Abstract
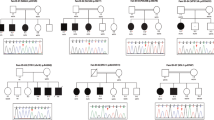
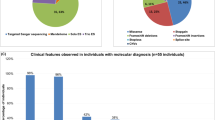
Autosomal recessive (AR) gene defects are the leading genetic cause of intellectual disability (ID) in countries with frequent parental consanguinity, which account for about 1/7th of the world population. Yet, compared to autosomal dominant de novo mutations, which are the predominant cause of ID in Western countries, the identification of AR-ID genes has lagged behind. Here, we report on whole exome and whole genome sequencing in 404 consanguineous predominantly Iranian families with two or more affected offspring. In 219 of these, we found likely causative variants, involving 77 known and 77 novel AR-ID (candidate) genes, 21 X-linked genes, as well as 9 genes previously implicated in diseases other than ID. This study, the largest of its kind published to date, illustrates that high-throughput DNA sequencing in consanguineous families is a superior strategy for elucidating the thousands of hitherto unknown gene defects underlying AR-ID, and it sheds light on their prevalence.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Shaw-Smith C, Redon R, Rickman L, Rio M, Willatt L, Fiegler H, et al. Microarray based comparative genomic hybridisation (array-CGH) detects submicroscopic chromosomal deletions and duplications in patients with learning disability/mental retardation and dysmorphic features. J Med Genet. 2004;41:241–8.
Gilissen C, Hehir-Kwa JY, Thung DT, van de Vorst M, van Bon BW, Willemsen MH, et al. Genome sequencing identifies major causes of severe intellectual disability. Nature. 2014;511:344–7.
Vissers LE, de Ligt J, Gilissen C, Janssen I, Steehouwer M, de Vries P, et al. A de novo paradigm for mental retardation. Nat Genet. 2010;42:1109–12.
Ropers HH. Genetics of early onset cognitive impairment. Annu Rev Genom Hum Genet. 2010;11:161–87.
Musante L, Ropers HH. Genetics of recessive cognitive disorders. Trends Genet. 2014;30:32–39.
de Ligt J, Willemsen MH, van Bon BW, Kleefstra T, Yntema HG, Kroes T, et al. Diagnostic exome sequencing in persons with severe intellectual disability. N Engl J Med. 2012;367:1921–9.
Rauch A, Wieczorek D, Graf E, Wieland T, Endele S, Schwarzmayr T, et al. Range of genetic mutations associated with severe non-syndromic sporadic intellectual disability: an exome sequencing study. Lancet. 2012;380:1674–82.
Fitzgerald TW, Gerety SS, Jones WD, van Kogelenberg M, King DA, McRae J, et al. Large-scale discovery of novel genetic causes of developmental disorders. Nature. 2015;519:223–8.
Lelieveld SH, Reijnders MR, Pfundt R, Yntema HG, Kamsteeg EJ, de Vries P, et al. Meta-analysis of 2104 trios provides support for 10 new genes for intellectual disability. Nat Neurosci. 2016;19:1194–6.
Hamdan FF, Srour M, Capo-Chichi JM, Daoud H, Nassif C, Patry L, et al. De novo mutations in moderate or severe intellectual disability. PLoS Genet. 2014;10:e1004772.
Krumm N, O’Roak BJ, Shendure J, Eichler EE. A de novo convergence of autism genetics and molecular neuroscience. Trends Neurosci. 2014;37:95–105.
Yuen RK, Thiruvahindrapuram B, Merico D, Walker S, Tammimies K, Hoang N, et al. Whole-genome sequencing of quartet families with autism spectrum disorder. Nat Med. 2015;21:185–91.
MacArthur DG, Balasubramanian S, Frankish A, Huang N, Morris J, Walter K, et al. A systematic survey of loss-of-function variants in human protein-coding genes. Science. 2012;335:823–8.
Iossifov I, O’Roak BJ, Sanders SJ, Ronemus M, Krumm N, Levy D, et al. The contribution of de novo coding mutations to autism spectrum disorder. Nature. 2014;515:216–21.
Ronemus M, Iossifov I, Levy D, Wigler M. The role of de novo mutations in the genetics of autism spectrum disorders. Nat Rev Genet. 2014;15:133–41.
