Abstract
Colorectal cancer (CRC) is one of the most common cancers, with an annual incidence of ~135,000 in the US, associated with ~50,000 deaths. Autosomal dominant polycystic kidney disease (ADPKD), associated with mutations disabling the PKD1 gene, affects as many as 1 in 1000. Intriguingly, some studies have suggested that individuals with germline mutations in PKD1 have reduced incidence of CRC, suggesting a genetic modifier function. Using mouse models, we here establish that loss of Pkd1 greatly reduces CRC incidence and tumor growth induced by loss of the tumor suppressor Apc. Growth of Pkd1−/−;Apc−/− organoids was reduced relative to Apc−/− organoids, indicating a cancer cell-intrinsic activity, even though Pkd1 loss enhanced activity of pro-oncogenic signaling pathways. Notably, Pkd1 loss increased colon barrier function, with Pkd1-deficient animals resistant to DSS-induced colitis, associated with upregulation of claudins that decrease permeability, and reduced T cell infiltration. Notably, Pkd1 loss caused greater sensitivity to activation of CFTR, a tumor suppressor in CRC, paralleling signaling relations in ADPKD. Overall, these data and other data suggest germline and somatic mutations in PKD1 may influence incidence, presentation, and treatment response in human CRC and other pathologies involving the colon.
Similar content being viewed by others
Introduction
Colorectal cancer (CRC) is one of the most common cancers, with an annual incidence of ~135,000 in the US, associated with ~50,000 deaths [1]. For the most common forms of CRC, mutational loss of the Adenomatous Polyposis Coli (APC) tumor suppressor gene, leading to WNT/β-catenin (CTNNB1) pathway activation, is followed by mutations activating KRAS and inactivating TP53 during progression [2, 3]. A number of modifier genes have been recognized as influencing CRC risk, tumor aggressiveness and response to therapy, and overall survival [4, 5]. For example, individuals with familial adenomatous polyposis (FAP), associated with truncating germline mutations in APC, or with Lynch syndrome, associated with germline mutations in DNA mismatch repair (MMR) genes, experience a higher incidence of CRC, and disease onset at an earlier age [6]. It is estimated that there are over 100 inherited gene variants modifying CRC risk [7].
This study was motivated by results from two studies investigating genes associated with a genetically inherited developmental syndrome. Autosomal dominant polycystic kidney disease (ADPKD, MIM #173900; # 613095) typically arises from germline mutations inactivating the PKD1 or PKD2 genes, and affects approximately 1 in 1000 individuals, and has a phenotype of epithelial cell overgrowth, resulting in extensive replacement of normal renal ductal and tubular structure with fluid-filled cysts. In a study comparing 10,146 kidney recipients with ADPKD, versus 107 339 kidney recipients without ADPKD, individuals with ADPKD had a reduced incidence of CRC (incidence rate ratio 0.72), as well as lower incidence of other cancers affecting the gastrointestinal tract [8]. Although this is a modest signal, it was obtained in a population where ~15% of the individuals were likely to have mutations in PKD2, associated with a mild disease course, and at least 30–40% of the individuals with PKD1 mutations would have inherited a single moderately hypomorphic allele rather than a homozygous or strongly inactivating mutation [9, 10]. Independently, a study investigating PKHD1 (mutated in autosomal recessive PKD (ARPKD), and encoding a protein, fibrocystin, which functionally interacts with PKD1), also found a lower than expected rate of CRC in mutation bearers (0.42% in 3603 controls versus 0.027% in 3767 patients with CRC; p = 0.0002) [11].
Although investigations of PKD1, PKD2, and PKHD1 typically focus on their roles in the kidney, ADPKD, and ARPKD have many extrarenal manifestations [12]. The PKD1 protein is expressed and active in many tissues, including the colonic epithelium [13]. Relevant to CRC, mutations in PKD1 have been shown to modulate the activity of the WNT/CTNNB1 and other pro-growth signaling pathways in the kidney, although the mechanism of interaction is not incompletely understood, and distinct assay systems have yielded conflicting results [14, 15]. Interestingly, in an analysis of 190 clinical CRC specimens, PKD1 was overexpressed in tumors relative to normal tissue, regardless of TNM stage, suggesting elevation at an early stage of tumorigenesis. PKD1 expression was highest in tissues with the deepest invasion and associated with poor survival [16]. Overexpression of PKD1 in CRC cell lines increased growth rate and epithelial-mesenchymal transition (EMT), enhancing migration and invasion [16]; whereas a mouse model null for the Pkd1 gene was embryonically lethal due to a cell migration defect [17].
Based on these suggestive studies, we have used mouse models to directly test the idea that genetic loss of Pkd1 reduces the risk of CRC arising from mutations in Apc, and if so, to explore relevant mechanisms. Based on this work, we define Pkd1 loss as tumor suppressive for CRC, based on cell-intrinsic activity in the colon epithelium. We further show that Pkd1 loss does not reduce CRC growth by inhibiting pro-proliferative pathways, but does reprogram cell-cell contacts in the colon epithelium so as to limit tissue damage and inflammation-associated proliferation.
Materials and methods
Mouse strains and drug treatments
The Institutional Animal Care and Use Committee (IACUC) of Fox Chase Cancer Center approved all experiments involving mice. To study DSS-induced colitis, we used tamoxifen-regulated-Cre+ expressed from the β-actin promoter in all tissues, crossed to Pkd1fl/fl [18, 19] mice, as in [20,21,22], with mice referred to as Pkd1−/− post-tamoxifen treatment. Mice lacking Cre (Pkd1fl/fl, referred to as Pkd+/+) were used as negative controls. Mice were injected intraperitoneally with tamoxifen (250 mg/kg body weight [BW], formulated in corn oil) on post-natal days P35 and P36 to induce loss of Pkd1. 10 weeks after injection, mice were treated with 2.5% dextran sodium sulfate (DSS) in drinking water for 5 days to induce acute colitis, and euthanized 2 days later. For some experiments, a FITC-dextran tracer (46944, Sigma-Aldrich, St. Louis, MO; 4 kDa, 0.6 mg/g body weight) was administered by oral gavage 3 hours prior to euthanasia. 300–400 μl of blood was collected immediately prior to euthanasia, and concentration of FITC-dextran was measured in the blood using a ProXpress Visible-UV-fluorescence 16-bit scanner (Perkin-Elmer, Waltham, MA). Colon tissues were collected for subsequent analysis.
To evaluate the role of Pkd1 in CRC, Pkd1fl/fl mice were crossed to Cdx-ERT-Cre+ mice bearing floxed Apc (Cdx2-ERT-Cre+;Apcfl/fl [23,24,25], with colon epithelium-preferential Cdx2 transgene expression, referred to as Apc−/− post-tamoxifen treatment), to generate Cdx2-ERT-Cre+;Apcfl/fl;Pkd1fl/wt and Cdx2-ERT-Cre+;Apcfl/fl;Pkd1fl/fl mice (designated Apc−/−;Pkd1+/− and Apc−/−;Pkd1−/−, respectively). These mice and control Apcfl/fl mice were injected intraperitoneally with tamoxifen [250 mg/kg body weight (BW),] at the age of 3 months to induce deletion of Apc and Pkd1 in the colon. Mice were euthanized 5 weeks later and colons collected for analysis.
