τ, FDR-adjusted P < 0.05, in ≥130 subsamples). Molecules more abundant in preterm neonates (negative correlation with gestational age) have previously been associated with health complications related to prematurity, such as cardiovascular diseases, diabetes and cognitive impairment, and in two out of three cases may be mediated through gut microbiota. Metabolite annotations for glutamine, phenylacetylglutamine, histidine, (cis)-urocanate, glutamate, tryptophan, kynurenine and kynurenic acid were retrieved from GNPS with a spectral similarity score (cosine score) ≥0.9, corresponding to a level 2 metabolite identification according to the Metabolomics Standard Initiative’s reporting standards,38 whereas chemical structural information of imidazole propionate and quinaldic acid was based on parent mass and substructure information retrieved from MS2LDA.
A total of 17 molecular families comprising 230 metabolites were found to be significantly overrepresented (P < 0.01) with metabolites correlating with gestational age (non-FDR-adjusted P < 0.01) by hypergeometric testing in a randomly selected subset of 148 samples with 10 samples per gestational week selected (except for week 30, where only eight samples were available) (Fig. 3). Chemical structure annotation could be retrieved for six molecular families (152 metabolites) and revealed that carnitine families mostly correlated positively with gestational age, whereas nucleotide, bile acid and spermine-related families correlated negatively with gestational age (Fig. 3, Supplementary Figs. 4–8). A family of structural analogues of penicillamine, likely a degradation product of penicillin, was found to correlate negatively with gestational age, while structural analogues of chlorhexidine, a common disinfectant, were found to correlate positively with gestational age. Comparable compound classes were also retrieved when looking across all samples. A total of 617 metabolites in the mass spectral molecular network of the full dataset (298 samples) had significantly more neighbouring metabolites with a spectral similarity score (cosine score) >0.7 significantly correlating with gestational age (FDR-adjusted P < 0.05, in ≥130 subsamples) than would be expected by chance (P < 0.05). These metabolites included amino and bile acids, carnitines, dipeptides, nucleotides, phosphatidylcholine-derived compounds, polyamines and xenobiotics (ampicillin, penicillamine disulfide and chlorhexidine) (Supplementary Data 2). Six metabolites (four unknown metabolites, N1-acetylspermine and a carnitine structural analogue) were identified by all three statistical approaches to be significantly associated with gestational age (Fig. 1d). Figure 4 shows a summary of a total of 53 mass spectral features significantly associated with gestational age either by univariate correlation analysis, hypergeometric testing at the molecular family level or LASSO regression, for which metabolite annotations could be retrieved from GNPS with a spectral similarity (cosine) score ≥0.9, corresponding to a level 2 metabolite identification according to the Metabolomics Standard Initiative’s reporting standards.38
Node colours represent correlation with gestational age (Kendall’s τ). Metabolites for which chemical structural annotation could be retrieved are indicated with grey shadowing. The thickness of the lines connecting the nodes represents tandem mass spectral similarity, implying high chemical structural similarity. Nodes with bold black borders indicate GNPS spectral library hits. Carnitines, bile acids, nucleotides and spermine were previously reported to be implicated in the maturation of the gastrointestinal tract or being affected by the gut microbiome. Penicillamine could be reflective of antibiotics use and chlorhexidine, a common skin disinfectant reflective of different sampling strategies across term and preterm neonates.
For illustration, a subset of all samples is shown, with 10 subsamples per week of gestational age selected (except for week 30, n = 8). Metabolite annotations retrieved from GNPS with a cosine score ≥0.9 are shown, corresponding to a level 2 metabolite identification according to the Metabolomics Standard Initiative’s reporting standards.38
Discussion
There is currently limited knowledge on the metabolomic status of preterm neonates and a deeper understanding hereof will help further elucidate the complex pathophysiology of prematurity.
In this methodological study, tools deployed on untargeted metabolomics data from newborn DBS were demonstrated to have great potential to address future research questions. While a number of studies have assessed variation of a few selected metabolites with gestational age (e.g. refs. 47,48,49,50) this study assesses full metabolome-wide changes of several thousand of metabolites related to prematurity in dried blood spots from newborn screening.5,51
We found that the metabolome of 2–3-day-old neonates is highly reflective of gestational age. Using statistical modelling, univariate correlation analysis at the individual metabolite level and hypergeometric testing at the molecular family level, we found that 83, 1459 and 617 metabolites, respectively, were significantly associated with gestational age.
