Abstract
The mammalian neocortex is organized into layers distinguished by the size, packing density, and connectivity of their constituent neurons. Many neuropsychiatric illnesses are complex trait disorders with etiologic factors converging on neuronal protein networks. Cortical pathology of neuropsychiatric diseases, such as schizophrenia, is often restricted to, or more pronounced in, certain cortical layers, suggesting that genetic vulnerabilities manifest with laminar specificity. Thus, the ability to investigate cortical layer-specific protein levels in human postmortem brain is highly desirable. Here, we developed and validated a laser capture microdissection–mass spectrometry (LCM-MS) approach to quantify over 200 proteins in cortical layers 3 and 5 of two cohorts of human subjects as well as a monkey model of postmortem interval. LCM-MS was readily implementable and reliably identified protein patterns that differed between cortical layers 3 and 5. Our findings suggest that LCM-MS facilitates the precise quantification of proteins within individual cortical layers from human postmortem brain tissue, providing a powerful tool in the study of neuropsychiatric disease.
Similar content being viewed by others
Introduction
The mammalian neocortex is organized into layers that can be distinguished by the size and packing density of their constituent neurons. This laminar pattern arises from distinct populations of neocortical projection neurons that mature and migrate in waves throughout embryonic development [1, 2]. Neurons in each cortical layer tend to differ in their patterns of connectivity [3, 4] and the features of their dendritic spines [5]. For example, dendritic spines on layer 2/3 pyramidal neurons are denser and more dynamic than are those on layer 5 pyramidal neurons and have different rates of formation and loss during adolescence and learning [5].
Many neuropsychiatric illnesses are complex trait disorders in which multiple etiologic factors converge on, and likely impair, specific neuronal protein networks [6]. For example, genetic studies of schizophrenia suggest the involvement of postsynaptic protein networks, consistent with other evidence implicating impairments in the formation and/or maintenance of synapses [7, 8]. However, studies in cortical tissue from subjects with schizophrenia have not observed uniform alterations in neuronal or synaptic structures across all cortical layers [9, 10]. Instead, these studies have revealed pathology that is restricted to, or more pronounced in, certain cortical layers, suggesting that genetic vulnerabilities in synaptic connectivity are manifest with laminar specificity [10, 11]. Thus, the ability to investigate the neuronal protein networks of specific cortical layers from human postmortem brain tissue is highly desirable in the study of human neuropsychiatric illness.
Laser capture microdissection (LCM) has been paired with transcriptomic approaches to interrogate messenger RNA (mRNA) expression in cortical layers and single cell populations captured from human postmortem brain tissue [11,12,13,14]. However, brain mRNA transcript levels are only one of several factors influencing protein levels, which can limit the ability to infer protein levels from mRNA expression [15,16,17], even within a single cell population [17]. Thus, a complementary approach that can accurately quantify multiple proteins within select layers is needed. Recent advances in mass spectrometry (MS) instrument sensitivity now allow for proteomic analysis of laser captured tissue sections and even single cells in suitable model systems [18, 19]. However, to date, laser capture microdissection–mass spectrometry (LCM-MS) has seen limited use in human postmortem brain tissue [20, 21] and a systematic investigation of the approach has not been reported.
The goal of the present study was to test the utility of LCM-MS for reproducible, multiplexed protein quantification within cortical layers in human postmortem brain tissue. We chose to utilize targeted MS, also known as selected reaction monitoring (SRM), with a [13C6] lysine-labeled brain proteome internal standard ([13C6]brain ISTD) for its throughput, sensitivity, and precision [22]. A critical potential technical concern in human postmortem tissue studies is postmortem interval (PMI), the time interval between subject death and preservation of the brain tissue. PMI is known to affect levels of many proteins and could also possibly alter the localization of proteins within layers, for example, by causing dendritic proteins to diffuse or traffic back to the cell body. To assess this and other potential confounds (e.g., brain pH), we quantified protein levels in cortical layers 3 and 5 from 10 human subjects with a range of PMIs as well as a monkey brain tissue model of PMI. We found that LCM-MS was readily implementable and the pattern of protein levels distinguished cortical layers 3 and 5 in both humans and monkeys. Within the range of PMIs studied, only modest effects of PMI on individual protein levels or localization were observed. Our findings are in line with previous reports that cortical layers are distinguished by their molecular composition [12, 23] and confirm that LCM-MS can facilitate the precise quantification of proteins within individual cortical layers from human postmortem brain tissue, providing a powerful tool in the study of layer-specific alterations in neuropsychiatric disease.
