Abstract
Sarcomatoid mesothelioma is an aggressive malignancy that can be challenging to distinguish from benign spindle cell mesothelial proliferations based on biopsy, and this distinction is crucial to patient treatment and prognosis. A novel deep learning based classifier may be able to aid pathologists in making this critical diagnostic distinction. SpindleMesoNET was trained on cases of malignant sarcomatoid mesothelioma and benign spindle cell mesothelial proliferations. Performance was assessed through cross-validation on the training set, on an independent set of challenging cases referred for expert opinion (‘referral’ test set), and on an externally stained set from outside institutions (‘externally stained’ test set). SpindleMesoNET predicted the benign or malignant status of cases with AUC’s of 0.932, 0.925, and 0.989 on the cross-validation, referral and external test sets, respectively. The accuracy of SpindleMesoNET on the referral set cases (92.5%) was comparable to the average accuracy of 3 experienced pathologists on the same slide set (91.7%). We conclude that SpindleMesoNET can accurately distinguish sarcomatoid mesothelioma from benign spindle cell mesothelial proliferations. A deep learning system of this type holds potential for future use as an ancillary test in diagnostic pathology.
Similar content being viewed by others
Introduction
Malignant mesothelioma is an uncommon aggressive malignancy arising from the mesothelial lining of serosal surfaces such as the pleura and peritoneum. Accurate distinction of this usually lethal disease from similar appearing benign reactive processes is critical for appropriate patient care. Diagnosis relies on tissue biopsy, which allows classification into three general subtypes of mesothelioma: epithelioid, sarcomatoid and biphasic [1]. Morphologic variants that may be considered subtypes of sarcomatoid mesothelioma include desmoplastic (a very paucicellular pattern), and transitional (sheets of cohesive plump elongated cells) forms [2, 3].
Sarcomatoid mesotheliomas are characterised by spindle cell morphology and are associated with the most dismal prognosis (median survival 4-5 months). Many benign reactive mesothelial proliferations, generally referred to as organizing pleuritis, also show spindle cell morphology and a variety of histologic patterns that often closely mimic the appearance of sarcomatoid mesotheliomas in terms of cellularity, cytologic atypia, and microscopic organization [4]. Although features such as necrosis or invasion of adjacent tissues can be diagnostic, these features are not present in every case of sarcomatoid mesothelioma and may not be captured in small biopsies. A variety of ancillary immunohistochemical (loss of BAP1 or MTAP) or molecular genetic tests (e.g. homozygous loss of CDKN2A) are also useful in separating sarcomatoid mesotheliomas from benign reactions [5], but these have limited sensitivity and are not available in many laboratories.
Most practicing pathologists encounter few sarcomatoid mesotheliomas, which contributes to the challenge of their diagnosis. In contrast, benign spindle cell mesothelial proliferations commonly occur in reaction to benign pleural effusions, such that questions regarding the benign or malignant nature of such proliferations are relatively frequent. These cases are often sent for expert consultation, which delays definitive diagnosis and the initiation of time-sensitive treatment regimens. Furthermore even expert review panels may not reach a consensus on diagnosis.
Artificial intelligence systems using deep learning have aimed to prognostically stratify and morphologically subclassify tumors already known to be malignant mesothelioma [2, 6], but have not yet been applied to distinguishing benign and malignant mesothelial proliferations. An artificial neural network that accurately estimates the likelihood of a spindle-cell mesothelial proliferation being benign or malignant could be a valuable ancillary test for guiding diagnostic decision making, similar to the role that immunohistochemistry plays in the diagnostic pathology workflow. Identifying areas most predictive of malignant or benign status may also provide insight into which morphologic features pathologists can use to distinguish these entities.
In this study we demonstrate proof-of-concept that a deep learning approach can distinguish benign and malignant spindle cell mesothelial proliferations with sufficient accuracy to be of clinical value, and we interrogate the features used for prediction by the trained neural network. We demonstrate that a neural network labeled “SpindleMesoNET” produces highly accurate predictions for cross-validation and external validation sets. We suggest avenues through which SpindleMesoNET could be used to complement pathologist interpretations for more accurate and definitive diagnosis of these challenging cases.
