Abstract
The γ-amino butyric acid (GABA)ergic system shapes the activation and function of immune cells. The present study was conducted to explore the regulation of GABA transporter (GAT)-2 on the differentiation of Th17 cells. Here we found that Th17 cells show higher abundance of GAT-2, and have distinct cellular metabolic signatures, such as the GABA shunt pathway, as compared to naïve T cells. GAT-2 deficiency had little effect on the metabolic signature in naïve T cells, but impaired the GABA uptake and GABA shunt pathway in Th17 cells. GAT-2 deficiency had little effect on T cell development and peripheral T cell homeostasis; however, its deficiency promoted Th17 cell differentiation in vitro. Mechanistically, GAT-2 deficiency promoted differentiation of Th17 cells through activation of GABA–mTOR signaling. In a mouse model of intestinal infection and inflammation, GAT-2 deficiency promoted Th17 responses. Collectively, GAT-2 deficiency promotes Th17 cell responses through activation of GABA–mTOR signaling.
Similar content being viewed by others
Introduction
The γ-amino butyric acid (GABA)ergic system, which includes GABA, GABA receptors, glutamate decarboxylase (GAD, the enzyme that synthetizes GABA), vesicular inhibitory amino acid transporter (VIAAT; the vesicular protein involved in GABA storage), GABA transporters (GATs) and GABA transaminase (GABA-T, the enzyme that catabolizes GABA), has inhibitory functions in the central nervous system of vertebrates, Increasing investigations are showing that immune cells also possess the GABAergic system.1 For example, GAD and the GABA receptor have been found in macrophages, and GAD, GATs, GABA-T, and GABA receptors are found in T cells.1 This promotes to investigate the regulatory function of GABAergic system in activation and function of immune cells. GABAergic agents directly inhibits the function of antigen-presenting cells to reduce subsequent inflammatory responses to antigens; thus, GABA ameliorates paralysis via inhibition of inflammation in mouse model of experimental autoimmune encephalomyelitis.2 Likewise, results from other investigations also support the conclusion that GABAergic system inhibits the immune responses.3,4,5 However, GABA promotes intestinal Th17 cell differentiation and interleukin-17 (IL-17) expression during enterotoxigenic Escherichia coli (ETEC) infection in piglets and mice.6 Thus, the effect of the GABAergic system in the immune system needs further investigations.
GATs terminate GABA signaling by mediating translocation of GABA from the extracellular to the intracellular space.7 There are four types of GAT: GAT-1 (SLC6A1), GAT-2 (SLC6A13), GAT-3 (SLC6A11), and BGT-1 (SLC6A12). Among them, GAT-1 and BGT-1 are detected in T cells.7 GAT-1−/− mice are more susceptible to experimental autoimmune encephalomyelitis, and show increased expression of inflammatory cytokines, including interferon-γ (IFN-γ), tumor necrosis factor-α, IL-6, IL-23 IL-17, and IL-12, because GAT-1 negatively regulates T cell activation and T cell-mediated responses.8,9 However, whether T cells express GAT-2, and the role of GAT-2 in T cell-mediated responses, especially Th17 responses, is still unknown. Thus, this study was designed to uncover this issue with in vitro model of Th17 cell differentiation, and with a conventional gene-targeting approach used to delete GAT-2 in mice.
Here, we found that there is significant metabolic profile between naïve T cells and Th17 cells, especially about GABA shunt pathway. GAT-2 deficiency has little effect on the metabolic signature in naïve T cells; however, GAT-2 deficiency impairs the GABA uptake and GABA shunt pathway in Th17 cells. GAT-2 deficiency has little effect on the T cell development and peripheral T cell homeostasis; however, it promotes Th17 cell differentiation. Mechanistically, GAT-2 deficiency promotes the differentiation of Th17 cells through activation of GABA–mTOR signaling. In a mouse model of intestinal infection and inflammation, GAT-2 deficiency promotes Th17 responses.
Materials and methods
Bacterial strains and antibodies
This study involved the use of an Escherichia coli F4-producing strain W25K (hereafter referred as ETEC; O149:K91, K88ac; LT, STb, EAST), which was isolated from a piglet with diarrhea,10 and cultured in LB medium. Citrobacter rodentium (DBS100) was cultured in LB medium. Antibodies against GAT-2 (Sc-7668) and Actin (Sc-47778) were purchased from Santa Cruz Biotechnology, Inc. (Dallas, Texas, USA). Antibodies against mTOR (#2972), p-mTOR (#5536), STAT-3 (#9139P), p-STAT-3 (#9131S), P65 (#6956S), p-P65 (#3033S) were purchased from Cell Signaling Technology (Danvers, MA, USA). Antibodies against GAT-1 (ab177483) and BGT (ab200676) came from Abcam (Cambridge, UK). Antibody against GAPDH (10494-1-AP) was purchased from Proteintech (Rosemont, IL, USA).
Mice
Rag1–/– mice were the gifts from Prof. Xinming Jia (Department of Immunology, Tongji University School of Medicine, Shanghai, China). Slc6a13–/– mice (in C57BL/6J genetic background) were generated by a conventional gene-targeting strategy, in which coding exons 2 were replaced with a PGK-neomycin-resistance cassette (Fig. 1a). Heterozygous Slc6a13+/– mice were bred to generate age- and sex-matched homozygous GAT2-ablated (Slc6a13–/–) and WT (Slc6a13–/–) mice. Genotyping PCR primers are listed in Table S1. Mice were conducted according to the guidelines of the Laboratory Animal Ethical Commission of the Chinese Academy of Sciences.