Kochinke K, Zweier C, Nijhof B, Fenckova M, Cizek P, Honti F, et al. Systematic phenomics analysis deconvolutes genes mutated in intellectual disability into biologically coherent modules. Am J Hum Genet. 2016;98:149–64.
Vissers LE, Gilissen C, Veltman JA. Genetic studies in intellectual disability and related disorders. Nat Rev Genet. 2016;17:9–18.
Kuss AW, Garshasbi M, Kahrizi K, Tzschach A, Behjati F, Darvish H, et al. Autosomal recessive mental retardation: homozygosity mapping identifies 27 single linkage intervals, at least 14 novel loci and several mutation hotspots. Hum Genet. 2011;129:141–8.
Najmabadi H, Motazacker MM, Garshasbi M, Kahrizi K, Tzschach A, Chen W, et al. Homozygosity mapping in consanguineous families reveals extreme heterogeneity of non-syndromic autosomal recessive mental retardation and identifies 8 novel gene loci. Hum Genet. 2007;121:43–8.
Hu H, Haas SA, Chelly J, Van Esch H, Raynaud M, de Brouwer AP, et al. X-exome sequencing of 405 unresolved families identifies seven novel intellectual disability genes. Mol Psychiatry. 2016;21:133–48.
Heinzen EL, Neale BM, Traynelis SF, Allen AS, Goldstein DB. The genetics of neuropsychiatric diseases: looking in and beyond the exome. Annu Rev Neurosci. 2015;38:47–68.
Durkin MS, Hasan ZM, Hasan KZ. Prevalence and correlates of mental retardation among children in Karachi, Pakistan. Am J Epidemiol. 1998;147:281–8.
Kaufman L, Ayub M, Vincent JB. The genetic basis of non-syndromic intellectual disability: a review. J Neurodev Disord. 2010;2:182–209.
Saad HA, Elbedour S, Hallaq E, Merrick J, Tenenbaum A. Consanguineous marriage and intellectual and developmental disabilities among Arab Bedouins Children of the Negev Region in Southern Israel: a pilot study. Front Public Health. 2014;2:3.
Weller M, Tanieri M, Pereira JC, Almeida Edos S, Kok F, Santos S. Consanguineous unions and the burden of disability: a population-based study in communities of Northeastern Brazil. Am J Hum Biol. 2012;24:835–40.
Monies D, Abouelhoda M, AlSayed M, Alhassnan Z, Alotaibi M, Kayyali H, et al. The landscape of genetic diseases in Saudi Arabia based on the first 1000 diagnostic panels and exomes. Hum Genet. 2017;136:921–39.
Fareed M, Afzal M. Estimating the inbreeding depression on cognitive behavior: a population based study of child cohort. PLoS ONE. 2014;9:e109585.
Hamamy H. Consanguineous marriages: preconception consultation in primary health care settings. J Community Genet. 2012;3:185–92.
Bittles AH, Black ML. Evolution in health and medicine Sackler colloquium: consanguinity, human evolution, and complex diseases. Proc Natl Acad Sci USA. 2010;107 Suppl 1:1779–86.
Schuurs-Hoeijmakers JHM, Hehir-Kwa JY, Pfundt R, van Bon BWM, de Leeuw N, Kleefstra T, et al. Homozygosity mapping in outbred families with mental retardation. Eur J Human Genet. 2011;19:597–601.
Schuurs-Hoeijmakers JHM, Vulto-van Silfhout AT, Vissers LELM, van de Vondervoort IIGM, van Bon BWM, de Ligt J, et al. Identification of pathogenic gene variants in small families with intellectually disabled siblings by exome sequencing. J Med Genet. 2013;50:802–11.
Akawi NA, Al-Jasmi F, Al-Shamsi AM, Ali BR, Al-Gazali L. LINS, a modulator of the WNT signaling pathway, is involved in human cognition. Orphanet J Rare Dis. 2013;8:87.
Najmabadi H, Hu H, Garshasbi M, Zemojtel T, Abedini SS, Chen W, et al. Deep sequencing reveals 50 novel genes for recessive cognitive disorders. Nature. 2011;478:57–63.