For drug treatment experiments, inactivation of the Apc and Pkd1 genes was induced in 3 months old Cdx2-ERT-Cre+;Apcfl/fl and Cdx2-ERT-Cre+;Apcfl/fl;Pkd1fl/fl mice. One week after tamoxifen injection, 3–4 mice of each genotype were randomly assigned to one of the following treatment groups: (1) Vehicle, (2) 5-FU (50 mg/kg) + LV (leucovorin, 90 mg/kg) + Irinotecan (24 mg/kg) – FOLFIRI, and (3) 5-FU (50 mg/kg) + LV (leucovorin 90 mg/kg) + Oxaliplatin (6 mg/kg) – FOLFOX. Drugs were administered once a week for 4 weeks intravenously, after which mice were euthanized and organs were collected for analysis.
qRT-PCR analysis of Pkd1 expression
Assessment of effective Cre targeting of Pkd1 is complicated by the presence of multiple Pkd1 pseudo-genes. We performed qRT-PCR probing for Pkd1 exon 1-2 (intact Pkd1) and exon 1-5 (deleted exons 2-3-4 in Pkd1 floxed mice) junctions. The oligonucleotide sequences used for PCR are as follows: CTGCCGCGTCAATTGCT; CCTATGTCCAGCGTCTGAAGTA (Pkd1 junction 1-2). CTGCCGCGTCAATTGCT; GCAGGGAGGAAGTAATATGGAAG (Pkd1 junction 1-5). Briefly, total RNA was isolated from tumor tissues or organoids using a Zymo Research Quick-RNA MicroPrep Kit (#R1050) and tested for quality on a Bioanalyzer (Agilent Technologies, Santa Clara, CA). RNA concentrations were determined with a NanoDrop spectrophotometer (Thermo Fisher Scientific, Waltham, MA). Total RNA prepared from tumor tissue was used for quantitative RT-PCR, with expression of sequences of interest normalized to that of the housekeeping gene 36B4.
Organoid isolation and culture
Organoid culturing was performed as in [26, 27]. Briefly, dissected colon lesions (~2–3 mm) were sequentially washed with cold PBS and cold chelation buffer on ice. Tissue was then incubated in digestion buffer (DMEM medium with 2.5% FBS, 1% collagenase, and 0.125% dispase) at 37 °C for 45 min followed by treatment with 1x accutase (#07922, StemCell Technologies, Vancouver, BC). Single cells were collected by centrifugation, resuspended in PBS, and embedded in growth factor-reduced Matrigel (CLS354230, Corning, Sigma-Aldrich, St. Louis, MO) and seeded in 24-well plates in culture medium (advanced DMEM/F12 (ADF) supplemented with penicillin/streptomycin, 10 mmol/L HEPES, Glutamax, 1x N2, 1x B27 (all from Invitroge]) and murine EGF (50 μg/ml, SRP3196, Sigma-Aldrich, St. Louis, MO). Medium was changed every 4 days during experiments.
For drug treatment experiments, organoids were treated with vehicle, a mixture of 10 μM lumacaftor + 3 μM ivacaftor, or 0.5 μM PRI-724 for 10 days. Alternatively, organoids were treated with vehicle, a mixture of 5-FU (9.6 μM) + LV (4.7 μM) + irinotecan (1 μM) – FOLFIRI, or 5-FU (9.6 μM) + LV (4.7 μM) + oxaliplatin (0.4 μM) – FOLFOX for 10 days. All drugs were purchased from MedChemExpress (Monmouth Junction, NJ). Cell or organoid viability was assessed by a CellTiterGlo luminescent cell viability assay (#G7570, Promega, Madison, WI, USA), using the manufacturer’s protocols.
Tissue preparation, histology, and quantitative analysis
Colon sections were fixed in 10% phosphate-buffered formaldehyde (formalin) for 24–48 h, dehydrated, and embedded in paraffin. 5 µm specimens were analyzed by hematoxylin and eosin (H&E, Sigma-Aldrich, St. Louis, MO), trichrome (HT10516, Sigma-Aldrich, St. Louis, MO), or by immunohistochemistry (IHC) using standard protocols. Slides were scanned using a Vectra Automated Quantitative Pathology Imaging System (Perkin Elmer, Waltham, MA). All reported data were verified by a board-certified pathologist, using a standard grading protocol. IHC staining of tissue sections for Ki-67 and CD45 was performed using #27309-I-AP (Ki-67, 1:1000 dilution) and antibodies from Proteintech (Rosemont, IL) and #70257S (CD45, 1:200 dilution, Cell Signaling, Beverly, MA). Expression levels of Ki-67 and CD45 were quantified using inForm software and H-score was calculated as reported previously [28]. Immunofluorescence-IHC using antibodies to Ki-67 (1:100, #27309-I-AP, Proteintech, Rosemont, IL), β-Catenin (1:100, 2698S Cell Signaling, Beverly, MA), ph-S675-β-Catenin (1:70, 4176S Cell Signaling, Beverly, MA), alpha-smooth muscle actin (α-SMA, 1:300, A2547 Sigma-Aldrich, St. Louis, MO), Cldn4 and Cldn7 (1:50 and 1:200, respectively, #16195-I-AP and #10118-I-AP, Proteintech, Rosemont, IL) was used for FFPE tissue sections and organoids embedded in Matrigel using protocols as in [28, 29]. Slides were counterstained with Vectashield mounting medium with DAPI (4′,6-diamidino-2-phenylindole) (H-1200, Vector Labs, Burlingame, CA) to visualize DNA.
Samples were imaged at room temperature (RT) using an SP8 confocal system equipped with an oil-immersion 363 objective with numerical aperture (NA) 1.4 (Leica Microsystems, Buffalo Grove, IL) and LASAF (Leica Application Suite Advanced Fluorescence) software, using NIH ImageJ Imaging Software to quantify area and MetaMorph 7.6.5 software (Molecular Devices, Union City, CA) for measurements of integrated optical density values.
Luminex assay
The Bio-Plex Pro Cell Signaling Akt Panel 8-Plex Assay kit (#LQ00006JK0K0RR, Bio-Rad Laboratories, Hercules, CA) was used. Tissue samples were lysed using T-PER buffer (ThermoScientific, Waltham, MA), protein concentration established using a BCA Protein Assay Kit (Thermo Scientific, Waltham, MA), and samples incubated overnight in a 96-well plate containing magnetic beads. Subsequently, protein signal was assessed using a Bio-Plex 200 system (Bio-Rad Laboratories, Hercules, CA). Results were processed using GraphPad Prism 6 (GraphPad Software, San Diego, CA) software.
RNA sequencing and data analysis
Total RNA was isolated from organoids using a Zymo Research Quick-RNA MicroPrep Kit (#R1050) and tested for quality on a Bioanalyzer (Agilent Technologies, Santa Clara, CA). RNA concentrations were determined with a NanoDrop spectrophotometer (Thermo Fisher Scientific, Waltham, MA). Total RNA (standard concentration ≥200 ng/μl, mass ≥10 μg, and RNA integrity number (RIN) ≥ 8.0) was sequenced by Novogene (Sacramento, CA). FastQC was used for read quality assessment (https://www.bioinformatics.babraham.ac.uk/projects/fastqc/). Sequence reads were aligned to the mouse mm10 genome using Tophat2 [30]. The Cufflinks algorithm was used to assemble and quantify transcripts [31], and Cuffdiff to statistically assess expression changes in quantified genes across different conditions [32]. Genes with a false discovery rate of 5% and a fold change of 1.5 were considered differentially expressed. The networks and upstream regulators analysis were generated using Ingenuity Pathway Analysis (IPA) (QIAGEN Inc., https://digitalinsights.qiagen.com/IPA).
