Abstract
Traditional pathology approaches have played an integral role in the delivery of diagnosis, semi-quantitative or qualitative assessment of protein expression, and classification of disease. Technological advances and the increased focus on precision medicine have recently paved the way for the development of digital pathology-based approaches for quantitative pathologic assessments, namely whole slide imaging and artificial intelligence (AI)–based solutions, allowing us to explore and extract information beyond human visual perception. Within the field of immuno-oncology, the application of such methodologies in drug development and translational research have created invaluable opportunities for deciphering complex pathophysiology and the discovery of novel biomarkers and drug targets. With an increasing number of treatment options available for any given disease, practitioners face the growing challenge of selecting the most appropriate treatment for each patient. The ever-increasing utilization of AI-based approaches substantially expands our understanding of the tumor microenvironment, with digital approaches to patient stratification and selection for diagnostic assays supporting the identification of the optimal treatment regimen based on patient profiles. This review provides an overview of the opportunities and limitations around implementing AI-based methods in biomarker discovery and patient selection and discusses how advances in digital pathology and AI should be considered in the current landscape of translational medicine, touching on challenges this technology may face if adopted in clinical settings. The traditional role of pathologists in delivering accurate diagnoses or assessing biomarkers for companion diagnostics may be enhanced in precision, reproducibility, and scale by AI-powered analysis tools.
Similar content being viewed by others
Introduction
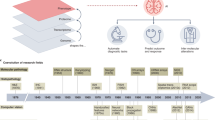
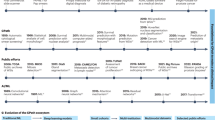
Pathology has historically played a crucial role in the drug development process, including preclinical research to facilitate target identification, define drug mechanism of action and pharmacodynamics, and enable toxicology assessments1,2. More recently, pathology has formed a bridge between drug discovery, translational, and clinical research programs that are striving to decipher disease pathophysiology in the context of the mechanism of action, patient selection, or patient stratification (Fig. 1)3,4. Such insights form the basis of novel hypotheses that can further be explored in drug discovery programs or applied to inform clinical trial design, thereby improving the probability of technical and regulatory success.
Dx diagnostic, EGFR epidermal growth factor receptor, FISH fluorescence in situ hybridization, HER2 human epidermal receptor 2, IHC immunohistochemistry, IVD in vitro diagnostic, MOA mechanism of action, NGS next-generation sequencing, PD pharmacodynamics, PD-L1 programmed death-ligand 1, TME tumor microenvironment.
Pathology-based assessments have been used to classify disease and determine efficacy in drug development across a variety of disease areas5,6,7. For example, during phase 2 trials for drug development in non-alcoholic steatohepatitis, the US Food and Drug Administration (FDA) considers evidence of efficacy on a histological endpoint to support initiation of phase 3 trials7. Additionally, pathological complete response (pCR) has been studied as a surrogate endpoint in patients with cancer for the prediction of long-term clinical benefit and favorable prognosis with the administration of neoadjuvant therapy8,9,10,11,12,13. More recently, pCR was associated with improved long-term efficacy in patients with human epidermal growth factor receptor 2 (HER2)-positive breast cancer treated with chemotherapy plus either intravenous or subcutaneous trastuzumab14. In the immuno-oncology (I-O) arena, immune-related pathologic response criteria have been applied retrospectively to surgical specimens from patients treated with immunotherapy in the neoadjuvant or advanced disease setting to predict survival in several tumor types15,16.
Immunohistochemistry (IHC) has been used to characterize biomarkers, such as programmed cell death-ligand 1 (PD-L1), and their association with clinical benefit. Traditional pathology techniques present several advantages, such as low cost, widespread availability, and application on formalin-fixed, paraffin-embedded (FFPE) tissue samples17, but challenges pertaining to differences in laboratory methods and subjective interpretation, particularly with the evaluation of immune cell staining, may lead to inter-observer variability18. This can produce inconsistency in diagnoses, which may impact treatment decisions19,20,21,22,23. While the use of IHC assays has led to better identification of patients who respond to I-O therapy24,25,26, there remains a need to more accurately quantify complex immune markers, including cell phenotypes in a spatial context, that require advanced quantitative tools to maximize the amount of information yielded from individual samples27,28.
Artificial intelligence (AI) applications in pathology improve quantitative accuracy and enable the geographical contextualization of data using spatial algorithms. Adding spatial metrics to IHC can improve the clinical value of biomarker identification approaches. For example, in a recent meta-analysis, the addition of spatial context to IHC, achieved using multiplex IHC and immunofluorescence (IF), was significantly better at predicting objective response to immune checkpoint inhibitors (ICIs) compared with gene expression profiling (GEP) or IHC alone28, indicating the need for more complex computational approaches to decipher the underlying biology and enhance clinical utility.
The development and integration of digital pathology and AI–based approaches provide substantive advantages over traditional methods, such as enabling spatial analysis while generating highly precise, unbiased, and consistent readouts that can be accessed remotely by pathologists29.
Advancing from traditional pathology to digital pathology
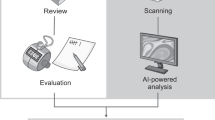
Efforts to overcome some of the challenges seen with traditional pathology methods have led to the development and adoption of complex, novel imaging systems and whole slide image (WSI) scanners that have enabled the transition of pathology into the digital era, also known as digital pathology. Within minutes, WSI scanners capture multiple images of entire tissue sections on the slide, which are digitally stitched together to generate a WSI that can be reviewed by a pathologist on a computer monitor (Fig. 2)30,31. Two scanners, Philips IntelliSite Pathology Solution (PIPS) (Philips, Amsterdam, Netherlands) and Leica Aperio AT2 DX System (Leica Biosystems, Buffalo Grove, Illinois, USA), are approved by the FDA for review and interpretation of digital surgical pathology slides prepared from biopsied tissue32,33.
There are many practical advantages to using these digital pathology image systems and solutions that would bring substantial benefits to translational and clinical research. These include the organization and storage of large amounts of data in a centralized location, integration of digital workflow software to help streamline processes and improve efficiency, convenient sharing of image data to enable cross-specialty worldwide remote communication, reduced testing turnaround time, and the generation of precise and highly reproducible tissue-derived readouts reducing inter-pathologist variability29,34,35,36,37. The increased speed and efficiency gained in image acquisition can enhance the downstream utilization options of traditional techniques such as hematoxylin and eosin (H&E), IHC, and in situ hybridization. These slides can be converted into a remotely available image within minutes and centrally reviewed by multiple pathologists from various sites29, with applications including education, research, consultation, and diagnostics29.
Recently, due to ongoing disruptions in relation to the COVID-19 pandemic, including remote working and restricted travel, digital pathology has been crucial in the continuation of clinical and academic research, as well as routine pathology services38. Without the need to transport glass slides and the ensuing logistical and safety concerns, central pathology review enables secure remote working38. Additionally, the utilization of digital images allows the generation of pixel-level pattern information, leading to expanded use of computational approaches that enable a quantitative analysis of WSIs39,40.
Improvements gained from digital pathology: quantitative analysis of the WSI
The use of digital image analysis in pathology can identify and quantify specific cell types quickly and accurately and can quantitatively evaluate histological features, morphological patterns, and biologically relevant regions of interest (e.g., tumoral or peritumoral areas, relationships between different immune cell populations, areas of expression, presence of metastasis)41,42. Quantitative image analysis tools also enable the capturing of data from tissue slides that may not be accessible during manual assessment via routine microscopy. Additionally, performing similar tasks manually can require significant time investment and can be prone to human error, such as counting fatigue43,44.
