Abstract
Although low-grade non-intestinal-type sinonasal adenocarcinoma (SNAC) is formally a diagnosis of exclusion defined by the absence of salivary or intestinal differentiation, most tumors in this category comprise a distinctive histologic group that are increasingly thought to derive from seromucinous glands. However, the molecular underpinnings of SNAC remain poorly understood, and it is unclear if diverse genetic alterations recently reported in isolated cases should delineate separate subgroups. This study aims to perform comprehensive evaluation of gene fusions and mutations and their histologic correlates in low-grade SNAC to clarify its pathogenesis and classification. We identified 18 non-intestinal-type SNAC that all displayed characteristic tubulopapillary architecture and low-grade cytology, although several cases had other unique histologic features and 3 showed intermixed high-grade areas. Among tumors stained with S100 protein, SOX10, and DOG1, 86% expressed at least one of these seromucinous markers. Of 17 cases with sufficient RNA or DNA available for analysis, likely oncogenic molecular alterations were identified in 76% of cases, most notably including CTNNB1 p.S33F mutations in 2 cases, concomitant BRAF p.V600E and AKT1 p.E17K mutations in 2 cases, and ETV6::NTRK3, PRKAR1A::MET, FN1::NRG1, and DNAJB1::PRKACA fusions in 1 case each. While tumors with most genetic alterations were histologically indistinguishable, cases with CTNNB1 mutations had intermixed squamoid morules and cases with BRAF and AKT1 mutations showed a myoepithelial cell population and prominent papillary to micropapillary architecture. Overall, these findings confirm previous reports of frequent seromucinous differentiation in low-grade SNAC. However, these tumors display striking molecular diversity with involvement of multiple kinase fusions, leading to frequent activation of signaling cascades including the MAPK pathway. While most genetic alterations are not associated with sufficiently distinctive histologic features to suggest separate classification, biphasic tumors with BRAF p.V600E mutations are more unique and may represent a distinctive subgroup.
Similar content being viewed by others
Introduction
The sinonasal tract can give rise to a diverse range of gland-forming neoplasms. Those tumors that do not fit into defined salivary tumor types or show intestinal differentiation have traditionally been classified as non-intestinal-type sinonasal adenocarcinomas (SNAC)- a category that is further partitioned into low-grade and high-grade groups1,2. Although this terminology suggests that non-salivary, non-intestinal SNAC is a diagnosis of exclusion, the majority of tumors in this category actually represent a histologically distinctive group of low-grade adenocarcinomas that demonstrate a recognizable mix of tubular and papillary architecture. While these tumors were historically thought to be of surface origin3,4,5, their consistent tubulopapillary morphology and immunohistochemical expression of putative seromucinous markers S100 protein, SOX10, and/or DOG1 has supported increasing consensus that they actually arise from seromucinous glands6,7,8,9. As such, the name seromucinous adenocarcinoma has been proposed to describe this distinctive group of SNAC7,8, although this terminology is not currently recommended by the World Health Organization (WHO) Classification of Head and Neck Tumours1.
Despite these well-defined histologic characteristics, the molecular underpinnings of low-grade SNAC are still not well understood, with somewhat piecemeal literature assessing for single alterations or documenting findings in individual cases. While ETV6::NTRK3 and ETV6::RET have been identified in several low-grade SNAC, these cases represent <10% of all tumors evaluated10,11,12. A SYN2::PPARG fusion has also been reported in a single case of low-grade SNAC, and an EGFR::ZNF267 fusion was identified in a case with overlapping features of seromucinous hamartoma and low-grade SNAC13,14. CTNNB1 activating mutations and BRAF V600E mutations have also been documented in two cases each, with distinctive histologic elements15,16. Some of these molecular findings have been proposed to define unique subgroups of SNAC10,11,12,17. However, no study has evaluated both gene fusions and mutations in low-grade SNAC to establish their frequency or histologic correlates. This study aims to perform comprehensive histologic and molecular analysis of a large series of non-intestinal-type SNAC to better understand the classification and pathogenesis of these unique neoplasms.
Materials and methods
Case selection
We identified 18 cases of non-intestinal-type SNAC from the authors’ consultation files and surgical pathology archives that had tissue available for molecular analysis. For inclusion in this study, tumors had to meet the 2017 WHO Classification of Head and Neck Tumours criteria for low-grade non-intestinal-type SNAC, including (1) no diagnostic features of any specific salivary tumor type or intestinal differentiation, (2) tubular and/or papillary architecture, (3) complex growth including back to back or cribriform glands with minimal intervening stroma, and (4) uniform nuclei1. As invasive growth is not part of this definition, degree of invasion was not considered as an inclusion criterion in this series. Tumors that harbored some higher grade components were included if recognizable low-grade areas that met the above criteria were also evident. All available hematoxylin and eosin (H&E) sections were reviewed for all cases, and histologic features were tabulated. Because the majority of cases were seen in consultation, detailed clinical and follow-up information was not available.