Materials, subjects, and methods
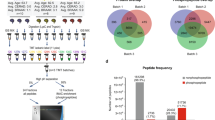
Figure 1 provides an overview of the experimental workflow.
Study overview. a Coronal blocks containing area 46 of the PFC were obtained from 10 male human subjects. Eight DLPFC tissue blocks simulating PMIs of 0, 6, 12, and 24 h were obtained from a single monkey. b Image one of a 12 µm section of human DLPFC after harvesting by LCM. The 4.8E7 µm3 of layer 3 and layer 5 (4E6 µm2 from a 12 µm section) were dissected from each human subject and each monkey PMI cortical block by LCM. Additionally, gray matter homogenates were prepared from the monkey blocks. c For each experiment (human cohort 1, human cohort 2, monkey LCM, and monkey homogenate), a pooled control was created within each experiment to control for variability in sample preparation and MS analysis and a different randomized sample order, keeping layers 3 and 5 from the same section together, was utilized at every experimental step: Addition of the [13C6] lysine-labeled neuronal proteome standard, trypsin digestion, and MS analysis. d All samples were analyzed by targeted MS
Human subjects
Brain specimens from male 10 subjects (Table 1) were obtained during autopsies conducted at the Allegheny County Medical Examiner’s Office after receiving consent from the next of kin. All procedures were approved by the University of Pittsburgh’s Committee for the Oversight of Research and Clinical Training Involving the Dead and Institutional Review Board for Biomedical Research. An independent panel of experienced clinicians made consensus Diagnostic and Statistical Manual of Mental Disorders Fourth Edition (DSM-IV) diagnoses for each subject on the basis of medical records and the results of structured interviews conducted with family members of the deceased [24]. Following brain removal at autopsy, the right hemisphere was dissected into 1 cm thick coronal blocks, flash-frozen in isopentane, and stored at −80 °C. Subjects were divided into two cohorts of 5: cohort 1 and cohort 2. Coronal blocks containing area 46 of the dorsolateral prefrontal cortex (DLPFC) were cut on a cryostat at 12 µM thickness. Sections were captured onto RNAse-free foil slides for microdissection (Leica Microsystems, Bannockburn, USA). The sections were stored up to 30 days at −80 °C until LCM. Cohorts 1 and 2 were processed separately at all stages including tissue blocking, slide mounting, LMC capture, targeted MS, raw data analysis, and initial statistical analysis.
Monkey
Tissue blocks contacting the DLPFC were obtained from one male long-tailed macaque as described previously [25, 26]. Briefly, the animal was sedated with ketamine hydrochloride, intubated, anesthetized with 1% halothane in 28% oxygen, placed in a stereotaxic apparatus, and a bilateral craniotomy over the frontal lobes was made. Eight 5 mm thick blocks from the DLPFC were removed (four from each hemisphere) and frozen immediately in isopentane (0 h PMI), or stored at room temperature in artificial cerebrospinal fluid for 6, 12, or 24 h (2 blocks at each time point) and then frozen (Table S1, Figure S1). From each block, 12 µM tissue sections were collected for LCM, and an additional 20 mg of gray matter was collected for extraction and analysis of total homogenate protein levels. Housing and experimental procedures were conducted in accordance with the guidelines of the US Department of Agriculture and the National Institutes of Health Guide for the Care and Use of Laboratory Animals and with the approval of the University of Pittsburgh Institutional Animal Care and Use Committee.
