Abstract
In rare cases, giant cell tumor of bone (GCTB) can undergo primary or secondary malignant transformation to malignant giant cell tumor of bone (MGCTB), but the details of the molecular alterations are still unclear. The present study aimed to elucidate the clinicopathologic and molecular features of MGCTBs based on immunohistochemistry, fluorescence in situ hybridization (FISH) and next generation sequencing (NGS) of nine MGCTBs (five primary and four secondary). Seven (78%) of 9 MGCTBs were immunohistochemically positive for H3.3 G34W. In two (22%) patients, although GCTB components were focally or diffusely positive for H3.3 G34W, their malignant components were entirely negative for H3.3 G34W, which was associated with heterozygous loss of H3F3A by FISH. NGS on four MGCTBs revealed pathogenic mutations in TP53 (n = 3), EZH2 (n = 1) and several other genes. Immunohistochemical analysis of the nine MGCTBs confirmed the p53 nuclear accumulation (n = 5) and loss of H3K27me3 expression (n = 3) and showed that they were mutually exclusive. In addition, four (80%) of five cases of pleomorphic or epithelioid cell-predominant MGCTBs were positive for p53, while three (75%) of four cases of spindle cell-predominant MGCTBs were negative for trimethylation at lysine 27 of histone 3 (H3K27me3). The results suggested that p53 alteration and dysfunction of histone methylation as evidenced by H3K27me3 loss may play an important role in the malignant progression of GCTB, and might contribute to the phenotype–genotype correlation in MGCTB. The combined histologic, immunohistochemical and molecular information may be helpful in part for the diagnosis of challenging cases.
Similar content being viewed by others
Introduction
Giant cell tumor of bone (GCTB) is an osteolytic neoplasm that usually develops in the metaphysis to epiphysis of a long bone (such as the femur or tibia) or in the axial skeleton (such as the spine or sacrum) of young to middle-aged adults1. GCTB has a unique histological feature consisting of mononuclear neoplastic cells, mononuclear stromal cells and osteoclast-like multinucleated giant cells1. Histone H3.3 is encoded by the H3F3A and H3F3B genes, and almost all GCTBs have a mutation of H3F3A p.G342. H3F3A p.G34W is the most frequent mutation in GCTB (~90%)3,4,5,6,7. Minor subsets (each <2%) have H3F3A p.G34L, p.G34M, p.G34R, or p.G34V mutation3,4,5,6. Biologically, the interaction between receptor activator of nuclear factor kappa-Β (RANK) on the osteoclasts and nuclear factor kappa-Β ligand (RANKL) secreted by neoplastic osteoblastic cells plays an important role for promotion of the process of osteoclast formation and activation, leading to acceleration of osteolysis8,9,10. Denosumab, an inhibitor of RANKL, is used as an effective therapeutic agent for patients with GCTB11,12. Denosumab therapy induces histological changes such as a decrease of osteoclasts and increase of woven bone and fibro-osseous tissue12. However, some population of GCTBs do not respond to denosumab, and biological or pathological factors predictive of denosumab efficacy have not been established13.
Clinically, GCTB behaves in a locally aggressive manner and sometimes metastasizes in keeping their conventional histology1. In rare instances, GCTB shows transformation to malignant giant cell tumor of bone (MGCTB)1. MGCTB is treated by wide resection, but the prognosis is unfavorable14. MGCTB is subgrouped into two categories: primary MGCTB and secondary MGCTB1,14. Primary MGCTB is a malignant bone tumor having an overt malignant component concurrent with a conventional giant cell tumor of bone (CGCTB) component. Secondary MGCTB is a malignancy that co-exists with recurrent CGCTB or that occurs in patients with a past history of CGCTB. Histologically, MGCTBs show various morphologies mimicking osteosarcoma (OS), undifferentiated pleomorphic sarcoma (UPS) and fibrosarcoma1. A subset of MGCTBs has been reported to reveal epithelioid features like carcinoma15. A small population of OS and UPS have been reported to be positive for anti-H3.3G34W antibody, suggesting that these cases may be identical to primary MGCTB at a molecular level5,16.
It is speculated that the transformation from GCTB to malignancy takes several years or decades, but the details of the morphological changes and molecular alterations along with the stepwise progression are still unclear3. Recent studies have reported that the H3F3A mutations can also be detected in MGCTB4,5. On the other hand, another study reported that some MGCTBs were negative for H3F3A mutations and H3.3 G34W immunohistochemistry (IHC), even though the paired CGCTB component was positive for H3F3A mutations16. The investigators speculated that this peculiar phenomenon might be explained in part by heterozygous loss of the H3F3A mutant allele (with concomitant preservation of the wild-type allele), based on the findings of fluorescence in situ hybridization (FISH)16. Several earlier reports suggested that TP53 mutation may be associated with malignant progression of GCTB17,18. A recent study reported that, in addition to H3F3A or H3F3B mutation, most MGCTBs showed features associated with TERT promoter mutations, suggesting a possible role of telomere dysfunction in the malignant transformation of GCTB19. As a nonrecurrent event, biallelic losses of histone lysine demethylase KDM4B or KDM6A were also identified19.