Molecular analysis
RNA sequencing (RNA-Seq) was attempted on all 18 cases and DNA next-generation sequencing (NGS) was attempted on 15 cases as described in detail elsewhere18. In brief, both RNA and DNA were isolated from 10 um whole-slide tissue sections using Qiagen AllPrep kits (Qiagen, Germantown, MD). A modified TruSight RNA Pan-Cancer kit (Illumina, San Diego, CA) was used to make a sequencing library containing all exons from 1425 cancer-related genes. Sequencing was performed on the NextSeq 550 (Illumina) with a minimum of 6,000,000 mapped reads. All fusions and variants were reviewed in the Integrated Genomics Viewer (Broad Institute, Cambridge, MA). The Star-Fusion algorithm was used to call fusions, and somatic variants were identified using databases including dbSNP and gnomAD.
Immunohistochemistry
The results of existing immunohistochemical stains were tabulated for each case. Antibodies used in the majority of cases included AE1/AE3 (clone pck-26; prediluted; Ventana Medical Systems, Tucson, AZ), CK7 (clone ov-tl; 1:500; Dako, Carpinteria, CA), CK20 (clone Ks20.8; prediluted; Dako), CDX2 (clone EPR2764Y; prediluted; Dako), S100 protein (clone 4C4.9; prediluted; Ventana Medical Systems), SOX10 (clone N-20; prediluted; BioCare Medical, Pacheco, CA), p63 (clone 4a4; prediluted; BioCare Medical), p40 (clone BC28; 1:100; BioCare Medical), DOG1 (clone SP31; prediluted; CellMarque/Sigma-Aldrich, St. Louis, MO), smooth muscle actin (SMA; clone 1A4; prediluted; Ventana Medical Systems), calponin (clone M3556; 1:500; Dako), and beta-catenin (clone 14; 1:1000; BD Biosciences, Franklin Lakes, NJ). In most cases, staining was performed using standard protocols on Ventana BenchMark Ultra autostainers (Ventana Medical Systems) in the presence of appropriate controls, and signals were visualized using the ultraView polymer detection kit (Ventana Medical Systems). In cases where few or no stains were performed at the time of diagnosis, tissue was prioritized for molecular testing over additional immunohistochemistry.
Results
Histologic findings
The low-grade SNAC all arose in the nasal cavities of 10 women and 8 men with a median age of 56 years (range 17–86 years). All tumors were composed of crowded, back to back, and confluent tubules, with variable areas of papillary architecture, composed of cuboidal to polygonal cells with a moderate amount of pale eosinophilic to amphophilic cytoplasm. However, a subset of cases had more unique histologic features intermixed. Three tumors demonstrated a diffuse oncocytic appearance, 2 had scattered squamoid morules, and 1 included a prominent population of goblet cells. Although most tumors appeared to be composed of only one cell type, 2 cases had distinct biphasic populations of basal and luminal cells evident on H&E sections. While all cases had areas of low-grade cytology, 1 case had a markedly elevated mitotic rate and 2 had zones of increased cytologic atypia and necrosis, 1 of which demonstrated overt sarcomatoid transformation. One case also arose in association with a respiratory epithelial adenomatoid hamartoma (REAH) but had areas of crowded, back-to-back tubules diagnostic of adenocarcinoma.
Immunohistochemical findings
Immunohistochemical findings are tabulated in Table 1. All cases tested were positive for AE1/AE3 (n = 8) and CK7 (n = 13) and negative for CK20 (n = 10) and CDX2 (n = 7). There were also 7 cases positive for S100 protein (50%), 7 cases positive for SOX10 (64%), and 6 cases positive for DOG1 (75%). Of 7 cases that had S100 protein, SOX10, and DOG1 staining all performed, 6 (86%) were positive for at least one of these markers; the 1 tumor negative for all 3 stains had an oncocytic appearance. The 2 cases with clear biphasic cell populations on H&E sections demonstrated positivity for p63 or p40, SMA, and calponin in a basal distribution, with S100 protein and SOX10 expression in both basal and luminal cells. Two additional cases that lacked a clear basal cell layer on H&E also displayed peripheral expression of p40 but were negative for SMA and calponin. The single tumor tested had diffuse nuclear and cytoplasmic beta-catenin positivity.