LCM harvest
Slides were stained for Nissl substance as previously described (Figure S2) [27, 28] and then immediately used for microdissection. Approximately 4.5E−6 µm2 of layer 3 or layer 5 each was laser microdissected (×5 magnification, Leica LMC6500) from each subject (Fig. 1a). Layers 3 and 5 in human neocortex were distinguished from the adjacent layers based on the following criteria. The borders of layer 3 are readily identified by the lower cell packing density and larger neuronal size bodies in layer 3 relative to layers 2 and 4. The border between layers 4 and 5 is also easily identified by the same features. The border between layers 5 and 6 is less sharp but the two layers are distinguished by the larger size and more uniform shape and orientation of pyramidal neurons in layer 5 relative to layer 6 (Figure S2). The resulting tissue fragments were collected into a protein extraction buffer (100 mM Tris-HCL, pH 7.4, 2% sodium dodecyl sulfate with protease and phosphatase inhibitors (SIGMA)) (Fig. 1b). For each tissue block from each brain, four tissue samples were harvested (two from layer 3 and two from layer 5). For the majority of subjects, all captures were taken from a single slide, though a minority required two slides to collect sufficient tissue. The order of capture was layer 3, layer 3, layer 5, layer 5. We have confirmed that layer capture order does not impact proteins levels (Figure S3). The harvest took ~90 min from stain to final collection for each slide.
[13C6]brain ISTD
The [13C6]brain ISTD is prepared by homogenizing cerebral cortex tissue from a Stable Isotope Labeling in Mammals (SILAM) mouse (Cambridge Isotopes). These animals are raised on a diet in which the only source of Lysine is 13C6 labeled, resulting in near complete (99%) labeling of the animal proteome in three generations. Labeling efficiency of each [13C6]brain ISTD preparation is confirmed prior to use. To account for species differences, SRMs are designed only for peptides that have 100% homology between humans and mice [22] taking advantage of the high level of synaptic protein sequence homology between these species [29]. We have previously demonstrated that when the same total protein amounts (as measured by micro bicinchoninic acid (BCA)) from [13C6]brain ISTD and human cortical homogenates are assayed, the resultant synaptic protein levels are highly similar [22], and we confirmed this for the protein levels in individual cortical layer captures (Figure S4).
Sample and pooled control preparation
A total of 2 μg of the [13C6]brain ISTD, reducing agent (Invitrogen), and lithium dodecyl sulfate loading buffer (Invitrogen) were added to each LCM capture. For homogenates, 10 μg of protein was mixed with 10 μg of [13C6]brain ISTD. These mixtures were then heated at 95 °C for 10 min. At this stage an aliquot was taken from each sample and the aliquots were combined to create a pooled control for each experiment to monitor for variability in sample preparation and MS analysis. Samples and pooled controls were processed together for targeted MS analysis by in-gel trypsin digestion as previously described [22]. Samples were then evaporated to completion, suspended in 12 µl H2O with 0.1% formic acid, and filtered with 0.22 µm Ultra free-MC filter cartridges (Millipore).
Mass spectrometry
Targeted MS analyses were conducted on a TSQ Quantiva triple stage quadrupole mass spectrometer (ThermoFisher Scientific) with an Ultimate 3000 HPLC (Dionex). Then, 3 µl (~0.75 μg protein) sample was loaded on to a Magic C18 column (Michrom) at 1 μl/min for 12 min and eluted at 400 nl/min over a 25 min gradient from 3–35% mobile phase B (acetonitrile containing 0.1% formic acid). Selected reaction monitoring transitions were timed using 1–1.5 min retention windows, depending on the number of SRMs to be assayed. Transitions were monitored, allowing for a cycle time of 1 s, resulting in a dynamic dwell time never falling below 10 ms. The MS instrument parameters were as follows: capillary temperature 275 °C, spray voltage 1350 V, and a collision gas of 1.4 mTorr (argon). The resolving power of the instrument was set to 0.7 Da (full width half maximum) for the first and third quadrupole. Data were acquired using a Chrom Filter peak width of 4.0 s (Fig. 1d).
Peptide selected reaction monitoring design and validation
The majority of selected reaction monitoring transitions for the peptides utilized in the present work were designed and validated in previous reports [22, 30]. Selected reaction monitoring transitions for additional synaptic peptides/proteins, using more recent spectral libraries, were designed and validated as previously described [22].