Alazami AM, Patel N, Shamseldin HE, Anazi S, Al-Dosari MS, Alzahrani F, et al. Accelerating novel candidate gene discovery in neurogenetic disorders via whole-exome sequencing of prescreened multiplex consanguineous families. Cell Rep. 2015;10:148–61.
Karaca E, Harel T, Pehlivan D, Jhangiani SN, Gambin T, Coban Akdemir Z, et al. Genes that affect brain structure and function identified by rare variant analyses of mendelian neurologic disease. Neuron. 2015;88:499–513.
Riazuddin S, Hussain M, Razzaq A, Iqbal Z, Shahzad M, Polla DL, et al. Exome sequencing of Pakistani consanguineous families identifies 30 novel candidate genes for recessive intellectual disability. Mol Psychiatry. 2017;22:1604–14.
Reuter MS, Tawamie H, Buchert R, Hosny Gebril O, Froukh T, Thiel C, et al. Diagnostic yield and novel candidate genes by exome sequencing in 152 consanguineous families with neurodevelopmental disorders JAMA Psychiatry. 2017;74:293–9.
Anazi S, Maddirevula S, Faqeih E, Alsedairy H, Alzahrani F, Shamseldin HE, et al. Clinical genomics expands the morbid genome of intellectual disability and offers a high diagnostic yield Mol Psychiatry. 2017;22:615–24.
Harripaul R, Vasli N, Mikhailov A, Rafiq MA, Mittal K, Windpassinger C, et al. Mapping autosomal recessive intellectual disability: combined microarray and exome sequencing identifies 26 novel candidate genes in 192 consanguineous families. Mol Psychiatry. 2017 Apr 11. https://doi.org/10.1038/mp.2017.60. [Epub ahead of print]
Saadat M, Ansari-Lari M, Farhud DD. Consanguineous marriage in Iran. Ann Hum Biol. 2004;31:263–9.
Jalal Abbasi-Shavazi M, McDonald P, Hosseini-Chavoshi M. Modernization or cultural maintenance: the practice of consanguineous marriage in Iran. J Biosoc Sci. 2008;40:911–33.
Najmabadi H, Ghamari A, Sahebjam F, Kariminejad R, Hadavi V, Khatibi T, et al. Fourteen-year experience of prenatal diagnosis of thalassemia in Iran. Community Genet. 2006;9:93–97.
Sloan-Heggen CM, Babanejad M, Beheshtian M, Simpson AC, Booth KT, Ardalani F, et al. Characterising the spectrum of autosomal recessive hereditary hearing loss in Iran. J Med Genet. 2015;52:823–9.
Kahrizi, K. and Najmabadi, H. 2015. Genetics of Recessive Cognitive Disorders. eLS. 1–21. Wiley Online Library.
Hu H, Wienker TF, Musante L, Kalscheuer VM, Kahrizi K, Najmabadi H, et al. Integrated sequence analysis pipeline provides one-stop solution for identifying disease-causing mutations. Hum Mutat. 2014;35:1427–35.
Richards S, Aziz N, Bale S, Bick D, Das S, Gastier-Foster J, et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med. 2015;17:405–24.
Keleman K, Kruttner S, Alenius M, Dickson BJ. Function of the Drosophila CPEB protein Orb2 in long-term courtship memory. Nat Neurosci. 2007;10:1587–93.
Siwicki KK, Ladewski L. Associative learning and memory in Drosophila: beyond olfactory conditioning. Behav Process. 2003;64:225–38.
Willemsen MH, Ba W, Wissink-Lindhout WM, de Brouwer AP, Haas SA, Bienek M, et al. Involvement of the kinesin family members KIF4A and KIF5C in intellectual disability and synaptic function. J Med Genet. 2014;51:487–94.
Whibley AC, Plagnol V, Tarpey PS, Abidi F, Fullston T, Choma MK, et al. Fine-scale survey of X chromosome copy number variants and indels underlying intellectual disability. Am J Hum Genet. 2010;87:173–88.