Analysis of publicly available datasets at cBioPortal
The Cancer Genome Atlas (TCGA) PanCancer dataset (594 samples), DFCI (Nature 2016, 619 samples), and Genentech (Nature 2012, 74 samples) datasets for CRC were accessed (July, 2022) and analyzed using tools available at cBioPortal for Cancer Genomics (http://www.cbioportal.org/). In some cases, datasets were merged to increase sample numbers and hence statistical power.
Statistical analysis
We used Wilcoxon Rank-sum two-tailed tests for pairwise comparisons and 1-way ANOVA for 2 or more group comparisons. Relationships between mutations and patient characteristics were assessed using Fisher exact tests. P-values < 0.05 were considered as statistically significant and data presented as mean and S.E.M. Statistical analyses were performed using GraphPad Prism 8 (GraphPad Software, San Diego, CA).
Results
Genetic deletion of Pkd1 reduces CRC tumorigenesis induced by loss of Apc
To directly assess whether loss of Pkd1 affects the incidence or presentation of colorectal cancer, we used an inducible CRC mouse model in which biallelic loss of a floxed allele of Apc (Apcfl/fl) is induced by CreERT2 (in which tamoxifen induces Cre-lox dependent targeted inactivation), with CreERT under the control of the Cdx2 promoter, and expressed specifically in the epithelium of the distal ileum, cecum, and colon (Cdx2ERT-Apcfl/fl mice; designated Apc−/− hereafter). These mice develop tumors primarily in the distal colon, which is physiologically similar to human CRC [25]. These mice were crossed to mice with a floxed Pkd1 allele [18,19,20], to compare tumor formation in Apc−/−, Apc−/−;Pkd1-/+, and Apc−/−;Pkd1−/− mice (Fig. 1A, Supp Fig. S1A).
A Experimental design. B Number of tumors in colon of Apc−/− (n = 10), Apc−/−;Pkd1+/− (n = 17), or Apc−/−;Pkd1−/− (n = 15) mice. C Representative image of colons from mice quantified in (B); also see Supp Fig. S1 for additional images. Scale bar, 500 μm. D Average size of adenomas at time of euthanasia, in genotypes indicated. E Representative H&E stained adenomas or polyps in mice of genotypes indicated. Magnification 4x. Scale bar, 500 μm. F Quantification of Ki-67 staining of lesions from genotypes noted, in proximal and distal colon; non-significant differences were observed based on genotype. *p < 0.05; **p < 0.01; ***p < 0.001, in all panels.
At five weeks after tamoxifen treatment, there was a highly significant reduction in the number of tumors of mice with heterozygous or homozygous loss of Pkd1 specifically in the same cells as those with loss of Apc (e.g., not in the normal colon epithelium). Inactivating Pkd1 recombination was validated using qRT-PCR probing for Pkd1 exon 1-2 (intact Pkd1) and exon 1-5 (deleted exons 2-3-4 in Pkd1 floxed mice) junctions (see Methods for greater detail). In many cases, reduction of tumor formation was more notable in the distal colon (Fig. 1B, C, Supp Fig. S1B). Quantification of tumor size indicated that tumors emerging in Apc−/−;Pkd1-/+, and Apc−/−;Pkd1−/− mice were significantly smaller than tumors in Apc−/− mice (Fig. 1D, E).
Histopathological assessment by a board-certified pathologist indicated no differences in grade among the distinct tumor genotypes, with all tumors appearing predominantly as adenomas, with rare well-differentiated adenocarcinomas. Ki-67 staining indicated that Pkd1 genotype did not significantly affect proliferation rate of cells within observed lesions in either the proximal or distal colon (Figs. 1F, S1C). In ADPKD, mutation of PKD1 is often associated with renal fibrosis, which contributes to scarring and loss of kidney function [33]. Trichrome staining indicated no differences in fibrosis in the CRC lesions of Apc−/− and Apc−/−;Pkd1−/− CRC mice, although a trend to higher deposition in the submucosal layers of the colons of Apc−/−/ Pkd1−/− mice (Fig. S2A). We also stained the lesions with alpha-smooth muscle actin (α-SMA) as a marker for a subset of activated myofibroblasts, which are important effectors of tissue fibrogenesis (Fig. S2B). Interestingly, α-SMA expression was expressed in the muscularis layer of the colons of Apc−/−/Pkd1−/− mice versus Apc−/− mice, and to a greater extent in the Apc−/−/Pkd1−/− lesions, paralleling results in ADPKD.
Reduced CRC incidence in Pkd1-deficient colons is partially mimicked in organoids
The reduced incidence of Pkd1-deficient tumors might solely reflect cell intrinsic consequences of Pkd1 loss in colon epithelial cells or might instead reflect the altered interaction of PKD1-deficient epithelial cells with the nascent tumor microenvironment. To address these possibilities, we treated Apc−/− and Apc−/−Pkd1−/− mice with tamoxifen, and after 5 weeks collected visible lesions and generated organoids in Matrigel (Fig. 2). Gross morphology of organoids and efficiency of seeded cells in forming organoids were not affected by Pkd1 genotype, but there was a delay in organoid formation with Apc−/−Pkd1−/− relative to Apc−/− cells (Fig. 2A, B), reflecting reduced growth rate. Overall, after 7 days in culture, Apc−/−Pkd1−/− organoids were significantly smaller than Apc−/− organoids on average (mean diameter 284.9 versus 184.7 microns; p < 0.0001) (Fig. 2C). In contrast to in vivo observations, Ki-67 staining of colonies indicated a somewhat reduced number of proliferating cells among those with the Apc−/−Pkd1−/− genotype after 10-14 days in culture (Fig. 2D, E). This difference may reflect the higher overall proliferative index occuring in vitro versus in vivo, or alternatively, some compensation in vivo for the antiproliferative effect of Pkd1 loss.
A Representative images of organoid growth in Matrigel. Scale bar, 50 μm. B Seeding efficiency of organoids based on genotype, based on a size threshold of 50 μm. C Average size of organoids from Apc−/− versus Apc−/−;Pkd1−/− genotypes after 7 days.. D, E Quantification (D) and representative images (E) of Ki-67 staining of organoids of genotypes indicated. Scale bar, 100 μm. ns, not significant; *p < 0.05; **p < 0.01; ***p < 0.001, ****p < 0.0001 in all panels.
Genetic deletion of Pkd1 elevates Wnt/Ctnnb1-dependent transcription
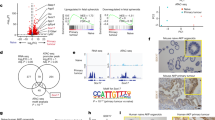
To gain insight into signaling differences induced by loss of Pkd1, performed RNAseq analysis on organoids prepared from the Apc−/− and Apc−/−Pkd1−/− genotypes to gain a more detailed insight into the transcriptional consequences of loss of Pkd1 specifically in Apc−/− colorectal epithelia. Loss of Pkd1 significantly increased the expression of 127 genes, and decreased the expression of 11 genes, based on a threshold of 1.5-fold-change and a p-value < 0.05 to establish significance (Fig. 3A, Supp Table S1). Surprisingly, Ingenuity pathway analysis (Fig. 3B) identified a complex set of changes, including upregulation of multiple pro-proliferative pathways, including WNT/CTNNB1, ERK1, and IGF1, as well as proinflammatory signaling pathways associated with IFNG and IL1A. However, antiproliferative pathways were also upregulated, including numerous transcripts regulated by TP53, indicating opposing signals. In addition, Luminex analysis of a panel of signaling proteins involved in mediating cell growth revealed no significant difference between the Apc−/− and Apc−/−Pkd1−/− genotypes (Supp Fig. S2C).