Data processing
Peak areas and area ratios were calculated within Skyline [31]. Raw files generated by targeted MS analysis were loaded into Skyline files containing target proteins/peptides/transitions. All individual selected reaction monitoring transitions and integration areas were manually inspected. Transitions for which the signal-to-noise ratio was below 3 were excluded from analysis. The ratios of the integrated areas for “light” endogenous peptides and “heavy” [13C6]brain ISTD peptides were calculated to obtain peptide measures using multiple transitions per peptide.
Statistics
To assess variabilities from sample preparation and targeted MS analysis, coefficients of variation (CVs) were calculated for each data cohort based on peptide values from the replicates of pooled samples (a measure of variability in sample preparation + targeted MS analysis) as well as based on peptide values from a single pooled sample analyzed multiple times (a measure of variably in MS analysis alone). Then, for multiple peptides mapped to the same protein, their mean abundance value, inversely weighted based on CV, was used as the protein-level measure for the downstream analysis. Specifically, the summation of two CVs was used: CV from the replicates of pooled samples and the CV from the multiple injections of a single pooled sample. For the two human cohorts, a linear regression model was fitted to model the correlation between each protein (log2 transformed) and PMI, for layer 3, layer 5 and the layer 5–3 difference, respectively. The p value for testing the regression slope (0 or not), as well as the corresponding false discovery rate adjusted q-values, were obtained for each protein. Similarly, for the monkey samples, a linear regression model was fitted to model the correlation between PMI and cortical levels and the layer 5–3 differences for each protein. Finally, we produced the Q–Q plot on p values from each analysis. Chi-Square 3 × 2 contingency table was used to compare overlap between significantly altered layer 5–3 proteins between human and monkey samples.
Unsupervised hierarchical clustering
Self-organized heat maps of proteins and subject by peptides values were prepared in Cluster 3.0 and Treeview [32, 33]. Values were log2 transformed, median centered by protein and subject, then normalized by peptide and subject. Un-centered correlation was used as the similarity metric and the clustering method was centroid linkage.
Raw and extracted datafiles
All raw files are available for download on chorus. Extracted light/heavy ratios for individual peptides are provided in Supplemental Tables 2–4.
Results
Peptide SRM validation
SRMs for peptides unique to 240 proteins passed quality control filters [22] during initial validation experiments in LCM cortical samples. Of these, 238 were deemed acceptable in human cohort 1, 213 in human cohort 2, and 222 in the monkey LCM and homogenate preparations (Tables S2–4).
Technical and biological variability
Instrument stability was assessed during each experiment via multiple injections of a single pooled control aliquot. Representative findings from the monkey whole cortex homogenates are presented in Figure S5A: mean CV was 0.1 with 75% of peptide CVs < 0.14. To assess reproducibility in sample preparation, four aliquots of a pooled control were independently subjected to trypsin digestion, desalting, and MS analysis in each experiment. Representative findings from human cohort 1 are presented in Figure S5B: mean CV for technical variance was 0.1 with 75% of peptide CVs < 0.13. Finally, variation in measured peptide levels across individual human subjects (which would comprise biological + sample preparation + instrument variance) ranged from a mean CV of 0.15 to 0.28 depending on cohort and layer (Figure S5C–F).
Postmortem interval effects
We first investigated the effects of PMI on protein levels in human tissue within and between layers. In each analysis, the levels of only 3–10% of proteins were nominally correlated with PMI (p < 0.05) and none passed multiple hypothesis testing (q < 0.1) (Figure S6A–F). Second, similar to human tissue, in monkey tissue protein levels did not vary with PMI; levels of only 3% of proteins were nominally correlated with PMI and none passed multiple hypothesis testing (Figure S7). Finally, as a measure of the stability of protein localization, we examined the correlation between layer 5–3 differences in monkeys; only 9% were nominally correlated with PMI and none passed multiple hypothesis testing (q < 0.1) (Figure S6B).
Brain pH effects
The layer 5–3 differences of only three proteins were nominally correlated with brain pH in the human subjects (p < 0.05). Likewise, only one protein was nominally correlated in layer 3 and one in layer 5 (q < 0.1). None passed multiple hypothesis testing (Table S9).