The present study aimed to further elucidate the clinicopathologic and molecular features of MGCTBs based on an exhaustive review of malignant bone tumors and analysis using next-generation sequencing (NGS) and IHC.
Materials and methods
Case selection
A total of nine cases of MGCTB were retrieved from the files of Department of Anatomic Pathology, Kyushu University Hospital. The nine cases were obtained by exhaustive review of a total of 132 cases of malignant bone tumors with immunohistochemical staining for H3.3 G34W (Supplementary Fig. 1, Table 1). Based on immunoreactivity for H3.3 G34W, three tumors were reappraised as primary MGCTB, including one of 80 tumors originally diagnosed as OS (Case 1, giant cell rich OS), two of 30 tumors originally diagnosed as UPS or formerly malignant fibrous histiocytoma (Cases 2 and 3) and none of 16 leiomyosarcomas. Among the total of nine tumors, six were MGCTBs which had been diagnosed during routine practice based on their clinical history and histological features (Cases 4–9). Because two of the tumors (Cases 4 and 5) were diagnosed as malignancies at 3–4 months after the initial diagnosis of GCTB, these cases presumably represent a sampling error of “hidden” primary MGCTB. Thus, we included both these tumors as primary MGCTBs in this study. Remaining four cases (Cases 6–9) had prior history of GCTB several years ago before the diagnosis of malignancy, consistent with the concept of secondary MGCTB. Finally, five cases of primary MGCTB (Cases 1–5) and four cases of secondary MGCTB (Cases 6–9) were enrolled in this study (Table 1). This study was conducted in accordance with the principles embodied in the Declaration of Helsinki, and was approved by the institutional review board of Kyushu University Hospital (Nos. 29–429, 29–625).
Immunohistochemistry
Formalin-fixed, paraffin-embedded (FFPE) samples were sectioned at a thickness of 3 μm. We used primary antibodies against H3.3 G34W (x400, rabbit monoclonal, RM263; RevMAb Biosciences, South San Francisco, CA), p53 (x500, mouse monoclonal, PAb1801; Calbiochem, San Diego, CA), trimethylation at lysine 27 of histone 3 (H3K27me3) (x200, rabbit monoclonal C36B11; Cell Signaling Technology, Danvers, MA), dimethylation at lysine 27 of histone 3 (H3K27me2) (x1000, rabbit polyclonal, ab24684; Abcam, Cambridge, UK), MLH1 (x50, mouse monoclonal, G168-15; BD Pharmingen, San Jose, CA), MSH6 (x100, rabbit monoclonal, EP49; Dako, Carpinteria, CA) and c-kit (x200, rabbit polyclonal, A4502; Dako). All of these antibodies except H3.3 G34W were examined to check the significance of candidate genes obtained by NGS data as described below. Epitope retrieval was performed in target retrieval solution (Dako) for 20 min at 98 °C by using a microwave. Each primary antibody was mounted on a tissue section for 1 hour at room temperature, and then the EnVision plus system (Dako) and 3,3’-diaminobenzidine tetrahydrochloride (DAB) were used to visualize the labeled antigens. p53 and c-kit were judged as positive when more than 10% of tumor cells showed immunoreactivity. Loss of expression for H3K27me2/3, MLH1 or MSH6 was defined as negative staining in almost all tumor nuclei and positive staining of endothelial cells and inflammatory cells as an internal positive control.
Sanger sequencing analysis for H3F3A and TERT promoter mutations
Polymerase chain reaction (PCR) and Sanger sequencing analysis were carried out to detect gene mutations in H3F3A. Genomic DNA was extracted from FFPE or snap-frozen samples. The primers and sequencing method were as described previously4. We also examined TERT promoter mutation using the following primer sets; forward 5ʹ-CTGCCCCTTCACCTTCCAG-3ʹ and reverse 5ʹ-AGCGCTGCCTGAAACTCG-3ʹ. H3F3A and TERT were not included in the NGS panel used in this study.
FISH for H3F3A
To examine H3F3A copy number alterations, we used an H3F3A/CEN1q Dual Color FISH Probe (GSP Laboratory, Kobe, Japan)16. FISH images were captured using an HS All-in-one Fluorescence Microscope - BZ-9000 series (BIOREVO) (Keyence, Osaka, Japan). We evaluated mononuclear tumor cells that had at least one evaluable signal and interpreted the signals as previously described16. We defined the H3F3A signal pattern as deletion when a mononuclear tumor cell contained CEN1q signals (green) whose number exceeded that of H3F3A signals (red); in the representative case, there was one signal of H3F3A (red) and two signals of CEN1q (green) in individual tumor cells.