Molecular findings
Results of molecular analysis are summarized in Table 2. Among the 18 low-grade SNAC, RNA was successfully amplified for analysis in 15 cases. Four of these cases (27%) demonstrated gene fusions including ETV6::NTRK3, PRKAR1A::MET, FN1::NRG1, and DNAJB1::PRKACA in 1 case each. DNA was also successfully amplified in 14 cases, of which putative oncogenic mutations were identified in 9 (64%), encompassing 2 cases with concomitant BRAF p.V600E and AKT1 p.E17K mutations, 2 cases with activating CTNNB1 p.S33F mutations, 1 case with biallelic NF1 p.S1262fs and c.888 + 1 G > T mutations, 1 case with TP53 p.K132M mutation, 1 case with CUL3 p.V427fs mutation, 1 case with RAF1 amplification, and 1 case with homozygous CDKN2A loss. The remaining cases had only variants of uncertain significance. Overall, of 17 cases with either RNA or DNA available for analysis, 13 cases (76%) harbored a likely oncogenic fusion or driver mutation. Pathways known to be activated by these gene alterations are also noted in Table 2, with 7 of the 13 alterations (54%) leading to MAPK pathway activation.
Tumors with gene fusions
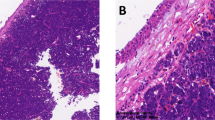
All 4 SNAC with gene fusions were almost entirely composed of crowded to confluent tubules with a moderate amount of eosinophilic to amphophilic cytoplasm and rare poorly-formed papillary excrescences. This pattern was exemplified by the case with DNAJB1::PRKACA fusion, which had no other exceptional features (Fig. 1A). While all four of these tumors largely had monotonous, centrally-placed, round to oval nuclei, the case with ETV6::NTRK3 fusion harbored areas of markedly increased mitotic activity (Fig. 1B). The case with FN1::NRG1 fusion arose in association with REAH, which showed prominent invaginations of surface epithelium and basement membrane material amongst less crowded tubules (Fig. 1C) but transitioned to more expansile areas of confluent growth diagnostic of adenocarcinoma (Fig. 1D). The most unusual morphology was seen in the case with PRKAR1A::MET fusion, which demonstrated a dominant population of goblet cells with well-defined intracytoplasmic mucin vacuoles and more abundant papillary structures (Fig. 1E) intermixed with focal areas of more conventional serous tubules (Fig. 1F). Notably, despite the goblet cells, this tumor was negative for CK20 and CDX2.
The tumor with DNAJB1::PRKACA fusion exemplified the crowded tubules of SNAC (A; case 4). The case with ETV6::NTRK3 fusion was notable for a markedly elevated mitotic rate despite otherwise low-grade cytology (B; case 3). Areas of invaginated surface epithelium lined by dense basement membrane material diagnostic of REAH (C; case 1) were adjacent to more confluent glands in a SNAC with FN1::NRG1 fusion (D; case 1). The tumor with PRKAR1A::MET fusion showed prominent goblet cells with well-defined intracytoplasmic mucin vacuoles (E; case 2) intermixed with more conventional serous tubules (F; case 2).
Tumors with CTNNB1 mutations
Both tumors that demonstrated activating p.S33F point mutations in the CTNNB1 gene were uniformly composed of crowded tubules with pale eosinophilic cytoplasm and round nuclei similar to other cases. However, both of these tumors also displayed scattered squamoid morules that ranged from small and focal in one case (Fig. 2A) to large and prominent in 1 case (Fig. 2B). Beta-catenin immunohistochemistry was available for one case and demonstrated diffuse nuclear and cytoplasmic positivity (Fig. 2C). S100 protein and SOX10 were also positive in both tumors.
The tumors with CTNNB1 activating mutations both were composed of crowded tubules with pale eosinophilic cytoplasm and round nuclei, with squamoid morules that ranged from small and focal (A; case 5) to large and prominent (B; case 6). The one case tested displayed diffuse nuclear and cytoplasmic beta-catenin positivity (C; case 6).
Tumors with BRAF and AKT1 mutations
The 2 tumors with concomitant BRAF p.V600E and AKT1 p.E17K mutations both had biphasic cell populations evident on H&E. The peripheral layer was composed of cuboidal cells with scant clear to pale eosinophilic cytoplasm and round nuclei (Fig. 3A). The central layer displayed larger cells with highly variable architecture, including both crowded tubules with areas of oncocytic change that were indistinguishable from most monophasic tumors (Fig. 3B) as well as complex micropapillary excrescences (Fig. 3C) and papillary fronds (Fig. 3D). By immunohistochemistry, the peripheral cells demonstrated a well-developed myoepithelial phenotype with positivity for p63 or p40 (Fig. 3E), S100 protein, SOX10, SMA, and calponin (Fig. 3F). The luminal cells were positive for AE1/AE3 and CK7 as well as S100 protein and SOX10. Notably, both cell populations were intimately intermixed and showed complex confluent growth throughout the tumor, suggesting that the tumor truly had biphasic elements and was not just colonizing existing myoepithelial cells. Interestingly, despite clear low-grade areas, both of these tumors also had high grade components, including nuclear enlargement, increased pleomorphism, and patchy tumor necrosis, and 1 tumor displayed overt sarcomatoid transformation (Fig. 3G) with areas of osteoid production.