Effects of cortical layer
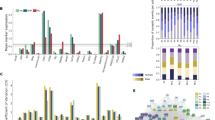
Peptide-level data were used to cluster subjects/layers. For each subject in the first human cohort, peptide/protein expression formed two clusters corresponding to layers 3 and 5 (Fig. 2a). We observed similar clustering in the second human cohort in all but one of the subjects (H1336, with a PMI of 18.4 h, Fig. 2b). In the monkey cohort, layers 3 and 5 from all but one tissue block clustered into two groups. The block in which layers 3 and 5 were not distinguished had the highest PMI, 24 h (Fig. 2c).
Unsupervised hierarchical clustering heat maps. Unsupervised hierarchical clustering was performed using Cluster 3 to organize subject/layers by peptide expression. a In human cohort 1, layers 3 and 5 segregated into two groups. b In human cohort 2, layers 3 and 5 segregated in all but one subject 1336 (PM 18.4). c In the monkey PMI cohort layers 3 and 5 segregated in all but one slab (PMI 24 h). d CALB1 log2 layer 3–5 difference in human and monkey cohorts. CALB1 levels are significantly elevated in layer 3 of both humans and monkeys, *q < 0.1
To increase power for comparisons between layers 3 and 5 of individual protein differences, human cohorts 1 and 2 were combined. In all, 66 proteins differed between layers 3 and 5 with a false discovery rate q < 0.1 (Table S5). Proteins elevated in layer 3 were enriched for terms relating to mitochondrial function (e.g., Mitochondrion p = 2.7E−7, Table S6), while proteins elevated in layer 5 were enriched for terms relating to axons (e.g., Intermediate Filament p = 4.5E−3, and Axon Development p = 4.6E−3, Table S6). Likewise, in the monkey tissue, 71 proteins differed between layers 3 and 5 (with a false discovery rate < 0.1, Table S7). Proteins elevated in layer 3 were enriched for terms relating to mitochondrial function (e.g., Mitochondrion p = 1.8E−4, Table S8), while proteins elevated in layer 5 were enriched for terms relating to axons (e.g., Neurofilament p = 4.4E−2, and Axon Development p = 1.2E−3, Table S8). The list of proteins with significant layer 5–3 differences in human tissue was highly similar to that in monkey tissue (38 proteins shared, p < 1E−5). Finally, in both human and monkey cohorts, the layer 3 protein marker CALB1 [34] was significantly elevated in layer 3 (Fig. 2d).
Discussion
The mammalian brain is a highly complex and structured proteome. Here we demonstrate an approach, LCM-MS, that is able to interrogate this proteome with high precision and spatial resolution, beyond cortical homogenates. The success of this approach likely reflects the complementary strengths of the techniques that we combined. Laser capture allows for the harvesting of cortical layers with exceptional precision and targeted MS performed on a triple quadrupole with stable isotope-labeled standards has a well-documented high level of precision in protein quantification [35].
We observed no significant correlations, and only a handful of nominally significant correlations, between protein levels or layer 5–3 protein differences and PMIs up to 24 h in both human tissue and the monkey PMI model. Further testing, however, is required to determine the maximum PMI (beyond 24 h) to which this approach can reliably be applied. Likewise, the order of layer collection, pH, and RNA integrity number (RIN) had no significant effect on protein levels or differences.
Unsupervised hierarchical clustering segregated layers 3 and 5 by protein expression in the first human cohort. In the second human cohort and the PMI monkey series, layers 3 and 5 segregated in all but one subject. Clustering effects were driven by the same proteins in monkey and humans, as evidenced by the highly significant overlap in layer 5–3 protein differences between species. Interestingly, the human and monkey subjects that did not segregate had PMI of 18.4 and 24 h, respectively. Thus, while individual protein measures were not significantly correlated with PMI, unsupervised hierarchical clustering of the proteomic data appears to be sensitive to higher PMIs. Nevertheless, even with these two high-PMI subjects included and a relatively small sample size, we detected robust differences in layer 5–3 protein levels in both the human (n = 10 subjects) and monkey cohorts (n = 8 blocks). Functional gene annotation analysis of both the human and monkey data revealed that mitochondrial protein levels are relatively greater in layer 3, perhaps reflecting a greater number of mitochondrial-rich parvalbumin neurons in layer 3, while axonal protein levels are relatively greater in layer 5, likely as a greater number of axons transverse layer 5 en-route to more superficial layers.