Expanding data capabilities: multiplex and multispectral imaging
Quantitative image analysis can also be used to generate high-content data through application to a technique known as multiplexing, which allows co-expression and co-localization analysis of multiple markers in situ with respect to the complex spatial context of tissue regions, including the stroma, tumor parenchyma, and invasive margin45,46. Current imaging metrics can utilize multispectral unmixing strategies to reveal co-expression patterns that define unique cell phenotypes and spatial relationships (Fig. 3)47.
a Monoplex slide stained for PD-L1, as seen on a monitor. b Monoplex slide stained for CD8. c Multiplex stain annotated using AI-based analysis allowing multiple marker identification. d Multispectral immunofluorescence. Legend indicates examples of possible phenotypes detected from this assay; however, many more are possible. CD cluster of differentiation, PD-1 programmed death-1, PD-L1 programmed death-ligand 1.
Automated classification of epithelial and immune cells and simultaneous marker analysis at the single-cell level has been conducted using prostate cancer, pancreatic adenocarcinoma, and melanoma tissue samples46,48,49. Application of this technique allowed identification of distinct T-cell populations and their spatial distributions and underscored the potential of immune markers to identify patients who may benefit from immunotherapy48,49.
While a highly multiplexed imaging platform can be used to understand intra- and inter-cellular signaling pathways by examining how phenotypically distinct cell populations are spatially distributed relative to one another, it is a time-consuming process applicable to a predefined region of interest50. However, as technology quickly advances, allowing digital evaluation of entire tissue slides, we are no longer confined to a region of interest37,51. The wealth of new information provided by these techniques has created a need for more consistent and reproducible interpretation of large and complex datasets, along with defining the interaction patterns between cell types and spatial context found in pathological images that define biological underpinnings37,52,53.
Advances in computational approaches: AI and machine learning
The need for data reproducibility and the increasing complexity of the analyses described above has led to the application of AI in pathology37,52,53. AI refers to a broad scientific discipline that involves using algorithms to train machines to extract information or features beyond human visual perception37,41,54. AI approaches are built to initially extract appropriate image representations and then to train a machine classifier for a particular segmentation, diagnostic, or prognostic task using a supervised or unsupervised approach37,41,54. The power of AI to analyze large amounts of data quickly can significantly speed up the discovery of novel histopathology features that may aid our understanding of or ability to predict how a patient’s disease will progress and how the patient will likely respond to a specific treatment37,39,55. In breast cancer, for example, unsupervised learning models have been used to generate histologic scores that can differentiate between low- and high-grade tumors and evaluate prognostically relevant morphological features from the epithelium and stroma of tissue samples to provide a score associated with the probability of overall survival56,57. The success of these AI-based approaches relies on the quality and quantity of the data used to train the algorithm, limiting the generalizability of these image analysis algorithms to larger or more complex datasets58.
Taking it further: deep learning networks
Deep learning takes machine learning a step further, using sophisticated, multilevel deep or convolutional neural networks (DNN or CNN) to create systems that perform feature classification from large datasets37,40,41,54. Figure 4 highlights key differences between machine learning and deep learning. The impact of applications of deep learning algorithms to IHC- and H&E-stained specimens have been well documented across many tumor types. These include grading prostate cancer59, identifying biomarkers for disease-specific survival in early-stage melanoma60, detection of invasive breast cancer regions on WSIs61,62, predicting response to chemoradiotherapy in locally advanced rectal cancer63, and identifying morphological features (nuclear shape, nuclear orientation, texture, tumor architecture, etc.) to predict recurrence in early-stage non-small cell lung cancer (NSCLC) from H&E slides64. Deep learning has also been used to construct entity-graph-based tissue representations, where cell morphology and topology are embedded within each node to effectively describe the phenotypical and structural properties of tissues and can be processed by graph neural networks (GNNs). GNNs therefore enhance the interpretability of pathological assessments gleaned from neural networks65,66.
It is important to compare AI-based interpretations with those of the pathologist to define the associated algorithm’s performance characteristics and utility. For example, when a CNN trained to classify melanoma samples was compared against manual scoring by histopathologists, the CNN was significantly superior in classifying images as malignant melanoma or benign nevi compared with manual assessment by histopathologists67. In the CAMELYON16 challenge, deep learning algorithms to detect breast cancer metastases in H&E-stained WSIs of lymph node sections performed similarly to the best performing pathologists under time constraints in detecting macrometastases and were better in detecting micrometastases68. However, it should be noted that the performance of any algorithm will depend on the task, due to the degree of accuracy required and the quality of the samples to be assessed37.
Another application of machine learning in the preclinical space is the assessment of tumor purity (TP). TP estimation, currently evaluated visually by pathologists, is used to ensure a signal is derived from cancer cells rather than other noncancerous cells that may be present in the TME based on tissue morphology when tissue is used to generate orthogonal data such as transcriptome or exome69,70. In a comparison of TP determined using AI (using deep learning algorithms generated on the PathAI platform) and manual estimates by pathologists, AI-assessed TP was found to be more accurate than visual assessment by pathologists71. Previous evidence has shown that immunosuppressive pathways are upregulated in patients with low TP, suggesting that low TP is associated with poor prognosis in some tumor types, including gastric cancer. Therefore, improved methods of evaluating TP may also aid in the identification of patients who may be suitable for immunotherapy72.
Given the amount of additional detail and insights that can be gained from combining WSI with machine learning algorithms, this technology can be readily applied to translational research. However, one major limitation of machine learning is the large amount of high-quality data required to develop these algorithms58. Data used for training need to be accurate and as complete as possible in order to maximize predictability and utility39. This can be challenging when histological data are obtained from various laboratories, leading to some variability due to factors such as differences in slide preparation (sectioning, fixation, staining, and mounting)73, scoring algorithms18, and inherent inter-observer variability74. These challenges become more apparent when more complex computational analytics methods are used for multiplexed imaging. Although AI could be used to overcome inter-reader variability across multiple institutions with the development of robust algorithms that take specific histological features of various tumors and subtypes into account75, further research is needed to fully understand the impact of these factors on the quality of AI data.
Applications of digital pathology in translational medicine
Enhancing our understanding of the TME
Tumor evolution and progression involve many complex cellulars and molecular interactions that are spatially and temporally regulated within the TME52. IHC can be used to gain insights into the composition of the TME by facilitating the identification of different cell types expressing a protein of interest and assessing the density and spatial distribution of specific biomarkers50. Digital pathology approaches, such as quantitative analysis of TILs, present an opportunity to gain greater insight into intra-tumor heterogeneity, spatial patterns of cell phenotypes, and the complex interactions between cancer and the immune system within the TME52,53. Image-based techniques can be used to determine immune cell responses to immunotherapy such as macrophage activation76 or lymphocyte infiltration by regulatory T cells (Tregs) into core tumor regions in solid tumors77, which may in turn have value as a predictive indicator for the effectiveness of ICIs. Favorable cancer prognosis has also been associated with factors in the TME, including high CD8+ TIL rates78,79. Recently, image analysis and AI methods have contributed to the development of novel approaches to concurrently assess multiple biomarkers in preclinical and exploratory studies, revealing complex interactions within the TME and providing the potential to improve cancer diagnosis and the selection of treatment regimens. Combining multiple techniques, such as multiplex IF, with image analysis has yielded important insights into specific immune cell populations, such as those in the TME of classical Hodgkin lymphoma, and their associations with PD-1/CTLA-4+/− T cells80. These studies require multiple large cohorts to add the scale and robustness necessary to gain these important insights, to elucidate relationships that may not be apparent to the human eye, and to help overcome observer bias that may mask potential biomarker signals.