Two tumors with concomitant BRAF p.V600E and AKT1 p.E17K mutations demonstrated biphasic populations of peripheral cuboidal cells with clear to pale eosinophilic cytoplasm (A; case 8) and central ductal cells that showed variable tubular (B; case 7), micropapillary (C; case 8), and papillary (D; case 7) architecture. The peripheral cells showed a true myoepithelial phenotype with positivity for both p40 (E; case 7) and calponin (F; case 7). Despite largely low-grade histology, both of these cases also showed high grade areas including overt sarcomatoid transformation in one case (G; case 8).
Tumors with other molecular alterations
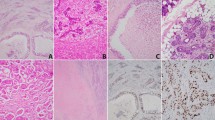
The remaining cases harbored various molecular findings in association with two discrete histologic patterns. Seven tumors displayed classic crowded tubules with a moderate amount of amphophilic to pale eosinophilic cytoplasm and round nuclei. This group included tumors with biallelic NF1 p.S1262fs and c.888 + 1 G > T mutations, TP53 p.K132M mutation, CUL3 p.V427fs mutation (Fig. 4A), RAF1 amplification (Fig. 4B), as well as 3 cases with no identifiable genetic alterations or which failed nucleic acid amplification (Fig. 4C). There were also 3 tumors that had an oncocytic appearance composed of back-to-back tubules and trabeculae of larger cells with abundant eosinophilic granular cytoplasm and prominent nucleoli. One of these tumors had homozygous CDKN2A loss and was composed of a monophasic population of confluent tubules (Fig. 4D). Interestingly, 2 of these tumors (Fig. 4E), both of which lacked identifiable mutations or fusions, had some degree of biphasic differentiation. Although not well-visualized on H&E sections, they harbored peripheral cells that were positive for p40 (Fig. 4F) and negative for other myoepithelial markers calponin and SMA as well as S100 and SOX10.
Classic tubulopapillary architecture with crowded, cytologically bland glands were seen in SNAC with CUL3 mutation (A; case 14), RAF1 amplification (B; case 15), and no detectable genetic alterations (C; case 16). Oncocytic change was also seen in SNAC with homozygous CDKN2A loss and a monophasic cell population (D; case 11) as well as cases with no detectable genetic alterations (E; case 9) and peripheral cells that expressed p40 (F; case 9).
Discussion
Low-grade non-intestinal-type SNAC is formally defined as a diagnosis of exclusion that encompasses any cytologically bland, gland-forming sinonasal tumor that lacks salivary or intestinal differentiation. In practice, however, almost all SNAC that fall into this category comprise a discrete group of tumors with recurrent tubulopapillary architecture that are increasingly regarded to be of seromucinous gland origin. Despite this well-defined histologic profile, the molecular underpinnings of low-grade SNAC are not well-understood. Recent reports of several different gene fusions and mutations in small numbers of cases have raised questions as to whether further molecular subtypes of SNAC should be defined. This study performed comprehensive RNA and DNA sequencing on a large series of low-grade non-intestinal-type SNAC to better understand their classification and pathogenesis.
First, our findings confirm previous reports of frequent seromucinous differentiation in low-grade SNAC, both at a histologic and immunohistochemical level. Despite some variable elements, all SNAC included in this series displayed histologic seromucinous features in the form of consistent tubulopapillary architecture, which traditionally has been regarded as analogous to the terminal tubules of the seromucinous glands6. While a subset of cases had oncocytic, mucinous, or even myoepithelial differentiation, these elements may actually recapitulate other cell types that are also normal constituents of seromucinous glands. Although comprehensive immunohistochemical analysis was not possible because we prioritized molecular testing in cases with limited tissue, 86% of SNAC that underwent staining for S100 protein, SOX10, and DOG1 expressed at least 1 of these putative seromucinous markers. These results are in keeping with previous reports of a recurrent seromucinous immunophenotype in low-grade SNAC7. Furthermore, 3 cases that showed seromucinous differentiation but harbored both low-grade and high-grade areas add to existing evidence that the seromucinous phenotype may transcend conventional grade-based grouping of non-intestinal-type SNAC7,19. These histological and immunohistochemical findings support previous proposals for reclassifying low-grade non-intestinal-type SNAC as seromucinous-type adenocarcinoma7,8. Paradoxically, despite their historical designation as “non-salivary,” SNAC with seromucinous features may even be conceptualized as specialized salivary tumors that exclusively arise in the sinonasal tract. However, it is essential to note that seromucinous differentiation is not specific for malignancy in the sinonasal tract, as it is also seen in seromucinous hamartomas and REAH. Both of these benign lesions not only can be difficult to distinguish from SNAC in limited samples, but also are occasionally seen adjacent to low-grade SNAC and may represent precursor lesions13,20. While one SNAC in this series was indeed associated with a REAH, all SNAC included here clearly displayed the crowded, back-to-back glands that define the presence of adenocarcinoma according to the 2017 WHO Classification of Head and Neck Tumours.