Our data are in line with prior findings of connectivity and molecular differences between cortical layers [12, 23] further supporting the idea that layer-specific neuronal protein network alterations, potentially unnoticed in tissue homogenate studies, could be present in etiologically complex psychiatric diseases and regulate specific synaptic and circuit pathologies. Additionally, our findings strongly suggest that LCM-MS is capable of observing these differences in appropriately powered and matched case–control experiments.
The studies described here were conducted by targeted mass spectrometry with a stable isotope-labeled standard on a triple quadrupole mass spectrometer. This approach has the advantage of exceptional precision, sensitivity, and throughput. However, we assessed ~1.5% of expressed genes in the human cortex (17,068 transcripts, Human Protein Atlas website [36]). As the field of mass spectrometry progresses, new generations of mass spectrometers with increased speed are continuously becoming available and quantitative approaches such as isobaric labeling and differential mass spectrometry have increased in both precision and their multiplexed capabilities. Indeed, two groups have recently reported the utilization of tandem mass tag (TMT) isobaric labeling with offline fractionation to quantify >10,000 proteins with precision matching that reported here [37, 38], albeit with less throughput. Alternatively, approaches such as multiplexed ion beam imaging [39] and mass cytometry [40] that tag antibodies with isotopically pure elements allow for exception sensitivity and precise quantification proteins on a smaller scale (~100 targets) and could be used to quantify markers of cell or synapse subtype population in cortical layer captures. Finally, LCM-MS of single cell type populations in postmortem brain tissue is likely feasible but, while allowing for layer specificity, could not assay dendritic synapses, a compartment of intense interest in many neuropsychiatric diseases. Likewise, cell sorting approaches are beginning to see use for microglia isolation from human postmortem brain tissue [41] and could in theory be applied to synaptosomes. While providing a powerful level of specificity at the cell body (or potentially synapse) level, this approach requires grams of tissue, precluding layer-specific analyses. Thus, LMC-MS approaches will likely find utility in future human postmortem brain studies to assess larger numbers of synaptic protein populations in smaller and smaller tissue samples while maintaining high levels of precision.
In conclusion, our data indicate that laser capture microdissection–quantitative mass spectrometry facilitates the precise quantification of proteins within individual cortical layers from human postmortem brain tissue. This tool will likely find utility in the study of a number of neuropsychiatric diseases and increase in power as instrumentation advances.
Funding and disclosure
This work was funded by the National Institutes of Health grants R01 MH071533, R03 MH108849, P50 MH103204, and K01 MH107756, as well as a NARSAD Young Investigator Award from the Brain and Behavior Research Foundation. DAL currently receives investigator-initiated research support from Pfizer and serves as a consultant in the areas of target identification and validation in Merck. MLM, DF, MG, ZS, DA, YD, NY, and RAS declare no competing interests. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institute of Mental Health, the National Institutes of Health, or the US Government.
References
Anderson SA, Eisenstat DD, Shi L, Rubenstein JL. Interneuron migration from basal forebrain to neocortex: dependence on Dlx genes. Science. 1997;278:474–6.
Ghashghaei HT, Lai C, Anton ES. Neuronal migration in the adult brain: are we there yet? Nat Rev Neurosci. 2007;8:141–51.
Hoftman GD, Datta D, Lewis DA. Layer 3 excitatory and inhibitory circuitry in the prefrontal cortex: developmental trajectories and alterations in schizophrenia. Biol Psychiatry. 2017;81:862–73.
Kim EJ, Juavinett AL, Kyubwa EM, Jacobs MW, Callaway EM. Three types of cortical layer 5 neurons that differ in brain-wide connectivity and function. Neuron. 2015;88:1253–67.
Tjia M, Yu X, Jammu LS, Lu J, Zuo Y. Pyramidal neurons in different cortical layers exhibit distinct dynamics and plasticity of apical dendritic spines. Front Neural Circuits. 2017;11:43.