A Volcano plot showing distribution of genes differentially expressed in organoids of the Apc−/−;Pkd1−/− versus Apc−/− genotypes, based on statistical significance (Y axis, p value) and log2 fold change (x axis). B Waterfall plot demonstrating upstream transcriptional regulators statistically significantly changed in Apc−/−;Pkd1−/− versus Apc−/− organoids. C Expanded volcano plot focused on genes above cutoffs for statistical significance and magnitude of change, from A. Gene names shown reflect those linked to CTNNB1 signaling as targets identified by Ingenuity (23 of 30 total genes above cut-off values) or as regulators, based on literature (in italics). D, E Immunofluorescence staining of lesions from Apc−/−;Pkd1−/− versus Apc−/− mice colons for β-catenin (D), at 10x (left images) and 40x (right images), with quantification of signal intensity (E).
Given the activation of the CTNNB1 pathway is a primary proliferation-promoting consequence of APC loss in colon crypts [34], and given the large number of upregulated transcripts associated with CTTNB1 (Fig. 3C), we further probed activity of this pathway. immunohistochemical (IHC) analysis of pre-malignant colon tissue, polyps, and adenomas from Apc−/−Pkd1−/− versus Apc−/− mice confirmed elevated CTNNB1 expression and active (S675-phosphorylated) CTNNB1 associated with the Apc−/−Pkd1−/− genotype, as well as in Apc−/−Pkd1−/− organoids (Fig. 3D, E, Supp Fig. S3A, B). In all cases, although there was not a marked shift of CTNNB1 from the periphery to the nucleus, the elevation of S675-phosphorylated CTNNB1 [35], was compatible with greater pathway activity and upregulation of CTNNB1-dependent transcripts.
Loss of Pkd1 increases colon barrier function
Intriguingly, loss of PKD1 function in ADPKD enables the epithelial monolayer to increase the transit of fluids into the cyst and to form a non-leaky barrier that can withstand high hydrostatic pressure within the cyst [36, 37]. Conversely, CRC risk is elevated by factors that reduce intestinal barrier function [38,39,40,41], which promotes increased inflammation in human patients and mouse models [23, 42]. In this process, designated Tumor-Elicited Inflammation (TEI), colonies of microbes adjacent to pre-malignant colon tissue gradually infiltrate the epithelial layer. Interaction of microbes with recruited myeloid cells and tumor cells elicits IL-23/IL-17 signaling that promotes tumor growth and initiates a feed-forward cycle, reprogramming of tight junctions to further reduce epithelial barrier permeability [23, 42, 43]. This was suggestive, because comparison of Apc−/−Pkd1−/− versus Apc−/− tumors identified a trend towards lower CD45-positive leukocyte infiltration in the Apc−/−Pkd1−/− genotype (Fig. 4A).
A Immunohistochemical staining of CD45 expression in colon lesions of Apc−/− versus Apc−/−;Pkd1−/− mice. Left—representative images, scale bars, 200 μm. Right—quantification. B Quantification of FITC-dextran levels in serum of dextran sodium sulfate (DSS)- or control water (Ctrl)-treated mice. C Representative H&E images of colon tissue from Pkd1wt versus Pkd1−/− mice treated with water or DSS. Scale bars, 100 μm. Measurements of mouse weight during treatment are located in Fig S4B. D, E Representative images (left) and quantification (right) of expression of CLDN4 (D) and CLDN7 (E) in wt versus Pkd1−/− colon epithelia. Scale bars 50 μm. F, G Representative images (left) and quantification (right) of CLDN4 and CLDN7 staining in organoids. Blue, DAPI; green, claudins. Scale bars 50 μm. *p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001 for all graphs.
To test the hypothesis that loss of Pkd1 influenced gut barrier function, positioning it to reduce TEI, we used treatment with dextran-sodium sulfate (DSS), an irritant that damages the gut epithelium, eliminating the protective mucin lining of the gut epithelium, and causing epithelial cell loss and tissue damage; reduction in epithelial barrier function typically sensitizes mice to DSS [44, 45]. We treated 15-week-old Pkd1+/+ (designated wild-type [wt] hereafter) or Pkd1−/− mice (with loss of Pkd1 induced in all tissues by tamoxifen injection at days P35 and P36) for 5 days with 2.5% DSS in drinking water, versus water-only controls. We then gave FITC-dextran tracer by oral gavage and bled mice after 3 h to collect serum and analyzed the appearance of FITC-dextran in the serum, as in [23] (Fig. 4B). Whereas DSS elevated serum levels 5-fold over those in untreated wt mice, Pkd1−/− mice showed only a 2-fold, statistically insignificant increase in FITC-dextran in serum, strongly suggesting reinforced barrier function. Further, immunohistochemical analysis of colonic tissue from vehicle- or DSS-treated wt and Pkd1−/− mice shows that PKD1 deficiency substantially reduces the tissue damage induced by response to DSS (Fig. 4C). In wt mice, DSS induced a complete loss of the organized mucosal layer, including loss of mucus-producing cells, irregularity and expansion of the submucosal layer, and evidence of infiltration of immune cells (arrows, Fig. 4C). In contrast, while some defects in the mucosal layer were observed in DSS-treated Pkd1−/− mice, they were much less severe in all mice. In addition, while DSS promoted the infiltration of CD45+ leukocytes, infiltration was limited to submucosal layer in DSS-treated animals lacking Pkd1, whereas much more pervasive leukocyte infiltration characterized DSS-treated mice with wt Pkd1 (Fig. S4A).
Loss of Pkd1 in the colon induces claudins that promote impermeable barriers
In developing a hypothesis for Pkd1 action, we considered the fact that mechanistically, Pkd1 loss in ADPKD produces cysts in part by upregulating claudins 4 and 7 (CLDN4, CLDN7) and others that similarly induce impermeable tight junctions, while downregulating those that permit barrier permeability [46, 47]. In human CRC, an early step in TEI is the post-transcriptional downregulation of CLDN7 and CLDN4 [23], which contributes to the penetration of bacteria and their inflammatory products [23]; loss of CLDN7 has been shown to promote colorectal inflammation and tumorigenesis [48]. Comparison of colon tissue from wt and Pkd1−/− mice indicated significantly elevated levels of CLDN4 and CLDN7 in Pkd1−/− colonic epithelia (Fig. 4D, E). Similar elevation of CLDN4 and CLDN7 expression was observed in Apc−/−Pkd1−/− versus Apc−/− organoids (Fig. 4F, G). Together with results from DSS administration, these data suggest loss of Pkd1 induces claudins to decrease permeability of the colon epithelium, restricting inflammation- and proliferation-promoting TEI.