Our findings also highlight marked genetic heterogeneity in these low-grade SNAC, including a prominent role for diverse kinase fusions. The SNAC in this series harbored 11 different oncogenic drivers, only 2 of which (CTNNB1 and BRAF/AKT1 mutations) were found in more than 1 case. Notably, 4 separate kinase fusions were present, accounting for almost a third of tumors. Of course, ETV6::NTRK3, as well as ETV6::RET, were previously identified in up to 10% of low-grade SNAC10,11,12. However, the other 3 fusions seen in this cohort have never been reported in this tumor type. DNAJB1::PRKACA was originally described as the defining molecular alteration in fibrolamellar hepatocellular carcinoma and was subsequently identified in intraductal oncocytic papillary neoplasm of the pancreas21,22,23,24. FN1::NRG1 and PRKAR1A::MET appear to be entirely novel fusions, although both NRG1 and MET are rarely rearranged with various partners in multiple tumor types, most commonly lung adenocarcinoma25,26,27,28. In combination with previous reports of SYN2::PPARG and EGFR::ZNF267 fusions in single cases13,14, these findings suggest that many different kinase fusions may serve as drivers for a significant subset of SNAC. The majority of SNAC that lacked fusions showed alterations in other common oncogenic drivers including CTNNB1, BRAF, AKT1, TP53, CDKN2A, NF1, and RAF1. While CUL3 is a less well-established driver, it has been reported as a tumor suppressor gene in esophageal squamous cell carcinoma and papillary renal cell carcinoma, where loss of function mutations are thought to contribute to oncogenesis via dysregulation of transcription factor NRF229,30, As such, we regard the loss of function frameshift mutation seen in 1 SNAC as potentially oncogenic as well. Because several of these fusions, BRAF p.V600E mutations, and RAF1 amplification represent drug targets, comprehensive molecular analysis could point to treatment options in rare SNAC that demonstrate aggressive behavior. Interestingly, most of these alterations also activate cellular signaling cascades, including the MAPK pathway in 54% of cases, suggesting some degree of unified pathogenesis despite this genetic variability. Regardless, the degree of heterogeneity at the molecular level indicates that a strategy relying on any single alteration for diagnosis of SNAC is impractical.
Given this diverse genetic landscape, we also question whether any single genetic alteration is sufficient to delineate subtypes of low-grade SNAC within the unified histologic spectrum. Specific gene fusions and mutations are increasingly regarded as pathognomonic for sinonasal and salivary gland entities. And indeed, ETV6::NTRK3 and ETV6::RET fusions have been proposed to define a separate category of “ETV6-rearranged low-grade SNAC”10,11,12. However, we find it impossible to objectively distinguish SNAC with ETV6::NTRK3 fusions, either in this series or published literature, from most other cases at a histologic level. In a background of multiple molecular alterations, it is difficult to confirm any fusion-defined subgroup of SNAC as unique without a correspondingly distinctive histologic profile. SNAC with CTNNB1 mutations have also been suggested to represent a separate group16,17. These tumors are more identifiable histologically with consistent formation of squamoid morules- a feature that was recognized as unique before their molecular underpinnings were documented7 and is common to other tumor types with beta-catenin alterations31,32,33,34,35. But they also display crowded tubules identical to other seromucinous-type SNAC. Moreover, squamoid morules have also been reported in the SNAC with SYN2::PPARG fusion14, casting doubt on the specificity of this finding for CTNNB1 mutations. Undoubtedly, detailed clinicopathologic and molecular analysis of a still-larger group of SNAC will be necessary to better establish whether subgroups can and should be carved out of this category. At this point, we favor that low-grade SNAC are more analogous to soft tissue or thyroid tumors, in which multiple genetic alterations produce similar histologic findings36,37, than molecularly-defined sinonasal entities.
Nevertheless, these findings do highlight unique cases of SNAC with well-developed myoepithelial elements, prominent papillary and micropapillary architecture, and recurrent BRAF p.V600E and AKT1 p.E17K mutations that may represent a more distinctive category. In addition to the 2 cases in our series, 4 very similar low-grade SNAC with biphasic cell populations and papillary growth patterns were previously reported20,38, of which 2 were also found to have BRAF p.V600E mutations15, suggesting that these tumors are a recurrent phenomenon. Even in a relatively small cohort, these biphasic, BRAF-mutant SNAC have a sufficiently unique histologic and molecular profile to suggest they may represent a distinctive subgroup. The myoepithelial differentiation in these tumors is particularly exceptional among SNAC. Only 1 other SNAC in the literature7 and 2 cases with oncocytic features in this series showed peripheral expression of p63 or p40 alone- a finding that likely reflects basal differentiation typical in oncocytic tumors rather than true myoepithelial elements39. The high-grade features in both cases with BRAF p.V600E mutation in this cohort, including overt sarcomatoid transformation in one case, are also notable. Of course, these unique characteristics call into question whether these BRAF-mutant tumors are really SNAC at all. The biphasic cell populations in these tumors have repeatedly raised the possibility that they should be classified as salivary epithelial-myoepithelial carcinoma (EMC)20,38. However, EMC is vanishingly rare in the sinonasal tract, generally lacks papillary and micropapillary architecture, and does not harbor recurrent BRAF mutations40,41,42, arguing against this diagnosis. These tumors also bear some resemblance to salivary sialadenoma papilliferum and oncocytic variant of intraductal carcinoma, which share biphasic populations and BRAF p.V600E mutations43,44,45,46. But they lack the surface squamous papillations of sialadenoma papilliferum and demonstrate more architectural diversity than oncocytic variant of intraductal carcinoma. Without a comfortable fit into existing salivary tumor types, these biphasic, BRAF-mutant tumors are probably best regarded as a distinctive group within the SNAC spectrum until more comprehensive analysis can assess whether they represent a unique entity.
