Abstract
Pancreatic neoplasms are heterogenous and have traditionally been classified by assessing their lines of cellular differentiation using histopathologic methods, particularly morphologic and immunohistochemical evaluation. These methods frequently identify overlapping differentiation along ductal, acinar, and neuroendocrine lines, raising diagnostic challenges as well as questions regarding the relationship of these neoplasms. Neoplasms with acinar differentiation, in particular, frequently show more than one line of differentiation based on immunolabeling. Genome methylation signatures, in contrast, are better conserved within cellular lineages, and are increasingly used to support the classification of neoplasms. We characterized the epigenetic relationships between pancreatoblastomas, acinar cell carcinomas (including mixed variants), pancreatic neuroendocrine tumors, solid pseudopapillary neoplasms, and pancreatic ductal adenocarcinomas using a genome-wide array platform. Using unsupervised learning approaches, pancreatic neuroendocrine tumors, solid pseudopapillary neoplasms, ductal adenocarcinomas, and normal pancreatic tissue samples all localized to distinct clusters based on their methylation profiles, whereas all neoplasms with acinar differentiation occupied a broad overlapping region located between the predominantly acinar normal pancreatic tissue and ductal adenocarcinoma clusters. Our data provide evidence to suggest that acinar cell carcinomas and pancreatoblastomas are similar at the epigenetic level. These findings are consistent with genomic and clinical observations that mixed acinar neoplasms are closely related to pure acinar cell carcinomas rather than to neuroendocrine tumors or ductal adenocarcinomas.
Similar content being viewed by others
Introduction
Pancreatic neoplasms including pancreatic ductal adenocarcinoma (PDAC), pancreatic neuroendocrine tumor (PanNET), pancreatoblastoma (PB), solid pseudopapillary neoplasm (SPN), acinar cell carcinoma (ACC) and mixed acinar carcinomas, comprise a heterogenous group of tumors with varied clinical behavior, ranging from indolent to highly aggressive disease course. The neoplasms recapitulate lines of differentiation seen in normal pancreatic parenchyma (ductal, acinar, neuroendocrine), and determination of the line of differentiation is a major basis of their classification (Fig. 1). Assessment of differentiation is based on a combination of histopathologic features, including morphology and immunohistochemistry (IHC). IHC has traditionally played an important role in defining the lines of differentiation, using protein markers specific for acinar (trypsin, chymostrypsin, lipase, BCL10), neuroendocrine (synaptophysin, chromogranin A), and ductal (glycoproteins) phenotypes1. SPNs are an exception, showing no definitive lineage differentiation based on morphologic or IHC markers that correspond to a non-neoplastic pancreatic cell type2,3.
A Pancreatic ductal adenocarcinoma showing glandular structures. B Pancreatic neuroendocrine tumor showing neuroendocrine features. C Acinar cell carcinoma showing acinar-like structures. D Solid pseudopapillary neoplasm showing loosely cohesive cells imparting a pseudopapillary appearance. E, F Pancreatoblastoma showing several areas of highly cellular tumor with solid arrangement, acinar-like structures and squamoid corpuscles (F).
Pancreatoblastomas and acinar cell carcinomas are regarded to be related neoplasms that share many histologic features (dense cellularity, solid and acinar patterns, monotonous nuclei with prominent nucleoli) and demonstrate acinar differentiation by immunohistochemistry, often with neuroendocrine differentiation as well. Accordingly, the distinction of acinar cell carcinoma and pancreatoblastoma can be challenging in some cases, especially on the basis of biopsies that may not sample the characteristic squamoid nests of pancreatoblastomas. Detailed guidance regarding this differential diagnosis is presented elsewhere1,4,5.