Sullivan PF, Daly MJ, O’Donovan M. Genetic architectures of psychiatric disorders: the emerging picture and its implications. Nat Rev Genet. 2012;13:537–51.
Harrison PJ, West VA. Six degrees of separation: on the prior probability that schizophrenia susceptibility genes converge on synapses, glutamate and NMDA receptors. Mol Psychiatry. 2006;11:981–3.
Banerjee A, Macdonald ML, Borgmann-Winter KE, Hahn CG. Neuregulin 1-erbB4 pathway in schizophrenia: from genes to an interactome. Brain Res Bull. 2010;83:132–9.
Moyer CE, Shelton MA, Sweet RA. Dendritic spine alterations in schizophrenia. Neurosci Lett. 2015;601:46–53.
Sweet RA, Fish KN, Lewis DA. Mapping synaptic pathology within cerebral cortical circuits in subjects with schizophrenia 2. Front Hum Neurosci. 2010;4:44.
Arion D, Horvath S, Lewis DA, Mirnics K. Infragranular gene expression disturbances in the prefrontal cortex in schizophrenia: signature of altered neural development? Neurobiol Dis. 2010;37:738–46.
Arion D, Unger T, Lewis DA, Mirnics K. Molecular markers distinguishing supragranular and infragranular layers in the human prefrontal cortex. Eur J Neurosci. 2007;25:1843–54.
Arion D, Corradi JP, Tang S, Datta D, Boothe F, He A, et al. Distinctive transcriptome alterations of prefrontal pyramidal neurons in schizophrenia and schizoaffective disorder. Mol Psychiatry. 2015;20:1397–405.
Altar CA, Jurata LW, Charles V, Lemire A, Liu P, Bukhman Y, et al. Deficient hippocampal neuron expression of proteasome, ubiquitin, and mitochondrial genes in multiple schizophrenia cohorts. Biol Psychiatry. 2005;58:85–96.
de Sousa Abreu R, Penalva LO, Marcotte EM, Vogel C. Global signatures of protein and mRNA expression levels. Mol Biosyst. 2009;5:1512–26.
Zhang B, Gaiteri C, Bodea LG, Wang Z, McElwee J, Podtelezhnikov AA, et al. Integrated systems approach identifies genetic nodes and networks in late-onset Alzheimer’s disease. Cell. 2013;153:707–20.
Sharma K, Schmitt S, Bergner CG, Tyanova S, Kannaiyan N, Manrique-Hoyos N, et al. Cell type- and brain region-resolved mouse brain proteome. Nat Neurosci. 2015;18:1819–31.
Mukherjee S, Rodriguez-Canales J, Hanson J, Emmert-Buck MR, Tangrea MA, Prieto DA, et al. Proteomic analysis of frozen tissue samples using laser capture microdissection. Methods Mol Biol. 2013;1002:71–83.
Heath JR, Ribas A, Mischel PS. Single-cell analysis tools for drug discovery and development. Nat Rev Drug Discov. 2016;15:204–16.
Nijholt DA, Stingl C, Luider TM. Laser capture microdissection of fluorescently labeled amyloid plaques from Alzheimer’s disease brain tissue for mass spectrometric analysis. Methods Mol Biol. 2015;1243:165–73.
Hashimoto M, Bogdanovic N, Nakagawa H, Volkmann I, Aoki M, Winblad B, et al. Analysis of microdissected neurons by 18O mass spectrometry reveals altered protein expression in Alzheimer’s disease. J Cell Mol Med. 2012;16:1686–700.
MacDonald ML, Ciccimaro E, Prakash A, Banerjee A, Seeholzer SH, Blair IA, et al. Biochemical fractionation and stable isotope dilution liquid chromatography-mass spectrometry for targeted and microdomain-specific protein quantification in human postmortem brain tissue. Mol Cell Proteom. 2012;11:1670–81.
Andressen C, Blumcke I, Celio MR. Calcium-binding proteins: selective markers of nerve cells. Cell Tissue Res. 1993;271:181–208.