Pouya AR, Abedini SS, Mansoorian N, Behjati F, Nikzat N, Mohseni M, et al. Fragile X syndrome screening of families with consanguineous and non-consanguineous parents in the Iranian population. Eur J Med Genet. 2009;52:170–3.
Hosseini-Chavoshi M, Abbasi-Shavazi MJ, Bittles AH. Consanguineous marriage, reproductive behaviour and postnatal mortality in contemporary Iran. Hum Hered. 2014;77:16–25.
Narasimhan VM, Hunt KA, Mason D, Baker CL, Karczewski KJ, Barnes MR, et al. Health and population effects of rare gene knockouts in adult humans with related parents. Science. 2016;352:474–7.
Khan MA, Rupp VM, Orpinell M, Hussain MS, Altmuller J, Steinmetz MO, et al. A missense mutation in the PISA domain of HsSAS-6 causes autosomal recessive primary microcephaly in a large consanguineous Pakistani family. Hum Mol Genet. 2014;23:5940–9.
Law R, Dixon-Salazar T, Jerber J, Cai N, Abbasi AA, Zaki MS, et al. Biallelic truncating mutations in FMN2, encoding the actin-regulatory protein Formin 2, cause nonsyndromic autosomal-recessive intellectual disability. Am J Hum Genet. 2014;95:721–8.
Larti F, Kahrizi K, Musante L, Hu H, Papari E, Fattahi Z, et al. A defect in the CLIP1 gene (CLIP-170) can cause autosomal recessive intellectual disability. Eur J Hum Genet. 2015;23:331–6.
Oladnabi M, Musante L, Larti F, Hu H, Abedini SS, Wienker T, et al. New evidence for the role of calpain 10 in autosomal recessive intellectual disability: identification of two novel nonsense variants by exome sequencing in Iranian families. Arch Iran Med. 2015;18:179–84.
Alakbarzade V, Hameed A, Quek DQ, Chioza BA, Baple EL, Cazenave-Gassiot A, et al. A partially inactivating mutation in the sodium-dependent lysophosphatidylcholine transporter MFSD2A causes a non-lethal microcephaly syndrome. Nat Genet. 2015;47:814–7.
Guemez-Gamboa A, Nguyen LN, Yang H, Zaki MS, Kara M, Ben-Omran T, et al. Inactivating mutations in MFSD2A, required for omega-3 fatty acid transport in brain, cause a lethal microcephaly syndrome. Nat Genet. 2015;47:809–13.
Iqbal Z, Willemsen MH, Papon MA, Musante L, Benevento M, Hu H, et al. Homozygous SLC6A17 mutations cause autosomal-recessive intellectual disability with progressive tremor, speech impairment, and behavioral problems. Am J Hum Genet. 2015;96:386–96.
Heidari A, Tongsook C, Najafipour R, Musante L, Vasli N, Garshasbi M, et al. Mutations in the histamine N-methyltransferase gene, HNMT, are associated with nonsyndromic autosomal recessive intellectual disability. Hum Mol Genet. 2015;24:5697–710.
Snijders Blok L, Madsen E, Juusola J, Gilissen C, Baralle D, Reijnders MR, et al. Mutations in DDX3X are a common cause of unexplained intellectual disability with gender-specific effects on Wnt signaling. Am J Hum Genet. 2015;97:343–52.
O’Rawe JA, Wu Y, Dorfel MJ, Rope AF, Au PY, Parboosingh JS, et al. TAF1 variants are associated with dysmorphic features, intellectual disability, and neurological manifestations. Am J Hum Genet. 2015;97:922–32.
Nellhaus G. Head circumference from birth to eighteen years. Practical composite international and interracial graphs. Pediatrics. 1968;41:106–14.
Davis JM, Keeney JG, Sikela JM, Hepburn S. Mode of genetic inheritance modifies the association of head circumference and autism-related symptoms: a cross-sectional study. PLoS One. 2013;8:e74940.
Liang H, Xiao G, Yin H, Hippenmeyer S, Horowitz JM, Ghashghaei HT. Neural development is dependent on the function of specificity protein 2 in cell cycle progression. Development. 2013;140:552–61.