Assessing treatment response: immune cell interactions in the TME
Digital pathology can also be used to gain insights into a receptor-ligand binding, as proximity may be indicative of receptor engagement and activation. For example, lymphocyte-activation gene 3 (LAG-3), expressed on exhausted T cells, principally interacts with major histocompatibility-II (MHC-II) molecules, expressed on the surface of antigen-presenting and tumor cells81,82. Spatial analysis in bladder and gastric cancer tumor cells has demonstrated that the density and proximity of LAG-3+ were significantly greater when associated with MHC II+ vs. MHC II− tumor cells, suggesting that LAG-3–expressing TILs may be preferentially located in proximity to MHC II+ tumor cells, allowing for LAG-3 activation and the inhibition of antitumor immunity83. The insights provided by digital pathology into the number and location of immune cells relative to tumor cells may provide information on immune response37,84, which could guide future treatment strategies. AI has also been used to quantify immune cells within the TME to define T-cell abundance and associated geographic localization in the tumor stroma, parenchyma, parenchyma-stromal interface, and invasive margin, which are then associated with transcriptomic factors to define underlying biological associations85.
Identifying genomic features
Additionally, AI-based approaches may find applications in translational medicine and clinical practice by predicting gene mutations from routine histopathology slides. With genomic tests being associated with high costs and high rates of failure due to stringent sample requirements86,87, AI may be particularly useful for evaluating genomic instability and the mutational landscape, with the possibility to assess pathologic and genomic features in conjunction with one another. A CNN trained with WSIs of H&E-stained hepatocellular carcinoma (HCC) tissue was used to predict the ten most common prognostic and mutated genes in HCC, with four of these (CTNNB1, FMN2, TP53, and ZFX4) correctly identified by the model88. Similar results were obtained when a DNN was trained to predict the most commonly mutated genes in lung adenocarcinoma, with 6 (STK11, EGFR, FAT1, SETBP1, KRAS, and TP53) being predicted from WSIs89. Deep learning has also been used to predict microsatellite instability (MSI) status from tumor tissue90. A CNN trained to classify MSI versus microsatellite stability was able to robustly distinguish features predictive of MSI in gastric and colorectal cancer samples90.
However, there are limitations to using AI for molecular classification. For example, current imaging techniques can only identify genetic variants when they directly impact tissue morphology, as described previously91. At the same time, AI algorithms cannot be applied in cases where actual variant allele frequencies of selected mutations can impact the classification and prognosis of individual diseases, such as hematologic myeloid neoplasms92.
Translating digital pathology into clinical practice
Potential for patient stratification
As a further application in translational medicine, digital pathology approaches have been used to predict response and identify patients most likely to respond to treatment. For example, studies have used spatial analysis to determine the response of patients with NSCLC to nivolumab therapy. These included training machine learning models to extract morphological details, such as the spatial arrangement of tumor nuclei and variance in shape and chromatin structure93, as well as the area and density of TILs and the proximity of TILs to each other and to tumor cells94. The features extracted from these models were able to distinguish patients who responded to nivolumab therapy93,94. In another example, digital image analysis was used to quantify CD8 and PD-L1 positive cell densities from patients treated with durvalumab across multiple tumor types95. Patients defined as positive for the CD8xPD-L1 composite signature had longer median survival compared with signature-negative patients, demonstrating the potential predictive value of digitally defined composite biomarkers.
AI and machine learning can also assist in classification and staging across various tumor types. A new approach to tumor subtyping has been developed based on a DNN (MesoNet) to predict OS of patients with mesothelioma from hematoxylin, eosin, and saffron stained WSIs, without any pathologist-provided annotations96. Results demonstrated that the model was more accurate in predicting patient survival than using current pathology practices and was able to identify regions contributing to patient outcomes96, suggesting that deep learning models can identify new features predictive of patient survival and potentially lead to new biomarker discoveries.
Application of digital pathology and AI algorithms in diagnostics
Biomarker research has been an area of particular interest in the I-O space due to its potential predictive value in some solid tumors25,26,97,98,99,100. ICIs, such as anti–PD-(L)1 and anti–cytotoxic T lymphocyte antigen-4, have been studied in multiple clinical trials, leading to improved prognosis for patients across various solid tumors101. Evidence has shown that PD-L1 expression may be indicative of response to ICI therapy in some tumor types25,26,97,98,99,100, while other studies have shown that patients demonstrated durable responses to ICIs regardless of PD-L1 expression3,102,103,104,105,106,107,108. Given the widespread clinical use of ICIs, predictive assays are needed to help stratify patients to determine who may benefit from such treatments.
While the use of these assays can help determine whether a patient will benefit from ICI therapy, biomarker identification, such as PD-L1 status, using tumor biopsies is challenging. Even when used by experienced pathologists, visual interpretation of PD-L1 using IHC is subjective and prone to error, which may contribute to inaccurate patient stratification. Digital scoring of PD-L1 expression can assist pathologists in overcoming these barriers by providing standardized metrics for biomarker assessment at single-cell resolution across whole tissue sections36.
Multiple studies have evaluated PD-L1 assessment using digital scoring and AI algorithms and have shown that digital-based techniques can perform better than or equal to manual pathological evaluation across various tumor types. A high correlation between AI and manual assessment of PD-L1 expression on tumor and immune cells has been observed in multiple CheckMate trials with samples from NSCLC, urothelial carcinoma, melanoma, and gastric cancer109,110,111. Furthermore, similar associations between PD-L1 expression and response to nivolumab have been reported between manual and digital scoring109,110. Using the combined positive score to assess PD-L1 expression on tumor and immune cells, digital image analyses and pathologists’ interpretations on stained slides (using the 22C3 pharmDx assay [Dako, Denmark]) demonstrated 33 (84.6%) of 39 cases had concordant results, and statistical analyses indicated that PD-L1 expression interpreted by pathologists or digital image analysis did not differ significantly for predicting responses to pembrolizumab112. Prospective clinical trials in colorectal cancer113 and NSCLC114 are also using digital image analyses to identify potential immune cell biomarkers within the TME.
The role of AI and machine learning in biomarker identification has been evaluated in studies outside of immunotherapy. For example, a DNN model (ConvNets) trained to automatically recognize cancer cell types were compared with conventional machine learning techniques. ConvNets achieved significantly higher accuracy than conventional algorithms, suggesting a role for computer-aided diagnosis to facilitate clinical decision-making115. Beyond oncology, AI and machine learning have been studied in the context of a morphological assessment of nonalcoholic steatohepatitis/nonalcoholic fatty liver disease and liver allograft fibrosis116,117. In these cases, AI-based methods were able to correctly reflect markers of steatotic severity116 and assess liver allograft fibrosis progression over time117.
Various platforms have been developed for the purpose of quantitative image analysis. Several have received FDA approval, including those used to detect HER2118. The goal of a HER2-directed image analysis platform is to detect and quantify HER2 membranous IHC staining of invasive breast cancer cells and to provide an accurate, precise, and reproducible quantitative HER2 result that can then be used to guide treatment decisions119. Digital image analysis has also been used to classify biological subtypes beyond HER2, including ER- and progesterone receptor (PR)–positive subtypes. Ahern et al demonstrated considerable overlap between unsupervised and supervised computational pathology platforms using image analysis to measure ER and PR expression in breast tumors between positive and negative groups, as classified by a pathologist120. While the supervised platform had a marginally higher performance than the unsupervised platform, both platforms provided meaningful results and may have important roles in future molecular epidemiology studies120.