NGS for genomic alterations
For DNA extraction, snap-frozen samples were available in 3 cases of MGCTB components (Cases 2, 6 and 7), and FFPE samples were used in the remaining 6 cases of MGCTB (Cases 1, 3, 4, 5, 8 and 9). Genomic DNA was extracted from the frozen tissue using a QIAamp DNA Mini Kit (Qiagen, Tokyo) or from the FFPE specimens using a DNAstorm™ FFPE (Cell Data Sciences, Fremont, CA). Library preparation was conducted by using an Ion AmpliSeq™ Comprehensive Cancer Panel for target sequencing of 409 genes (Thermo Fisher Scientific, Waltham, MA) and an Ion AmpliSeq™ Library Kit Plus (Thermo Fisher Scientific). Emulsion PCR and enrichment was conducted by using an Ion OneTouch™ 2 System (Thermo Fisher Scientific). We conducted sequencing by using the Ion PGM™System (Thermo Fisher Scientific) and Ion 318™ Chip Kit v2 BC (Thermo Fisher Scientific). We followed the manufacturer’s instructions for all procedures. The obtained raw sequence data were processed by using Ion Reporter software (Thermo Fisher Scientific; https://ionreporter.thermofisher.com/ir/). Filtering parameters were set as follows: coverage ≥20; allele frequency ≥0.05; location: exon and splice site; variant effect: missense, nonsense, unknown, non-frameshift indel, frameshift indel and splice variant. The pathogenic mutations listed in the Catalog of Somatic Mutation in Cancer (COSMIC; https://cancer.sanger.ac.uk/cosmic) were retrieved.
Statistics
We conducted statistical analysis by Fisher’s exact test using the JMP statistical software package (version 14; SAS Institute, Cary, NC).
Results
Clinicopathological findings
Clinicopathological findings of the nine cases of MGCTB are summarized in Table 1 and Supplementary Fig. 2. The patients consisted of seven men and two women, aged from 27 to 81 years (average age: 57). The main locations of the tumor were the distal epiphysis of femur in five cases, the proximal epiphysis of tibia in two cases and the iliac bone in two cases. The tumor size ranged from 2.2 to 10.1 cm (average: 5.2 cm). As outlined in the Case Selection section, three of five cases of primary MGCTB had been initially diagnosed as giant cell-rich OS (n = 1; Case 1) or UPS (n = 2; Cases 2 and 3), while two of them (Cases 4 and 5) were diagnosed as malignancies at 3–4 months after the initial diagnosis of GCTB on open biopsy or curettage specimens. For the four cases of secondary MGCTBs, the term to progress to malignancy ranged from two to 27 years (Cases 6–9).
As for the therapy for initial GCTB, four patients (Cases 5 and 7–9) underwent curettage and 1 (Case 6) underwent marginal resection. The remaining one patient (Case 4) received denosumab therapy after open biopsy; however, the tumor became bigger suggesting clinically poor response to denosumab. In all 6 of these patients (Cases 4–9), no radiation therapy had been conducted for initial GCTB lesions. For malignant tumors, four (Cases 2 and 6–8) and two (Cases 4 and 9) patients underwent wide resection and marginal resection, respectively. In two patients (Cases 1 and 5), chemotherapy was conducted after the diagnosis of malignancy by open biopsy or curettage specimens. In the remaining one patient (Case 3), information about treatment and follow-up data was not available.
The survival data were available in eight cases. The follow-up term from the malignant change ranged from five months to 19 years (average: approximately seven years). Neither recurrence nor metastasis was recognized in the six patients over at least 11 months of follow-up (Cases 2, 4 and 6–9). Two patients died due to metastatic spread of MGCTB at 25 months and five months (Cases 1 and 5, respectively).
Histological findings and H3F3A gene status in GCTB and MGCTB
In 8 patients (Case 1, 2 and 4–9), the diagnosis of MGCTB was made based on the findings that histologically confirmed CGCTB component was present previously (Cases 4–9) and/or concurrently (Cases 1, 2, 4 and 9) (Table 2). In one patient (Case 3), although CGCTB component was not present in any samples, H3.3 G34W immunoexpression and H3F3A p.G34W mutation were detected in the malignant tumor, and thus we included this case in MGCTBs in this study.
The representative histological appearance, H3.3 G34W expression and H3F3A gene sequence in CGCTBs are shown in Supplementary Fig. 3. CGCTB components were composed of mononuclear cells and osteoclast-like multinucleated giant cells. Mononuclear cells showed nuclear expression of H3.3 G34W by IHC. The H3F3A p.G34W mutation was confirmed by Sanger sequencing.
Histological findings of the malignant components of MGCTBs are shown in Fig. 1 and Table 2. The nine cases of MGCTBs exhibited various combinations of cellular morphology, including spindle (n = 4), pleomorphic (n = 5) and epithelioid (n = 5) features, and there was often more than one histological pattern in a single tumor. The predominant patterns in individual cases were spindle (n = 4), pleomorphic (n = 2) and epithelioid (n = 3), although it was difficult to clearly distinguish these histological patterns (especially pleomorphic vs epithelioid cell type). The spindle-cell subtype showed hypercellular fascicular proliferation of relatively uniform, atypical spindle-shaped cells (Fig. 1a). The pleomorphic subtype showed a haphazard proliferation of atypical polygonal cells, mimicking UPS (Fig. 1b). The epithelioid subtype showed sheet-like proliferation of atypical cells with enlarged round to oval-shaped nuclei and various amounts of eosinophilic cytoplasm (Fig. 1c). In one case (Case 2), there was a focal component of signet ring cell-like malignant cells with vacuolated cytoplasm and eccentric nuclei (Fig. 1d).