On average only 2–5% of metabolites can be chemically structurally annotated in untargeted LC-MS/MS-based metabolomics studies.52 Using a combination of different computational metabolomics tools, we were able to retrieve chemical structural information at a broad level for nearly 60% of the data collected, thus representing a major advance in biochemical interpretation. Furthermore, evaluating changes at the level of chemically structurally related molecular families (rather than individual metabolites) through hypergeometric testing is a novel approach, which allows us to understand metabolic changes of groups of metabolites changing consistently, but modestly across samples. This approach increases the chance for novel discoveries and a potentially better understanding of pathobiological processes as these metabolites would be missed in univariate approaches.
We identified amino acid-derived metabolites, dipeptides, polyamines, nucleotides, lipids (bile acids, carnitines and phosphatidylcholine-derived compounds), sugars and treatment-related compounds, such as penicillamine disulfide, cefuroxime and caffeine as significantly correlating with gestational age. Similarly, LASSO regression identified peptides, a polyamine and an amino acid (isoleucine) as most influential in estimating gestational age. Hypergeometric testing at the molecular family level additionally revealed that carnitine, bile acid, polyamine, nucleotide, antibiotic, phosphatidylcholine-derived compounds and chlorhexidine molecular families are most strongly associated with gestational age.
Our findings are in agreement with previous studies reporting significant differences across premature and term born children in carnitine and amino acid profiles.47,48,49,50 In addition, we highlight many other metabolite classes associated with gestational age, which in future studies may contribute to a better biochemical understanding of prematurity and pathophysiological mechanisms behind the development of prematurity-associated disorders.
Some of the metabolites highlighted in our study have, for example, previously been reported to be implicated by the gut microbiome (amino and bile acids, carnitines and phosphatidylcholine-derived compounds), involved in gut maturation (polyamines) or affecting gut microbial composition (antibiotics and diet-derived nucleotides)40,41,42,43,53,54,55,56,57 (Fig. 2 and Supplementary Discussion). In very recent studies, some of these microbiome-derived metabolites have been shown to be associated with a diverse range of pathophysiologies related to preterm birth (Fig. 2). There is a well-established increased risk for early- and also late-term complications to being born prematurely, so we can intuitively assume that risk biomarkers may be present in early-life samples in prematurely born children. In our study, we identified increased relative amounts of some microbial catabolites in preterm children who have been linked to well-known late-term complications of prematurity, such as phenylacetylglutamine (associated with increased risk for cardiovascular disease40,58,59) or imidazole propionate (associated with type 2 diabetes.42,60) Similarly, we found that kynurenic acid, a catabolite of tryptophan metabolism, is negatively correlated with gestational age, with higher relative abundances observed in preterm infants. Gut microbiota were shown to play an important role in tryptophan metabolism41,53 and high levels of kynurenic acid in the central nervous system have been associated with schizophrenia and cognitive impairment.61 Corroborating with these findings, previous studies have found tryptophan and bile acid metabolic pathways affected by gestational age in urine metabolomics samples of children at week 4 of age.62
Similarly, ophthalmic acid and 4-hydroxynonenal may be biomarkers of oxidative stress.44,45,46 Interestingly, ophthalmic acid was here found to be positively associated with gestational age, whereas 4-hydroxynonenal was found to be negatively associated with gestational age with the highest relative abundance observed in prematurely born children. Elevated levels of 4-hydroxynonenal have been proposed to be implicated in pathophysiological processes of a number of diseases, including cancer, diabetes, cardiovascular, inflammatory and neurodegenerative complications.46
Although this was a methodological study and not designed to establish a direct causal or pathophysiological link between a specific marker and a later occurring complication, we may speculate that the observed difference in the neonatal metabolome may reflect an increased risk of early- or late-term complications on an individual level.
Common clinical practice, such as antibiotics, use seems furthermore reflected in the neonatal metabolome and significantly related to gestational age. Antibiotic-related metabolites (penicillamine disulfide and cefuroxime) and caffeine correlated negatively with gestational age, which reflects that preterm neonates are at increased risk of infection and often require broad-spectrum antibiotics from birth onwards.8 Caffeine is the most commonly used medication for the treatment of apnoea of prematurity.63 Antibiotics are known to have profound effects on gut microbial communities, and modified metabolic activity of the antibiotic-altered gut microbiome has been shown to result in decreased faecal levels of dipeptides and increased levels of primary bile acids and sugar alcohols in mice.57 In agreement with this finding, we here observed decreased blood levels of dipeptides and increased levels of primary bile acids and sugars, which could possibly result from an increased antibiotic use in prematurely born children. Similarly, gestational age-dependent differential abundance of nucleotides or phosphatidylcholine-derived compounds among others may be reflective of differential feeding patterns. Preterm neonates are often fed parenterally, while term neonates are more likely to be breastfed.