Addressing consistency issues for application in clinical practice
There are several published resources for pathologists as well as for physicians, including guidelines, position papers, and directives relating to digital pathology30,31,37,121,122,123,124,125,126,127,128. These include detailed information on the handling of digital images in nonclinical121 and clinical122 settings, technical aspects and performance standards for WSI devices122,123,124, validation and quality assurance of digital pathology systems for nonclinical125 and clinical use30,122,126, AI concepts and best practices37,127, tutorials on using deep learning frameworks for image analysis128, and reimbursement considerations129. For example, the College of American Pathologists provides comprehensive guidelines to laboratories on validating their own WSI systems for clinical use, including emulation of the real-world environment, sample set size, establishing concordance using intra-observer variability, and documentation, among others30.
The performance of AI applications in digital pathology is largely dependent on the size and quality of the dataset used to train an algorithm41. Digital images used for training purposes should be obtained from multiple staining batches, scanners, and institutions to ensure generalizability. Such datasets should be curated by pathologists, ensuring that representative images have been obtained at an appropriate magnification and that all regions of interest are comprehensively annotated depending on the diagnostic application41. Crucially, the validation of AI algorithms developed for clinical purposes increases the concordance between manual and digital pathology interpretations. The role of pathologists in the validation step is equally important in order to ensure that datasets represent the sample type of interest (e.g., H&E-stained FFPE section), encompass the entirety of a glass slide, and are big enough to reveal potential interpretational discrepancies, as well as to evaluate the accuracy and performance of the algorithm30,119.
Adoption of digital pathology and AI: challenges and future considerations
Despite the advantages of incorporating digital pathology into the clinical setting, challenges remain (Table 1). Value determination and reimbursement structures for digital pathology are lacking. This leaves value interpretation, investment, and cost savings considerations up to individual laboratories, which is difficult and a substantial hinderance to widespread adoption. Image analysis platforms have been shown to provide prognostic value, such as risk classification in patients with colon cancer130. However, these are offered as single-site, standalone tests, thereby limiting their applicability to the wider pathology community. Studies that have evaluated the adoption of complete digital pathology workflows have shown increases in efficiency and operational utility131.
Technical concerns related to reproducibility, interpretability, the accuracy of competing devices, financial costs of processing hardware, and regulatory approvals that must accompany studies of clinical utility all represent barriers to adoption132. Some level of error with digital pathology is anticipated to be present at this point, and approaches that combine algorithm performance with manual validation, with margins of error similar to or stricter than those used for manual pathology, are likely to be the standard moving forward. This approach has already been tested in routine diagnostics, whereby pathologists interacted directly with an AI platform to conduct IHC-based intrinsic subtyping of breast cancers. The AI platform, both alone and working in consort with pathologists, was significantly more accurate in determining subtypes133. Additionally, translation and adoption into clinical practice will depend on algorithms being validated across many patient cohorts utilizing data not included in the training set. This will require large amounts of data to be acquired from multiple laboratories in order to assure the broad applicability required in a clinical setting39,134.
While there have been instances of AI being used in the clinical trial setting, most have been observational studies135. Techniques that take into account variations in real-world practice and can influence decision-making need to be evaluated in interventional studies to ascertain true clinical value134. Although a protocol for the development of a reporting guideline and risk of bias tool has been published136, no official guidelines are available yet on the numbers of annotations, images, and laboratories needed to capture the variation seen in the real-world. Additional statistical studies will be required for application to properly determine the optimal processes and workflows to ensure full implementation of this technology in clinical practice39. Algorithms would also be subject to periodic quality assurance (eg, when a new staining protocol is introduced), similar to how assays are revalidated when there is a change in workflow or procedure137. Various quality control (QC) techniques can be used to overcome preanalytical issues such as variations in slide preparation, origin, and scanner type. One approach is to train individual models of the same architecture to recognize specific variables73. Other approaches, such as combining image metrics in a QC application138, or transformation of image patches with synthetically generated artifacts139, can be used to train an algorithm to recognize different types of histological artifacts. Other unforeseen hurdles may exist once these systems are in place, including unfamiliarity with a new system and associated need for training, technical support, security, monitoring, and software integration140,141. In the US, software solutions should be developed under the FDA’s Quality System Regulation and Good Machine Learning Practices. However, artificial neural networks have been described as “black boxes”, whereby data can be difficult to interpret, which may lead to regulatory concerns, as image features are extracted in ways that are difficult for a human to understand127,142. Despite the challenges, the efficiency gains, such as faster results and higher throughput, are key motivators for pathologists to adopt digital pathology.
The benefits of AI can be seen across all stages of the drug development process and in the clinical setting143. One of the first applications of AI in the clinical setting is likely to be assessing multiple IHC I-O markers within a single tissue section. Application of image analysis to multiplexed IHC–stained samples offers accelerated scan times while increasing accuracy and productivity by automatically measuring parameters that may be hard to reliably achieve by eye47.
In the evolving field of digital pathology, a strategy towards the implementation of digital pathology may involve several phases culminating in the adoption of digitized images and AI technology in the clinic. A first step involves demonstrating the reliability of digital pathology with a biomarker that has shown clinical utility with manual pathology, such as approved complementary diagnostics. For example, using PD-L1 expression, which has demonstrated clinical utility across a range of tumor types26,100,144, would allow digital pathology readouts to be compared directly with manual pathology data and clinical outcomes. In this phase, pathologists would maintain a role in QC, but with improved efficiency. Data from the evaluation of such biomarkers with digital pathology could then be used in applications to the FDA for companion diagnostic status. Subsequent steps would introduce digital pathology as a diagnostic with novel biomarkers, with the aim of demonstrating the clinical utility of the biomarker with digital quantification. This phase would require the development of AI-based software for use in prospective clinical trials to evaluate the selected biomarker for patient stratification or selection. The next phase, and the long-term goal of digital pathology, would be to establish deep learning AI models trained using large quantities of data39 that can predict patient response and stratify patients using only WSIs.
Conclusions
The current advances in digital pathology offer practical advantages over manual pathology, including enhanced accuracy and precision, the ability for digital images to be uploaded and reviewed remotely by multiple pathologists, and the acquisition and processing of large and complex datasets. Within immuno-oncology, a deeper understanding of the complexity and underlying mechanisms of the TME can be achieved with the help of AI and machine learning, where datasets can be consistently analyzed and validated for application across many large cohorts, which may have implications for drug development and clinical trial design. AI and machine learning can then be utilized within the clinic to describe clinical and pathologic features across multiple patient samples. These advances will not only facilitate the entry of more precise I-O therapies, but also ultimately improve diagnostic, prognostic, and predictive clinical decision-making in cancer treatment.
References
Jubb, A. M., Koeppen, H. & Reis-Filho, J. S. Pathology in drug discovery and development. J. Pathol. 232, 99–102 (2014).
Kramer, J. A. The application of discovery toxicology and pathology towards the design of safer pharmaceutical lead candidates. Nat. Rev. Drug Discov. 6, 636–649 (2007).
Carbone, D. P. et al. First-line nivolumab in stage IV or recurrent non-small-cell lung cancer. N. Engl. J. Med. 376, 2415–2426 (2017).
Nagtegaal, I. D., West, N. P., van Krieken, J. H. J. M. & Quirke, P. Pathology is a necessary and informative tool in oncology clinical trials. J. Pathol. 232, 185–189 (2014).
Lehmann, B. D. & Pietenpol, J. A. Identification and use of biomarkers in treatment strategies for triple‐negative breast cancer subtypes. J. Pathol. 232, 142–150 (2013).