Vandeputte DA, Meije CB, van Dartel M, Leenstra S, IJ-K H, Das PK, et al. GOA, a novel gene encoding a ring finger B-box coiled-coil protein, is overexpressed in astrocytoma. Biochem Biophys Res Commun. 2001;286:574–9.
Parkel S, Lopez-Atalaya JP, Barco A. Histone H3 lysine methylation in cognition and intellectual disability disorders. Learn Mem. 2013;20:570–9.
Conway E, Healy E, Bracken AP. PRC2 mediated H3K27 methylations in cellular identity and cancer. Curr Opin Cell Biol. 2015;37:42–48.
Frerker N, Wagner L, Wolf R, Heiser U, Hoffmann T, Rahfeld JU, et al. Neuropeptide Y (NPY) cleaving enzymes: structural and functional homologues of dipeptidyl peptidase 4. Peptides. 2007;28:257–68.
Ono S, Yoshiura K, Kinoshita A, Kikuchi T, Nakane Y, Kato N, et al. Mutations in PRRT2 responsible for paroxysmal kinesigenic dyskinesias also cause benign familial infantile convulsions. J Hum Genet. 2012;57:338–41.
Iossifov I, Ronemus M, Levy D, Wang Z, Hakker I, Rosenbaum J, et al. De novo gene disruptions in children on the autistic spectrum. Neuron. 2012;74:285–99.
Basel-Vanagaite L, Attia R, Yahav M, Ferland RJ, Anteki L, Walsh CA, et al. The CC2D1A, a member of a new gene family with C2 domains, is involved in autosomal recessive non-syndromic mental retardation. J Med Genet. 2006;43:203–10.
Mir A, Kaufman L, Noor A, Motazacker MM, Jamil T, Azam M, et al. Identification of mutations in TRAPPC9, which encodes the NIK- and IKK-beta-binding protein, in nonsyndromic autosomal-recessive mental retardation. Am J Hum Genet. 2009;85:909–15.
Kishi N, MacDonald JL, Ye J, Molyneaux BJ, Azim E, Macklis JD. Reduction of aberrant NF-kappaB signalling ameliorates Rett syndrome phenotypes in Mecp2-null mice. Nat Commun. 2016;7:10520.
Gamsiz ED, Sciarra LN, Maguire AM, Pescosolido MF, van Dyck LI, Morrow EM. Discovery of rare mutations in autism: elucidating neurodevelopmental mechanisms. Neurotherapeutics. 2015;12:553–71.
Coba MP, Komiyama NH, Nithianantharajah J, Kopanitsa MV, Indersmitten T, Skene NG, et al. TNiK is required for postsynaptic and nuclear signaling pathways and cognitive function. J Neurosci. 2012;32:13987–99.
Bayes A, van de Lagemaat LN, Collins MO, Croning MD, Whittle IR, Choudhary JS, et al. Characterization of the proteome, diseases and evolution of the human postsynaptic density. Nat Neurosci. 2011;14:19–21.
Suhl JA, Chopra P, Anderson BR, Bassell GJ, Warren ST. Analysis of FMRP mRNA target datasets reveals highly associated mRNAs mediated by G-quadruplex structures formed via clustered WGGA sequences. Hum Mol Genet. 2014;23:5479–91.
Ouwenga RL, Dougherty J. Fmrp targets or not: long, highly brain-expressed genes tend to be implicated in autism and brain disorders. Mol Autism. 2015;6:16.
Gilman SR, Iossifov I, Levy D, Ronemus M, Wigler M, Vitkup D. Rare de novo variants associated with autism implicate a large functional network of genes involved in formation and function of synapses. Neuron. 2011;70:898–907.
O’Roak BJ, Deriziotis P, Lee C, Vives L, Schwartz JJ, Girirajan S, et al. Exome sequencing in sporadic autism spectrum disorders identifies severe de novo mutations. Nat Genet. 2011;43:585–9.
Basel-Vanagaite L, Taub E, Halpern GJ, Drasinover V, Magal N, Davidov B, et al. Genetic screening for autosomal recessive nonsyndromic mental retardation in an isolated population in Israel. Eur J Hum Genet. 2007;15:250–3.
Krawczak M, Barnes R. How obedience of marriage rules may counteract genetic drift. J Community Genet. 2010;1:23–28.
