Abstract
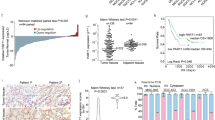
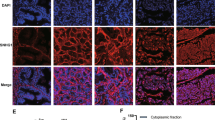
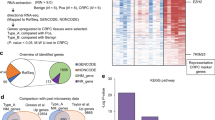
Long noncoding RNAs (lncRNAs) play pivotal roles in cancer development and progression, and some function in a highly cancer-specific manner. However, whether the cause of their expression is an outcome of a specific regulatory mechanism or nonspecific transcription induced by genome reorganization in cancer remains largely unknown. Here, we investigated a group of lncRNAs that we previously identified to be aberrantly expressed in prostate cancer (PC), called TPCATs. Our high-throughput real-time PCR experiments were integrated with publicly available RNA-seq and ChIP-seq data and revealed that the expression of a subset of TPCATs is driven by PC-specific transcription factors (TFs), especially androgen receptor (AR) and ETS-related gene (ERG). Our in vitro validations confirmed that AR and ERG regulated a subset of TPCATs, most notably for EPCART. Knockout of EPCART was found to reduce migration and proliferation of the PC cells in vitro. The high expression of EPCART and two other TPCATs (TPCAT-3-174133 and TPCAT-18-31849) were also associated with the biochemical recurrence of PC in prostatectomy patients and were independent prognostic markers. Our findings suggest that the expression of numerous PC-associated lncRNAs is driven by PC-specific mechanisms and not by random cellular events that occur during cancer development. Furthermore, we report three prospective prognostic markers for the early detection of advanced PC and show EPCART to be a functionally relevant lncRNA in PC.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J, Jemal A. Global cancer statistics, 2012. CA Cancer J Clin. 2015;65:87–108.
Yu J, Yu J, Mani R, Cao Q, Brenner CJ, Cao X, et al. An integrated network of androgen receptor, polycomb, and TMPRSS2-ERG gene fusions in prostate cancer progression. Cancer Cell. 2010;17:443–54.
Chen Y, Chi P, Rockowitz S, Iaquinta PJ, Shamu T, Shukla S, et al. ETS factors reprogram the androgen receptor cistrome and prime prostate tumorigenesis in response to PTEN loss. Nat Med. 2013;19:1023–9.
Pomerantz MM, Li F, Takeda DY, Lenci R, Chonkar A, Chabot M, et al. The androgen receptor cistrome is extensively reprogrammed in human prostate tumorigenesis. Nat Genet. 2015;47:1346.
Kron KJ, Murison A, Zhou S, Huang V, Yamaguchi TN, Shiah YJ, et al. TMPRSS2-ERG fusion co-opts master transcription factors and activates NOTCH signaling in primary prostate cancer. Nat Genet. 2017;49:1336–45.
Tomlins SA, Rhodes DR, Perner S, Dhanasekaran SM, Mehra R, Sun X, et al. Recurrent fusion of TMPRSS2 and ETS transcription factor genes in prostate cancer. Science. 2005;310:644–8.
Cancer Genome Atlas Research Network. The molecular taxonomy of primary prostate cancer. Cell. 2015;163:1011–25.
Cerveira N, Ribeiro FR, Peixoto A, Costa V, Henrique R, Jerónimo C, et al. TMPRSS2-ERG gene fusion causing ERG overexpression precedes chromosome copy number changes in prostate carcinomas and paired HGPIN lesions. Neoplasia. 2006;8:826–32.
Perner S, Mosquera J, Demichelis F, Hofer MD, Paris PL, Simko J, et al. TMPRSS2-ERG fusion prostate cancer: an early molecular event associated with invasion. Am J Surg Pathol. 2007;31:882–8.
Tomlins SA, Laxman B, Varambally S, Cao X, Yu J, Helgeson BE. et al. Role of the TMPRSS2-ERG gene fusion in prostate cancer. Neoplasia. 2008;10:177–88.