Overall, we found more metabolites in preterm neonates when compared to term neonates. Within-sample metabolite diversity, on the other hand, was found to differ only marginally across different categories of prematurity. Increased metabolite richness in preterm neonates could be reflective of increased medication in prematurely born children. Alternatively, it is known that intestinal permeability is higher in preterm compared to term neonates.64 Increased metabolite richness could thus also be reflective of increased intestinal permeability. The relatively constant within-sample metabolite diversity across different categories of prematurity would be in agreement with this hypothesis (more versus more diverse metabolites).
Our data reveal that metabolomic profiling of neonatal DBS in combination with recently developed computational metabolome mining methods offers a powerful tool for monitoring gestational age-dependent metabolic degree of prematurity in neonates and may contribute to an improved biochemical understanding of pathophysiological mechanisms behind the development of prematurity-associated disorders. Some of the metabolites here identified as significantly related to gestational age have previously been described to be related to the gut microbiome and short- as well as long-term health complications related to preterm birth. This finding is suggestive that catabolites of microbial metabolism are detectable in the neonatal blood as early as 2–3 days of life, and untargeted DBS metabolomic analyses in combination with computational tools applied here may offer a powerful tool to decipher complex biochemical underpinnings of prematurity-associated disorders.
Limitations
This study draws strength from being based on a cohort of samples retrieved from the Danish National Biobank and thus being collected prospectively as part of the National Newborn Screening Programme. Therefore, a systematic inclusion bias is not likely to have influenced the study. Although we detected a large diversity of metabolites, metabolites extracted and detected in our study are inherently reflective of metabolites targeted during neonatal screening of inborn diseases. Furthermore, only information provided on each newborn screening specimen, including gender, gestational age in weeks, age at sampling, multiplicity and mother’s age were available to us in this methodological study. Data on other covariates, such as delivery mode, parenteral nutrition, maternal characteristics or later development of disease, were not available. A multi-omics approach and extensive prospective sampling would be needed to establish a direct link between metabolic markers and later development of prematurity-associated disorders. Maternal disorders and maternal medication that may pass the placenta barrier could potentially impact the results, and comorbidities such as being born small for gestational age may equally be a confounding factor.65,66 More samples would allow for more statistical power, whereas an independent test and training data set would allow for generalization of the findings outside this study. Lastly, all metabolite identifications described here are putative, corresponding to a level 2–3 identification according to the Metabolomics Standard Initiative’s reporting standards. Further studies would be needed for unambiguous chemical structural identification. Although putative chemical structural information could be retrieved from a significant amount of metabolic features in this study, chemical structural information of many features related to gestational age remains unknown. Further development in the field of computational metabolomics, and increasing contributions to community-based metabolomics platforms such as GNPS will play a key role in improving our understanding of biochemical underpinnings of diverse pathophysiologies in the coming years.
Conclusions
Our data demonstrate that the neonatal metabolome is strongly reflective of gestational age. Some of the metabolites here found to be significantly associated with gestational age have previously been related to the gut microbiome or maturation and short- as well as long-term complications of preterm birth. This finding suggests that catabolites of microbial metabolism are detectable in the neonatal blood as early as 2–3 days of life, and neonatal dried blood metabolomics may offer a powerful tool to decipher complex biochemical underpinnings of prematurity-associated disorders. We show that metabolomic profiling of neonatal DBS in combination with recently developed computational metabolome mining methods offers a powerful tool for monitoring metabolic maturation in preterm neonates. Further studies will be needed to establish direct causal or pathophysiological links between marker metabolites and later occurring disorders related to preterm birth.
References
Saigal, S. & Doyle, L. W. An overview of mortality and sequelae of preterm birth from infancy to adulthood. Lancet 371, 261–269 (2008).
Blencowe, H. et al. National, regional, and worldwide estimates of preterm birth rates in the year 2010 with time trends since 1990 for selected countries: a systematic analysis and implications. Lancet Lond. Engl. 379, 2162–2172 (2012).
Nosarti, C. et al. Preterm birth and psychiatric disorders in young adult life. Arch. Gen. Psychiatry 69, 610-617 (2012).