The influence of Slc6a13 deficiency on T cell development in the thymus and peripheral lymphocyte profile. a Slc6a13 gene locus and targeting strategy. Exons 2 were replaced with a PGK-neomycin neomycin cassette. Forward and reverse genotyping PCR primers for amplifying wild-type (WT) allele (P1 and P2) and knockout (KO) allele (P3 and P4) are indicated. b Genotyping PCR to detect WT (+/+), Slc6a13 heterozygous (+/–) and homozygous (–/–) knockout mice. c Immunoblotting to detect the expression of GAT-2 in the liver of Slc6a13+/+ and Slc6a13–/– mice. d Flow cytometry analysis of thymocyte subpopulations of Slc6a13+/+ and Slc6a13–/– mice. Numbers indicate the percentage of CD4+ (lower right) and CD8+ (upper left) single-positive (SP), CD4+CD8+ double-positive (DP, upper right), and CD4–CD8– double- negative (DN, lower left) thymocytes. Right: Frequency of the indicated thymocyte subpopulations in Slc6a13+/+ and Slc6a13–/– mice with statistical analysis. e Flow cytometry analysis of CD3+CD4+ T cells, and CD3+CD8+ T cells in the spleen of Slc6a13+/+ and Slc6a13–/– mice. Data are representative of two independent experiments with 3–4 mice per group. Data in d and e were analyzed with unpaired t-test and shown as the means ± SD
ETEC infection in mice
Mice were orally gavaged with 108 CFUs of ETEC. At 6 h post infection, mice were sacrificed to collect the jejunum, and the samples were stored at −80 °C until processing. For Th17 cell population analysis, the mesenteric lymph node (MLN) samples were collected at 5 days post infection.
Citrobacter rodentium infection of mice
Mice (6 weeks of age) were orally gavaged with 108 CFUs of Citrobacter rodentium (DBS100). The colon and spleen samples from all groups were collected for further analysis at 7 days post infection. For Th17 cell population analysis, the MLN samples were collected at 10 days post infection.
Lipopolysaccharide challenge in mice
Mice were challenged with lipopolysaccharide (LPS) by intraperitoneal injection with dosage of 10 mg/kg. At 48 h of challenge, mice were sacrificed to collect the jejunum and lung.
T cell transfer model of infection
Naive T cells from WT or Slc6a13–/–mice were isolated and transferred via i.v. injection to sex-matched B6 Rag1−/− recipient mice (5 × 106 cells/mouse). Then, mice were infected with ETEC or Citrobacter rodentium at following day, and the tissues were collected as described above.
T cell differentiation
Naive CD4+ T cells were isolated from mouse splenocytes using a CD4+CD62L+ T cell isolation kit II (Miltenyi Biotec, purity >95%). For Th17 differentiation, cells were stimulated with anti-CD3 (2 μg/ml, 100313; Biolegend) and anti- CD28 (2.0 μg/ml, 102111; Biolegend) supplemented with 5 ng/ml TGF-β1 (7666-MB-005; RD), 20 ng/ml IL- 6 (575702; Biolegend), 10 μg/ml anti-IFN-γ (505812; Biolegend), and 10 μg/ml anti-IL-4 (504107; Biolegend) for 1–3 days. For Th1 differentiation, cells were stimulated with anti-CD3 and anti-CD28 supplemented with 10 ng/ml IL-12 (210-12-2; Peprotech) and 10 μg/ml anti-IL-4. For Th2 differentiation, cells were stimulated with anti-CD3 and anti-CD28 supplemented with 10 ng/ml IL-4 (214-14-5; Peprotech), and 10 μg/ml anti-IFN-γ. For Treg differentiation, cells were stimulated with anti-CD3 and anti-CD28 supplemented with TGF-β1 (5 ng/ml), IL-2 (10 ng/ml, 212-12-5; Peprotech), anti-IFN-γ (10 μg/ml) and anti-IL-4 (10 μg/ml). In some experiments, bicuculline, rapamycin, LPS, and GABA was added with indicated concentrations.
Flow cytometry analysis
Lymphocytes isolated from mouse thymus, MLN, and spleen were stained with cell surface markers of CD3 (FITC-CD3, 100203; Biolegend), CD4 (PE-CD4, 100407; Biolegend), CD8 (APC-CD4, 100711; Biolegend). For the analysis of intracellular cytokine, CD4+ T cells were stimulated with PMA (50 ng/ml), ionomycin (1 μg/ml), and monensin (3 μg/ml) for 5 h. After staining with cell surface markers, intracellular cytokine staining was performed with a fixation and permeabilization kit (eBioscience) and IL-17A Ab (APC-IL17A, 506195; Biolegend), IL-4 Ab (APC-IL4, 504105; Biolegend), IFN-γ Ab (FITC- IFN-γ, 505805; Biolegend), or Foxp3 Ab (FITC- Foxp3, 11-5773-80; eBioscience) in accordance with the manufacturer’s instructions. Flow cytometry was performed on a FACSCalibur (BD Biosciences) and data were analyzed using the FlowJo Software (Tree Star).
Metabolite profiling analysis
Metabolite concentrations in T cells were quantified using liquid chromatography triple-quadrupole mass spectrometry (LC-QqQ-MS),11,12 which is targeted metabolomics simultaneously analyzed up to 206 metabolites (Supplementary File 1).
Tissue histological examination
This was performed using hematoxylin and eosin (H&E) staining. Briefly, mouse lung samples were fixed with 4% paraformaldehyde-PBS overnight, and then dehydrated and embedded in paraffin blocks. Section of 5 μm was cut for histological analysis. The sections were deparaffinized and hydrated, and then stained with H&E. Five mice were studied from each group. The data collectors were unaware of the treatment status of the examined slides. Inflammatory index were scored separately from 0 (lesion absent) to 3 (severe lesion) based on a previous method.13
RT-PCR
Real-time PCR was performed according to our previous study.6 Primers (Supplementary Table 1) were selected according to previous references. β-Actin was used as an internal control to normalize target gene transcript levels.
Immunoblotting
Immunoblotting was performed according to our previous study.6,14 Signal intensity was digitally quantified and normalized to actin or GAPDH protein abundance.
Counting of bacteria
Intestinal tissues were homogenized in saline and then serial diluted and plated on MacConkey agar for ETEC and Citrobacter rodentium. Bacteria were enumerated after 16 h of growth at 37 °C. The counts were further verified by PCR with specific primers.
Statistical analyses
Data shown are the means ± the standard deviation (SD) or standard error of the mean (SEM). Data were statistically analyzed according to our previous papers.6,15 Data between two groups were analyzed by unpaired t-test (Prism 6.0) if the data are in Gaussian distribution and have equal variance, or by unpaired t-test with Welch’s correction (Prism 6.0) if the data are in Gaussian distribution but show unequal variance, or by non-parametric test (Mann–Whitney U test, Prism 6.0) if the data are not normally distributed. The Gaussian distribution of data was analyzed by D’Agootino-Pearson omnibus normality test (Prism 6.0) and Kolmogorov–Smirnov test (Prism 6.0). The variance of data was analyzed by Brown–Forsythe test (Prism 6.0). Differences with P < 0.05 were considered significant.