Potts, S. J. et al. Evaluating tumor heterogeneity in immunohistochemistry-stained breast cancer tissue. Lab. Invest. 92, 1342–1357 (2012).
Food and Drug Administration, Center for Drug Evaluation and Research. (2018) Noncirrhotic Nonalcoholic Steatohepatitis With Liver Fibrosis: Developing Drugs for Treatment Guidance for Industry, U.S. Department of Health and Human Services (ed) https://www.fda.gov/regulatory-information/search-fda-guidance-documents/noncirrhotic-nonalcoholic-steatohepatitis-liver-fibrosis-developing-drugs-treatment
Paluch-Shimon, S. et al. High efficacy of pre-operative trastuzumab combined with paclitaxel following doxorubicin & cyclophosphamide in operable breast cancer. Acta Oncol. 47, 1564–1569 (2008).
Coratazar, P. et al. Pathological complete response and long-term clinical benefit in breast cancer: the CTNeoBC pooled analysis. Lancet 384, 164–172 (2014).
Maas, M. et al. Long-term outcome in patients with a pathological complete response after chemoradiation for rectal cancer: a pooled analysis of individual patient data. Lancet Oncol. 11, 835–844 (2010).
Martin, S. T., Heneghan, H. M. & Winter, D. C. Systematic review and meta-analysis of outcomes following pathological complete response to neoadjuvant chemoradiotherapy for rectal cancer. Br. J. Surg. 99, 918–928 (2012).
Petrillo, M. et al. Prognostic role and predictors of complete pathologic response to neoadjuvant chemotherapy in primary unresectable ovarian cancer. Am. J. Obstet. Gynecol. 11, 632.e631–638 (2014).
Food and Drug Administration, Oncology Center of Excellence, Center for Drug Evaluation and Research, Center for Biologics Evaluation and Research. (2020) Pathologic complete response in neoadjuvant treatment of high-risk early-stage breast cancer: use as an endpoint to support accelerated approval, U.S. Department of Health and Human Services (ed)
Jackisch, C. et al. HannaH phase III randomised study: Association of total pathological complete response with event-free survival in HER2-positive early breast cancer treated with neoadjuvant-adjuvant trastuzumab after 2 years of treatment-free follow-up. Eur. J. Cancer 62, 62–75 (2016).
Cottrell, T. R. et al. Pathologic features of response to neoadjuvant anti-PD-1 in resected non-small-cell lung carcinoma: a proposal for quantitative immune-related pathologic response criteria (irPRC). Ann. Oncol. 29, 1853–1860 (2018).
Stein, J. E. et al. Pan-tumor pathologic scoring of response to PD-(L)1 blockade. Clin. Cancer Res. 26, 545–551 (2020).
Laurinavicius, A., Plancoulaine, B., Herlin, P. & Laurinaviciene, A. Comprehensive immunohistochemistry: digital, analytical and integrated. Pathobiology 83, 156–163 (2016).
Tsao, M. S. et al. PD-L1 immunohistochemistry comparability study in real-life clinical samples: results of Blueprint phase 2 project. J. Thorac. Oncol. 13, 1302–1311 (2018).
de Matos, L. L., Trufelli, D. C., de Matos, M. G. L. & da Silva Pinhal, M. A. Immunohistochemistry as an important tool in biomarkers detection and clinical practice. Biomark. Insights 5, 9–20 (2010).
Netto, G. J., Eisenberger, M. & Epstein, J. I. Interobserver variability in histologic evaluation of radical prostatectomy between central and local pathologists: findings of TAX 3501 multinational clinical trial. Urology 77, 1155–1160 (2011).
Gown, A. M. Diagnostic immunohistochemistry: what can go wrong and how to prevent it. Arch. Pathol. Lab. Med. 140, 893–898 (2016).
Brunnström, H. et al. PD-L1 immunohistochemistry in clinical diagnostics of lung cancer: inter-pathologist variability is higher than assay variability. Mod. Pathol. 30, 1411–1421 (2017).
Buisseret, L. et al. Reliability of tumor-infiltrating lymphocyte and tertiary lymphoid structure assessment in human breast cancer. Mod. Pathol. 30, 1204–1212 (2017).
Borghaei, H. et al. Nivolumab versus docetaxel in advanced nonsquamous non-small-cell lung cancer. N. Engl. J. Med. 373, 1627–1639 (2015).
Garon, E. B. et al. Pembrolizumab for the treatment of non–small-cell lung cancer. N. Engl. J. Med. 372, 2018–2028 (2015).
Schmid, P. et al. Atezolizumab and nab-paclitaxel in advanced triple-negative breast cancer. N. Engl. J. Med. 379, 2108–2121 (2018).
Rimm, D. L. et al. A prospective, multi-institutional, pathologist-based assessment of 4 immunohistochemistry assays for PD-L1 expression in non-small cell lung cancer. JAMA Oncol. 3, 1051–1058 (2017).
Lu, S. et al. Comparison of biomarker modalities for predicting response to PD-1/PD-L1 checkpoint blockade: a systematic review and meta-analysis. JAMA Oncol. 5, 1195–1204 (2019).
Mroz, P., Parwani, A. V. & Kulesza, P. Central pathology review for phase III clinical trials: the enabling effect of virtual microscopy. Arch. Pathol. Lab. Med. 137, 492–495 (2013).
Pantanowitz, L. et al. Validating whole slide imaging for diagnostic purposes in pathology: guideline from the College of American Pathologists Pathology and Laboratory Quality Center. Arch. Pathol. Lab. Med. 137, 1710–1722 (2013).
Zarella, M. D. et al. A practical guide to whole slide imaging: a white paper from the Digital Pathology Association. Arch. Pathol. Lab. Med. 143, 222–234 (2019).
Food and Drug Administration. (2017) IntelliSite Pathology Solution (PIPS, Philips Medical Systems) https://www.fda.gov/drugs/resources-information-approved-drugs/intellisite-pathology-solution-pips-philips-medical-systems (accessed 29/9/2020).
Food and Drug Administration. (2019) 510(k) Summary Aperio AT2 DX System, U.S. Department of Health and Human Services (ed)
Nam, S. et al. Introduction to digital pathology and computer-aided pathology. J. Pathol. Transl. Med. 54, 125–134 (2020).
Pell, R. et al. The use of digital pathology and image analysis in clinical trials. J. Pathol. Clin. Res. 5, 81–90 (2019).
Koelzer, V. H., Sirinukunwattana, K., Rittscher, J. & Mertz, K. D. Precision immunoprofiling by image analysis and artificial intelligence. Virchows Arch. 474, 511–522 (2019).
Aeffner, F. et al. Introduction to digital image analysis in whole-slide imaging: a white paper from the digital pathology association. J. Pathol. Inf. 10, 9 (2019).
Browning, L. et al. Digital pathology and artificial intelligence will be key to supporting clinical and academic cellular pathology through COVID-19 and future crises: the PathLAKE consortium perspective. J. Clin. Pathol. 74, 443–447 (2021).
Serag, A. et al. Translational AI and deep learning in diagnostic pathology. Front. Med. 6, 185 (2019).
Srinidhi, C. L., Ciga, O. & Martel, A. L. Deep neural network models for computational histopathology: a survey. Med. Image Anal. 67, 101813 (2021).
Bera, K., Schalper, K. A., Rimm, D. L., Velcheti, V. & Madabhushi, A. Artificial intelligence in digital pathology – new tools for diagnosis and precision oncology. Nat. Rev. Clin. Oncol. 16, 703–715 (2019).
Tumeh, P. C. et al. Liver metastasis and treatment outcome with anti-PD-1 monoclonal antibody in patients with melanoma and NSCLC. Cancer Immunol. Res. 5, 417–424 (2017).
