Abstract
p53 immunohistochemistry has long been proposed for the separation of benign from malignant mesothelial proliferations, with the older literature suggesting that any degree of positivity supported a diagnosis of mesothelioma. However, using modern immunohistochemistry platforms in other organ systems, notably gynecologic tumors, it has become clear that p53 staining can represent wild-type protein, and only specific staining patterns (absent, overexpression, or cytoplasmic expression) are indicative of a TP53 mutation. We applied these principles to two tissue microarrays containing 94 mesotheliomas and 66 reactive mesothelial proliferations. Seven/65 (11%) epithelioid mesotheliomas showed aberrant staining (four absent and three overexpression patterns) as did 5/29 (17%) of sarcomatoid mesotheliomas (all overexpression patterns). We sequenced the TP53 gene (exons 2–11) in five of the epithelioid and three of the sarcomatoid cases with aberrant staining as well as 12 epithelioid and eight sarcomatoid mesotheliomas with wild-type staining. All three sarcomatoid cases with aberrant staining showed mutated TP53, as did three of the epithelioid cases; in two of the epithelioid cases no mutation was detected, most likely because of large deletions not detected by this assay. In contrast, none of the 20 mesotheliomas with wild-type staining contained mutated TP53. We conclude that absent or overexpression p53 staining patterns can be used as a marker of a malignant vs. a benign mesothelial proliferation. The sensitivity of p53 staining by itself is low, but here addition of p53 to BAP1/MTAP staining increased sensitivity from 72 to 81% for epithelioid and 38 to 50% for sarcomatoid mesotheliomas.
Similar content being viewed by others
Introduction
Distinguishing benign and malignant mesothelial proliferations can be difficult based on histology alone. Existing immunohistochemical markers that may aid in this distinction have limited sensitivity, fueling interest in the characterization of potential new markers. p53 is a tumor suppressor commonly altered in malignancy and was one of the earlier markers investigated in mesothelioma. However, driver mutations in p53 are now understood to have a diversity of effects on protein abundance and localization that were not considered in early p53 studies (see below and Discussion).
p53 is involved in initiating apoptosis, DNA damage repair and cell cycle arrest. Genetic alterations that result in the absence of p53 protein, impair the nuclear localization of p53, or have a dominant negative effects on p53 function, may contribute to tumorigenesis1. p53 missense mutations often confer degradation resistance, resulting in the accumulation of p53 protein. Such accumulation may result in strong and diffuse p53 staining on immunohistochemistry (IHC).
Recent studies have considered intense p53 staining in >80% of tumor cells (overexpression pattern), complete loss of p53 staining (absent or null pattern), or diffuse cytoplasmic rather than nuclear p53 staining (cytoplasmic pattern) as an indication of aberrant p53 protein1,2,3. In contrast, early studies of p53 immunostaining in mesothelioma used low thresholds for positivity (e.g., cases with staining of any intensity in >10% of tumor cells were considered “positive”)4,5. However, a considerable proportion of cases with p53 staining in >10% of tumor cells had survival beyond that expected for patients with mesothelioma6, casting doubt on the specificity of >10% p53 staining as a marker of malignancy. More recently, genetic alterations affecting TP53 have been found in only 6–29% of mesotheliomas7,8,9,10,11,12,13, a fraction much lower than the 58% or greater proportion reported in some early studies to have positive p53 staining14.
It remains unclear what proportion of mesotheliomas and reactive mesothelial proliferations actually have TP53 alterations and whether p53 IHC can be a good surrogate marker when using modern criteria. Here we assessed p53 immunostaining in benign and malignant mesothelial proliferations and correlated results with TP53 sequencing data.
Methods
Patient samples and tissue microarray construction
This study was approved by the University of British Columbia Research Ethics Board. The cohort includes retrospectively identified samples from the archives of Vancouver General Hospital and the consult files of AC. We included 65 epithelioid mesotheliomas, 29 sarcomatoid mesotheliomas, 41 reactive epithelioid mesothelial proliferations, and 25 reactive spindle cell mesothelial proliferations. The diagnoses of mesothelioma were confirmed with appropriate clinical follow-up and IHC. Tissue microarrays (TMAs) were built on a receiver paraffin block using duplicate 0.6 mm cores from each case.