a Spindle cell subtype. Typical spindle cells proliferate in a fascicular pattern. b Pleomorphic subtype. Large atypical polygonal cells with hyperchromatic nuclei are noted. c Epithelioid cell subtype. Large atypical cells having eosinophilic cytoplasm, rounded to oval nuclei and distinct nucleoli proliferate in a sheet-like pattern. d Tumor cells with signet ring cell-like appearance with vacuolated cytoplasm are focally seen. e The malignant cells are immunohistochemically positive for H3.3 G34W (the same case as Fig. 1c).
In addition to malignant components, there were also CGCTB components within a single tumor in four cases (Cases 1, 2, 4 and 9) (Table 2). The proportion of co-existing CGCTB component was about 30–70%. In one patient (Case 4), foci of immature bone formation with bland-looking cells were also seen in the tumor resected after denosumab therapy, suggesting a partial response to denosumab in the CGCTB component.
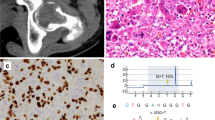
Most cases were positive for H3.3 G34W IHC and H3F3A p.G34W mutation in both CGCTB and MGCTB (Fig. 1e, Table 2). In a MGCTB (Case 7), despite the positivity for H3.3 G34W IHC, the H3F3A sequence data were not available, probably due to the insufficient quality of DNA extracted from the FFPE specimen. Notably, two (22%) of nine MGCTBs (Cases 5 and 6) were entirely negative for H3.3 G34W IHC in the malignant component, while their preceding CGCTBs were focally or diffusely positive for H3.3 G34W IHC in Cases 5 and 6, respectively (Table 2, Figs. 2a and b). In addition, H3F3A mutation was not detectable by Sanger sequencing (Fig. 2c), and heterozygous loss of one allele of H3F3A was detected by FISH in these two malignant cases (Fig. 2d).
a Atypical spindle tumor cells proliferate in (a) fascicular pattern. b The sarcomatous cells are negative for H3.3 G34W immunohistochemistry. c H3F3A mutation is not detected by Sanger sequencing. d Heterozygous loss of H3F3A is revealed by FISH. In most individual tumor cells, one red signal of H3F3A and two green signals are observed.
Atypical neoplastic cells in GCTB
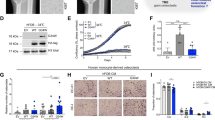
During the review of MGCTB cases, we also noticed the existence of microscopic foci of atypical neoplastic cells sparsely distributed within the preceding CGCTB components in three of nine (33%) patients (Cases 4, 5 and 9) (Table 2, Supplementary Fig. 2, Fig. 3). These atypical neoplastic cells were not identified within the overt malignant components in any cases of MGCTB. The atypical neoplastic cells were characterized by epithelioid feature with pale eosinophilic cytoplasm, rounded nuclei, vesicular chromatin and prominent nucleoli (Figs. 3a and e). The nucleus was 2–3 times larger than that of typical mononuclear neoplastic cells, but smaller than that of apparent malignant cells (Figs. 3c and g). Immunohistochemically, the atypical neoplastic cells were positive for H3.3 G34W in two of three cases (Cases 4 and 9).
Case 4: The atypical neoplastic cells are surrounded by multinucleated giant cells (a), and atypical cells are positive for p53 (b). In the malignant tumor, epithelioid malignant cells proliferate in a sheet-like pattern (c) and are positive for p53 (d). Case 5: A vague microscopic nodule of atypical neoplastic cells is shown (e). A few multinucleated giant cells are associated with this lesion. H3K27me3 is preserved in these atypical cells (f). In the malignant tumor, spindle sarcoma cells proliferate in hypercellular fascicles (g), and show loss of H3K27me3 (h). H3K27me2 is also deficient (inset).
As a comparison, we reviewed 86 cases of CGCTB which had not transformed to malignancy after the surgery, and only one (1.2%) case harbored focal atypical neoplastic cells (Supplementary Fig. 4). The prevalence of atypical cells in CGCTBs without malignant change (1/86 cases; 1.2%) was significantly lower than that in the CGCTB component of MGCTBs (3/8 cases; 37.5%) (p < 0.001) (Supplementary Table 1).
Genomic alterations, immunohistochemical validations and histological correlation
The library preparation for target sequencing succeeded in a total of five samples, consisting of four MGCTBs (Cases 2, 4, 6 and 7) and 1 CGCTB (Case 4). The target sequencing revealed alterations of several genes, and the mutated genes estimated as “pathogenic” by COSMIC were related to receptor tyrosine kinase (FGFR3, FGFR4, KIT, MET and ROS1), cell proliferation/cell cycle (TP53, PTCH1, BIRC3, EP400 and FBXW7), epigenetic modification (EZH2 and ASXL1) and DNA repair (MLH1, MSH6, PARP1 and TRRAP) (Fig. 4). Although the mutations in EZH2, KIT, MLH1 and MSH6 were non-recurrent among MGCTBs, the fact that these genes as well as TP53 were responsible for the pathogenesis of several kinds of cancer attracted our attention20,21, and thus we conducted an immunohistochemical analysis for validation (Supplementary Fig. 5). The EZH2 p.D185H-mutated MGCTB (n = 1, Case 2) showed loss of H3K27me3 and H3K27me2, and the TP53 p.K382Nfs*40 or p.V73Wfs*50-mutated MGCTB (n = 3, Cases 4, 6 and 7) showed nuclear accumulation of p53.