Barisoni, L., Lafata, K. J., Hewitt, S. M., Madabhushi, A. & Balis, U. G. J. Digital pathology and computational image analysis in nephropathology. Nat. Rev. Nephrol. 16, 669–685 (2020).
Neltner, J. H. et al. Digital pathology and image analysis for robust high-throughput quantitative assessment of Alzheimer disease neuropathologic changes. J. Neuropathol. Exp. Neurol. 71, 1075–1085 (2012).
Dixon, A. R. et al. Recent developments in multiplexing techniques for immunohistochemistry. Expert Rev. Mol. Diagn. 15, 1171–1186 (2015).
Blom, S. et al. Systems pathology by multiplexed immunohistochemistry and whole-slide digital image analysis. Sci. Rep. 7, 15580 (2017).
Stack, E. C., Wang, C., Roman, K. A. & Hoyt, C. C. Multiplexed immunohistochemistry, imaging, and quantitation: a review, with an assessment of Tyramide signal amplification, multispectral imaging and multiplex analysis. Methods 70, 46–58 (2014).
Carstens, J. L. et al. Spatial computation of intratumoral T cells correlates with survival of patients with pancreatic cancer. Nat. Commun. 8, 15095 (2017).
Feng, Z. et al. Multispectral imaging of formalin-fixed tissue predicts ability to generate tumor-infiltrating lymphocytes from melanoma. J. Immunother. Cancer 3, 47 (2015).
Taube, J. M. et al. The Society for Immunotherapy of Cancer statement on best practices for multiplex immunohistochemistry (IHC) and immunofluorescence (IF) staining and validation. J. Immunother. Cancer 8, e000155 (2020).
Lopès, A. et al. Deciphering the immune microenvironment of a tissue by digital imaging and cognition network. Sci. Rep. 8, 16692 (2018).
Heindl, A., Nawaz, S. & Yuan, Y. Mapping spatial heterogeneity in the tumor microenvironment: a new era for digital pathology. Lab. Invest. 95, 377–384 (2015).
Yuan, J. et al. Novel technologies and emerging biomarkers for personalized cancer immunotherapy. J. Immunother. Cancer 4, 3 (2016).
Vamathevan, J. et al. Applications of machine learning in drug discovery and development. Nat. Rev. Drug Discov. 18, 463–477 (2019).
Barsoum, I., Tawedrous, E., Faragalla, H. & Yousef, G. M. Histo-genomics: digital pathology at the forefront of precision medicine. Diagnosis 6, 203–212 (2019).
Ganesan S., et al. Computerized histologic image-based risk score (IbRiS) classifier for ER+ breast cancer. Cancer Res. 69, 3046 (2009).
Beck, A. H. et al. Systematic analysis of breast cancer morphology uncovers stromal features associated with survival. Sci. Transl. Med. 3, 108ra113 (2011).
Tizhoosh, H. R. & Pantanowitz, L. Artificial intelligence and digital pathology: challenges and opportunities. J. Pathol. Inf. 9, 38 (2018).
Bulten, W. et al. Epithelium segmentation using deep learning in H&E-stained prostate specimens with immunohistochemistry as reference standard. Sci. Rep. 9, 864 (2019).
Kulkarni, P. M. et al. Deep learning based on standard H&E images of primary melanoma tumors identifies patients at risk for visceral recurrence and death. Clin. Cancer Res. 26, 1126–1134 (2020).
Cruz-Roa, A. et al. Accurate and reproducible invasive breast cancer detection in wholeslide images: a deep learning approach for quantifying tumor extent. Sci. Rep. 7, 46450 (2017).
Araújo, T. et al. Classification of breast cancer histology images using Convolutional Neural Networks. PLoS One 12, e0177544 (2017).
Zhang, F. et al. Predicting treatment response to neoadjuvant chemoradiotherapy in local advanced rectal cancer by biopsy digital pathology image features. Clin. Transl. Med. 10, e110 (2020).
Wang, X. et al. Prediction of recurrence in early stage non-small cell lung cancer using computer extracted nuclear features from digital H&E images. Sci. Rep. 7, 13543 (2017).
Pati P. et al. (2021) Hierarchical Graph Representations in Digital Pathology: Cornell University.
Adnan M., Kalra S. & Tizhoosh H. R. (2020) Representation Learning of Histopathology Images using Graph Neural Networks: Cornell University.
Hekler, A. et al. Deep learning outperformed 11 pathologists in the classification of histopathological melanoma images. Eur. J. Cancer 118, 91–96 (2019).
Ehteshami Bejnordi, B. et al. Diagnostic assessment of deep learning algorithms for detection of lymph node metastases in women with breast cancer. JAMA 318, 2199–2210 (2017).
Azimi V. et al. Breast cancer histopathology image analysis pipeline for tumor purity estimation. Proc. IEEE Int. Symp. Biomed. Imaging. 1137–1140 (2017).
Smits, A. J. J. et al. The estimation of tumor cell percentage for molecular testing by pathologists is not accurate. Mod. Pathol. 27, 168–174 (2014).
Lee G., Srinivasan S., Kalinava N., Sasson A. & Baxi V. Artificial intelligence-based tumor purity assessment of digitized histology samples in multiple tumor types from clinical trials of nivolumab. J. Immunother. Cancer 7, abstr. P738 (2019).
Gong, Z., Zhang, J. & Guo, W. Tumor purity as a prognosis and immunotherapy relevant feature in gastric cancer. Cancer Med. 9, 9052–9063 (2020).
Schmitt, M. et al. Hidden variables in deep learning digital pathology and their potential to cause batch effects: prediction model study. J. Med. Internet Res. 23, e23436 (2021).
Gavrielides M. A. et al. Pathologist Concordance for Ovarian Carcinoma Subtype Classification and Identification of Relevant Histologic Features Using Microscope and Whole Slide Imaging: A Multisite Observer Study. Arch. Pathol. Lab. Med. https://doi.org/10.5858/arpa.2020-0579-OA (2021). epub ahead of print.
Taube, J. M. et al. Multi-institutional TSA-amplified multiplexed immunofluorescence reproducibility evaluation (MITRE) study. J. Immunother. Cancer 9, e002197 (2021).
Pavillon, N., Hobro, A. J., Akira, S. & Smith, N. I. Noninvasive detection of macrophage activation with single-cell resolution through machine learning. Proc. Natl. Acad. Sci. USA 115, E2676–E2685 (2018).
Sun, R. et al. A radiomics approach to assess tumour-infiltrating CD8 cells and response to anti-PD-1 or anti-PD-L1 immunotherapy: an imaging biomarker, retrospective multicohort study. Lancet Oncol. 19, 1180–1191 (2018).
Sato, E. et al. Intraepithelial CD8+ tumor-infiltrating lymphocytes and a high CD8+/regulatory T cell ratio are associated with favorable prognosis in ovarian cancer. Proc. Natl. Acad. Sci. USA 102, 18538–18543 (2005).
Gao, Q. et al. Intratumoral balance of regulatory and cytotoxic T cells is associated with prognosis of hepatocellular carcinoma after resection. J. Clin. Oncol. 25, 2586–2593 (2007).
Patel, S. S. et al. The microenvironmental niche in classic Hodgkin lymphoma is enriched for CTLA-4–positive T cells that are PD-1–negative. Blood 134, 2059–2069 (2019).
Ostroumov, D., Fekete-Drimusz, N., Saborowski, M., Kuhnel, F. & Woller, N. CD4 and CD8 T lymphocyte interplay in controlling tumor growth. Cell. Mol. Life Sci. 75, 689–713 (2018).