Glantz LA, Lewis DA. Decreased dendritic spine density on prefrontal cortical pyramidal neurons in schizophrenia. Arch General Psychiatry. 2000;57:65–73.
Curley AA, Arion D, Volk DW, Asafu-Adjei JK, Sampson AR, Fish KN, et al. Cortical deficits of glutamic acid decarboxylase 67 expression in schizophrenia: clinical, protein, and cell type-specific features. Am J Psychiatry. 2011;168:921–9.
Hayes TL, Cameron JL, Fernstrom JD, Lewis DA. A comparative analysis of the distribution of prosomatostatin-derived peptides in human and monkey neocortex. J Comp Neurol. 1991;303:584–99.
Datta D, Arion D, Roman KM, Volk DW, Lewis DA. Altered expression of ARP2/3 complex signaling pathway genes in prefrontal layer 3 pyramidal cells in schizophrenia. Am J Psychiatry. 2017;174:163–71.
Datta D, Arion D, Corradi JP, Lewis DA. Altered expression of CDC42 signaling pathway components in cortical layer 3 pyramidal cells in schizophrenia. Biol Psychiatry. 2015;78:775–85.
Bayes A, van de Lagemaat LN, Collins MO, Croning MD, Whittle IR, Choudhary JS, et al. Characterization of the proteome, diseases and evolution of the human postsynaptic density. Nat Neurosci. 2011;14:19–21.
MacDonald ML, Ding Y, Newman J, Hemby S, Penzes P, Lewis DA, et al. Altered glutamate protein co-expression network topology linked to spine loss in the auditory cortex of schizophrenia. Biol Psychiatry. 2015;77:959–68.
MacLean B, Tomazela DM, Shulman N, Chambers M, Finney GL, Frewen B, et al. Skyline: an open source document editor for creating and analyzing targeted proteomics experiments. Bioinformatics. 2010;26:966–8.
Eisen MB, Spellman PT, Brown PO, Botstein D. Cluster analysis and display of genome-wide expression patterns. Proc Natl Acad Sci USA. 1998;95:14863–8.
Page RD. TreeView: an application to display phylogenetic trees on personal computers. Comput Appl Biosci CABIOS. 1996;12:357–8.
Hayes TL, Lewis DA. Nonphosphorylated neurofilament protein and calbindin immunoreactivity in layer III pyramidal neurons of human neocortex. Cereb Cortex. 1992;2:56–67.
Method of the Year 2012, Nat Methods. 2013;10:1. https://www.nature.com/articles/nmeth.2329
Uhlen M, Fagerberg L, Hallstrom BM, Lindskog C, Oksvold P, Mardinoglu A, et al. Proteomics. Tissue-based map of the human proteome. Science. 2015;347:1260419.
Ping L, Duong DM, Yin L, Gearing M, Lah JJ, Levey AI, et al. Global quantitative analysis of the human brain proteome in Alzheimer’s and Parkinson’s Disease. Sci Data. 2018;5:180036.
Kim DK, Park J, Han D, Yang J, Kim A, Woo J, et al. Molecular and functional signatures in a novel Alzheimer’s disease mouse model assessed by quantitative proteomics. Mol Neurodegener. 2018;13:2.
Angelo M, Bendall SC, Finck R, Hale MB, Hitzman C, Borowsky AD, et al. Multiplexed ion beam imaging of human breast tumors. Nat Med. 2014;20:436–42.
Ornatsky O, Baranov VI, Bandura DR, Tanner SD, Dick J. Multiple cellular antigen detection by ICP-MS. J Immunol Methods. 2006;308:68–76.
Mizee MR, Miedema SS, van der Poel M, Adelia, Schuurman KG, van Strien ME, et al. Isolation of primary microglia from the human post-mortem brain: effects of ante- and post-mortem variables. Acta Neuropathol Commun. 2017;5:16.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
MacDonald, M.L., Favo, D., Garver, M. et al. Laser capture microdissection–targeted mass spectrometry: a method for multiplexed protein quantification within individual layers of the cerebral cortex. Neuropsychopharmacol 44, 743–748 (2019). https://doi.org/10.1038/s41386-018-0260-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41386-018-0260-0