Carver BS, Tran J, Gopalan A, Chen Z, Shaikh S, Carracedo A, et al. Aberrant ERG expression cooperates with loss of PTEN to promote cancer progression in the prostate. Nat Genet. 2009;41:619–24.
Schmitt AM, Chang HY. Long noncoding RNAs in cancer pathways. Cancer Cell. 2016;29:452–63.
Martens-Uzunova ES, Böttcher R, Croce CM, Jenster G, Visakorpi T, Calin GA. Long noncoding RNA in prostate, bladder, and kidney cancer. Eur Urol. 2014;65:1140–51.
Sahu A, Singhal U, Chinnaiyan AM. Long noncoding RNAs in cancer: from function to translation. Trends Cancer. 2015;1:93–109.
Hessels D, Klein Gunnewiek JMT, van Oort I, Karthaus HFM, van Leenders, et al. DD3(PCA3)-based molecular urine analysis for the diagnosis of prostate cancer. Eur Urol. 2003;44:16.
Prensner JR, Zhao S, Erho N, Schipper M, Iyer MK, Dhanasekaran SM, et al. RNA biomarkers associated with metastatic progression in prostate cancer: a multi-institutional high-throughput analysis of SChLAP1. Lancet Oncol. 2014;15:1469–80.
Shukla S, Zhang X, Niknafs YS, Xiao L, Mehra R, Cieslik M, et al. Identification and validation of PCAT14 as prognostic biomarker in prostate cancer. Neoplasia. 2016;18:489–99.
White NM, Zhao SG, Zhang J, Rozycki EB, Dang HX, McFadden SD, et al. Multi-institutional analysis shows that low PCAT-14 expression associates with poor outcomes in prostate cancer. Eur Urol. 2017;71:257–66.
Prensner JR, Iyer MK, Balbin OA, Dhanasekaran SM, Cao Q, Brenner JC, et al. Transcriptome sequencing identifies PCAT-1, a Novel lincRNA implicated in prostate cancer progression. Nat Biotechnol. 2011;29:742–9.
Ylipää A, Kivinummi K, Kohvakka A, Annala M, Latonen L, Scaravilli M, et al. Transcriptome sequencing reveals PCAT5 as a Novel ERG-regulated long noncoding RNA in prostate cancer. Cancer Res. 2015;75:4026–31.
Boormans JL, Porkka K, Visakorpi T, Trapman J. Confirmation of the association of TMPRSS2(exon 0):ERG expression and a favorable prognosis of primary prostate cancer. Eur Urol. 2011;60:183–4.
Chawla K, Tripathi S, Thommesen L, Lægreid A, Kuiper M. TFcheckpoint: a curated compendium of specific DNA-binding RNA polymerase II transcription factors. Bioinformatics. 2013;29:2519–20.
Yang L, Lin C, Jin C, Yang JC, Tanasa B, Li W, et al. lncRNA-dependent mechanisms of androgen-receptor- regulated gene activation programs. Nature. 2013;500:598.
Takayama K, Horie-Inoue K, Katayama S, Suzuki T, Tsutsumi S, Ikeda K, et al. Androgen-responsive long noncoding RNA CTBP1-AS promotes prostate cancer. EMBO J. 2013;32:1665–80.
Zhang A, Zhao JC, Kim J, Fong K, Yang YA, Chakravarti D, et al. LncRNA HOTAIR enhances the androgen-receptor-mediated transcriptional program and drives castration-resistant prostate cancer. Cell Rep. 2015;13:209–21.
Zhang Y, Pitchiaya S, Cieślik M, Niknafs YS, Tien JC-, Hosono Y, et al. Analysis of the androgen receptor-regulated lncRNA landscape identifies a role for ARLNC1 in prostate cancer progression. Nat Genet. 2018;50:814–24.
The ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature. 2012;489:57–74.