More recently, molecular characterization of the neoplasms has buttressed traditional classification schemes and enabled identification of subcategories as well as opportunities for targeted therapeutic interventions. Molecularly, PDAC is driven by mutations in TP53, CDKN2A, SMAD4, and the RAS-MAPK pathway, most commonly KRAS, and rarely by gene fusions6,7,8. ACCs, PBs, and SPNs are all associated with WNT pathway alterations, although to different degrees. SPNs are almost universally associated with mutations in CTNNB1 and WNT pathway inactivation3,9. PBs harbor CTNBB1 mutations in approximately 90% of cases and APC mutations in the remaining cases, along with near universal loss of heterozygosity (LOH) of chromosome arm 11p10. ACCs show a more varied mutational landscape with subgroups being driven by gene fusions in the MAPK pathway (BRAF, RAF1, and RET), alterations in DNA repair pathway genes (BRCA1/2, ATM, mismatch repair genes), and less frequently mutations affecting CTNNB1 or APC11,12,13. PanNETs are associated with MEN1 mutations, alterations in PI3K/Akt/mTOR pathway genes (e.g., PIK3CA, PTEN, TSC1, TSC2, VHL), and telomere maintenance genes (DAXX, ATRX)14,15,16,17.
Although pancreatic neoplasms may show unequivocal lines of differentiation, they often have overlapping and mixed features at the histopathologic, immunohistochemical, and molecular level, complicating diagnostic workup and raising questions about their origins and pathogenesis. For example, although SPNs share molecular alterations of the WNT pathway with ACCs and PBs, they show no evidence of acinar differentiation, or differentiation of any other known lineage. Acinar neoplasms, on the other hand, may show evidence of multiple lines of differentiation by IHC, leading to their designation as mixed carcinomas when they contain additional neuroendocrine or ductal components that constitute greater than 30% of the neoplasm. The second component is most commonly neuroendocrine (mixed acinar neuroendocrine carcinoma), rarely ductal (mixed acinar ductal carcinoma) or, even rarer, both ductal and neuroendocrine components (mixed acinar neuroendocrine ductal carcinoma)1. This has important therapeutic implications given the distinct clinical management of ACCs and PanNETs, and the relationship of the neuroendocrine component of mixed carcinomas to PanNETs is unclear. Studies characterizing the molecular and clinical features of mixed acinar neuroendocrine carcinomas and PanNETs have generally concluded that mixed acinar neuroendocrine carcinomas share clinical and molecular features of ACCs rather than PanNETs and should be regarded as ACCs for clinical management5,11,18.
DNA methylation is an important cellular process involved in gene regulation during development and neoplastic transformation. A given cancer cell’s methylation state is shaped by normal developmental processes of tissue differentiation as well as oncogenic transformation19. In this way, the methylome serves as a ‘fingerprint’ for the cell’s epigenetic history and lineage. Genome-wide methylation profiling on large tissue cohorts has leveraged this dynamic for diagnostic purposes in tissue-of-origin identification, tumor classification, and cancer screening across cancer types20,21,22,23,24,25. Previous studies have demonstrated distinct genome-wide methylation signatures of PDACs, PanNETs, and ACCs and global hypomethylation of PB genomes10,26. Building on these prior studies, here we sought to explore the epigenetic relationships between a broader group of acinar neoplasms, normal pancreas, and other solid tumors of the pancreas.
Materials and methods
Cases
We selected 99 cases comprising 9 normal pancreas tissue (NPT) samples, 38 PanNETs (12 WHO grade 1; 15 grade 2; 2 grade 3), 11 PDACs, 12 SPNs, 13 ACCs (7 pure; 3 mixed acinar neuroendocrine carcinomas; 2 mixed acinar ductal carcinomas; 1 mixed acinar neuroendocrine ductal carcinoma), and 16 PBs. All samples were obtained from the pathology archives of Memorial Sloan Kettering Cancer Center (MSK) apart from seven PBs that were obtained from St. Jude Children’s Research Hospital.
All cases were subjected to central pathology review to confirm the initial diagnosis. Regarding the diagnosis of acinar neoplasms, the cases in this study were classified as pancreatoblastomas if they arose in children and demonstrated at least some squamoid nests, which were usually easily identified and well formed, although in this age group, more poorly circumscribed nests or larger cells were accepted to represent squamoid nests. In adults, only cases with numerous, well formed squamoid nests were accepted as pancreatoblastomas. All other cases were regarded as acinar cell carcinomas or mixed acinar carcinomas based on the extent of immunolabeling for acinar and neuroendocrine markers. Intracellular or abundant stromal mucin was regarded as evidence of ductal differentiation in mixed acinar ductal carcinomas.