Loss of Pkd1 in colon epithelial cells influences response to targeted and cytotoxic therapies
To gain insight into the mechanism by which Pkd1 loss reduced tumor growth, we considered that in ADPKD, Pkd1 loss activates the cystic fibrosis transmembrane receptor (CFTR), contributing to polarized ion secretion [49]. Notably, CFTR has been reported to act as a tumor suppressor in CRC [50], with cystic fibrosis patients prone to elevated CRC risk [51], and loss of CFTR previously shown to reduce expression of CLDN7 [52]. To gain insight into the role of CFTR signaling in PKD1 activity in the colon, we compared the growth of Apc−/−Pkd1−/− versus Apc−/− organoids treated with a CFTR-activating drug combination lumacaftor/ivacaftor (L/I) [53]. We benchmarked these results to the effect of Apc−/−Pkd1−/− versus Apc−/− organoids treated with the WNT pathway inhibitor PRI-724 [54], which inhibits the interaction between CTNNB1 and its transcriptional coactivator CBP, and should be active in all CRC arising from an Apc−/− genotype. Both drugs were very effective in reducing organoid growth, indicating the importance of both WNT activity and CFTR inhibition for allowing colon cell growth (Fig. 5A, B). Interestingly, while Apc−/− organoids responded more strongly to inhibition of CTNNB1 than activation of CFTR, Apc−/−Pkd1−/− organoids responded comparably to both drugs, suggesting greater importance of CFTR activation in regulating growth of organoids lacking Pkd1.
A, B Representative images (A) and quantitation (B) of organoid size following treatment with vehicle, the CFTR activator pair lumacaftor/ivacaftor (L/I), or the CTNNB1 inhibitor PRI-724 (PRI), based on Apc−/− versus Apc−/−;Pkd1−/− genotype. Scale bars on (A), 50 μm. C Experimental design to assess in vivo response to FOLFIRI or FOLFOX. D, E Representative image of colons from mice treated with FOLFOX and FOLFIRI; gross presentation (D) and H&E-stained tissue (E). F, G Quantification of number of lesions based on gross assessment of number in colon (F) and area, based on histopathological assessment (G) following treatment with vehicle, FOLFIRI, or FOLFOX, in genotypes indicated. H Quantitation of viability based on CellTiterGlo of organoid cultures following treatment with vehicle, FOLFIRI, or FOLFOX, in indicated genotypes, with data normalized to vehicle-treated Apc−/− organoids. *p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001 for all graphs; in (B), ###p < 0.001; compared to Apc−/− vehicle.
Standard of care treatment for CRC typically involves used of DNA-damaging cytotoxic chemotherapies, FOLFIRI and FOLFOX [55]. To determine if Pkd1 loss affected response to these drugs, we induced loss of Apc and Pkd1 with tamoxifen treatment of 12-week-old mice, and commencing at week 13, treated mice weekly with either vehicle, or with standard DNA-damaging cytotoxic chemotherapies for CRC, FOLFIRI and FOLFOX (Fig. 5C). Both treatments were effective at eliminating >60% of lesions relative to vehicle in both Apc−/− and >Apc−/−Pkd1−/− mice (Fig. 5D–G). Given the lower number and size of lesions occurring in Apc−/−Pkd1−/− mice, almost no adenomas were observed in these animals following drug treatments. In contrast to results with targeted inhibitors, treatment of organoids with FOLFIRI or FOLFOX was similarly effective in reducing organoid growth in both genotypes (Fig. 5H).
PKD1 mutation and expression profiles in human CRC
We analyzed the profile of PKD1 mutations in CRC datasets reported in cBioPortal, to determine if PKD1-mutated tumors had any specific distinguishing features (Fig. 6). In the 603 CRC samples for which data was available, 497 tumors were microsatellite stable (MSS), and 106 had microsatellite instability (MSI) (82.4 versus 17.6%) (Fig. 6A). Among the 546 PKD1 wt tumors, 473 (87%) were MSS; in contrast, among the 57 tumors with non-synonymous mutations in PKD1 and with annotation as MSS or MSI, only 24 were MSS (42%), indicating a significant bias to occurrence in MSI. Typically, for both the MSI and MSS cohorts, tumors with PKD1 mutations were associated with higher TMB than tumors with wild-type (wt) PKD1 (Fig. 6B, C). Based on rules for determining degree of damaging effect established through studies of ADPKD-associated mutations [10], approximately 1/3 of the PKD1 mutations were predicting to be non-damaging, which may suggest a passenger role. However, overall, PKD1-mutated tumors tended to be observed at earlier stages than PKD1 tumors (although the observation did not reach statistical significance due the limited number of PKD1-mutated tumors) (Fig. 6D). The age range of diagnosis for PKD1-mutated tumors was similar to that of PKD1 wt tumors (Fig. 6E), as was the distribution by sex (Fig. 6F). We also examined distribution of PKD1 mutations among tumors of distinct histology, including mucinous versus non-mucinous (Fig. 6G). Although non-mucinous tumors predominated in both cohorts, a significantly higher percentage PKD1-mutated tumors were likely to be mucinous than were PKD1 wild-type tumors.
A Proportion of PKD1 wild type (wt) versus mutated (mut) human CRC specimens, separately analyze based on microsatellite stability status (MSI vs. MSS). B Association of PKD1 mutational status with TMB in MSI and MSS samples. C Distribution of PKD1 mutations normalized by 1000 mutations per sample. D Relationship of PKD1 mutation status and disease stage at diagnosis. E Relationship of PKD1 mutations versus age of patient at diagnosis. F Segregation of PKD1 mutations by sex in MSI versus MSS tumors. G Relationship between PKD1 mutation and tumor histology in CRCs. *p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001 for all graphs. H Range of PKD1 mRNA expression in 524 CRC samples reported in cBioPortal. I Overall survival following initial treatment of CRC tumors expressing top quartile (red) or bottom quartile (blue) PKD1 mRNA expression. Note, datasets with survival information do not include MSS/MSI status. J Distribution of mRNA expression levels in CRC tumors analyzed at stages I–IV. K Expression of PKD1 mRNA in tumors of mucinous versus non-mucinous histology.
PKD1 mRNA expression varied over a significant range (~12-fold) in human CRCs (Fig. 6H). Notably, comparison of CRC with PKD1 expression in the highest quartile versus the lowest quartile showed significantly lower overall survival after treatment in individuals with tumors expressing the higher levels of PKD1 (Fig. 6I). Analysis of PKD1 expression by tumor stage indicated no significant difference in PKD1 expression at distinct stages of CRC (Fig. 6J), suggesting the survival differences were not attributable to higher PKD1 expression in later stage tumors. Given the differences seen in the frequency of PKD1 mutations in mucinous versus non-mucinous tumors, we also investigated mRNA expression differences for PKD1 in these histologies, but noted no differences (Fig. 6K).
Discussion
As major conclusions, this study for the first time directly demonstrates that an intact Pkd1 gene supports early stages of CRC formation, and that loss of PKD1 is tumor suppressive in adenoma formation mediated by loss of APC in a mouse model. These tumor suppressive activities of PKD1 loss are not linked to suppression of pro-proliferative signaling by CTNNB1 or other oncogenic pathways, but associated with increased colon barrier function, increased expression of claudins that mediate increased colon epithelial barrier function, and decreased leukocyte infiltration. Although loss of PKD1 does not differentially affect the response of organoids or tumors to cytotoxic regimens, the lower baseline of tumors in Apc−/−Pkd1−/− mice results in a more effective reduction in total tumor burden following treatment with these agents. Finally, analysis of public human data suggest PKD1 mutations are more likely to occur in MSI versus MSS CRC, and to be found in mucinous versus non-mucinous tumors, differing from the distribution of PKD1 wt CRC; further, expression of higher levels of PKD1 is associated with poorer outcomes for CRC recurrence and survival.