We additionally examined TERT promoter mutation by Sanger sequencing. One MGCTB (Case 5) and one CGCTB (Case 9) had C318T and C201T mutations at 214 bp and 97 bp upstream of the TERT transcription start site, respectively (Supplementary Table 2).
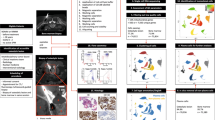
Next, we expanded the immunohistochemical staining to all nine MGCTBs, and detected the p53 nuclear accumulations in five cases (Cases 3, 4, 6, 7 and 9) and dual H3K27me3/2 losses in three cases (Cases 2, 5 and 8) in a mutually exclusive manner (Fig. 5a). Among the 2 MGCTB cases with loss of H3.3 G34W expression (H3F3A heterozygous loss), one showed p53 positivity and TP53 mutation (Case 6), and the other showed H3K27me3/2 loss (Case 5).
a The predominant cellular subtype of the malignant component (pleomorphic, epithelioid and spindle) and immunohistochemical results of H3K27me3 loss, p53 positivity and H3.3 G34W loss are shown by colored tiles. H3K27me3 loss (indicated by red) and p53 positivity (blue) are mutually exclusive. There are tendencies of correlation between H3K27me3 loss and spindle cell morphology (Cases 2, 5 and 8) and between p53 and epithelioid cell morphology (Cases 3, 4 and 7). There are two cases with H3.3 G34W loss (green) (Cases 5 and 6). b The p53 positivity (blue), H3K27me3 loss (red) and H3.3 G34W loss (green) in the CGCTB, atypical neoplastic cell and malignant components in individual cases are shown. Gray tiles indicate the absence of corresponding components. In 2 cases, p53 positivity is seen both in atypical neoplastic cell and malignant components (Cases 4 and 9). H3K27me3 loss is present only in the malignant components (Cases 2, 5, and 8) but not in atypical cell and CGCTB components. H3.3 G34W loss is present only in the malignant component in 1 patient (Case 6), whereas it is present throughout the course in another patient (Case 5).
The molecular alterations were then compared with the histological findings (Fig. 5a). Most cases (4/5; 80%) of pleomorphic or epithelioid cell-predominant MGCTB were positive for p53 and vice versa (Figs. 3c, d and 5a). In contrast, most cases (3/4; 75%) of spindle cell-predominant MGCTB were H3K27me3-deficient; conversely, all three cases of H3K27me3-deficient MGCTB were of spindle cell type (Figs. 3g, h and 5a).
We further compared the expression status of p53, H3K27me3 and H3.3 G34W in each component of individual cases (Fig. 5b). All CGCTB components examined were negative for p53, whereas atypical neoplastic cells and matched malignant components were positive for p53 in two patients (Cases 4 and 9) (Figs. 3a-d and 5b). H3K27me3 expression was retained in all CGCTB and atypical neoplastic cell components, and H3K27me3 loss was seen only in malignant components (Cases 2, 5 and 8) (Figs. 3e-h and 5b). There were 2 cases with eventual loss of H3.3 G34W expression in malignant components (Cases 5 and 6); H3.3 G34W was diffusely positive in the benign component but entirely negative in malignant component in Case 6, whereas H3.3 G34W was focally positive in the benign component and entirely lost in both atypical neoplastic cells and malignant component in Case 5.
Discussion
The molecular mechanisms of malignant progression of GCTB have been investigated. Some earlier studies reported the TP53 gene mutation, p53 overexpression17,18 and gene amplification of CCND1 and MET22 in a subset of MGCTB. Recently, Fittall et al. reported frequent telomere dysfunction associated with TERT mutation in MGCTB19. TERT promoter mutations such as C228T and C250T are reported to upregulate the transcription of TERT in various types of cancer including glioma and thyroid carcinoma, and C228T mutation has been reported in MGCTB19. In our series, small subset of GCTBs had TERT promoter C318T or C201T mutations; however, the biological significance of those mutations is still unclear whether those mutations are pathogenic or not.