Immunohistochemistry protocols and interpretation
We used 4 µm sections from TMAs and performed p53 IHC on an Omnis Autostainer (Dako, Santa Clara CA, USA) using the DO-7 clone (Dako #M7001) at 1:500 for 20 min. Heat induced epitope retrieval used high pH buffer for 30 min, mouse-rabbit linker was applied for 10 min, and polymer applied for 20 min. BAP1 and MTAP IHC was performed as described previously15.
The interpretation of the p53 IHC was done according to previously established and validated criteria in the gynecologic pathology literature. Wild-type (WT) pattern was defined as low intensity nuclear staining ≥1% and <80%, while complete absence of staining (with positively staining background tissue as an internal control) and strong nuclear or cytoplasmic staining (overexpressor) in ≥80% of tumor cells were considered aberrant, as per p53 staining interpretation in gynecological malignancies1,3,16. The p53 staining pattern observed in TMA sections was subsequently confirmed on whole slide sections for all cases on which TP53 sequencing was performed.
TP53 sequencing
DNA for TP53 sequencing was extracted from formalin-fixed, paraffin-embedded (FFPE) tumor tissue sections using the standard QIAamp FFPE DNA Extraction Kit (Qiagen, Germantown, MD, USA) protocol. Primer sets for Sanger sequencing and next-generation sequencing (NGS) strategies can be found in Supplementary Table 1. Primers were designed to cover the entire coding sequence of TP53 (exon 2 to 11). PCR products were amplified using QuantStudio 6 Flex Real-Time PCR System (ThermoFisher Scientific, Waltham, MA, USA) with 2.5 ng of DNA. Amplicons were pooled on a per sample basis and each sample pool was barcoded with unique indexes. Following indexing, all samples were pooled equimolar for sequencing on a MiSeq Instrument (Illumina, San Diego, CA, USA) using a 300 cycle v2 sequencing kit. Mutations were called across primer sets and manually verified in bam files to ensure at least two (of three) amplicons contained the variant of interest. All variants were validated by Sanger sequencing. Predicted effects of missense variants on protein function were assessed using the International Agency for Research on Cancer TP53 Database (http://p53.iarc.fr/TP53GeneVariations.aspx), and splice site alterations were interpreted according to data in the ClinVar database16. Only pathogenic variants are reported.
Results
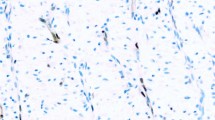
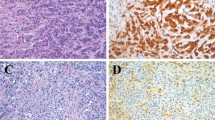
The distribution of staining by percent of positive cells and staining intensity is shown in Supplementary Table 2. Aberrant p53 immunostaining was identified using TMA sections in 7 (11%) out of 65 epithelioid mesotheliomas and 5 (17%) of 29 sarcomatoid mesotheliomas (Table 1). Of the epithelioid mesotheliomas, four had complete loss of p53 staining and three had strong diffuse nuclear staining. Of the aberrantly staining sarcomatoid mesotheliomas, all five had strong diffuse nuclear staining. An example of the various staining patterns is shown in Fig. 1. No cytoplasmic staining was identified in any mesothelioma. None of the 66 benign reactive mesothelial proliferations (41 epithelioid and 25 spindle cell) showed aberrant p53 staining; all showed low intensity (ie, wild type) positivity in varying fractions of cells (Supplementary Table 2). Thus, while the specificity of aberrant p53 staining was 100% for malignant versus benign mesothelial proliferations, the sensitivity for epithelioid mesothelioma was only 11%, and sensitivity for sarcomatoid mesothelioma only 17% (this calculation is made on the assumption that the IHC staining is an accurate surrogate for genomic analysis, as our results described below suggest).