Methylation profiling
For methylation analysis, genomic DNA was extracted from representative formalin-fixed paraffin-embedded tissue sections for each of the 99 samples. Next, 250 ng of genomic DNA was subjected to bisulfite conversion and processed on the Illumina (San Diego, CA) methylationEPIC/850k platform according to manufacturer’s instructions. Data analysis was performed using R version 3.5.2. Sample loading and probe filtering was done using the ChAMP R package version 2.12.4 and normalization was done using the preprocessIllumina() function in the R minfi package version 1.28.327,28. Methylation level was measuring using beta values29. Sample and data quality were assessed by internal controls and the ChAMP pipeline, which filters out failed probes, XY CpG probes, cross-reactive probes, multi-hit probes, and SNP-related probes. Among the internal controls are a set of negative control probes designed in such a way that they do not hybridize to any target DNA and therefore are used to quantify background noise. The controls enable the calculation of a p value metric for each probe that quantifies the degree to which the probe’s signal differs from that of background noise (i.e., it is the probability of observing the probe’s signal under the assumption that it generated by noise). A probe was defined as a failure if the p value was greater than 0.001. If greater than 10% of probes in a sample failed (the default ChAMP parameter), then the sample was deemed a failure and excluded from downstream analysis.
Data analysis
Three unsupervised approaches were employed, T-distributed stochastic neighborhood embedding (t-SNE) clustering, Uniform Manifold Approximation and Projection (UMAP) clustering, and hierarchical clustering. Dimensionality reduction was performed using two methods. First, t-SNE using the Rtsne package version 0.15 on the top 10,000 most variable CpGs by variance with the following non-default parameters: perplexity = 10, max_iter = 5000, and theta = 030. Second, UMAP was performed using the umap package version 0.2.5.0 with default parameters31. Unsupervised hierarchical clustering and heatmap generation were obtained using the pheatmap R package version 1.0.12 by Ward’s linkage and Euclidean distance.
Differentially methylated probes (DMPs) vis-à-vis NPT were identified using ChAMP via the champ.DMP function with default parameters. The methylationEPIC platform is not a reliable quantitative assay. Although the beta value is the ratio of methylated signal to total signal, a beta value of, for example, 40% at a particular CpG site does not equate to 40% of the DNA in the sample being methylated at that site. Beta values are influenced by a number of factors including experimental variation, true biological methylation state, probe characteristics, sample type (frozen, FFPE, etc.), and tissue heterogeneity32,33. Because of these sources of variation, false positives may be detected when standard statistical methods used for differential methylation analyses identify differences in beta values between groups that are statistically significant, but don’t account for variation on the platform or tumor purity. For example, although the mean beta values of 0.70 and 0.90 for a CpG site between two groups may be statistically significant, that difference is likely not biologically meaningful because they both indicate methylated status and the difference may be attributed to experimental variation and/or tissue heterogeneity.
To further scrutinize the results, an independent cutoff-based method was utilized to quantify CpG sites with biologically meaningful differences between tumors and normal tissue. This method excluded statistically significant CpG sites if the mean beta values between groups did not span a biologically meaningful cutoff. Enumeration was performed across varying cutoff levels to visualize the difference across biologically relevant cutoffs and a precise enumeration of DMPs was performed using the cutoff of 0.3, a well-recognized boundary between methylated and unmethylated status, to determine global methylation differences32,33.
Results
Epigenetic relationship
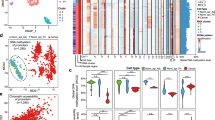
Three PBs were excluded from downstream analysis due to poor quality control metrics. These three samples demonstrated a high number of probe failure (17%, 31%, and 50%), likely due to the old age of the samples (greater than 30 years old). After probe filtering, a total of 649,896 high-quality CpG sites remained for analysis across the 96 samples. The three different unsupervised clustering approaches showed similar findings. Both dimensionality reduction methods (Fig. 2A, B) revealed that PanNETs, SPNs, PDACs and NPT localized to distinct well-delineated clusters based on their methylation profiles while the acinar neoplasms (ACCs and PBs) occupied a broad overlapping region located between the NPT and PDAC clusters. One SPN localized to the PDAC region by UMAP clustering. Re-review of this case showed relatively low tumor content and a high amount of fibrotic tissue, which may explain the epigenetic finding. Four PBs and one ACC localized closer to the PDAC cluster by both t-SNE and UMAP. Review of these cases showed that they all had treatment-related effects characterized by foamy histiocytes, hemorrhage, necrosis, and fibrosis while the other PBs did not. No mixed tumors localized to either the PDAC or PanNET clusters, despite having immunohistochemical evidence of ductal or neuroendocrine differentiation.