Linja MJ, Savinainen KJ, Saramäki OR, Tammela TL, Vessella RL, Visakorpi T. Amplification and overexpression of androgen receptor gene in hormone-refractory prostate cancer. Cancer Res. 2001;61:3550–5.
White NM, Cabanski CR, Silva-Fisher JM, Dang HX, Govindan R, Maher CA. Transcriptome sequencing reveals altered long intergenic non-coding RNAs in lung cancer. Genome Biol. 2014;15:429.
Su X, Malouf GG, Chen Y, Zhang J, Yao H, Valero V, et al. Comprehensive analysis of long non-coding RNAs in human breast cancer clinical subtypes. Oncotarget. 2014;5:9864–76.
Yan X, Hu Z, Feng Y, Hu X, Yuan J, Zhao SD, et al. Comprehensive genomic characterization of long non-coding RNAs across human cancers. Cancer Cell. 2015;28:529–40.
Perner S, Demichelis F, Beroukhim R, Schmidt FH, Mosquera J, Setlur S, et al. TMPRSS2:ERG fusion-associated deletions provide insight into the heterogeneity of prostate cancer. Cancer Res. 2006;66:8337–41.
Nam RK, Sugar L, Wang Z, Yang W, Kitching R, Klotz LH, et al. Expression of TMPRSS2:ERG gene fusion in prostate cancer cells is an important prognostic factor for cancer progression. Cancer Biol Ther. 2007;6:40–45.
Weischenfeldt J, Simon R, Feuerbach L, Schlangen K, Weichenhan D, Minner S, et al. Integrative genomic analyses reveal an androgen-driven somatic alteration landscape in early-onset prostate cancer. Cancer Cell. 2013;23:159–70.
Böttcher R, Hoogland AM, Dits N, Verhoef EI, Kweldam C, Waranecki P, et al. Novel long non-coding RNAs are specific diagnostic and prognostic markers for prostate cancer. Oncotarget. 2015;6:4036–50.
Waltering KK, Helenius MA, Sahu B, Manni V, Linja MJ, Jänne OA, et al. Increased expression of androgen receptor sensitizes prostate cancer cells to low levels of androgens. Cancer Res. 2009;69:8141–9.
Davis CA, Hitz BC, Sloan CA, Chan ET, Davidson JM, Gabdank I, et al. The encyclopedia of DNA elements (ENCODE): data portal update. Nucleic Acids Res. 2018;46:D794–801.
Urbanucci A, Sahu B, Seppälä J, Larjo A, Latonen LM, Waltering KK, et al. Overexpression of androgen receptor enhances the binding of the receptor to the chromatin in prostate cancer. Oncogene. 2012;31:2153–63.
Acknowledgements
This study was supported by grants from the Academy of Finland (TV 317755, MN 310829, and LL 317871), Sigrid Juselius Foundation (TV and LL), Cancer Society of Finland, Business Finland, the Finnish Cultural Foundation (AK), the European Union’s Horizon 2020 (MS, TransPot - 721746), Norwegian Cancer Society grant (AU 198016-2018), Research collegium of the University of Tampere/IASR (KK). The authors want to thank Jenni Jouppila, Paula Kosonen, Riina Kylätie, Päivi Martikainen, Hanna Selin, and Marika Vähä-Jaakkola for their technical assistance and Tampere Imaging Facility (TIF) for their service. The results published here are in part based upon data generated by The Cancer Genome Atlas project (dbGaP Study Accession: phs000178.v9.p8) established by the NCI and NHGRI. Information about TCGA can be found at http://cancergenome.nih.gov. We acknowledge ENCODE Consortium and the ENCODE production laboratories for generating the DNase-seq data.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Kohvakka, A., Sattari, M., Shcherban, A. et al. AR and ERG drive the expression of prostate cancer specific long noncoding RNAs. Oncogene 39, 5241–5251 (2020). https://doi.org/10.1038/s41388-020-1365-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-020-1365-6