Atzori, L., Antonucci, R., Barberini, L., Griffin, J. L. & Fanos, V. Metabolomics: a new tool for the neonatologist. J. Matern. Fetal Neonatal Med J. Eur. Assoc. Perinat. Med Fed. Asia Ocean Perinat. Soc. Int. Soc. Perinat. Obstet. 22(Suppl. 3), 50–53 (2009).
Carter, R. A., Pan, K., Harville, E. W., McRitchie, S. & Sumner, S. Metabolomics to reveal biomarkers and pathways of preterm birth: a systematic review and epidemiologic perspective. Metabolomics 15, 124 (2019) https://doi.org/10.1007/s11306-019-1587-1.
Fettweis, J. M. et al. The vaginal microbiome and preterm birth. Nat. Med. 25, 1012–1021 (2019).
Tamburini, S., Shen, N., Wu, H. C. & Clemente, J. C. The microbiome in early life: implications for health outcomes. Nat. Med. 22, 713–722 (2016).
Henderickx, J. G. E., Zwittink, R. D., van Lingen, R. A., Knol, J. & Belzer, C. The preterm gut microbiota: an inconspicuous challenge in nutritional neonatal care. Front. Cell Infect. Microbiol. 9, 85 (2019).
Robertson, C. et al. Incidence of necrotising enterocolitis before and after introducing routine prophylactic Lactobacillus and Bifidobacterium probiotics. Arch. Dis. Child Fetal Neonatal Ed. fetalneonatal-2019-317346 (2019).
Kolevzon, A., Gross, R. & Reichenberg, A. Prenatal and perinatal risk factors for autism: a review and integration of findings. Arch. Pediatr. Adolesc. Med. 161, 326–333 (2007).
Sharon, G. et al. Human gut microbiota from autism spectrum disorder promote behavioral symptoms in mice. Cell 177, 1600–1618.e17 (2019).
Bastiaanssen, T. F. S., Cowan, C. S. M., Claesson, M. J., Dinan, T. G. & Cryan, J. F. Making sense of … the microbiome in psychiatry. Int. J. Neuropsychopharmacol. 22, 37–52 (2019).
Lu, J. & Claud, E. C. Connection between gut microbiome and brain development in preterm infants. Dev. Psychobiol. 61, 739–751 (2019).
Romano-Keeler, J. & Weitkamp, J.-H. Maternal influences on fetal microbial colonization and immune development. Pediatr. Res. 77, 189–195 (2015).
Ma, B. et al. Microbial biomarkers of intestinal barrier maturation in preterm infants. Front. Microbiol. 9, 2755 (2018).
de Goffau, M. C. et al. Human placenta has no microbiome but can contain potential pathogens. Nature 572, 329–334 (2019).
Aagaard, K. et al. The placenta harbors a unique microbiome. Sci. Transl. Med. 6, 237ra65–237ra65 (2014).
Korpela, K. et al. Intestinal microbiota development and gestational age in preterm neonates. Sci. Rep. 8, 1–9 (2018).
La Rosa, P. S. et al. Patterned progression of bacterial populations in the premature infant gut. Proc. Natl Acad. Sci. USA 111, 12522–12527 (2014).
Wilmanski, T. et al. Blood metabolome predicts gut microbiome α-diversity in humans. Nat. Biotechnol. 37, 1217–1228 (2019).
Quinn, R. A. et al. Molecular networking as a drug discovery, drug metabolism, and precision medicine strategy. Trends Pharm. Sci. 38, 143–154 (2017).
Freeman, J. D. et al. State of the science in dried blood spots. Clin. Chem. 64, 656–679 (2018).
van der Hooft, J. J. J., Wandy, J., Barrett, M. P., Burgess, K. E. V. & Rogers, S. Topic modeling for untargeted substructure exploration in metabolomics. Proc. Natl Acad. Sci. USA 113, 13738–13743 (2016).
Wang, M. et al. Sharing and community curation of mass spectrometry data with Global Natural Products Social Molecular Networking. Nat. Biotechnol. 34, 828–837 (2016).
da Silva, R. R. et al. Propagating annotations of molecular networks using in silico fragmentation. PLoS Comput. Biol. 14, e1006089 (2018).
Ernst, M. et al. MolNetEnhancer: enhanced molecular networks by integrating metabolome mining and annotation tools. Metabolites 9, 144 (2019).
Nørgaard-Pedersen, B. & Hougaard, D. M. Storage policies and use of the Danish Newborn Screening Biobank. J. Inherit. Metab. Dis. 30, 530–536 (2007).