Unsupervised hierarchical clustering (Fig. 2C) demonstrated similar clade segregation.
Differential epigenetic profiles
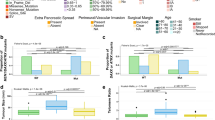
Next, we sought to explore the degree of global hyper- and hypo-methylation that occurred across tumor types relative to NPT. Differentially methylated probe analysis initially detected 233,787 (PB), 191,183 (ACC), 206,947 (PDAC), 278,666 (SPN), and 176,293 (PanNet) differentially methylated CpG probes (DMPs), the majority of which were false positives and were not biologically meaningful. Cutoff-based filtering (Fig. 3A) and scrutinization of beta value cutoffs that were more biologically meaningful (performing counts at a cutoff of beta = 0.3) showed that PDACs (35,968) and SPNs (34,042) demonstrated the highest number of DMPs with respect to NPT; ACCs (16,113) demonstrated the lowest number, and PanNETs (21,561) and PBs (21,126) showed a similar and intermediate number, respectively. The analysis also revealed the degree of hyper- and hypo-methylation of the tumors with respect to NPT (Fig. 3B). At the same cutoff, PDACs exhibited the highest degree of hypermethylation relative to NPT (75.2% of DMPs were hypermethylated with respect to NPT), followed by PanNETs (60.4%), SPNs (51.8%), ACCs (48.1%), and PBs (35.9%).
A Number of differential probes between NPT and pancreatic tumors. Counts are defined as number of sites that are statistically significant and span the cutoffs on the x-axis. Biologically meaningful cutoffs are generally considered around 0.15–0.3. SPN and PDAC showed the greatest difference, following by PanNET and PB, followed by ACC with the least. B Number of differential probes spanning cutoffs split by hypomethylated state or hypermethylated state relative to NPT. PDAC show the highest degree of hypermethylation while others have similar changes.
Discussion
Reassuringly, the epigenetic signatures of the pancreatic neoplasms in this study recapitulated established diagnostic classifications that divide them into acinar, ductal, and neuroendocrine groups; indeed, it also illustrated that the still elusive SPNs are epigenetically distinct from all other pancreatic neoplasms. By three unsupervised learning methods, acinar neoplasms (PBs and ACCs) demonstrated similar epigenetic signatures, a finding consistent with morphologic characteristics but at odds with some immunohistochemical features and the varied molecular drivers reported in these neoplasms. Not surprisingly, acinar neoplasms clustered near the predominantly acinar NPT. Interestingly, one SPN sample with low tumor purity clustered with the PDAC group, suggesting that UMAP clustering may be more sensitive to sample quality issues than t-SNE. Of clinical importance, all but one mixed acinar neoplasm (which had close proximity to the PDAC cluster) shared a clear epigenetic signature with the pure ACCs, implying that mixed acinar neoplasms are epigenetically acinar, rather than neuroendocrine or ductal. This is in line with prior findings and adds epigenetic evidence to the established molecular and clinicopathologic similarities between mixed and pure ACCs9. Importantly, mixed acinar neuroendocrine carcinomas clustered separately from PanNETs, reinforcing that the presence of neuroendocrine differentiation at the immunohistochemical level in a mixed acinar neuroendocrine carcinoma does not equate a biological similarity with PanNETs. The consistent presence of acinar differentiation in PBs based on IHC is also reflected in a close relationship of these entities based on the methylation profiles. Tumors with low purity or treatment effect localized closer to the PDAC region, highlighting the importance of high tumor purity in methylation-based analyses. Grade 1 and 2 PanNETs all clustered together, consisting with these well differentiated neoplasms being closely related. Indeed, grade 3 PanNETs also clustered in the same region, although sample size was limited to make definitive conclusions.
This study had several limitations. First, on account of their rarity, an in-depth analysis of the mixed acinar cell tumors was not possible due to sample size limitations. Second, unlike other more expensive and labor-intensive platforms, the array-based platform is not a quantitative assay and is generally better at qualitative analysis. Due to this the cutoff-based method was used to further scrutinize results from standard statistical tests that result in high false positives.