Extensive studies of functional changes in cell and tissue organization in the context of ADPKD have demonstrated an extremely pleiotropic activity, in which mutations reducing PKD1 expression or function produce phenotypes similar to those observed in tumors and described as the “hallmarks of cancer”, albeit typically in attenuated form [56]. However, in spite of these many similarities in phenotypes and signaling, ADPKD cysts are absolutely distinguished from tumors in their retention of a monolayer growth habit. This distinction argues for a strong PKD1-dependent inhibitory signal restricting invasive growth. Stabilization of impermeable tight junctions (TJs)—an essential feature of cyst pathogenesis mediated by claudin reprogramming [47]—is a strong candidate for such an inhibitory signal, as loss of TJs is the first step in the process of EMT that triggers invasion. Specifically, our data is compatible with the idea that loss of Pkd1 activates a CFTR-CLDN7 signaling axis to restrict TEI, based on data in this study and independent publications showing enhanced intestinal permeability and colon inflammation following inactivation of Cftr and Cldn7 [46, 48, 52, 57, 58]. In future work, it would be of interest to compare the activity of Pkd1 mutation in restricting CRC formation in azoxymethane/DSS-induced tumorigenesis models [59] to the effect observed here in models based on mutation of Apc.
There are numerous differences between the biology of ADPKD and CRC. In one notable example, in normal kidney tissues and in ADPKD, PKD1 function is typically linked to the action of the protein on cilia [60, 61]; in contrast, few if any colon epithelial cells are ciliated [62, 63]. In ADPKD, signaling pathways activated in renal cysts by loss of PKD1 include many known to be activated and growth-promoting in cancer including WNT/CTNNB1, PI3K/AKT, mTOR/S6K, RAF/MEK/ERK, SRC, MYC, AURKA, and others [56, 64, 65]. Some prior studies of PKD1 have documented a role of overexpressed PKD1 in activating of some of these signaling pathways in CRC and other cancer cells [16, 66]. Other pathways influenced by loss of PKD1 in ADPKD include planar cell polarity (non-canonical WNT) and the Hippo contact-inhibition associated pathways [67, 68]; these also are relevant to CRC etiology (e.g., [69]).
This study does not assign a single specific signaling mechanism by which loss of PKD1 inhibits CRC, although it excludes inhibition of WNT/CTNNB1 as one possibility, based on elevation of this pathway in Pkd1-mutated murine adenomas or in human tumors bearing somatic PKD1 mutations. A curious element of this study is the fact that CTNNB1 total levels are elevated, and WNT/CTNNB1-dependent transcript levels are elevated, but we are unable to detect a shift in distribution from cell periphery to nucleus for the CTNNB1 protein. This suggests that the increase in CTNNB1-dependent transcription reflects the overall increase in the levels of cellular CTNNB1, but not a specific activation of the pathway. In contrast, there is little evidence for PKD1 mutations stimulating activity of growth-inhibitory signaling proteins, with the exception of CFTR. A point bearing further study is the relationship of Pkd1 mutation and reduction of proliferation (Ki-67 staining), which is striking in organoids, but less notable in vivo in this study. One explanation might be the fact that while loss of Pkd1 is growth-inhibitory in a cell-autonomous manner in vitro, factors in the nascent tumor microenvironment—perhaps mediated by immune system components—ameliorate this effect in vivo. In this context, it is intriguing that trichrome and α-SMA staining of lesions indicates altered collagen deposition patterns and increased numbers of myofibroblasts associated with the Apc−/−Pkd1−/− genotype. Myofibroblasts and their secreted extracellular matrix can have significant effects on tumor growth, immunological response, and response to treatment [70]. The fact that leukocyte infiltration patterns are altered and reduced by absence of Pkd1, both in lesions and following treatment with DSS, may in part be influenced indirectly by changes in the tumor microenvironment—a topic of considerable interest for future investigations.
Finally, earlier studies have connected PKD1 mutations and CRC risk [8], and also suggested that elevated levels of PKD1 in late-stage tumors and tumor-derived cell lines are associated with more invasive phenotypes due to increased EMT and migration, and worse survival [16]. By reinforcing and extending these earlier studies, this study raises the possibility that not only the risk, but also the presentation and clinical response of CRC—and potentially inflammatory bowel disease—may differ in individuals with ADPKD versus the general population. The tendency of PKD1 mutations to occur in MSI tumors identified here is of interest. Given the distinct biology of MSS tumors, which typically (~80%) originate with APC mutations, and MSI tumors, where APC mutations are less common and other activating mutations are more prevalent [71], it is possible that PKD1 mutation and restriction of TEI is selectively detrimental in the MSS sequence of tumor formation and progression; although additional work is required to determine whether the relationship is causal or correlative. Estimates of the rate of APDKD in the population suggest as many as 1 in 1000 individuals may have inherited gene variants damaging PKD1 or its partner, PKD2 [12, 72]; reflecting ~300,000 individuals in the United States. Data from this study suggests that studies examining the features of diseases affecting the gastrointestinal tract in the ADPKD population would be merited.
Data availability
RNA sequencing data have been deposited in the NCBI database (url pending). Other primary data are available via application to the corresponding author.
References
American_Cancer_Society. Colorectal cancer facts and figures 2014–2016. Atlanta: American Cancer Society I; 2014.
Fearon ER, Vogelstein B. A genetic model for colorectal tumorigenesis. Cell. 1990;61:759–67.
Jones S, Chen WD, Parmigiani G, Diehl F, Beerenwinkel N, Antal T, et al. Comparative lesion sequencing provides insights into tumor evolution. Proc Natl Acad Sci USA. 2008;105:4283–8.
Glaire MA, Brown M, Church DN, Tomlinson I. Cancer predisposition syndromes: lessons for truly precision medicine. J Pathol. 2017;241:226–35.
Scarpa A, Chang DK, Nones K, Corbo V, Patch AM, Bailey P, et al. Whole-genome landscape of pancreatic neuroendocrine tumours. Nature. 2017;543:65–71.
Ma H, Brosens LAA, Offerhaus GJA, Giardiello FM, de Leng WWJ, Montgomery EA. Pathology and genetics of hereditary colorectal cancer. Pathology. 2018;50:49–59.
Huyghe JR, Bien SA, Harrison TA, Kang HM, Chen S, Schmit SL, et al. Discovery of common and rare genetic risk variants for colorectal cancer. Nat Genet. 2019;51:76–87.
Wetmore JB, Calvet JP, Yu AS, Lynch CF, Wang CJ, Kasiske BL, et al. Polycystic kidney disease and cancer after renal transplantation. J Am Soc Nephrol. 2014;25:2335–41.
Cornec-Le Gall E, Torres VE, Harris PC. Genetic complexity of autosomal dominant polycystic kidney and liver diseases. J Am Soc Nephrol. 2018;29:13–23.
Heyer CM, Sundsbak JL, Abebe KZ, Chapman AB, Torres VE, Grantham JJ, et al. Predicted mutation strength of nontruncating PKD1 mutations aids genotype-phenotype correlations in autosomal dominant polycystic kidney disease. J Am Soc Nephrol. 2016;27:2872–84.
Ward CJ, Wu Y, Johnson RA, Woollard JR, Bergstralh EJ, Cicek MS, et al. Germline PKHD1 mutations are protective against colorectal cancer. Hum Genet. 2011;129:345–9.
Cornec-Le Gall E, Alam A, Perrone RD. Autosomal dominant polycystic kidney disease. Lancet. 2019;393:919–35.