Shown are (A) wild type staining (B) absent staining (null) and (C) strong diffuse staining (overexpressor) in epithelioid mesotheliomas, and (D) wild type and (E) strong diffuse staining (overexpressor) in sarcomatoid mesotheliomas. 200×. Absent or strong diffuse staining confirms that the process is malignant. Wild-type low intensity staining cannot be used to separate benign from malignant.
In comparison, BAP1 and MTAP were lost in 31/54 (57%) and 3/26 (12%) of the epithelioid mesotheliomas, and 16/54 (30%) and 9/26 (35%) of the sarcomatoid mesotheliomas, respectively (14 cases did not have interpretable tissue remaining for BAP1 and MTAP IHC). Of the seven epithelioid mesotheliomas with aberrant p53 staining, five retained both BAP1 and MTAP staining, and two showed loss of MTAP but retention of BAP1 staining. Of the five sarcomatoid mesotheliomas with aberrant p53 staining, three retained BAP1 and MTAP, one had loss of BAP1 and MTAP staining, and one did not have tissue remaining. The combined sensitivity of p53, BAP1, and MTAP IHC was therefore 81% for epithelioid mesothelioma and 50% for sarcomatoid mesotheliomas. This represents a modest increase over the 72% and 38% sensitivity of combined BAP1 and MTAP IHC for epithelioid and sarcomatoid mesothelioma, respectively.
We then sought to confirm that our interpretation of p53 IHC correctly identified cases with TP53 mutations. We performed whole slide p53 IHC on the eight aberrantly staining samples with available tissue and on 20 randomly selected mesotheliomas with WT p53 staining. In all cases the whole slide staining pattern was consistent throughout the section and concordant with the TMA results (Supplementary Fig. 1).
We then sequenced all coding exons for TP53 (exon 2 to 11) in the cases on which whole sections had been performed. TP53 mutations were identified in all 4 sequenced mesotheliomas that had strong diffuse p53 staining (three sarcomatoid and one epithelioid, Table 2). TP53 mutations were also identified in 2 of the 4 sequenced mesotheliomas with complete loss of p53 staining (all epithelioid, Table 2). Variant allele frequencies ranged from 0.283 to 0.551. The identified mutations included three missense mutations predicted to impact protein function, a splice site mutation and a frameshift mutation. One sarcomatoid mesothelioma with strong diffuse p53 staining had an in-frame trinucleotide deletion with unknown effects on protein function. The missense mutations affecting protein function were all identified in cases with strong diffuse p53 staining, whereas the splice site and frameshift mutations were identified in cases with loss of p53 staining. The case with a frameshift mutation also had a missense variant on the same allele that was not predicted to affect function. No TP53 mutations were detected in the 20 sequenced mesotheliomas with WT p53 staining (8 sarcomatoid and 12 epithelioid).
The concordance between p53 IHC and TP53 sequencing was excellent, with only two disagreements among 29 cases (7%). Both of the latter cases had an absent p53 pattern, but no mutation was identified using our sequencing assay. As the sequencing methodology used would not detect deletions of one or more exons (typically associated with the absent p53 pattern) the discordance described in these two cases was not unexpected17,18.
Discussion
p53 was among the earliest immunohistochemical markers investigated for distinguishing benign and malignant mesothelial proliferations. While studies performed between 1992 and 2003 showed promising results (e.g., pooled data from ten studies showed 91% specificity and 58% sensitivity)14, these studies had considerable variability in results: the proportion of cases with positive staining ranged from 25 to 97% for mesothelioma and 0 to 82% for benign reactive mesothelial proliferations5,19,20,21,22,23,24,25,26 Variation in technical factors and interpretation likely contributed to this wide range of results, with most studies considering either any staining or staining in a small proportion of cells (e.g., >10%5,6) to be a “positive” result suggestive of malignancy. Poor correlation with clinical outcomes ultimately suggested that these interpretations of p53 staining resulted in inadequate specificity6, and in all likelihood, most of the reported staining in these publications was wild type or a mixture of mostly wild type with a few cases carrying mutations.