EZH2 is a methyltransferase that is responsible for the methylation activity of polycomb repressive complex 2 (PRC2)20,23. PRC2 also contains EED and SUZ12, and the complex plays a crucial role in histone modification by methylating H3K27 to H3K27me2 and H3K27me323. Mutation in EZH2, EED or SUZ12 impairs PRC2 function, leading to loss of expression of H3K27me2 and H3K27me3. Malignant peripheral nerve sheath tumor (MPNST) frequently harbors mutation in EED or SUZ12 and often shows loss of expression of H3K27me2 and H3K27me323,24,25. In the current study of MGCTB, EZH2 p.D185H mutation was identified in one case by NGS, and three cases showed complete loss of both H3K27me2 and H3K27me3 by IHC. Previous reports have suggested that the EZH2 p.D185H mutation confers pathogenicity20,21, and the dual losses of H3K27me2/3 strongly support the idea that PRC2 dysfunction may be present in a subset of MGCTB. In addition, H3K27me3 expression was retained in the preceding CGCTB components, suggesting that dysfunction of PRC2, including EZH2 mutation, may occur in the step of malignant transformation of GCTB.
As for histology, the 3 MGCTBs with H3K27me3 loss were of predominantly spindle cell morphology, resembling fibrosarcoma or MPNST. Notably, a recent study has reported that a subset of dedifferentiated chondrosarcomas showed loss of H3K27me3 immunoexpression26, and histologically such sarcomas had a spindle cell pattern resembling MPNST26. Collectively, the results suggest that there might be a common phenotype–genotype correlation (spindle cell morphology and H3K27me3 loss/PRC2 dysfunction) among MPNST, MGCTB and dedifferentiated chondrosarcoma; however, the detailed molecular mechanism underlying the morphogenesis is still unclear. In addition, the contributions of H3K27me3 loss and H3.3 G34 mutation to cellular proliferation and differentiation should be further elucidated27. From the view point of diagnostic markers, these previous and our present results suggest that H3K27me3 is no longer a specific marker for MPNST.
Consistent with previous reports, in our series, TP53 mutation was detected in three cases by NGS, and 5 cases showed abnormal nuclear accumulation of p53 by IHC in the malignant component but not in the CGCTB component. These results support a possible role of TP53 alteration in the malignant transformation of GCTB. As for histology, most p53-positive MGCTBs showed epithelioid or pleomorphic morphology. In addition, it is of note that the p53 positivity and H3K27me3 loss were mutually exclusive. Similarly, according to the previous reports, epithelioid MPNSTs typically show preserved expression of H3K27me324, and are often positive for p5328,29. The exact reason for this phenomenon of mutual exclusivity is still unclear. One possible explanation is that each of H3K27me3 loss (PRC2 dysfunction) and p53 alteration may be a strong driver, with either event being sufficient for the progression of tumors in both MGCTB and MPNST.
The prevalence of H3F3A mutation in MGCTB is controversial. Earlier studies analyzing small numbers of cases reported that H3F3A G34 mutation was present in most MGCTBs [reviewed in ref. 3]. Recently, Yoshida et al. found that a subset of MGCTB lacked H3F3A mutation and H3.3 G34W immunoreactivity, even though the matched CGCTB components were H3.3 G34W-positive16. In addition, they demonstrated heterozygous loss of H3F3A by FISH in MGCTB, and suggested that the loss of the mutant allele of H3F3A may be responsible for this peculiar phenomenon. Similarly, in the current study, two of nine cases of clinicopathologically diagnosed MGCTB showed absence of H3F3A mutation, disappearance of H3.3 G34W expression and heterozygous loss of H3F3A in the malignant components, in contrast to the complete or focal presence of H3.3 G34W expressions in the GCTB components (Cases 5 and 6). The two cases showed similar spindle cell-predominant morphology but otherwise different features; i.e., one case showed p53 positivity and an indolent clinical course (Case 6), whereas the other case showed H3K27me3 loss and rapid progression resulting in death (Case 5). Thus, the clinicopathological and molecular biological significance of loss of the H3F3A mutant allele in MGCTB remains unclear, and further study is needed.
On rare occasion, GCTBs show unusual histological features making it difficult to draw a sharp line between benign and malignancy30,31. In the current study, we identified focal atypical neoplastic cells in biopsy or curette specimens of GCTB preceding to overt malignancy in 3 of 9 patients. These atypical neoplastic cells had epithelioid appearance with eosinophilic cytoplasm, enlarged round nuclei, vesicular chromatin and distinct nucleoli. Of note is that the morphological feature of these atypical neoplastic cells seems different from that of symplastic change in GCTB; the latter is estimated as degenerative atypia as shown by nuclear pleomorphism and smudged nuclear chromatin but rare mitotic activity (mimicking degenerative atypia in ancient schwannoma)30. Based on our results, the atypical neoplastic cells with epithelioid feature seemingly have intermediate features between conventional and malignant GCTBs in terms of morphology and molecular abnormalities. In addition, the three cases were diagnosed as overt malignancy shortly (three months, four months and two years) after the first presentation of GCTB with focal atypia (Supplementary Fig. 2). Thus, these atypical neoplastic cells might represent lower grade malignancy or premalignant lesion, and such a morphological spectrum of atypia can make the diagnosis of MGCTB very difficult, if not impossible, on limited tissue. In our opinion, these atypical cells should not be immediately diagnosed as malignancy in the practical diagnosis, because they are very limited microscopic lesions within CGCTB, and their histological impression is quite different from that of overt malignancy. In addition, it cannot yet be conclusively stated that the presence of atypical neoplastic cells drastically increases the risk of the following sarcomatous overgrowth, because only a few cases were examined in the current study. Moreover, there was no direct proof that atypical neoplastic cells proceeded to malignancy because these atypical neoplastic cells were not identified within the overt malignant components in any cases. However, the fact that a GCTB with focal atypia (Case 4) did not fully respond to denosumab therapy suggests that the clinical management of “atypical GCTB” should be done much more cautiously. Since the diagnosis of MGCT was made only a few months later in patients with “atypical GCTB” (Cases 4 and 5), the presence of atypical neoplastic cell in limited biopsy tissue should raise suspicion for more clear cut MGCTB in unsampled areas of the tumor. If seen in a curetted specimen, all of the tissue should be evaluated to rule out the possibility of MGCTB areas.