Pluskal, T., Castillo, S., Villar-Briones, A. & Orešič, M. MZmine 2: modular framework for processing, visualizing, and analyzing mass spectrometry-based molecular profile data. BMC Bioinform. 11, 395 (2010).
Anderson, M. J. A new method for non-parametric multivariate analysis of variance: non-parametric manova for ecology. Austral. Ecol. 26, 32–46 (2001).
Kendall, M. G. A new measure of rank correlation. Biometrika 30, 81–93 (1938).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B 57, 289–300 (1995).
R Core Team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, Vienna, 2019).
Van Rossum, G. & Drake, F. L. Python Reference Manual (PtyhonLabs, Virginia, 2001).
Watrous, J. et al. Mass spectral molecular networking of living microbial colonies. Proc. Natl Acad. Sci. USA 109, E1743–E1752 (2012).
Nothias, L. F. et al. Feature-based Molecular Networking in the GNPS Analysis Environment. Nat. Methods 17, 905–908 (2020).
Mohimani, H. et al. Dereplication of peptidic natural products through database search of mass spectra. Nat. Chem. Biol. 13, 30–37 (2017).
Djoumbou Feunang, Y. et al. ClassyFire: automated chemical classification with a comprehensive, computable taxonomy. J. Cheminformatics 8, 61 (2016).
Sumner, L. W. et al. Proposed minimum reporting standards for chemical analysis: Chemical Analysis Working Group (CAWG) Metabolomics Standards Initiative (MSI). Metabolomics 3, 211–221 (2007).
Wang, M. et al. Mass spectrometry searches using MASST. Nat. Biotechnol. 38, 23–26 (2020).
Bogiatzi, C. et al. Metabolic products of the intestinal microbiome and extremes of atherosclerosis. Atherosclerosis 273, 91–97 (2018).
Dehhaghi, M., Kazemi Shariat Panahi, H. & Guillemin, G. J. Microorganisms, tryptophan metabolism, and kynurenine pathway: a complex interconnected loop influencing human health status. Int. J. Tryptophan Res. 12, https://doi.org/10.1177/1178646919852996 (2019).
Koh, A. et al. Microbially produced imidazole propionate impairs insulin signaling through mTORC1. Cell 175, 947–961.e17 (2018).
Wang, Z. et al. Gut flora metabolism of phosphatidylcholine promotes cardiovascular disease. Nature 472, 57–63 (2011).
Soga, T. et al. Differential metabolomics reveals ophthalmic acid as an oxidative stress biomarker indicating hepatic glutathione consumption. J. Biol. Chem. 281, 16768–16776 (2006).
Zarkovic, N. 4-Hydroxynonenal as a bioactive marker of pathophysiological processes. Mol. Asp. Med. 24, 281–291 (2003).
Shoeb, M., Ansari, N., Srivastava, S. & Ramana, K. 4-Hydroxynonenal in the pathogenesis and progression of human diseases. Curr. Med. Chem. 21, 230–237 (2013).
Wilson, K. et al. Accurate prediction of gestational age using newborn screening analyte data. Am. J. Obstet. Gynecol. 214, 513.e1–513.e9 (2016).
Wilson, K. et al. Metabolomics of prematurity: analysis of patterns of amino acids, enzymes, and endocrine markers by categories of gestational age. Pediatr. Res. 75, 367–373 (2014).
Jelliffe-Pawlowski, L. L., Norton, M. E., Baer, R. J., Santos, N. & Rutherford, G. W. Gestational dating by metabolic profile at birth: a California cohort study. Am. J. Obstet. Gynecol. 214, 511.e1–511.e13 (2016).
Ryckman, K. K., Berberich, S. L. & Dagle, J. M. Predicting gestational age using neonatal metabolic markers. Am. J. Obstet. Gynecol. 214, 515.e1–515.e13 (2016).
Wilson, L. A. et al. Postnatal gestational age estimation via newborn screening analysis: application and potential. Expert Rev. Proteom. 16, 727–731 (2019).
da Silva, R. R., Dorrestein, P. C. & Quinn, R. A. Illuminating the dark matter in metabolomics. Proc. Natl Acad. Sci. USA 112, 12549–12550 (2015).
Roager, H. M. & Licht, T. R. Microbial tryptophan catabolites in health and disease. Nat. Commun. 9, 3294 (2018).
Lloyd-Price, J. et al. Multi-omics of the gut microbial ecosystem in inflammatory bowel diseases. Nature 569, 655–662 (2019).