Results
Slc6a13 deficiency does not affect T cell development in the thymus and peripheral T cell homeostasis
In this study, we employed a conventional gene-targeting approach to delete GAT-2 in mice (Fig. 1a–c). Slc6a13–/– mice were born at expected Mendelian ratios and did not show obvious abnormalities in growth or survival (data not shown). These mutant animals also had normal thymocyte development, as revealed by the comparable frequencies of thymocyte subpopulations in the Slc6a13+/+ and Slc6a13–/– mice (Fig. 1d). To examine the role of GAT-2 in regulating peripheral T cell homeostasis, we examined the frequency of T cells in the spleen of the 8 weeks of Slc6a13+/+ and Slc6a13–/– mice. The frequencies and absolute numbers of CD3+, CD3+CD4+, and CD3+CD8+ cells were also largely comparable between Slc6a13+/+ and Slc6a13–/– mice (Fig. 1e). Also, the frequencies and absolute numbers of CD4+IFN-γ+, CD4+IL-4+, CD4+IL-17+, CD4+Foxp3+ cells did not differ between Slc6a13+/+ and Slc6a13–/– mice (Fig. 2a). Then, we analyzed the expression of cytokines in the jejunum of the Slc6a13+/+ and Slc6a13–/– mice. Slc6a13–/– mice had higher expression of Il-4, but similar levels of expression of Il-1β, Il-6, and Tgf-β1, Il-17, as compared to Slc6a13+/+ mice (Fig. 2b). Slc6a13–/– mice had a moderate decrease in the expression of Il-23 (Fig. 2b), as compared to Slc6a13+/+ mice. Collectively, these data show that slc6a13 deficiency does not affect T cell development in the thymus and peripheral T cell homeostasis.
Peripheral T cell subtypes in the spleen and expression of Th17-related cytokines in the jejunum. a Flow cytometry analysis of the frequency and absolute number of CD4+INF-γ+, CD4+IL-4+, CD4+IL-17+, CD4+Foxp3+ T cells in the spleen of Slc6a13+/+ and Slc6a13–/– mice. Data are representative of two independent experiments with 3–4 mice per group. Data about CD4+Foxp3+ T cells were analyzed by unpaired t-test, while others by Mann–Whitney test. Data were shown as the means ± SD. b mRNA expression of IL-1β, IL-4, IL-6, IL-17, IL-23, and TGF-β1 in the jejunum of Slc6a13+/+ and Slc6a13–/– mice. Data are representative of two independent experiments with 5–6 mice per group. Data were analyzed by unpaired t-test and shown as the means ± SEM
Slc6a13 deficiency inhibits GABA transport and cellular GABA shunt pathway during Th17 cell differentiation
Previous investigation shows that GABA affects Th17 differentiation; thus, we compared the expression of GAT-2 in naïve T cells and Th17 cells. The protein abundance of GAT-2 was hardly detected in naïve T cells, while it had a high abundance in Th17 cells (Fig. 3a), suggesting GAT-2 is involved in GABA transport during Th17 cell differentiation. GAT-2 expression was also detected in Th1, Th2, and Treg cells (Supplementary Figure 1). Then, we compared the cellular level of GABA and other metabolites between naïve T cells and Th17 cells with metabolomics. There was a significant difference in the cellular metabolites profile between naïve T cells and Th17 cells after principal component analysis (PCA) and orthogonal partial least-squares discriminant analysis (OPLS-DA) (Supplementary Figure 2A). Among 53 different metabolites, 9 metabolites (IMP, kynurenine, lactic acid, tyrosine, vitamin B2, myo-inositol, tryptophan, aspartic acid, and pseudouridine) decreased, while 44 metabolites, including GABA, increased in Th17 cells, as compared to naïve T cells (Fig. 3b). After KEGG (Kyoto Encyclopedia of Genes and Genomes) pathway analysis, the differentiation of Th17 cells affected five metabolic pathways, including alanine, aspartate and glutamate metabolism, arginine and proline metabolism, purine metabolism, glycine, serine and threonine metabolism, and riboflavin metabolism (Fig. 3c). Notably, the metabolites in the GABA shunt pathway were increased in Th17 cells, including glutamine, GABA, succinate, and fumarate (Fig. 3c). These results suggest that GAT-2 expression and cellular GABA shunt pathway increase during the differentiation of Th17 cells from naïve T cells.
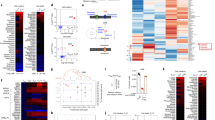
GAT-2 deficiency alters the cellular metabolic pathway in Th17 cells. a The expression of GAT-2 in naïve T cells and Th17 cells. Representative of two independent experiments. b Heatmap analysis of different metabolites in Th17 cells after metabolomics analysis. N = 3 for naïve T cells, while n = 5 for Th17 cells. c Metabolic pathway analysis about the different metabolites between Th17 cells and naïve T cells. The altered metabolic pathways are marked with name. Metabolites with red color increase in Th17 cells. d Heatmap analysis of different metabolites in naïve T cells from Slc6a13+/+ and Slc6a13–/– mice. N = 3 for each group. e Metabolic pathway analysis about the different metabolites found in d. f The levels of GABA in the medium of Th17 cells. Each point represents one sample. g Heatmap analysis of different metabolites in Th17 cells from Slc6a13+/+ and Slc6a13–/– mice. N = 5 for each group. h Metabolic pathway analysis about the different metabolites found in g. Metabolites with blue color decrease in Th17 cells from Slc6a13–/– mice
Next, we compared the metabolites in Slc6a13–/–and Slc6a13+/+ naïve T cells. There was a significant difference in the metabolic profile between naïve T cells from Slc6a13–/– mice and those from Slc6a13+/+ mice (Supplementary Figure 2B). Deoxyinosine, methylguanidine, and pseudouridine were decreased, while adenine, carnitine, thiamine, guanidinobutyric acid, and choline were increased in naïve T cells from Slc6a13–/– mice (Fig. 3d). After KEGG pathway analysis, thiamine metabolism was found to be different between two groups (Fig. 3e). For Th17 cells, the levels of amino acids in the medium were first analyzed after 3 days of differentiation between two groups. A higher level of GABA was found in the medium of Th17 cells from Slc6a13–/– mice, as compared to those from Slc6a13+/+ mice (Fig. 3f). There was no difference in the levels of other amino acids (Supplementary Figure 3), suggesting that GAT-2 deficiency only affects GABA transport. Then, the cellular metabolites in Th17 cells were compared between those differentiated from Slc6a13–/– naïve T cells and those from Slc6a13+/+ naïve T cells. A significant difference about the cellular metabolites profile was found between two groups (Supplementary Figure 2C). Vitamin B2 increased, while 40 metabolites, such as GABA, glutamine, and succinate, decreased in Slc6a13–/– Th17 cells (Fig. 3g). Alanine, aspartate and glutamate metabolism, arginine and proline metabolism, glycerophospholipid metabolism, and riboflavin metabolism were affected in Th17 cells after GAT-2 deficiency (Fig. 3h). Notably, the metabolites in GABA shunt pathway including glutamine, GABA, succinate, and fumarate, were decreased in Slc6a13–/– Th17 cells (Fig. 3h). These results suggest that GAT-2 deficiency inhibits GABA transport and cellular GABA shunt pathway during the differentiation of Th17 cells from naïve T cells.