Solinas, C., Migliori, E., De Silva, P. & Willard-Gallo, K. LAG3: The biological processes that motivate targeting this immune checkpoint molecule in human cancer. Cancers 11, 1213 (2019).
Hedvat, C. V. et al. Quantitative spatial profiling of lymphocyte-activation gene 3 (LAG-3)/major histocompatibility complex class II (MHC II) interaction in gastric and urothelial tumors. Ann. Oncol. 30, V809 (2019).
Steele, K. E. et al. Measuring multiple parameters of CD8+ tumor-infiltrating lymphocytes in human cancers by image analysis. J. Immunother. Cancer 6, 20 (2018).
Szabo P. M. et al. CD8+ T cells in tumor parenchyma and stroma by image analysis (IA) and gene expression profiling (GEP): potential biomarkers for immuno-oncology (I-O) therapy. J. Clin. Oncol. 37, abstr. 2592 (2019).
Feliubadalo, L. et al. Benchmarking of whole exome sequencing and ad hoc designed panels for genetic testing of hereditary cancer. Sci. Rep. 7, 37984 (2017).
Ascierto, P. A., Bifulco, C., Palmieri, G., Peters, S. & Sidiropoulos, N. Preanalytic variables and tissue stewardship for reliable next-generation sequencing (NGS) clinical analysis. J. Mol. Diagn. 21, 756–767 (2019).
Chen, M. et al. Classification and mutation prediction based on histopathology H&E images in liver cancer using deep learning. NPJ Precis. Oncol 4, 14 (2020).
Coudray, N. et al. Classification and mutation prediction from non-small cell lung scancer histopathology images using deep learning. Nat. Med. 24, 1559–1567 (2018).
Kather, J. N. et al. Deep learning can predict microsatellite instability directly from histology in gastrointestinal cancer. Nat. Med. 25, 1054–1056 (2019).
Shamai, G. et al. Artificial intelligence algorithms to assess hormonal status from tissue microarrays in patients with breast cancer. JAMA Netw. Open 2, e197700 (2019).
Sallman, D. A. & Padron, E. Integrating mutation variant allele frequency into clinical practice in myeloid malignancies. Hematol. Oncol. Stem Cell Ther. 9, 89–95 (2016).
Wang X. et al. Computer extracted features of cancer nuclei from H&E stained tissues of tumor predicts response to nivolumab in non-small cell lung cancer. J. Clin. Oncol. 36, 12061 (2018).
Barrera C. et al. Computer-extracted features relating to spatial arrangement of tumor infiltrating lymphocytes to predict response to nivolumab in non-small cell lung cancer (NSCLC). J. Clin. Oncol. 36, abstr. 12115 (2018).
Althammer, S. et al. Automated image analysis of NSCLC biopsies to predict response to anti-PD-L1 therapy. J. Immunother. Cancer 7, 121 (2019).
Courtiol, P. et al. Deep learning-based classification of mesothelioma improves prediction of patient outcome. Nat. Med. 25, 1519–1525 (2019).
Fehrenbacher, L. et al. Atezolizumab versus docetaxel for patients with previously treated non-small-cell lung cancer (POPLAR): a multicentre, open-label, phase 2 randomised controlled trial. Lancet 387, 1837–1846 (2016).
Herbst, R. S. et al. Pembrolizumab versus docetaxel for previously treated, PD-L1-positive, advanced non-small-cell lung cancer (KEYNOTE-010): a randomised controlled trial. Lancet 387, 1540–1550 (2016).
Reck, M. et al. Pembrolizumab versus chemotherapy for PD-L1-positive non-small-cell lung cancer. N. Engl. J. Med. 375, 1823–1833 (2016).
Rosenberg, J. E. et al. Atezolizumab in patients with locally advanced and metastatic urothelial carcinoma who have progressed following treatment with platinum-based chemotherapy: a single-arm, multicentre, phase 2 trial. Lancet 387, 1909–1920 (2016).
Yan, X. et al. Prognostic factors for checkpoint inhibitor based immunotherapy: an update with new evidences. Front. Pharm. 9, 1050 (2018).
Langer, C. J. et al. Carboplatin and pemetrexed with or without pembrolizumab for advanced, non-squamous non-small-cell lung cancer: a randomised, phase 2 cohort of the open-label KEYNOTE-021 study. Lancet Oncol. 17, 1497–1508 (2016).
Antonia, S. J. et al. Durvalumab after chemoradiotherapy in stage III non–small-cell lung cancer. N. Engl. J. Med. 377, 1919–1929 (2017).
Bellmunt, J. et al. Pembrolizumab as second-line therapy for advanced urothelial carcinoma. N. Engl. J. Med. 376, 1015–1026 (2017).
Gulley, J. L. et al. Avelumab for patients with previously treated metastatic or recurrent non-small-cell lung cancer (JAVELIN Solid Tumor): dose-expansion cohort of a multicentre, open-label, phase 1b trial. Lancet Oncol. 18, 599–610 (2017).
Horn, L. et al. Nivolumab versus docetaxel in previously treated patients with advanced non-small-cell lung cancer: two-year outcomes from two randomized, open-label, phase III trials (CheckMate 017 and CheckMate 057). J. Clin. Oncol. 35, 3924–3933 (2017).
Rittmeyer, A. et al. Atezolizumab versus docetaxel in patients with previously treated non-small-cell lung cancer (OAK): a phase 3, open-label, multicentre randomised controlled trial. Lancet 389, 255–265 (2017).
Janjigian, Y. Y. et al. CheckMate-032 study: efficacy and safety of nivolumab and nivolumab plus ipilimumab in patients with metastatic esophagogastric cancer. J. Clin. Oncol. 36, 2836–2844 (2018).
Duan C. et al. Association of digital and manual quantification of tumor PD-L1 expression with outcomes in nivolumab-treated patients. Cancer Res. 80, abstr. 2017 (2020).
Beck A. et al. An empirical framework for validating artificial intelligence-derived PD-L1 positivity predictions applied to urothelial carcinoma. J. Immunother. Cancer 7, abstr. P730 (2019).
Baxi V. et al. Artificial intelligence–powered retrospective analysis of PD-L1 expression in nivolumab trials of advanced non-small cell lung cancer. J. Immunother. Cancer 7, abstr. O65 (2019).
Kim, H.-N. et al. PD-L1 expression in gastric cancer determined by digital image analyses: pitfalls and correlation with pathologist interpretation. Virchows Arch. 476, 243–250 (2020).
ClinicalTrials.gov. Bethesda (MD) National Library of Medicine Identifier NCT03698461, Treatment of colorectal liver metastases with immunotherapy and bevacizumab (CLIMB); 2018 Oct 9 (accessed 24/8/2021). Available from: https://clinicaltrials.gov/ct2/show/NCT03698461.
ClinicalTrials.gov. Bethesda (MD) National Library of Medicine Identifier NCT03853187, Imaging tumor-infiltrating T-cells in non-small cell lung cancer (Donan); 2019 Feb 25 (accessed 24/8/2021). Available from: https://clinicaltrials.gov/ct2/show/NCT03853187.
Vandenberghe, M. E. et al. Relevance of deep learning to facilitate the diagnosis of HER2 status in breast cancer. Sci. Rep. 7, 45938 (2017).
Nativ, N. I. et al. Automated image analysis method for detecting and quantifying macrovesicular steatosis in hematoxylin and eosin-stained histology images of human livers. Liver Transpl. 20, 228–236 (2014).