Shaheen R, Patel N, Shamseldin H, Alzahrani F, Al-Yamany R, AL A, et al. Accelerating matchmaking of novel dysmorphology syndromes through clinical and genomic characterization of a large cohort. Genet Med. 2016;18:686–95.
Power RA, Kyaga S, Uher R, MacCabe JH, Langstrom N, Landen M, et al. Fecundity of patients with schizophrenia, autism, bipolar disorder, depression, anorexia nervosa, or substance abuse vs their unaffected siblings. JAMA Psychiatry. 2013;70:22–30.
Casanova JL, Conley ME, Seligman SJ, Abel L, Notarangelo LD. Guidelines for genetic studies in single patients: lessons from primary immunodeficiencies. J Exp Med. 2014;211:2137–49.
Zhang B, Spreafico M, Zheng C, Yang A, Platzer P, Callaghan MU, et al. Genotype-phenotype correlation in combined deficiency of factor V and factor VIII. Blood. 2008;111:5592–5600.
Ropers HH. New perspectives for the elucidation of genetic disorders. Am J Hum Genet. 2007;81:199–207.
Soukarieh O, Gaildrat P, Hamieh M, Drouet A, Baert-Desurmont S, Frebourg T, et al. Exonic splicing mutations are more prevalent than currently estimated and can be predicted by using in silico tools. PLoS Genet. 2016;12:e1005756.
Lee M, Roos P, Sharma N, Atalar M, Evans TA, Pellicore MJ, et al. Systematic computational identification of variants that activate exonic and intronic cryptic splice sites. Am J Hum Genet. 2017;100:751–65.
Lupianez DG, Spielmann M, Mundlos S. Breaking TADs: how alterations of chromatin domains result in disease. Trends Genet. 2016;32:225–37.
Huang L, Jolly LA, Willis-Owen S, Gardner A, Kumar R, Douglas E, et al. A noncoding, regulatory mutation implicates HCFC1 in nonsyndromic intellectual disability. Am J Hum Genet. 2012;91:694–702.
Bohning D, Rocchetti I, Alfo M, Holling H. A flexible ratio regression approach for zero-truncated capture-recapture counts. Biometrics. 2016;72:697–706.
Acknowledgements
We are grateful to all patients and families for their participation in this study, S. Banihashemi and Kh. Jalalvand for technical support and Gabriele Eder for assisting us with the preparation of the manuscript. We would also like to thank the NHLBI GO Exome Sequencing Project and its ongoing studies, as well as the Exome Aggregation Consortium and the groups that provided exome variant data for comparison. A full list of contributing groups can be found at http://exac.broadinstitute.org/about. This research was supported by the European Union through FP7 project GENCODYS, grant no 241995 (organizer: Hans van Bokhoven), the Max Planck Innovation Funds and the Ministry of Health and Medical Education, Islamic Republic Iran. Additional support was received from the Iranian National Science Foundation (grant no.s 92038458 and 92035782), the Iranian National Elite Foundation and the Iranian Science Elite Federation.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Hu, H., Kahrizi, K., Musante, L. et al. Genetics of intellectual disability in consanguineous families. Mol Psychiatry 24, 1027–1039 (2019). https://doi.org/10.1038/s41380-017-0012-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41380-017-0012-2
This article is cited by
-
In silico analysis of mutation spectrum of Ehlers–Danlos, osteogenesis imperfecta, and cutis laxa overlapping phenotypes in Iranian population
Egyptian Journal of Medical Human Genetics (2024)
-
Bi-allelic truncating variants in CASP2 underlie a neurodevelopmental disorder with lissencephaly
European Journal of Human Genetics (2024)
-
New insights into the clinical and molecular spectrum of the MADD-related neurodevelopmental disorder
Journal of Human Genetics (2024)
-
Novel compound heterozygous ABCA2 variants cause IDPOGSA, a variable phenotypic syndrome with intellectual disability
Journal of Human Genetics (2024)
-
Label free quantitative proteomic profiling of serum samples of intellectually disabled young patients revealed dysregulation of complement coagulation and cholesterol cascade systems
Metabolic Brain Disease (2024)