Slc6a13 deficiency promotes Th17 cell differentiation through GABA–mTOR signaling
Besides GAT-2, T cells also express other transporters for GABA transport, including GAT-1 and GAT-4. There was no alteration about the protein abundance of GAT-1 and GAT-4 in the Th17 cells after GAT-2 deficiency (Fig. 4a–c); thus, GAT-2 deficiency results in increase of GABA level in the medium (Fig. 3f), but decrease of GABA level in Th17 cells (Fig. 3g). To define the effects of GAT-2 deficiency on Th17 cell differentiation, Slc6a13–/– and Slc6a13+/+ naïve T cells were isolated and cultured under Th17 cell differentiation condition, and then the levels of IL-17 in the medium and percentages of Th17 cells were compared. There were higher levels of IL-17 in Slc6a13–/– Th17 cells than Slc6a13+/+ Th17 cells (Fig. 4g). Also, the percentages of Th17 cells from Slc6a13–/– naïve T cells were higher than those from Slc6a13+/+ naïve T cells (Fig. 4h). A previous study has shown that GABA promotes intestinal IL-17 expression through mTOR signaling,6 and GABA signaling has been reported to affect the activation of cellular NF-κB and STAT signaling;16,17 thus, this study then compared the activation of cellular signaling pathways related to Th17 differentiation, including mTOR, p65, and STAT-3.18,19 Slc6a13 deficiency promoted the activation of p65 and mTOR in the Th17 cells, while had little effect on the activation of STAT-3 (Fig. 4a, d–f). These results suggested that Slc6a13 deficiency promotes Th17 cell differentiation may be through GABA–mTOR and p65 pathway.
GAT-2 deficiency promotes Th17 cell differentiation and activation of p65 and mTOR in Th17 cells. a Immunoblotting to detect the protein abundance of GAT-1, BGT-1, p65, p-p65, STAT-3, p-STAT-3, mTOR, and p-mTOR in Th17 cells differentiated from Slc6a13+/+ and Slc6a13–/– naïve T cells. N = 5 for Slc6a13–/– mice, while n = 6 for Slc6a13+/+ mice. b–f Statistically analysis the relative abundance of GAT-1 (b), BGT-1 (c), and the activation of p65 (d), mTOR (e), and STAT-3 (f) between two groups. Data were analyzed by unpaired t-test and shown as the means ± SD. g The secretion of IL-17 from Th17 cells differentiated from Slc6a13+/+ and Slc6a13–/– naïve T cells. Each point represents one sample. Data were analyzed by Mann–Whitney test and shown as the means ± SD. h Flow cytometry analysis of percentages of Th17 cells differentiated from Slc6a13+/+ and Slc6a13–/– naïve T cells. Each point represents one sample. Data were analyzed by Mann–Whitney test and shown as the means ± SD
To validate this, bicuculline (a GABA receptor specific inhibitor) was added to inhibit GABA signaling during Th17 cell differentiation from Slc6a13–/– naïve T cells. Bicuculline significantly decreased the percentages of Th17 cells (Fig. 5a, b), and inhibited the secretion of IL-17 from Th17 cells (Fig. 5c), indicating that Slc6a13 deficiency promotes Th17 cell differentiation through GABA signaling. Indeed, GABA treatment in Slc6a13+/+ naïve T cells promoted Th17 cell differentiation (Fig. 5d), while bicuculline treatment inhibited the differentiation of Th17 cells from WT naïve T cells (Fig. 5e). Notably, bicuculline inhibited the activation of mTOR in Th17 cells (Fig. 5f, g); however, bicuculline had little effect on the activation of p65 and STAT-3 (Fig. 5h–j), suggesting that Slc6a13 deficiency promotes Th17 cell differentiation through GABA–mTOR signaling. To test this further, rapamycin was added to inhibit mTOR signaling during Th17 cell differentiation from Slc6a13–/– naïve T cells. Rapamycin significantly lowered the percentages of Th17 cells (Fig. 5a, b), inhibited the secretion of IL-17 from Th17 cells (Fig. 5c) and the activation of mTOR in Th17 cells (Fig. 5f, g); however, rapamycin had little effect on the activation of p65 and STAT-3 in Th17 cells (Fig. 5h–j). Similarly, rapamycin treatment also decreased the percentages of Th17 cells from Slc6a13+/+ naïve T cells (Fig. 5e). Collectively, these results show that Slc6a13 deficiency promotes Th17 cell differentiation through GABA–mTOR signaling.