Interpretation of p53 immunostaining has since evolved with increasingly sensitive IHC systems and with the understanding of the different staining patterns indicative of TP53 mutations. This new approach, however, had never been validated in a cohort of mesotheliomas. Our reevaluation of p53 staining using modern scoring criteria found aberrant p53 staining to have a specificity of 100% for mesothelioma vs. benign mesothelial proliferations, but sensitivities of only 11% for epithelioid mesothelioma and 17% for sarcomatoid mesothelioma. The low sensitivity of aberrant p53 staining limits its diagnostic utility as a stand-alone test, particularly given the higher sensitivity of other markers used for this purpose. For example, BAP1 IHC has a sensitivity of ~70% for epithelioid mesothelioma and <20% for sarcomatoid mesothelioma, MTAP IHC has a sensitivity of 33–65% for epithelioid mesothelioma and up to 86% for sarcomatoid mesothelioma, and PD-L1 has a sensitivity of 46–71% for sarcomatoid mesothelioma27,28. Here the addition of p53 to BAP1/MTAP IHC increased sensitivity for epithelioid mesothelioma from 72 to 81%, and for sarcomatoid mesothelioma from 38 to 50%. These increases are fairly small, but p53 IHC does provide an answer in some cases and is potentially useful, since many laboratories routinely run p53 IHC.
The frequency of TP53 mutations we identified is in keeping with sequencing studies of mesotheliomas identifying TP53 mutations in only 6–29% of cases7,8,9,10,11,12,13. Also consistent with our findings, the TP53 mutations identified in prior sequencing studies tended to be ~60–70% missense mutations7,8,10, with sarcomatoid mesothelioma having slightly more frequent TP53 mutations than epithelioid mesothelioma (i.e., 12% of sarcomatoid vs. 5% of epithelioid)13.
Large sequencing studies of malignant pleural mesotheliomas showed that TP53 mutations are associated with a significantly shorter overall survival8,13,29. However these sequencing assays are slow, expensive, and usually not available to clinicians and pathologists, unlike p53 IHC which is readily available in many laboratories. Until now p53 IHC was considered unreliable but our results show that p53 IHC can be an efficient surrogate for TP53 mutation status. We propose that p53 IHC could be used to further study the role of TP53 mutations in mesotheliomas and, potentially, as a prognostic tool. However, we caution that accurate interpretation of p53 IHC requires appropriate optimization of the assay, and delayed fixation leads to loss of p53 staining. The identification of wild-type p53 staining in benign background tissue is valuable for ensuring that the staining pattern in tumor cells is not due to technical errors, and as pointed out by Kobel et al.30 in regard to gynecologic malignancies, the staining must be considerably more intense than the background wild-type staining to be counted as positive.
The two cases in our series without detected TP53 mutations may have undetected mutations outside of the sequenced region (exon 2–11 only) or large deletions spanning the TP53 locus that were not detectable using our assay. Alternatively, p53 protein may be undetectable secondary to alterations affecting its upstream regulators. For instance, ~20% of mesotheliomas have detectable levels of MDM231, an E3 ubiquitin ligase whose activity promotes p53 degradation and may result in loss of p53 staining11.
As has been shown for other malignancies16, we found that strong diffuse p53 staining was associated with missense mutations (which may prevent ubiquitination and degradation of p53), whereas loss of p53 staining was associated with frameshift or splice site mutations (i.e., loss of function mutations). In theory, transcripts with TP53 loss of function mutations towards the 3′ end of the gene may escape nonsense mediated decay, and the resulting truncated protein may still be recognizable by p53 antibody. Thus, not all cases with loss of function TP53 mutations may have loss of p53 staining3. This mechanism is thought to underlie the only 76% sensitivity of absent p53 staining for loss of function mutations in ovarian carcinoma3. Nonetheless, both cases with loss of function TP53 mutations in our study had absent p53 staining.