Although the diagnostic workup is generally straightforward, the distinct demarcation of the tumors by epigenetic signature could conceivably enable an ancillary diagnostic classifier as has been done with other tumor classes19,20. The study also provides a window into the epigenetic changes associated with oncogenic transformation. PDAC and PanNETs tend to undergo global hypermethylation, PBs hypomethylation, and ACCs and SPNs possess approximately equal degree of hyper and hypo-methylation changes. This study sheds light on the origin and differentiation of these neoplasms and demonstrates the utility of high-throughput epigenetic platforms in refining tumor classification.
Data availability
Methylation data used in this study are available from the corresponding author on reasonable request.
References
WHO Classification of Tumors Editorial Board. Digestive system tumours, WHO classification of tumours series, 5th edn, Vol. 1, (International Agency for Research of Cancer, Lyon (France), 2019).
Calvani, J. et al. Solid pseudopapillary neoplasms of the pancreas do not express major pancreatic markers in pediatric patients. Hum. Pathol. 83, 29–35 (2019).
La Rosa, S. & Bongiovanni, M. Pancreatic solid pseudopapillary neoplasm: key pathologic and genetic features. Arch. Pathol. Lab. Med. 144, 829–837 (2020).
Hruban RH, Pitman MB, Klimstra DS. Tumors of the pancreas, AFIP atlas of tumor pathology series 4. (American Registry of Pathology, Washington DC, USA, 2007).
Wood, L. D. & Klimstra, D. S. Pathology and genetics of pancreatic neoplasms with acinar differentiation. Semin Diagn Pathol. 31, 491–497 (2014).
Cancer Genome Atlas Research Network. Integrated genomic characterization of pancreatic ductal adenocarcinoma. Cancer Cell 32, 185–203 e113 (2017).
Cocco, E., Scaltriti, M. & Drilon, A. NTRK fusion-positive cancers and TRK inhibitor therapy. Nat. Rev. Clin. Oncol. 15, 731–747 (2018).
O’Reilly, E. M. & Hechtman, J. F. Tumour response to TRK inhibition in a patient with pancreatic adenocarcinoma harbouring an NTRK gene fusion. Ann. Oncol. 30, viii36–viii40 (2019).
Selenica, P. et al. Solid pseudopapillary neoplasms of the pancreas are dependent on the Wnt pathway. Mol. Oncol. 13, 1684–1692 (2019).
Isobe, T. et al. Integrated molecular characterization of the lethal pediatric cancer pancreatoblastoma. Cancer Res. 78, 865–876 (2018).
Chmielecki, J. et al. Comprehensive genomic profiling of pancreatic acinar cell carcinomas identifies recurrent RAF fusions and frequent inactivation of DNA repair genes. Cancer Disco 4, 1398–1405 (2014).
Chou, A. et al. RET gene rearrangements occur in a subset of pancreatic acinar cell carcinomas. Mod. Pathol. 33, 657–664 (2020).
Prall, O. W. J. et al. RAF1 rearrangements are common in pancreatic acinar cell carcinomas. Mod. Pathol. 33, 1811–1821 (2020).
Shi, C. & Klimstra, D. S. Pancreatic neuroendocrine tumors: pathologic and molecular characteristics. Semin. Diagn. Pathol. 31, 498–511 (2014).
Asa, S. L. et al. Molecular pathology of well-differentiated gastro-entero-pancreatic neuroendocrine tumors. Endocr. Pathol. 32, 169–191 (2021).
Mafficini, A. & Scarpa, A. Genomic landscape of pancreatic neuroendocrine tumours: the International Cancer Genome Consortium. J. Endocrinol. 236, R161–R167 (2018).
Scarpa, A. et al. Whole-genome landscape of pancreatic neuroendocrine tumours. Nature 543, 65–71 (2017).
Stelow, E. B., Shaco-Levy, R., Bao, F., Garcia, J. & Klimstra, D. S. Pancreatic acinar cell carcinomas with prominent ductal differentiation: mixed acinar ductal carcinoma and mixed acinar endocrine ductal carcinoma. Am. J. Surg. Pathol. 34, 510–518 (2010).
Bergman, Y. & Cedar, H. DNA methylation dynamics in health and disease. Nat. Struct. Mol. Biol. 20, 274–281 (2013).