GAT-2 deficiency promotes the differentiation of Th17 cells through GABA–mTOR signaling. a, b Flow cytometry analysis of Th17 cells after bicuculline and rapamycin treatment. Naïve T cells were isolated from Slc6a13+/+ and Slc6a13–/– mice and cultured in Th17 differentiation medium. Bicuculline (50 μM) and rapamycin (10 μM) were added in the indicated group. Data are representative of two independent experiments with 4–6 replicates in each time. c Bicuculline and rapamycin inhibits the secretion of IL-17 from Slc6a13–/– Th17 cells. Data are representative of two independent experiments with 4–5 replicates in each time. d Flow cytometry analysis of Slc6a13+/+ Th17 cells after GABA treatment. Data are representative of two independent experiments with 4–6 replicates in each time. e Flow cytometry analysis of Slc6a13+/+ Th17 cells after bicuculline and rapamycin treatment. Bicuculline (50 μM) and rapamycin (10 μM) were added in the indicated group. Data are representative of two independent experiments with 4–6 replicates in each time. f Immunoblotting to detect the protein abundance of mTOR and p-mTOR in Slc6a13–/– Th17 cells. g Statistically analysis the activation of mTOR among three groups. h Immunoblotting to detect the protein abundance of p65, p-p65, STAT-3, and p-STAT-3 in Slc6a13–/– Th17 cells. i, j Statistically analysis the activation of p65 (i), and STAT-3 (j) among three groups. Data were analyzed by unpaired t-test and shown as the means ± SD (a–j)
Slc6a13 deficiency promotes Th17 responses against C. rodentium and ETEC infection
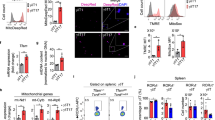
In the context of C. rodentium infection, Slc6a13–/– mice had a reduced burden of C. rodentium in the colon, as compared to the Slc6a13+/+ mice (Fig. 6a). A higher frequency and a higher number of CD4+IL-17+ cells in the spleen were observed in Slc6a13–/– mice, as compared to the Slc6a13+/+ mice (Fig. 6b), though the frequencies and absolute numbers of CD4+IFN-γ+, CD4+IL-4+, and CD4+Foxp3+ cells were comparable between Slc6a13+/+ and Slc6a13–/– mice (Fig. 6c). Notably, naïve T cells from the spleen of Slc6a13–/–-infected mice were more likely to differentiate into CD4+IL-17+ T cells under Th17 polarization conditions, as compared to cells from Slc6a13+/+-infected mice (Fig. 6d). Then, this study characterized the expression of cytokines related to Th17 responses in the colon of Slc6a13+/+- and Slc6a13–/–-infected mice. Although there was little change in the expression of Il-6 and Il-23 between Slc6a13+/+ and Slc6a13–/– mice, Slc6a13–/– mice had higher expression of Il-1β, Il-17, and Tgf-β1 in the colon, as compared to the Slc6a13+/+ mice (Fig. 6e). Also, Slc6a13–/– infected mice had higher percentages of Th17 cells in the MLN than those in Slc6a13+/+ mice (Fig. 6f). Similar to C. rodentium infection, ETEC-infected Slc6a13–/– mice had lower bacterial burdens, as compared to Slc6a13+/+ mice (Fig. 7a). Also, Slc6a13–/– mice had higher expression of Il-17 in the jejunum and higher percentage of Th17 cells in MLN after ETEC infection, compared to the Slc6a13+/+ mice (Fig. 7b–d). In the LPS-challenged mouse model, Slc6a13–/– mice had higher expression of Il-17 and Ifn-γ in the lung and jejunum, as compared to the Slc6a13+/+ mice (Fig. 7e, f). The expression of Il-22 was up-regulated in the lung, but was down-regulated in the jejunum in Slc6a13–/– mice after LPS challenge (Fig. 7e, f). The inflammatory indices in the lung were also higher in Slc6a13–/– mice after LPS challenge, as compared to the Slc6a13+/+ mice (Fig. 7g, h). To further validate the roles of Slc6a13 deficiency in Th17 cell responses during inflammation, LPS was added to the Th17 differentiation medium in both Slc6a13+/+ naïve T cells and Slc6a13–/– naïve T cells, and the percentages of Th17 cells were analyzed after 5 days of differentiation. After LPS challenge, the Slc6a13–/– group had higher percentages of Th17 cells compared to the WT group (Fig. 7i, j). Collectively, mice with Slc6a13 deficiency have higher Th17 responses during intestinal infection and LPS challenge.
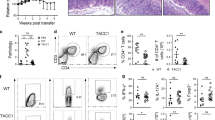
Slc6a13 deficiency promotes Th17 responses against C. rodentium infection. a Bacterial counting of C. rodentium in the colon in indicated models. b Flow cytometry analysis of the frequency and absolute number of CD4+IL-17+ cells in the spleen of Slc6a13+/+ and Slc6a13–/– mice after 7 days of C. rodentium infection. c Flow cytometry analysis of the frequency and absolute number of CD4+INF-γ+, CD4+IL-4+, CD4+Foxp3+ T cells in the spleen of Slc6a13+/+ and Slc6a13–/– mice after 7 days of C. rodentium infection. d Intracellular staining of the expression of IL-17 by CD4+ T cells cultured under the Th17-inducing conditions for 24 h. Naïve CD4+ T cells were isolated from the spleen of Slc6a13+/+ (WT) and Slc6a13–/– (KO) mice after 7 days of C. rodentium infection and cultured under the Th17-inducing conditions for 24 h. e mRNA expression of IL-1β, IL-6, IL-17, IL-23, and TGF-β1 in the colon of Slc6a13+/+ and Slc6a13–/– mice after 7 days of C. rodentium infection. f Flow cytometry analysis of the frequency number of CD4+IL-17+ cells in the MLN of Slc6a13+/+ and Slc6a13–/– mice after 10 days of C. rodentium infection. Data are representative of two independent experiments with 5–6 mice per group. Data were analyzed by unpaired t-test and shown as the means ± SD (a–d, f) or SEM (e)