Based on the above findings, we propose a hypothetical model for malignant progression of GCTB (Fig. 6). TP53 alteration and H3K27me3 loss may play a role in the malignant progression of GCTB by mutually exclusive manner. H3F3A mutant allelic loss and/or as-yet-unknown molecular events might also participate in the progression. Conceptually, “atypical GCTB” might exist at the intermediate step between benign and malignancy, but its histological definition, reproducibility and clinicopathological significance are not yet established. Further studies with larger number of cases with extensive review of histology and clinical course as well as molecular analyses are necessary.
According to the previous reports, aside from H3F3 mutations, CGCTB seems to show a relatively low somatic mutation burden, because no or only a few gene alterations were found in addition to the H3F3A mutation in CGCTBs2,19,31. In the current study, some mutations were found in one case of CGCTB, but the vast majority of them did not overlap with the mutated genes in previous reports19,31. Recently, Sangatsuda et al. reported that H3.3 G34R-mutant gliomas had a characteristic methylation signature which was shared with H3.3 G34W-mutated osteosarcoma32. From these findings, epigenomic alterations linked to H3.3 G34 mutation rather than somatic mutations might play a role in the pathogenesis of GCTB, although at present this is only speculation.
In conclusion, our results suggest that p53 alteration and dysfunction of histone methylation as evidenced by H3K27me3 loss may play an important role in the malignant progression of GCTB, and may contribute to the phenotype–genotype correlation in MGCTB. The combined histologic, immunohistochemical and molecular information may be helpful for diagnostically challenging settings.
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request.
References
WHO Classification of Tumours Editorial Board. Soft tissue and bone tumours. Lyon(France): International Agency for Research on Cancer, 2020
Behjati, S. et al. Distinct H3F3A and H3F3B driver mutations define chondroblastoma and giant cell tumor of bone. Nat Genet 45, 1479–1482 (2013).
Yamamoto, H., Ishihara, S., Toda, Y. & Oda, Y. Histone H3.3 mutation in giant cell tumor of bone: an update in pathology. Med Mol Morphol 53, 1–6 (2020).
Yamamoto, H. et al. Diagnostic utility of histone H3.3 G34W, G34R, and G34V mutant-specific antibodies for giant cell tumors of bone. Hum Pathol 73, 41–50 (2018).
Amary, F. et al. H3F3A (Histone 3.3) G34W Immunohistochemistry: a reliable marker defining benign and malignant giant cell tumor of bone. Am J Surg Pathol 41, 1059–1068 (2017).
Lüke, J. et al. H3F3A mutation in giant cell tumour of the bone is detected by immunohistochemistry using a monoclonal antibody against the G34W mutated site of the histone H3.3 variant. Histopathology 71, 125–133 (2017).
Presneau, N. et al. Diagnostic value of H3F3A mutations in giant cell tumour of bone compared to osteoclast-rich mimics. J Pathol Clin Res 1, 113–123 (2015).
Atkins, G. J. et al. Expression of osteoclast differentiation signals by stromal elements of giant cell tumors. J Bone Min Res 15, 640–649 (2000).
Huang, L., Xu, J., Wood, D. J. & Zheng, M. H. Gene expression of osteoprotegerin ligand, osteoprotegerin, and receptor activator of NF-κB in giant cell tumor of bone: Possible involvement in tumor cell-induced osteoclast-like cell formation. Am J Pathol 156, 761–767 (2000).
Atkins, G. J. et al. Osteoprotegerin inhibits osteoclast formation and bone resorbing activity in giant cell tumors of bone. Bone 28, 370–377 (2001).
Thomas, D. et al. Denosumab in patients with giant-cell tumour of bone: an open-label, phase 2 study. Lancet Oncol 11, 275–280 (2010).
Branstetter, D. G. et al. Denosumab induces tumor reduction and bone formation in patients with giant-cell tumor of bone. Clin Cancer Res 18, 4415–4424 (2012).
Errani, C. et al. Denosumab may increase the risk of local recurrence in patients with giant-cell tumor of bone treated with curettage. J Bone Jt Surg 100, 496–504 (2018).
Bertoni, F., Bacchini, P. & Staals, E. L. Malignancy in giant cell tumor of bone. Cancer 97, 2520–2529 (2003).