Venturi, C. et al. Dynamics of allograft fibrosis in pediatric liver transplantation. Am. J. Transpl. 14, 1648–1656 (2014).
Food and Drug Administration. (2021) List of cleared or approved companion diagnostic devices (in vitro and imaging tools) https://www.fda.gov/medical-devices/in-vitro-diagnostics/list-cleared-or-approved-companion-diagnostic-devices-in-vitro-and-imaging-tools (accessed 2/4/2020).
Bui, M. M. et al. Quantitative image analysis of human epidermal growth factor receptor 2 immunohistochemistry for breast cancer: guideline from the College of American Pathologists. Arch. Pathol. Lab. Med. 143, 1180–1195 (2019).
Ahern, T. P. et al. Continuous measurement of breast tumour hormone receptor expression: a comparison of two computational pathology platforms. J. Clin. Pathol. 70, 428–434 (2017).
Tuomari, D. L. et al. Society of toxicologic pathology position paper on pathology image data: compliance with 21 CFR parts 58 and 11. Toxicol. Pathol. 35, 450–455 (2007).
Pantanowitz, L. et al. American Telemedicine Association clinical guidelines for telepathology. J. Pathol. Inf. 5, 39 (2014).
Food and Drug Administration, Center for Devices and Radiological Health, Division of Molecular Genetics and Pathology, Molecular Pathology and Cytology Branch. (2016) Technical Performance Assessment of Digital Pathology Whole Slide Imaging Devices: Guidance for Industry and Food and Drug Administration Staff, U.S. Department of Health and Human Services (ed)
Garcia-Rojo, M. International clinical guidelines for the adoption of digital pathology: a review of technical aspects. Pathobiology 83, 99–109 (2016).
Long, R. E. et al. Scientific and Regulatory Policy Committee (SRPC) paper: validation of digital pathology systems in the regulated nonclinical environment. Toxicol. Pathol. 41, 115–124 (2013).
Lowe A. et al. (2011) Validation of digital pathology in a healthcare environment. Madison, WI: Digital Pathology Association.
Abels, E. et al. Computational pathology definitions, best practices, and recommendations for regulatory guidance: a white paper from the Digital Pathology Association. J. Pathol. 249, 286–294 (2019).
Janowczyk, A. & Madabhushi, A. Deep learning for digital pathology image analysis: a comprehensive tutorial with selected use cases. J. Pathol. Inf. 7, 29 (2016).
Hipp J. et al. (2017) Digital Pathology Resource Guide. 7. Northfield, IL: College of American Pathologists.
Pages, F. et al. International validation of the consensus Immunoscore for the classification of colon cancer: a prognostic and accuracy study. Lancet 391, 2128–2139 (2018).
Retamero, J. A., Aneiros-Fernandez, J. & Del Moral, R. G. Complete digital pathology for routine histopathology diagnosis in a multicenter hospital network. Arch. Pathol. Lab Med. 144, 221–228 (2020).
Acs, B. & Rimm, D. L. Not just digital pathology, intelligent digital pathology. JAMA Oncol. 4, 403–404 (2018).
Bodén, A. C. S. et al. The human-in-the-loop: an evaluation of pathologists’ interaction with artificial intelligence in clinical practice. Histopathology 79, 210–218 (2021).
van der Laak, J., Litjens, G. & Ciompi, F. Deep learning in histopathology: the path to the clinic. Nat. Med. 27, 775–784 (2021).
Dong, J. et al. Clinical trials for artificial intelligence in cancer diagnosis: a cross-sectional study of registered trials in ClinicalTrials.gov. Front. Oncol. 10, 1629 (2020).
Collins, G. S. et al. Protocol for development of a reporting guideline (TRIPOD-AI) and risk of bias tool (PROBAST-AI) for diagnostic and prognostic prediction model studies based on artificial intelligence. BMJ Open 11, e048008 (2021).
US Federal Government. (2019) Code of Federal Regulations 42 – Public Health. In § 493.1273 – Standard: Histopathology: Government Publishing Office.
Janowczyk, A., Zuo, R., Gilmore, H., Feldman, M. & Madabhushi, A. HistoQC: an open-source quality control tool for digital pathology slides. JCO Clin. Cancer Inf. 3, 1–7 (2019).
Schömig-Markiefka B. et al. Quality control stress test for deep learning-based diagnostic model in digital pathology. Mod. Pathol. epub ahead of print (2021). https://www.nature.com/articles/s41379-021-00859-x
Randell, R., Ruddle, R. A. & Treanor, D. Barriers and facilitators to the introduction of digital pathology for diagnostic work. Stud. Health Technol. Inf. 216, 443–447 (2015).
Hanna, M. G. et al. Whole slide imaging equivalency and efficiency study: experience at a large academic center. Mod. Pathol. 32, 916–928 (2019).
Duran, J. M. & Jongsma, K. R. Who is afraid of black box algorithms? On the epistemological and ethical basis of trust in medical AI. J. Med. Ethics 47, 329–335 (2021).
Potts, S. J. Digital pathology in drug discovery and development: multisite integration. Drug Discov. Today 14, 935–941 (2009).
Büttner, R. et al. Programmed Death-Ligand 1 Immunohistochemistry Testing: A Review of Analytical Assays and Clinical Implementation in Non-Small-Cell Lung Cancer. J. Clin. Oncol. 35, 3867–3876 (2017).
Patidar S. (2018) Machine learning vs deep learning https://dzone.com/articles/comparison-between-deep-learning-vs-machine-learni (accessed 17/12/2020).
Gong, C. et al. Quantitative characterization of CD8+ T cell clustering and spatial heterogeneity in solid tumors. Front. Oncol. 8, 649 (2019).
Hamilton, P. W. et al. Digital pathology and image analysis in tissue biomarker research. Methods 70, 59–73 (2014).
Research Institute. (2019) Expensive, labour-Intensive, time-consuming: how researchers are overcoming barriers in machine learning https://medium.com/@1nst1tute/expensive-labour-intensive-time-consuming-how-researchers-overcome-barriers-in-machine-learning-4f686b2a1979 (accessed 16/12/2020).
Funding
This review was funded by Bristol Myers Squibb. Writing and editorial assistance were provided by Emily Motola, PharmD, and Matthew Weddig of Spark Medica Inc, funded by Bristol Myers Squibb, according to Good Publication Practice guidelines.
Author information
Authors and Affiliations
Contributions
All authors contributed to the writing and approval of this paper.
Corresponding author
Ethics declarations
Competing interests
V.B. and R.E. are employees of and own stock in Bristol Myers Squibb. M.M. is an employee of PathAI. S.S. is the CEO of Centessa Pharmaceuticals.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Baxi, V., Edwards, R., Montalto, M. et al. Digital pathology and artificial intelligence in translational medicine and clinical practice. Mod Pathol 35, 23–32 (2022). https://doi.org/10.1038/s41379-021-00919-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41379-021-00919-2
This article is cited by
-
Quantitative assessment of H&E staining for pathology: development and clinical evaluation of a novel system
Diagnostic Pathology (2024)
-
Association of the pathomics-collagen signature with lymph node metastasis in colorectal cancer: a retrospective multicenter study
Journal of Translational Medicine (2024)
-
Augmented interpretation of HER2, ER, and PR in breast cancer by artificial intelligence analyzer: enhancing interobserver agreement through a reader study of 201 cases
Breast Cancer Research (2024)
-
Flow cytometric analysis for Ki67 assessment in formalin-fixed paraffin-embedded breast cancer tissue
BMC Biology (2024)
-
Impact of functional electrical stimulation on nerve-damaged muscles by quantifying fat infiltration using deep learning
Scientific Reports (2024)