Slc6a13 deficiency promotes Th17 responses against ETEC infection and LPS challenge. a Bacterial counting of ETEC in the jejunum in indicated models. b Relative mRNA expression of IL-17 in mouse jejunum samples after 6 h of ETEC infection. Data shown are representative of two independent experiments with n = 3–6 in each experiment. Data were analyzed by unpaired t-test and shown as the means ± SEM. c, d Flow cytometry analysis of the frequency of CD4+IL-17+ cells in the MLN of Slc6a13+/+ and Slc6a13–/– mice after 5 days of ETEC infection. Data were analyzed by Mann–Whitney test and shown as the means ± SD. e, f Relative mRNA expression of indicated cytokines in mouse lung or jejunum samples after 48 h of LPS challenge in indicted mouse model (n = 10). Data were analyzed by unpaired t-test and shown as the means ± SEM. g, h The inflammation in the lung was analyzed with H&E staining after 48 h of LPS challenge in the indicted mouse model. Data were analyzed by unpaired t-test and shown as the means ± SD. i, j Flow cytometry analysis of Th17 cells after LPS treatment. Naïve T cells were isolated from Slc6a13+/+ and Slc6a13–/– mice and cultured in Th17 differentiation medium. LPS were used with 1 μg/ml. Data were analyzed by unpaired t-test and shown as the means ± SEM. Data shown are representative of two independent experiments with n = 3–5 in each experiment
Slc6a13 deficiency promotes Th17 responses in T cell adoptive transfer model of intestinal infection
To validate the intrinsic effect of GAT-2 on T cells, an adoptive transfer experiment was conducted in mice. Naive T cells from Slc6a13+/+ and Slc6a13–/– mice were isolated and then transferred to Rag1–/– mice, which are infected with C. rodentium or ETEC after 24 h post transfer. After C. rodentium infection, although there was no difference about the percentages of IL-17+ cells within CD4+ cells in the spleen of Slc6a13+/+ CD4+ cell-transferred Rag1–/– mice and Slc6a13–/– CD4+ cell-transferred Rag1–/– mice (Supplementary Figure 4A and B), the percentages of IL-17+ cells within CD4+ cells in the MLN of Rag1–/– mice transferred with Slc6a13–/– CD4+ cells were higher than those in Rag1–/– mice transferred with Slc6a13+/+ CD4+ cells (Fig. 8a, b). Similarly, the percentages of IL-17+ cells within CD4+ cells in the MLN of ETEC-infected Rag1–/– mice transferred with Slc6a13–/– CD4+ cells were higher than those in ETEC-infected Rag1–/– mice transferred with Slc6a13+/+ CD4+ cells (Fig. 8c, d), although no difference was found in the spleen (Supplementary Figure 4C and D). Summarily, these data suggest the intrinsic effect of GAT-2 on Th17 cell responses.
Slc6a13 deficiency promotes intestinal Th17 responses in T cell transfer model of infection. a, b Flow cytometry analysis of Th17 cells in the MLN after C. rodentium infection. Naïve T cells were isolated from Slc6a13+/+ and Slc6a13–/– mice and transferred into Rag1–/– mice before C. rodentium infection, and the CD4+IL-17+ cells analyzed at 10 days post infection. Data were analyzed by Mann–Whitney test and shown as the means ± SD. N = 2 in WT to Rag1–/– group, and 3 in GAT-2–/– to Rag1–/– group. c, d Flow cytometry analysis of Th17 cells in the MLN after ETEC infection. Naïve T cells were isolated from Slc6a13+/+ and Slc6a13–/– mice and transferred into Rag1–/– mice before ETEC infection, and the CD4+IL-17+ cells analyzed at 5 days post infection. Data were analyzed by Mann–Whitney test and shown as the means ± SD. N = 3 in WT to Rag1–/– group, and 5 in GAT-2–/– to Rag1–/– group
Discussion
Th17 cells provide protection against bacterial infection, and are also associated with the development of autoimmune diseases because of the recruitment of cells in the granulocyte lineage, especially neutrophils.20 Under the stimulation of TGF-β and IL-6, naïve T cells differentiate into Th17 cells, which produce the signature cytokine IL-17, but also secrete IL-17F, IL-21, IL-22, and IL-23.21 The expression of transcription factor retinoic acid receptor-related orphan receptor-γt (RORγt) and the activation of STAT-3 and mTOR are the characteristics for Th17 differentiation.19,22,23,24,25 There are also metabolic characteristics for Th17 cells, such as glycolysis, glutaminolysis, and fatty acid synthesis, compared to naïve T cells dependent on the tricarboxylic acid (TCA) cycle and oxidative phosphorylation (OXPHOS).26 However, the metabolic pathways that shape the differentiation of Th17 cells are not fully known. We compared the cellular metabolites in Th17 cells and naïve T cells, and found that alanine, aspartate and glutamate metabolism, arginine and proline metabolism, purine metabolism, glycine, serine and threonine metabolism, and riboflavin metabolism are altered during Th17 cell differentiation. This suggests that the metabolites in these pathways affect the T cell function and the differentiation of Th17 cells. Arginine has been shown to affect the cellular metabolism and survival of T cells,27 while serine promotes the effector T cell expansion through serine, glycine, one-carbon metabolic network.28 Interestingly, this study found that the GABA shunt pathway increases in Th17 cells, suggesting it may regulate the activation and differentiation of T cells, similar to macrophages.29 Other known pathways that define Th17 cells, such as glycolysis and pentose phosphate pathway, are not found in this study. The possible explanation is that this study used LC-QqQ-MS to analyze the metabolites in T cells, which is targeted metabolomics to analyze only 206 metabolites, rather than all metabolites in T cells.