We conclude that when using modern criteria, aberrant p53 staining is present in a much smaller proportion of epithelioid and sarcomatoid mesotheliomas than was previously reported. While aberrant p53 staining is highly specific for malignant versus benign mesothelial proliferations and highly sensitive for TP53 mutations, the diagnostic utility of p53 IHC is limited by the low frequency of p53 alterations in mesothelioma; however, p53 IHC can be a useful adjunct to BAP1 or MTAP1 IHC.
Data availability
All data generated or analyzed during this study are included in this published article and the Supplementary Data Files.
References
Singh, N. et al. p53 immunohistochemistry is an accurate surrogate for TP53 mutational analysis in endometrial carcinoma biopsies. J. Pathol. 250, 336–345 (2020).
Köbel, M. et al. An immunohistochemical algorithm for ovarian carcinoma typing. Int. J. Gynecol. Pathol. 35, 430–441 (2016).
Köbel, M. et al. Optimized p53 immunohistochemistry is an accurate predictor of TP53 mutation in ovarian carcinoma. J. Pathol. Clin. Res. 2, 247–258 (2016).
Hasteh, F., Lin, G. Y., Weidner, N. & Michael, C. W. The use of immunohistochemistry to distinguish reactive mesothelial cells from malignant mesothelioma in cytologic effusions. Cancer Cytopathol. 118, 90–96 (2010).
Mangano, W. E., Cagle, P. T., Churg, A., Vollmer, R. T. & Roggli, V. L. The diagnosis of desmoplastic malignant mesothelioma and its distinction from fibrous pleurisy: a histologic and immunohistochemical analysis of 31 cases including p53 immunostaining. Am. J. Clin. Pathol. 110, 191–199 (1998).
Churg, A. & Galateau-Salle, F. The separation of benign and malignant mesothelial proliferations. Arch. Pathol. Lab. Med. 136, 1217–1226 (2012).
Hmeljak, J. et al. Integrative molecular characterization of malignant pleural mesothelioma. Cancer Discov. 8, 1548–1565 (2018).
Markowitz, P. et al. Genomic characterization of malignant pleural mesothelioma and associated clinical outcomes. Cancer Treat. Res. Commun. 25, 100232 (2020).
Pagano, M. et al. Mutational profile of malignant pleural mesothelioma (MPM) in the phase II RAMES Study. Cancers 12, https://doi.org/10.3390/cancers12102948 (2020).
Campanella, N. C. et al. Mutational profiling of driver tumor suppressor and oncogenic genes in Brazilian malignant pleural mesotheliomas. Pathobiology 87, 208–216 (2020).
Yang, H., Xu, D., Schmid, R. A. & Peng, R.-W. Biomarker-guided targeted and immunotherapies in malignant pleural mesothelioma. Ther. Adv. Med. Oncol. 12, 1758835920971421 (2020).
Hung, Y. P. et al. Molecular characterization of diffuse malignant peritoneal mesothelioma. Mod. Pathol. 33, 2269–2279 (2020).
Quetel, L. et al. Genetic alterations of malignant pleural mesothelioma: association with tumor heterogeneity and overall survival. Mol. Oncol. 14, 1207–1223 (2020).
King, J., Thatcher, N., Pickering, C. & Hasleton, P. Sensitivity and specificity of immunohistochemical antibodies used to distinguish between benign and malignant pleural disease: a systematic review of published reports. Histopathology 49, 561–568 (2006).
Salisbury, T. & Churg, A. CD146 immunohistochemical staining for the separation of benign from malignant mesothelial proliferations. Virchows Arch. https://doi.org/10.1007/s00428-021-03077-7 (2021).
Köbel, M. et al. The biological and clinical value of p53 expression in pelvic high-grade serous carcinomas. J. Pathol. 222, 191–198 (2010).
Landrum, M. J. et al. ClinVar: improving access to variant interpretations and supporting evidence. Nucleic Acids Res. 46, D1062–D1067 (2018).
Tessier-Cloutier, B. et al. Major p53 immunohistochemical patterns in in situ and invasive squamous cell carcinomas of the vulva and correlation with TP53 mutation status. Mod. Pathol. 33, 1595–1605 (2020).