Machinami, R. et al. Carcinosarcomatous malignancy, osteosarcoma and squamous cell carcinoma, in giant cell tumor of the right distal femur. Pathol Res Pr 204, 583–588 (2008).
Yoshida, K. I. et al. Absence of H3F3A mutation in a subset of malignant giant cell tumor of bone. Mod Pathol 32, 1751–1761 (2019).
Oda, Y. et al. Secondary malignant giant-cell tumour of bone: Molecular abnormalities of p53 and H-ras gene correlated with malignant transformation. Histopathology 39, 629–637 (2001).
Okubo, T. et al. p53 mutations may be involved in malignant transformation of giant cell tumor of bone through interaction with GPX1. Virchows Arch 463, 67–77 (2013).
Fittall, M. W. et al. Drivers underpinning the malignant transformation of giant cell tumour of bone. J Pathol 252, 433–440 (2020).
Cohen, A. S. et al. Weaver syndrome-associated EZH2 protein variants show impaired histone methyltransferase function in vitro. Hum Mutat 37, 301–307 (2016).
Burgos S., et al. Novel EZH2 mutation in a patient with secondary B-cell acute lymphocytic leukemia after deletion 5q myelodysplastic syndrome treated with lenalidomide: a case report. Medicine (Baltimore) (2019) https://doi.org/10.1097/MD.0000000000014011
Saâda, E. et al. CCND1 and MET genomic amplification during malignant transformation of a giant cell tumor of bone. J Clin Oncol 29, 86–89 (2011).
Lee, W. et al. PRC2 is recurrently inactivated through EED or SUZ12 loss in malignant peripheral nerve sheath tumors. Nat Genet 46, 1227–1232 (2014).
Prieto-Granada, C. N. et al. Loss of H3K27me3 expression is a highly sensitive marker for sporadic and radiation-induced MPNST. Am J Surg Pathol 40, 479–489 (2016).
Marchione, D. M. et al. Histone H3K27 dimethyl loss is highly specific for malignant peripheral nerve sheath tumor and distinguishes true PRC2 loss from isolated H3K27 trimethyl loss. Mod Pathol 32, 1434–1446 (2019).
Makise, N. et al. H3K27me3 deficiency defines a subset of dedifferentiated chondrosarcomas with characteristic clinicopathological features. Mod Pathol 32, 435–445 (2019).
Khazaei, S. et al. H3.3 G34W promotes growth and impedes differentiation of osteoblast-like mesenchymal progenitors in giant cell tumor of bone. Cancer Disco 10, 1968–1987 (2020).
McMenamin, M. E. & Fletcher, C. D. M. Expanding the spectrum of malignant change in schwannomas. Am J Surg Pathol 25, 13–25 (2001).
Izycka-Swieszewska, E. et al. Epithelioid malignant peripheral nerve sheath tumor involving maxillary sinus. Neuropathology 25, 341–345 (2005).
Sarungbam, J. et al. Symplastic/pseudoanaplastic giant cell tumor of the bone. Skelet Radio 45, 929–935 (2016).
Ogura, K. et al. Highly recurrent H3F3A mutations with additional epigenetic regulator alterations in giant cell tumor of bone. Genes Chromosom Cancer 56, 711–718 (2017).
Sangatsuda, Y. et al. Base-resolution methylomes of gliomas bearing histone H3.3 mutations reveal a G34 mutant-specific signature shared with bone tumors. Sci Rep. 10, 16162 (2020).
Acknowledgements
We appreciate the technical support from the Department of Pathology, Kyushu University. We also thank the Research Support Center, Graduate School of Medical Science, Kyushu University. The English used in this manuscript was revised by KN International (http://www.kninter.com/). This work was supported by JSPS KAKENHI Grant Number 19H03444.
Author information
Authors and Affiliations
Contributions
S.I., H.Y. and Y.O. designed this stydy. S.I., Y.T., M.Y., Y.I., Y.S., K.K., I.K., Y.Y., K.K., T.F., N.S., M.E., Y.M., Y.K., and Y.N. collected the data and sample. S.I., T.I. and T.Y. conducted the experiments. S.I. and H.Y. performed histological evaluation of the samples. S.I. and H.Y. wrote the manuscript. Y.O. supervised the experiments.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval
The present study was approved by Kyushu University Committee of Bioethics (approval no. 29–429 and 29–625).
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Ishihara, S., Yamamoto, H., Iwasaki, T. et al. Histological and immunohistochemical features and genetic alterations in the malignant progression of giant cell tumor of bone: a possible association with TP53 mutation and loss of H3K27 trimethylation. Mod Pathol 35, 640–648 (2022). https://doi.org/10.1038/s41379-021-00972-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41379-021-00972-x
This article is cited by
-
Giant Cell Tumor of Mandible : A Case Report
Indian Journal of Otolaryngology and Head & Neck Surgery (2024)
-
Blue lacy matrix in giant cell tumour of bone with or without denosumab therapy
Virchows Archiv (2023)
-
In situ sequence-specific visualization of single methylated cytosine on tissue sections using ICON probe and rolling-circle amplification
Histochemistry and Cell Biology (2023)