In this study, we found that the cellular level of GABA and protein abundance of GAT-2 increased during Th17 cell differentiation, indicating that they are associated with its activation and differentiation. Similarly, a previous study reported that resting lymphocytes express GAT-1 whereas activated lymphocytes express at least one of the GAT-1 and GAT-2.7 GAT-1 is negatively involved in T cell activation and T cell-mediated responses because GAT-1 deficiency promotes activation of NF-κB signaling pathway and phosphorylation of Jnk.8,9 In this study, GAT-2 deficiency is found to promote Th17 cell differentiation in vitro, and IL-17 expression in mouse model of intestinal infection and inflammation. Notably, the T cell-specific effects of GAT-2 are also validated with a T cell adoptive transfer model of infection. GAT-2 deficiency increases the level of GABA in the medium, and affects the cellular metabolism in Th17 cells, especially, the defect of the GABA shunt pathway. However, the influence of GAT-2 deficiency on other metabolic pathways of Th17 cells, like glycolysis and OXPHOS, remains to know. This suggests that GAT-2 deficiency promotes the differentiation Th17 cells may through cellular GABA. In mouse and piglet model of intestinal infection, GABA promotes the differentiation of Th17 cells and expression of IL-17 through the mTORC1 pathway.6 Similarly, this study found that GAT-2 deficiency promotes Th17 cell differentiation through GABA–mTOR signaling. However, whether GAT-2 deficiency promotes Th17 cell differentiation through other metabolic pathways or signaling pathways is unknown, given that GAT-2 deficiency affects arginine and proline metabolism, glycerophospholipid metabolism, and riboflavin metabolism in Th17 cells. Also, in the mouse model of infection, GAT-2 deficiency promotes the expression of intestinal TGF-β1 and IL-1β, both of which have vital roles in Th17 differentiation.30,31,32 Thus, GAT-2 deficiency might promote Th17 responses in vivo by affecting the cytokine milieu for Th17 cell differentiation. Another interesting study also showed that BGT-1 is present in T cells.2 However, it is unknown whether BGT-1 also affects T cell-mediated responses in a manner similar to that of GAT-1 and -2.
In this study, the deficiency of GAT-2 has little effect on the T cell development and peripheral T cell homeostasis. Indeed, the expression of GAT-2 is hardly detected on naïve T cells, and there is little change in the metabolic pathway of naïve T cells after GAT-2 deficiency, except thiamine metabolism. Similarly, GAT-1 deficiency has little effect on the ratio of CD4+/CD8+ thymocytes, and the number of CD4+ cells and CD8+ cells in the spleen.9 However, it is interesting to know that whether GAT-2 deficiency affects other immune cells, such as macrophages. Besides to GAD and GABA receptor,1 macrophages also express GAT-2 and BGT-1 (unpublished data). GAT-2 is also detected in other subtypes of T cells; thus, it remains to know whether GAT-2 regulates the differentiation of other subtypes of T cells, including Th1 cells, Th2, Th9, Th22, and Treg cells. Interestingly, previous study found that GAT-1 deficiency promotes the Th1 responses in the mouse model of experimental autoimmune encephalomyelitis.8
Notably, GABA shows anti-inflammatory action through inhibiting of major inflammatory events. For example, GABA signaling ameliorates the inflammation in experimental autoimmune encephalomyelitis.2 Oral GABA treatment ameliorates the inflammatory process both in non-obese diabetic mice33 and in a mouse model of rheumatoid arthritis.34 Also, GABAA agonists are beneficial in experimental encephalomyelitis2 and allergic asthma,35 and GABAB agonists exhibit a positive effect in dermatitis models.36 This study shows that GABA signaling promotes the differentiation of Th17 cells in vitro, and Th17 cell responses in the mouse model of intestinal infection and inflammation. Indeed, our previous study also found that GABA increases intestinal Il-17 expression in the context of ETEC infection, while GABA synthetic enzyme blockers (l-allyglycine or semicarbazide) and GABA receptor antagonists (bicuculline or CGP-35348) inhibit intestinal Il-17 expression during ETEC infection.6 There are various possible reasons for this discrepancy, such as animal model. We hypothesize that the influence of GABAergic system on different immune cells (e.g., macrophages, T cells, and B cells), or even on different subtypes of immune cells (e.g., Th1, Th17, and Treg cells), differs. Also, the component of GABAergic system regulates the function of immune system may be through other mechanisms, rather than the GABA signaling.
In conclusion, there is significant metabolic profile between naïve T cells and Th17 cells, especially in the GABA shunt pathway. GAT-2 deficiency has little effect on the metabolic pathways in naïve T cells, and has little effect on the T cell development and peripheral T cell homeostasis. However, GAT-2 deficiency impairs the GABA uptake and GABA shunt pathway in Th17 cells, and promotes the Th17 cell differentiation through the activation of GABA–mTOR signaling. In the mouse model of intestinal infection and inflammation, GAT-2 deficiency promotes the Th17 responses.
References
Wu, C. et al. The immunological function of GABAergic system. Front. Biosci. (Landmark Ed.) 22, 1162–1172 (2017).
Bhat, R. et al. Inhibitory role for GABA in autoimmune inflammation. Proc. Natl. Acad. Sci. USA 107, 2580–2585 (2010).
Prud’homme, G. J., Glinka, Y. & Wang, Q. Immunological GABAergic interactions and therapeutic applications in autoimmune diseases. Autoimmun. Rev. 14, 1048–1056 (2015).
Crowley, T., Cryan, J. F., Downer, E. J. & O’Leary, O. F. Inhibiting neuroinflammation: the role and therapeutic potential of GABA in neuro-immune interactions. Brain Behav. Immun. 54, 260–277 (2016).
Yocum, G. T. et al. GABAA receptor alpha4-subunit knockout enhances lung inflammation and airway reactivity in a murine asthma model. Am. J. Physiol. Lung Cell Mol. Physiol. 313, L406–L415 (2017).
Ren, W. et al. Intestinal microbiota-derived GABA mediates interleukin-17 expression during enterotoxigenic Escherichia coli infection. Front Immunol. 7, 685 (2016).
Dionisio, L., Jose De, R. M., Bouzat, C. & Esandi Mdel, C. An intrinsic GABAergic system in human lymphocytes. Neuropharmacology 60, 513–519 (2011).
Wang, Y. et al. Gamma-aminobutyric acid transporter 1 negatively regulates T cell-mediated immune responses and ameliorates autoimmune inflammation in the CNS. J. Immunol. 181, 8226–8236 (2008).
Wang, Y. et al. Gamma-aminobutyric acid transporter 1 negatively regulates T cell activation and survival through protein kinase C-dependent signaling pathways. J. Immunol. 183, 3488–3495 (2009).
Ren, W. et al. Draft genome sequence of enterotoxigenic Escherichia coli strain W25K. Genome Announc. 2, https://doi.org/10.1128/genomeA.00593-14 (2014).
Du, Y. et al. Rapid discovery of cyclopamine analogs from Fritillaria and Veratrum plants using LC-Q-TOF-MS and LC-QqQ-MS. J. Pharm. Biomed. Anal. 142, 201–209 (2017).