Attanoos, R. L., Griffin, A. & Gibbs, A. R. The use of immunohistochemistry in distinguishing reactive from neoplastic mesothelium. A novel use for desmin and comparative evaluation with epithelial membrane antigen, p53, platelet-derived growth factor-receptor, P-glycoprotein and Bcl-2. Histopathology 43, 231–238 (2003).
Roberts, F., Harper, C. M., Downie, I. & Burnett, R. A. Immunohistochemical analysis still has a limited role in the diagnosis of malignant mesothelioma. A study of thirteen antibodies. Am. J. Clin. Pathol. 116, 253–262 (2001).
Cury, P. M., Butcher, D. N., Corrin, B. & Nicholson, A. G. The use of histological and immunohistochemical markers to distinguish pleural malignant mesothelioma and in situ mesothelioma from reactive mesothelial hyperplasia and reactive pleural fibrosis. J. Pathol. 189, 251–257 (1999).
Kafiri, G. et al. p53 expression is common in malignant mesothelioma. Histopathology 21, 331–334 (1992).
Mayall, F. G., Goddard, H. & Gibbs, A. R. p53 immunostaining in the distinction between benign and malignant mesothelial proliferations using formalin-fixed paraffin sections. J. Pathol. 168, 377–381 (1992).
Ramael, M. et al. Immunoreactivity for p53 protein in malignant mesothelioma and non-neoplastic mesothelium. J. Pathol. 168, 371–375 (1992).
Cagle, P. T., Brown, R. W. & Lebovitz, R. M. p53 immunostaining in the differentiation of reactive processes from malignancy in pleural biopsy specimens. Hum. Pathol. 25, 443–448 (1994).
Esposito, V. et al. p53 immunostaining in differential diagnosis of pleural mesothelial proliferations. Anticancer Res. 17, 733–736 (1997).
Churg, A. & Naso, J. R. The separation of benign and malignant mesothelial proliferations: new markers and how to use them. Am. J. Surg. Pathol. 44, e100–e112 (2020).
Derakhshan, F., Ionescu, D., Cheung, S. & Churg, A. Use of programmed death ligand-1 (PD-L1) staining to separate sarcomatoid malignant mesotheliomas from benign mesothelial reactions. Arch. Pathol. Lab. Med. 144, 185–188 (2020).
Bueno, R. et al. Comprehensive genomic analysis of malignant pleural mesothelioma identifies recurrent mutations, gene fusions and splicing alterations. Nat. Genet. 48, 407–416 (2016).
Köbel, M. et al. Interpretation of p53 immunohistochemistry in endometrial carcinomas: toward increased reproducibility. Int. J. Gynecol. Pathol. 38(Suppl 1), S123–S131 (2019).
Mairinger, F. D. et al. Mdm2 protein expression is strongly associated with survival in malignant pleural mesothelioma. Future Oncol. 10, 995–1005 (2014).
Funding
This study was supported by the University of British Columbia Department of Pathology Residency Training Program.
Author information
Authors and Affiliations
Contributions
A.C. conceived of the study and contributed to data analysis and paper review and revision. J.R.N., J.S., and B.T.C. performed data analysis and drafted the paper. D.G.H. and A.C. provided technical and material support. All authors read and approved the final paper.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Ethics approval
This study was approved by the University of British Columbia Research Ethics Board.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Naso, J.R., Tessier-Cloutier, B., Senz, J. et al. Significance of p53 immunostaining in mesothelial proliferations and correlation with TP53 mutation status. Mod Pathol 35, 77–81 (2022). https://doi.org/10.1038/s41379-021-00920-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41379-021-00920-9
This article is cited by
-
Mesothelioma: morphologic and immunohistochemical findings
Die Pathologie (2024)
-
Clinical and molecular validation of BAP1, MTAP, P53, and Merlin immunohistochemistry in diagnosis of pleural mesothelioma
Modern Pathology (2022)
-
Micro-RNA-215 and -375 regulate thymidylate synthase protein expression in pleural mesothelioma and mediate epithelial to mesenchymal transition
Virchows Archiv (2022)