Wargo, M. J. & Meadows, J. A. Carnitine in bacterial physiology and metabolism. Microbiology 161, 1161–1174 (2015).
Singh, J., Metrani, R., Shivanagoudra, S. R., Jayaprakasha, G. K. & Patil, B. S. Review on bile acids: effects of the gut microbiome, interactions with dietary fiber, and alterations in the bioaccessibility of bioactive compounds. J. Agric. Food Chem. 67, 9124–9138 (2019).
Theriot, C. M. et al. Antibiotic-induced shifts in the mouse gut microbiome and metabolome increase susceptibility to Clostridium difficile infection. Nat. Commun. 5, 3114 (2014).
Crump, C. et al. Association of preterm birth with risk of ischemic heart disease in adulthood. JAMA Pediatr. 173, 736–743 (2019).
Poesen, R. et al. Microbiota-derived phenylacetylglutamine associates with overall mortality and cardiovascular disease in patients with CKD. J. Am. Soc. Nephrol. 27, 3479–3487 (2016).
Kajantie, E., Osmond, C., Barker, D. J. P. & Eriksson, J. G. Preterm birth—a risk factor for type 2 diabetes? Diabetes Care 33, 2623–2625 (2010).
Rossi, F., Miggiano, R., Ferraris, D. M. & Rizzi, M. The synthesis of kynurenic acid in mammals: an updated kynurenine aminotransferase structural KATalogue. Front. Mol. Biosci. 6, 7 (2019).
Hill, C. J. et al. Evolution of gut microbiota composition from birth to 24 weeks in the INFANTMET Cohort. Microbiome 5, 4 (2017).
Abdel-Hady, H., Nasef, N., Shabaan, A. E. & Nour, I. Caffeine therapy in preterm infants. World J. Clin. Pediatr. 4, 81–93 (2015).
Taylor, S. N., Basile, L. A., Ebeling, M. & Wagner, C. L. Intestinal permeability in preterm infants by feeding type: mother’s milk versus formula. Breastfeed. Med. 4, 11–15 (2009).
Bahado-Singh, R. O. et al. Artificial intelligence and the analysis of multi-platform metabolomics data for the detection of intrauterine growth restriction. PLoS ONE 14, e0214121 (2019).
Sovio, U. et al. A maternal serum metabolite ratio predicts fetal growth restriction at term. Nat. Med. 26, 348–353 (2020).
Acknowledgements
This research has been conducted using the Danish National Biobank resource, and supported by the Novo Nordisk Foundation. The study was sponsored by the Lundbeck Foundation, grant numbers R102-A9118 and R155-2014-1724.
Author information
Authors and Affiliations
Contributions
M.E. performed statistical analysis and chemical structural annotation, interpreted the data and drafted the manuscript. S.R. performed statistical analysis, contributed to the interpretation of the data and the drafting of the manuscript. U.L.-T. contributed to data interpretation and the drafting of the manuscript. A.B. and S.S.L. conceptualized and designed the study and acquired the data. J.C. contributed to data interpretation and the drafting of the manuscript. A.B., M.N., T.W. and P.B.M. obtained funding. D.M.H. conceptualized and designed the study, obtained funding and together with A.S.C. takes full responsibility for compliance with data sharing policies and ethical approval of the study. A.S.C. conceptualized and designed the study and contributed to the interpretation of the data and drafting of the manuscript and together with D.M.H. takes full responsibility for compliance with data sharing policies and ethical approval of the study. All authors critically revised the manuscript for important intellectual content.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Ernst, M., Rogers, S., Lausten-Thomsen, U. et al. Gestational age-dependent development of the neonatal metabolome. Pediatr Res 89, 1396–1404 (2021). https://doi.org/10.1038/s41390-020-01149-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41390-020-01149-z
This article is cited by
-
Fast mass spectrometry search and clustering of untargeted metabolomics data
Nature Biotechnology (2024)
-
Metabolomic profiling of intrauterine growth-restricted preterm infants: a matched case–control study
Pediatric Research (2023)
-
Effect of common pregnancy and perinatal complications on offspring metabolic traits across the life course: a multi-cohort study
BMC Medicine (2023)
-
Cord blood metabolites and rapid postnatal growth as multiple mediators in the prenatal propensity to childhood overweight
International Journal of Obesity (2022)
-
Gaining a deeper understanding of social determinants of preterm birth by integrating multi-omics data
Pediatric Research (2021)