Geng L, Segal Y, Pavlova A, Barros EJ, Lohning C, Lu W, et al. Distribution and developmentally regulated expression of murine polycystin. Am J Physiol. 1997;272:F451–459.
Conduit SE, Hakim S, Feeney SJ, Ooms LM, Dyson JM, Abud HE, et al. Beta-catenin ablation exacerbates polycystic kidney disease progression. Hum Mol Genet. 2019;28:230–44.
Wuebken A, Schmidt-Ott KM. WNT/beta-catenin signaling in polycystic kidney disease. Kidney Int. 2011;80:135–8.
Gargalionis AN, Korkolopoulou P, Farmaki E, Piperi C, Dalagiorgou G, Adamopoulos C, et al. Polycystin-1 and polycystin-2 are involved in the acquisition of aggressive phenotypes in colorectal cancer. Int J Cancer. 2015;136:1515–27.
Outeda P, Huso DL, Fisher SA, Halushka MK, Kim H, Qian F, et al. Polycystin signaling is required for directed endothelial cell migration and lymphatic development. Cell reports. 2014;7:634–44.
Piontek K, Menezes LF, Garcia-Gonzalez MA, Huso DL, Germino GG. A critical developmental switch defines the kinetics of kidney cyst formation after loss of Pkd1. Nat Med. 2007;13:1490–5.
Piontek KB, Huso DL, Grinberg A, Liu L, Bedja D, Zhao H, et al. A functional floxed allele of Pkd1 that can be conditionally inactivated in vivo. J Am Soc Nephrol. 2004;15:3035–43.
Nikonova AS, Plotnikova OV, Serzhanova V, Efimov A, Bogush I, Cai KQ, et al. Nedd9 restrains renal cystogenesis in Pkd1-/- mice. Proc Natl Acad Sci USA. 2014;111:12859–64.
Nikonova AS, Deneka AY, Kiseleva AA, Korobeynikov V, Gaponova A, Serebriiskii IG, et al. Ganetespib limits ciliation and cystogenesis in autosomal-dominant polycystic kidney disease (ADPKD). FASEB J. 2018;32:2735–46.
Seeger-Nukpezah T, Proia DA, Egleston BL, Nikonova AS, Kent T, Cai KQ, et al. Inhibiting the HSP90 chaperone slows cyst growth in a mouse model of autosomal dominant polycystic kidney disease. Proc Natl Acad Sci USA. 2013;110:12786–91.
Grivennikov SI, Wang K, Mucida D, Stewart CA, Schnabl B, Jauch D, et al. Adenoma-linked barrier defects and microbial products drive IL-23/IL-17-mediated tumour growth. Nature. 2012;491:254–8.
Hinoi T, Loda M, Fearon ER. Silencing of CDX2 expression in colon cancer via a dominant repression pathway. J Biol Chem. 2003;278:44608–16.
Hinoi T, Akyol A, Theisen BK, Ferguson DO, Greenson JK, Williams BO, et al. Mouse model of colonic adenoma-carcinoma progression based on somatic Apc inactivation. Cancer Res. 2007;67:9721–30.
Billips LG, Petitte D, Landreth KS. Bone marrow stromal cell regulation of B lymphopoiesis: interleukin-1 (IL-1) and IL-4 regulate stromal cell support of pre-B cell production in vitro. Blood. 1990;75:611–9.
Boj SF, Hwang CI, Baker LA, Chio II, Engle DD, Corbo V, et al. Organoid models of human and mouse ductal pancreatic cancer. Cell. 2015;160:324–38.
Deneka AY, Kopp MC, Nikonova AS, Gaponova AV, Kiseleva AA, Hensley HH, et al. Nedd9 restrains autophagy to limit growth of early stage non-small cell lung cancer. Cancer Res. 2021;81:3717–26.
Debnath J, Muthuswamy SK, Brugge JS. Morphogenesis and oncogenesis of MCF-10A mammary epithelial acini grown in three-dimensional basement membrane cultures. Methods. 2003;30:256–68.
Trapnell C, Pachter L, Salzberg SL. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics. 2009;25:1105–11.
Trapnell C, Williams BA, Pertea G, Mortazavi A, Kwan G, van Baren MJ, et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat Biotechnol. 2010;28:511–5.
Trapnell C, Hendrickson DG, Sauvageau M, Goff L, Rinn JL, Pachter L. Differential analysis of gene regulation at transcript resolution with RNA-seq. Nat Biotechnol. 2013;31:46–53.
Fragiadaki M, Macleod FM, Ong ACM. The controversial role of fibrosis in autosomal dominant polycystic kidney disease. Int J Mol Sci. 2020;21:8936.
van de Wetering M, Sancho E, Verweij C, de Lau W, Oving I, Hurlstone A, et al. The beta-catenin/TCF-4 complex imposes a crypt progenitor phenotype on colorectal cancer cells. Cell. 2002;111:241–50.
Zhu G, Wang Y, Huang B, Liang J, Ding Y, Xu A, et al. A Rac1/PAK1 cascade controls beta-catenin activation in colon cancer cells. Oncogene. 2012;31:1001–12.
Grantham JJ. Polycystic kidney disease: neoplasia in disguise. Am J Kidney Dis. 1990;15:110–6.
Grantham JJ, Nair V, Winklhoffer F. Cystic diseases of the kidney. In: Brenner BM, editors. Brenner & rector’s the kidney. Philadelphia: WB Saunders Company; 2000. p. 1699–730.
Choi CR, Bakir IA, Hart AL, Graham TA. Clonal evolution of colorectal cancer in IBD. Nat Rev Gastroenterol Hepatol. 2017;14:218–29.
Gonzalez N, Prieto I, Del Puerto-Nevado L, Portal-Nunez S, Ardura JA, Corton M, et al. 2017 update on the relationship between diabetes and colorectal cancer: epidemiology, potential molecular mechanisms and therapeutic implications. Oncotarget. 2017;8:18456–85.
Park J, Morley TS, Kim M, Clegg DJ, Scherer PE. Obesity and cancer-mechanisms underlying tumour progression and recurrence. Nat Rev Endocrinol. 2014;10:455–65.
Rasool S, Kadla SA, Rasool V, Ganai BA. A comparative overview of general risk factors associated with the incidence of colorectal cancer. Tumour biology : the journal of the International Society for Oncodevelopmental Biology and Medicine. 2013;34:2469–76.
Wang K, Kim MK, Di Caro G, Wong J, Shalapour S, Wan J, et al. Interleukin-17 receptor a signaling in transformed enterocytes promotes early colorectal tumorigenesis. Immunity. 2014;41:1052–63.
Taniguchi K, Wu LW, Grivennikov SI, de Jong PR, Lian I, Yu FX, et al. A gp130-Src-YAP module links inflammation to epithelial regeneration. Nature. 2015;519:57–62.
Meira LB, Bugni JM, Green SL, Lee CW, Pang B, Borenshtein D, et al. DNA damage induced by chronic inflammation contributes to colon carcinogenesis in mice. J Clin Invest. 2008;118:2516–25.
Suzuki R, Kohno H, Sugie S, Nakagama H, Tanaka T. Strain differences in the susceptibility to azoxymethane and dextran sodium sulfate-induced colon carcinogenesis in mice. Carcinogenesis. 2006;27:162–9.
Gunzel D, Yu AS. Claudins and the modulation of tight junction permeability. Physiol Rev. 2013;93:525–69.