Cai, Y. P., Weng, K., Guo, Y., Peng, J. & Zhu, Z. J. An integrated targeted metabolomic platform for high-throughput metabolite profiling and automated data processing. Metabolomics 11, 1575–1586 (2015).
Wu, C. et al. Transcriptomic analysis on responses of murine lungs to Pasteurella multocida infection. Front. Cell Infect. Microbiol. 7, 251 (2017).
Ren, W. et al. Proteome analysis for the global proteins in the jejunum tissues of enterotoxigenic Escherichia coli-infected piglets. Sci. Rep. 6, 25640 (2016).
Ren, W. et al. Melatonin alleviates weanling stress in mice: involvement of intestinal microbiota. J. Pineal Res., https://doi.org/10.1111/jpi.12448 (2017).
Shu, Q. et al. GABAB R/GSK-3beta/NF-kappaB signaling pathway regulates the proliferation of colorectal cancer cells. Cancer Med. 5, 1259–1267 (2016).
Paul, A. M. et al. GABA transport and neuroinflammation are coupled in multiple sclerosis: regulation of the GABA transporter-2 by ganaxolone. Neuroscience 273, 24–38 (2014).
Ren, W. et al. mTORC1 signaling and IL-17 expression: defining pathways and possible therapeutic targets. Eur. J. Immunol. 46, 291–299 (2016).
Nagai, S., Kurebayashi, Y. & Koyasu, S. Role of PI3K/Akt and mTOR complexes in Th17 cell differentiation. Ann. NY Acad. Sci. 1280, 30–34 (2013).
Miossec, P. & Kolls, J. K. Targeting IL-17 and TH17 cells in chronic inflammation. Nat. Rev. Drug Discov. 11, 763–776 (2012).
Wilson, N. J. et al. Development, cytokine profile and function of human interleukin 17-producing helper T cells. Nat. Immunol. 8, 950–957 (2007).
Ivanov, I. I., Zhou, L. & Littman, D. R. Transcriptional regulation of Th17 cell differentiation. Semin. Immunol. 19, 409–417 (2007).
Shabgah, A. G., Fattahi, E. & Shahneh, F. Z. Interleukin-17 in human inflammatory diseases. Post. Dermatol. Alergol. 31, 256–261 (2014).
McGeachy, M. J. & Cua, D. J. Th17 cell differentiation: the long and winding road. Immunity 28, 445–453 (2008).
Kim, J. S. et al. Natural and inducible TH17 cells are regulated differently by Akt and mTOR pathways. Nat. Immunol. 14, 611–618 (2013).
Ren, W. et al. Amino-acid transporters in T-cell activation and differentiation. Cell Death Dis. 8, e2757 (2017).
Geiger, R. et al. L-Arginine modulates T cell metabolism and enhances survival and anti-tumor activity. Cell 167, 829–842 e813 (2016).
Ma, E. H. et al. Serine is an essential metabolite for effector T cell expansion. Cell Metab. 25, 345–357 (2017).
Tannahill, G. M. et al. Succinate is an inflammatory signal that induces IL-1beta through HIF-1alpha. Nature 496, 238–242 (2013).
Ghoreschi, K. et al. Generation of pathogenic T(H)17 cells in the absence of TGF-beta signalling. Nature 467, 967–971 (2010).
Mangan, P. R. et al. Transforming growth factor-beta induces development of the T(H)17 lineage. Nature 441, 231–234 (2006).
Veldhoen, M., Hocking, R. J., Atkins, C. J., Locksley, R. M. & Stockinger, B. TGFbeta in the context of an inflammatory cytokine milieu supports de novo differentiation of IL-17-producing T cells. Immunity 24, 179–189 (2006).
Tian, J. et al. Gamma-aminobutyric acid inhibits T cell autoimmunity and the development of inflammatory responses in a mouse type 1 diabetes model. J. Immunol. 173, 5298–5304 (2004).
Tian, J., Yong, J., Dang, H. & Kaufman, D. L. Oral GABA treatment downregulates inflammatory responses in a mouse model of rheumatoid arthritis. Autoimmunity 44, 465–470 (2011).
Munroe, M. E., Businga, T. R., Kline, J. N. & Bishop, G. A. Anti-inflammatory effects of the neurotransmitter agonist Honokiol in a mouse model of allergic asthma. J. Immunol. 185, 5586–5597 (2010).
Duthey, B. et al. Anti-inflammatory effects of the GABA(B) receptor agonist baclofen in allergic contact dermatitis. Exp. Dermatol. 19, 661–666 (2010).
Acknowledgements
The authors are grateful for the help of Mr. Lou from Shanghai Biotree Biotech Co., Ltd. (Shanghai, China) in the high-thoughput targeted metabolomics analysis. This work was supported by the National Natural Science Foundation of China (Grant Nos. 31872365, 31873010 and 31790411), The Natural Science Foundation of the Jiangsu Higher Education Institutions of China (Grant No. 18KJB310019) and State Scholarship Fund of China Scholarship Council. The sponsor has no role in the study design, data collection, analysis, and interpretation of data.
Author information
Authors and Affiliations
Contributions
W.R., B.T., G.Z. and Y.Y. designed the experiments; W.R., Y.L., X.D. and Y.J. conducted the experiments; J.Y. and Y.X. helped with animal experiments; J.D. and X.J. provided Rag1 knockout mice; Z.L. and G.Y. provided help in flowcytometery analysis; W.R., Y.L., X.D. and Y.J. analyzed the data; W.R. wrote the manuscript; and J.D., G.Z., C.Z., P.H., J.L. and Y.Y. revised the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ren, W., Liao, Y., Ding, X. et al. Slc6a13 deficiency promotes Th17 responses during intestinal bacterial infection. Mucosal Immunol 12, 531–544 (2019). https://doi.org/10.1038/s41385-018-0111-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41385-018-0111-7
This article is cited by
-
Gut microbiota bridges dietary nutrients and host immunity
Science China Life Sciences (2023)
-
Glutamine metabolism in Th17/Treg cell fate: applications in Th17 cell-associated diseases
Science China Life Sciences (2021)
-
Targeting T-cell oxidative metabolism to improve influenza survival in a mouse model of obesity
International Journal of Obesity (2020)