In summary, this study reinforces emerging consensus that low-grade non-intestinal-type SNAC is a largely unified histologic group and confirms previous reports of seromucinous differentiation in a majority of cases. Comprehensive molecular analysis also confirms that these low-grade SNAC are a molecularly heterogeneous category, with a particularly prominent role for diverse kinase fusions, although many of these variable alterations may lead to similar downstream signaling amplification including MAPK pathway activation. Tumors with most fusions or mutations did not display sufficiently distinctive histologic features to support separate classification in this relatively limited cohort. However, SNAC that have recurrent BRAF p.V600E mutations, true myoepithelial elements, and prominent papillary and micropapillary architecture may represent a more unique category. Further clinical, histologic, immunohistochemical, and molecular evaluation of larger groups of SNAC will allow for better understanding of its pathogenesis, diagnosis, and potential subgroups in the future.
Data availability
Most data generated or analyzed during this study are included in this published article. Additional data is available from the corresponding author on reasonable request.
References
Stelow, E. B., Brandwein-Gensler, M., Franchi, A., Nicolai, P. & Wenig, B. M. in WHO Classification of Head and Neck Tumours (eds El-Naggar, N. et al.) 24–26 (International Agency for Research on Cancer, 2017).
Stelow, E. B., Laco, J., Leivo, I. & Skalova, A. in WHO Classification of Head and Neck Tumours (ed. WHO Classification of Tumours Editorial Board) (International Agency for Research on Cancer, 2022).
Gnepp, D. R. & Heffner, D. K. Mucosal origin of sinonasal tract adenomatous neoplasms. Mod. Pathol. 2, 365–371 (1989).
Manning, J. T. & Batsakis, J. G. Salivary-type neoplasms of the sinonasal tract. Ann. Otol. Rhinol. Laryngol. 100, 691–694 (1991).
Skalova, A. et al. Sinonasal tubulopapillary low-grade adenocarcinoma. Histopathological, immunohistochemical and ultrastructural features of poorly recognised entity. Virchows Arch. 443, 152–158 (2003).
Kleinsasser, O. Terminal tubulus adenocarcinoma of the nasal seromucous glands. A specific entity. Arch. Otorhinolaryngol. 241, 183–193 (1985).
Purgina, B., Bastaki, J. M., Duvvuri, U. & Seethala, R. R. A subset of sinonasal non-intestinal type adenocarcinomas are truly seromucinous adenocarcinomas: a morphologic and immunophenotypic assessment and description of a novel pitfall. Head Neck Pathol. 9, 436–446 (2015).
Yom, S. S. et al. Genetic analysis of sinonasal adenocarcinoma phenotypes: distinct alterations of histogenetic significance. Mod. Pathol. 18, 315–319 (2005).
Ozolek, J. A., Barnes, E. L. & Hunt, J. L. Basal/myoepithelial cells in chronic sinusitis, respiratory epithelial adenomatoid hamartoma, inverted papilloma, and intestinal-type and nonintestinal-type sinonasal adenocarcinoma: an immunohistochemical study. Arch. Pathol. Lab. Med. 131, 530–537 (2007).
Andreasen, S., Kiss, K., Melchior, L. C. & Laco, J. The ETV6-RET gene fusion is found in ETV6-rearranged low-grade sinonasal adenocarcinoma without NTRK3 involvement. Am. J. Surg. Pathol. 42, 985–988 (2018).
Andreasen, S. et al. ETV6 gene rearrangements characterize a morphologically distinct subset of sinonasal low-grade non-intestinal-type adenocarcinoma: a novel translocation-associated carcinoma restricted to the sinonasal tract. Am. J. Surg. Pathol. 41, 1552–1560 (2017).
Zhai, C. W., Yuan, C. C. & Wang, S. Y. [ETV6-rearranged low-grade sinonasal non-intestinal-type adenocarcinoma: a clinicopathological analysis]. Zhonghua Bing Li Xue Za Zhi 50, 55–59 (2021).