Moran, S. et al. Epigenetic profiling to classify cancer of unknown primary: a multicentre, retrospective analysis. Lancet Oncol. 17, 1386–1395 (2016).
Capper, D. et al. DNA methylation-based classification of central nervous system tumours. Nature 555, 469–474 (2018).
Liu, M. C. et al. Sensitive and specific multi-cancer detection and localization using methylation signatures in cell-free DNA. Ann. Oncol. 31, 745–759 (2020).
Figueroa, M. E. et al. DNA methylation signatures identify biologically distinct subtypes in acute myeloid leukemia. Cancer Cell 17, 13–27 (2010).
Wu SP, et al. DNA methylation-based classifier for accurate molecular diagnosis of bone sarcomas. JCO Precis. Oncol. 2017, PO.17.00031 (2017).
Glass, J. L. et al. Epigenetic identity in AML depends on disruption of nonpromoter regulatory elements and is affected by antagonistic effects of mutations in epigenetic modifiers. Cancer Disco 7, 868–883 (2017).
Jäkel, C. et al. Genome-wide genetic and epigenetic analyses of pancreatic acinar cell carcinomas reveal aberrations in genome stability. Nat. Commun. 8, 1323 (2017).
Tian, Y. et al. ChAMP: updated methylation analysis pipeline for Illumina BeadChips. Bioinformatics 33, 3982–3984 (2017).
Aryee, M. J. et al. Minfi: a flexible and comprehensive Bioconductor package for the analysis of Infinium DNA methylation microarrays. Bioinformatics 30, 1363–1369 (2014).
Du, P. et al. Comparison of Beta-value and M-value methods for quantifying methylation levels by microarray analysis. BMC Bioinform. 11, 587 (2010).
van der Maaten, L. J. P. Accelerating t-SNE using tree-based algorithms. J. Mach. Learn Res. 15, 3221–3245 (2014).
McInnes L, Healy J, Melville J. Uniform manifold approximation and projection for dimension reduction. Preprint at https://arxiv.org/abs/1802.03426 (2018).
Warden, C. D. et al. COHCAP: an integrative genomic pipeline for single-nucleotide resolution DNA methylation analysis. Nucleic Acids Res. 41, e117 (2013).
Benhamida, J. K. et al. Reliable clinical MLH1 promoter hypermethylation assessment using a high-throughput genome-wide methylation array platform. J. Mol. Diagn. 22, 368–375 (2020).
Funding
Support for this study was provided by the Melamed Family Foundation. In addition, this study was partially funded by the NCI under the MSK Cancer Center Support Grant (P30 CA008748). J.S.R.-F. and B.W. are funded in part by the Breast Cancer Research Foundation and National Institutes of Health/ National Cancer Institute P50 CA247749 01 grant.
Author information
Authors and Affiliations
Contributions
D.S.K. performed study concept, design, and histopathologic review; J.K.B. performed study design, experiment planning, writing, data analysis, and revision of the paper; M.V., A.T., L.W., A.B., P.S., M.H.A.R., M.S. performed sample selection and acquisition; L.V. performed sample analyses; B.W., J.S.R.-F. provided technical support and interpretation of data; T.E.L., D.A.M. performed manuscript writing, review; K.O. performed histopathologic review. All authors read and approved the final paper.
Corresponding author
Ethics declarations
Competing interests
J.S.R.-F. reports receiving personal/consultancy fees from Goldman Sachs, REPARE Therapeutics and Paige.AI, membership of the scientific advisory boards of VolitionRx, REPARE Therapeutics and Paige.AI, membership of the Board of Directors of Grupo Oncoclinicas, and ad hoc membership of the scientific advisory boards of Roche Tissue Diagnostics, Ventana Medical Systems, Novartis, Genentech and InVicro, outside the scope of this study. B.W. reports ad hoc membership of the scientific advisory board of REPARE Therapeutics, outside the scope of this study.
Ethics approval/consent to participate
This study was approved by the Institutional Review Board.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Benhamida, J.K., Vyas, M., Tanaka, A. et al. Pancreatoblastomas and mixed and pure acinar cell carcinomas share epigenetic signatures distinct from other neoplasms of the pancreas. Mod Pathol 35, 956–961 (2022). https://doi.org/10.1038/s41379-021-00989-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41379-021-00989-2