Yu AS, Kanzawa SA, Usorov A, Lantinga-van Leeuwen IS, Peters DJ. Tight junction composition is altered in the epithelium of polycystic kidneys. J Pathol. 2008;216:120–8.
Tanaka H, Takechi M, Kiyonari H, Shioi G, Tamura A, Tsukita S. Intestinal deletion of Claudin-7 enhances paracellular organic solute flux and initiates colonic inflammation in mice. Gut. 2015;64:1529–38.
Ikeda M, Fong P, Cheng J, Boletta A, Qian F, Zhang XM, et al. A regulatory role of polycystin-1 on cystic fibrosis transmembrane conductance regulator plasma membrane expression. Cell Physiol Biochem. 2006;18:9–20.
Than BL, Linnekamp JF, Starr TK, Largaespada DA, Rod A, Zhang Y, et al. CFTR is a tumor suppressor gene in murine and human intestinal cancer. Oncogene. 2016;35:4179–87.
Scott P, Anderson K, Singhania M, Cormier R. Cystic fibrosis, CFTR, and colorectal cancer. Int J Mol Sci. 2020;21:2891.
De Lisle RC. Disrupted tight junctions in the small intestine of cystic fibrosis mice. Cell Tissue Res. 2014;355:131–42.
Deeks ED. Lumacaftor/ivacaftor: a review in cystic fibrosis. Drugs. 2016;76:1191–201.
Osawa Y, Oboki K, Imamura J, Kojika E, Hayashi Y, Hishima T, et al. Inhibition of cyclic adenosine monophosphate (cAMP)-response element-binding protein (CREB)-binding protein (CBP)/beta-catenin reduces liver fibrosis in mice. EBioMedicine. 2015;2:1751–8.
Saltz LB, Cox JV, Blanke C, Rosen LS, Fehrenbacher L, Moore MJ, et al. Irinotecan plus fluorouracil and leucovorin for metastatic colorectal cancer. Irinotecan Study Group. N Engl J Med. 2000;343:905–14.
Seeger-Nukpezah T, Geynisman DM, Nikonova AS, Benzing T, Golemis EA. The hallmarks of cancer: relevance to the pathogenesis of polycystic kidney disease. Nat Rev Nephrol. 2015;11:515–34.
Xu C, Wang K, Ding YH, Li WJ, Ding L. Claudin-7 gene knockout causes destruction of intestinal structure and animal death in mice. World J Gastroenterol. 2019;25:584–99.
Zhu L, Han J, Li L, Wang Y, Li Y, Zhang S. Claudin family participates in the pathogenesis of inflammatory Bowel diseases and colitis-associated colorectal cancer. Front Immunol. 2019;10:1441.
Snider AJ, Bialkowska AB, Ghaleb AM, Yang VW, Obeid LM, Hannun YA. Murine model for colitis-associated cancer of the colon. Methods Mol Biol. 2016;1438:245–54.
Hu J, Harris PC. Regulation of polycystin expression, maturation and trafficking. Cell Signal. 2020;72:109630.
Pan J, Seeger-Nukpezah T, Golemis EA. The role of the cilium in normal and abnormal cell cycles: emphasis on renal cystic pathologies. Cell Mol Life Sci. 2012;70:1849–74.
Paul C, Tang R, Longobardi C, Lattanzio R, Eguether T, Turali H, et al. Loss of primary cilia promotes inflammation and carcinogenesis. EMBO Rep. 2022;23:e55687.
Rocha C, Papon L, Cacheux W, Marques Sousa P, Lascano V, Tort O, et al. Tubulin glycylases are required for primary cilia, control of cell proliferation and tumor development in colon. EMBO J. 2014;33:2247–60.
Korobeynikov V, Deneka AY, Golemis EA. Mechanisms for nonmitotic activation of Aurora-A at cilia. Biochem Soc Trans. 2017;45:37–49.
Lal M, Song X, Pluznick JL, Di Giovanni V, Merrick DM, Rosenblum ND, et al. Polycystin-1 C-terminal tail associates with beta-catenin and inhibits canonical Wnt signaling. Hum Mol Genet. 2008;17:3105–17.
Papavassiliou KA, Zoi I, Gargalionis AN, Koutsilieris M. Polycystin-1 affects cancer cell behaviour and interacts with mTOR and Jak signalling pathways in cancer cell lines. J Cell Mol Med. 2019;23:6215–27.
Happe H, van der Wal AM, Leonhard WN, Kunnen SJ, Breuning MH, de Heer E, et al. Altered Hippo signalling in polycystic kidney disease. J Pathol. 2011;224:133–42.
Luyten A, Su X, Gondela S, Chen Y, Rompani S, Takakura A, et al. Aberrant regulation of planar cell polarity in polycystic kidney disease. J Am Soc Nephrol. 2010;21:1521–32.
Harvey KF, Zhang X, Thomas DM. The Hippo pathway and human cancer. Nat Rev Cancer. 2013;13:246–57.
Sahai E, Astsaturov I, Cukierman E, DeNardo DG, Egeblad M, Evans RM, et al. A framework for advancing our understanding of cancer-associated fibroblasts. Nat Rev Cancer. 2020;20:174–86.
Lieu CH, Golemis EA, Serebriiskii IG, Newberg J, Hemmerich A, Connelly C, et al. Comprehensive genomic landscapes in early and later onset colorectal cancer. Clin Cancer Res. 2019;25:5852–8.
Lanktree MB, Haghighi A, Guiard E, Iliuta IA, Song X, Harris PC, et al. Prevalence estimates of polycystic kidney and liver disease by population sequencing. J Am Soc Nephrol. 2018;29:2593–600.
Acknowledgements
We are grateful for support from Dr. Margret Einarson of the Fox Chase Cancer Center (FCCC) High Throughput Screening facility, and Dr. Kathy Cai of the FCCC Histopathology Facility. We thank Drs. Ilya Serebriiskii and Kerry Campbell for helpful discussions and comments. We thank Ms. Anna Lilly for assistance with immunofluorescence analysis. We are grateful to Dr. Gregory Germino, NIDDK, for the gift of the floxed Pkd1 mice. The authors and study received support from NIH R01s DK108195 and CA228187, the William Wikoff Smith Charitable Trust and S10 OD021754 (to EAG); by NIH T32 CA009035 (to AN); by a William J. Avery Postdoctoral Fellowship from Fox Chase Cancer Center (to AD); by Marie Skłodowska-Curie grant No 896865 from the European Union’s Horizon 2020 research and innovation program (to RT), NIH R01CA227629 and CA218133 (to SIG) and by the NCI Core Grant P30 CA006927 (to Fox Chase Cancer Center).
Author information
Authors and Affiliations
Contributions
ASN, and AYD were responsible for data acquisition, analysis, interpreting results, writing the manuscript draft and figures. FNS and SP were responsible for data acquisition, extracting and analyzing data and interpreting results. RT and AAK were responsible for initial experimental design, data acquisition and interpretation. YZ was responsible for RNA seq data analysis. DBF was responsible for pathology data verification and interpretation. EN and SIG contributed to initial conceptualization, interpreting results, and provided feedback on the manuscript. EAG was responsible for initial conceptualization, funds acquisition, data interpretation and verification, writing the manuscript, and updating reference lists.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Nikonova, A.S., Deneka, A.Y., Silva, F.N. et al. Loss of Pkd1 limits susceptibility to colitis and colorectal cancer. Oncogenesis 12, 40 (2023). https://doi.org/10.1038/s41389-023-00486-y
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41389-023-00486-y