Baneckova, M. et al. Immunohistochemical and genetic analysis of respiratory epithelial adenomatoid hamartomas and seromucinous hamartomas: are they precursor lesions to sinonasal low-grade tubulopapillary adenocarcinomas? Hum. Pathol. 97, 94–102 (2020).
Soon, G. S. T., Chang, K. T. E., Kuick, C. H. & Petersson, F. A case of nasal low-grade non-intestinal-type adenocarcinoma with aberrant CDX2 expression and a novel SYN2-PPARG gene fusion in a 13-year-old girl. Virchows Arch. 474, 619–623 (2019).
Franchi, A. et al. Low prevalence of K-RAS, EGF-R and BRAF mutations in sinonasal adenocarcinomas. Implications for anti-EGFR treatments. Pathol. Oncol. Res. 20, 571–579 (2014).
Villatoro, T. M. & Mardekian, S. K. Two cases of sinonasal non-intestinal-type adenocarcinoma with squamoid morules expressing nuclear beta-Catenin and CDX2: a curious morphologic finding supported by molecular analysis. Case Rep. Pathol. 2018, 8741017 (2018).
Stelow, E. B. & Mills, S. E. Low-Grade Tubulo-Acinar Sinonasal Adenocarcinoma: a specific entity with recurrent beta-catenin mutations. Mod. Pathol. 33, 1230 (2020).
Bishop, J. A. et al. Sclerosing Polycystic “Adenosis” of Salivary Glands: A Neoplasm characterized by PI3K pathway alterations more correctly named Sclerosing Polycystic Adenoma. Head Neck Pathol. 14, 630–636 (2020).
Stelow, E. B., Jo, V. Y., Mills, S. E. & Carlson, D. L. A histologic and immunohistochemical study describing the diversity of tumors classified as sinonasal high-grade nonintestinal adenocarcinomas. Am. J. Surg. Pathol. 35, 971–980 (2011).
Jo, V. Y., Mills, S. E., Cathro, H. P., Carlson, D. L. & Stelow, E. B. Low-grade sinonasal adenocarcinomas: the association with and distinction from respiratory epithelial adenomatoid hamartomas and other glandular lesions. Am. J. Surg. Pathol. 33, 401–408 (2009).
Graham, R. P. et al. DNAJB1-PRKACA is specific for fibrolamellar carcinoma. Mod. Pathol. 28, 822–829 (2015).
Honeyman, J. N. et al. Detection of a recurrent DNAJB1-PRKACA chimeric transcript in fibrolamellar hepatocellular carcinoma. Science 343, 1010–1014 (2014).
Singhi, A. D. et al. Recurrent rearrangements in PRKACA and PRKACB in Intraductal Oncocytic Papillary Neoplasms of the Pancreas and Bile Duct. Gastroenterology 158, 573–582 e572 (2020).
Vyas, M. et al. DNAJB1-PRKACA fusions occur in oncocytic pancreatic and biliary neoplasms and are not specific for fibrolamellar hepatocellular carcinoma. Mod. Pathol. 33, 648–656 (2020).
Jonna, S. et al. Detection of NRG1 gene fusions in solid tumors. Clin. Cancer Res. 25, 4966–4972 (2019).
Lee, M. et al. MET alterations and their impact on the future of non-small cell lung cancer (NSCLC) targeted therapies. Expert Opin. Ther. Targets 25, 249–268 (2021).
Liu, S. V. NRG1 fusions: Biology to therapy. Lung Cancer-J. Iaslc. 158, 25–28 (2021).
Stransky, N., Cerami, E., Schalm, S., Kim, J. L. & Lengauer, C. The landscape of kinase fusions in cancer. Nat. Commun. 5, 4846 (2014).
Lin, D. C. et al. Identification of distinct mutational patterns and new driver genes in oesophageal squamous cell carcinomas and adenocarcinomas. Gut 67, 1769–1779 (2018).
Ooi, A. et al. CUL3 and NRF2 mutations confer an NRF2 activation phenotype in a sporadic form of papillary renal cell carcinoma. Cancer Res. 73, 2044–2051 (2013).
He, C. et al. Pyloric Gland Adenoma (PGA) of the Gallbladder: a unique and Distinct Tumor from PGAs of the Stomach, Duodenum, and Pancreas. Am. J. Surg. Pathol. 42, 1237–1245 (2018).
Nakatani, Y. et al. Aberrant nuclear localization and gene mutation of beta-catenin in low-grade adenocarcinoma of fetal lung type: up-regulation of the Wnt signaling pathway may be a common denominator for the development of tumors that form morules. Mod. Pathol. 15, 617–624 (2002).
Saegusa, M. & Okayasu, I. Frequent nuclear beta-catenin accumulation and associated mutations in endometrioid-type endometrial and ovarian carcinomas with squamous differentiation. J. Pathol. 194, 59–67 (2001).
Tanaka, Y. et al. Significance of aberrant (cytoplasmic/nuclear) expression of beta-catenin in pancreatoblastoma. J. Pathol. 199, 185–190 (2003).
Xu, B. et al. Cribriform-morular variant of papillary thyroid carcinoma: a pathological and molecular genetic study with evidence of frequent somatic mutations in exon 3 of the beta-catenin gene. J. Pathol. 199, 58–67 (2003).
Chu, Y. H. et al. Clinicopathologic features of kinase fusion-related thyroid carcinomas: an integrative analysis with molecular characterization. Mod. Pathol. 33, 2458–2472 (2020).
Kao, Y. C. et al. Soft tissue tumors characterized by a wide spectrum of kinase fusions share a lipofibromatosis-like neural tumor pattern. Genes Chromosomes Cancer 59, 575–583 (2020).
Franchi, A. et al. Low-grade salivary type tubulo-papillary adenocarcinoma of the sinonasal tract. Histopathology 48, 881–884 (2006).
McHugh, J. B. et al. p63 immunohistochemistry differentiates salivary gland oncocytoma and oncocytic carcinoma from metastatic renal cell carcinoma. Head Neck Pathol. 1, 123–131 (2007).
Chiosea, S. I., Miller, M. & Seethala, R. R. HRAS mutations in epithelial-myoepithelial carcinoma. Head Neck Pathol. 8, 146–150 (2014).
El Hallani, S. et al. Epithelial-Myoepithelial Carcinoma: Frequent Morphologic and Molecular Evidence of Preexisting Pleomorphic Adenoma, Common HRAS Mutations in PLAG1-intact and HMGA2-intact Cases, and Occasional TP53, FBXW7, and SMARCB1 Alterations in High-grade Cases. Am. J. Surg. Pathol. 42, 18–27 (2018).
Urano, M. et al. Diagnostic significance of HRAS Mutations in Epithelial-Myoepithelial Carcinomas Exhibiting a Broad Histopathologic Spectrum. Am. J. Surg. Pathol. 43, 984–994 (2019).
Bishop, J. A. et al. Oncocytic intraductal carcinoma of salivary glands: a distinct variant with TRIM33-RET fusions and BRAF V600E mutations. Histopathology 79, 338–346 (2021).
Chen, S. et al. Sialadenoma papilliferum: clinicopathologic, Immunohistochemical, molecular analyses of new five cases and review of the literature. Diagn. Pathol. 16, 22 (2021).
Hsieh, M. S. et al. Salivary Sialadenoma Papilliferum Consists of Two Morphologically, Immunophenotypically, and Genetically Distinct Subtypes. Head Neck Pathol. 14, 489–496 (2020).
Nakaguro, M. et al. Histopathological evaluation of minor salivary gland papillary-cystic tumours: focus on genetic alterations in sialadenoma papilliferum and intraductal papillary mucinous neoplasm. Histopathology 76, 411–422 (2020).
Funding
This study was funded in part by the Jane B. and Edwin P. Jenevein, MD Endowment for Pathology at UT Southwestern Medical Center.
Author information
Authors and Affiliations
Contributions
L.M.R. and J.A.B. designed the study and prepared the manuscript. L.M.R., J.A.B., L.D.R.T., J.S.G.H., N.R.L., M.W.M., and T.M.S. contributed tumor samples and data analysis. J.G. analyzed sequencing data and interpreted the results. All authors read and approved the final paper.
Corresponding author
Ethics declarations
Competing interests
N.R.L. receives funding from Merck and holds stock in Navigen Pharmaceuticals, neither of which are related to the present study. The other authors have no competing interests to declare.
Ethics approval and consent to participate
This study was approved by the Institutional Review Board at UT Southwestern Medical Center.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Rooper, L.M., Thompson, L.D.R., Gagan, J. et al. Low-grade non-intestinal-type sinonasal adenocarcinoma: a histologically distinctive but molecularly heterogeneous entity. Mod Pathol 35, 1160–1167 (2022). https://doi.org/10.1038/s41379-022-01068-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41379-022-01068-w
This article is cited by
-
ETV6::NTRK3-associated papillary adenocarcinoma: let us play it by ear
Virchows Archiv (2024)
-
High-grade non-intestinal type sinonasal adenocarcinoma with ETV6::NTRK3 fusion, distinct from secretory carcinoma by immunoprofile and morphology
Virchows Archiv (2023)
-
NKX3.1 Expression and Molecular Characterization of Secretory Myoepithelial Carcinoma (SMCA): Advancing the Case for a Salivary Mucous Acinar Phenotype
Head and Neck Pathology (2023)
-
Nasal and sinonasal tumors formed by atypical adenomatous lesions arising in respiratory epithelial adenomatoid hamartoma/seromucinous hamartoma
Virchows Archiv (2023)
-
Investigation of -PRKACA/-PRKACB fusion genes in oncocytic tumors of the pancreatobiliary and other systems
Virchows Archiv (2022)