Materials and methods
Whole slide image acquisition and annotation
This project was approved by the University of British Columbia Research Ethics Board (REB# H18-03646). All cases were reviewed by a pathologist with extensive experience in mesothelioma diagnosis (AC) to confirm that available histologic, immunohistochemical and clinical evidence was sufficient for a definitive diagnosis of malignant mesothelioma or a benign process. Hematoxylin and eosin-stained (H&E) slides were scanned on an Aperio AT2 scanner with 40x objective magnification.
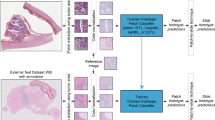
Three sets of cases were used, a training/cross-validation set, a referral test set, and an externally stained test set, each containing both benign and malignant cases. Cases for the training/cross-validation set were retrospectively identified from the archives of Vancouver General Hospital (VGH) and the consultation files of AC (Supplemental Table 1). The malignant training/cross-validation cases (n = 58) included 49 pleural biopsies, 3 pleural decortications, 1 lung biopsy, 1 lung lobectomy and chest wall excision, 1 hilar nodule excision, 1 paratracheal mass biopsy, 1 omental biopsy, and 1 peritoneal nodule biopsy. These cases included 16 sarcomatoid, 15 desmoplastic, 5 mixed sarcomatoid and desmoplastic, 20 biphasic, and 2 transitional mesotheliomas. Areas with a high confidence of malignancy, including cellular spindle cell areas, areas of fat invasion, muscle invasion, necrosis and transitional architecture were annotated by consensus of two pathologists. Cytologic atypia was generally not used, since it’s often difficult to discern in these tumors. Annotations excluded any areas of epithelioid mesothelioma and areas of hemorrhage, fibrin, and granulation tissue.
The benign training/cross-validation cases (n = 81) contained spindle cell mesothelial proliferations with an appearance such that a non-expert pathologist might consider malignant mesothelioma in the differential diagnosis. The benign cases included 48 samples of pleura (28 biopsies, 16 decortications and 4 biopsy and decortication) and 14 lung wedge resections from patients with a history of pleural effusion, empyema, fibrothorax, hemothorax, pneumothorax, restrictive lung disease, lung abscess, suspected tuberculosis or graft-vs-host disease. We also included 4 samples of omentum and 2 samples of pelvic peritoneum from patients with adnexal or endometrial lesions, 8 samples of normal lung from lung carcinoma resections, and 5 hernia sacs.
The referral test set was obtained from the consultation files of AC, and consisted entirely of cases referred for expert opinion regarding whether they showed a benign or malignant spindle cell mesothelial proliferation (n = 40). None of these cases was used in the training/cross-validation set so that SpindleMesoNET had no exposure to these cases prior to testing. The 19 benign referral test set cases were samples of pleura (14 biopsies, 4 decortications, and 1 pleurectomy) and the 21 malignant referral test set cases were pleural biopsies of 8 sarcomatoid, 4 desmoplastic, 3 mixed sarcomatoid and desmoplastic, 4 biphasic and 2 transitional mesotheliomas. Two of the referral test set cases were externally stained and the remainder were recuts stained in-house. The referral set cases were also evaluated by three very experienced pathologists, including two specialty trained in pulmonary pathology, who were independently asked to predict the benign or malignant status of each case, based solely on the histology of the same slide evaluated by SpindleMesoNET.
The externally stained test set cases consisted of 25 externally stained slides of pleural sarcomatoid mesothelioma and 14 benign pleural spindle cell mesothelial proliferations from two other institutions, selected to be good representations of benign and malignant histology.
All cases were confirmed as benign or malignant by followup and the mesothelial nature of the process confirmed by appropriate immunostains.
Image processing
Areas of good quality tissue on the whole slide image (WSI) were segmented, as described previously [7]. In brief, the WSI was downsampled by 16 from its maximum magnification using OpenSlide [8] (v3.4.1) and converted to greyscale. In-focus tissue was identified using a Laplacian filter applied with scikit-image [9]. The image was converted to a binary mask using Otsu’s method [10] and morphological closing was performed using a 50 μm ellipse-shaped element. Cases with either <8% or >70% recognised tissue (indicating a likely error in segmentation based on initial review of the tissue masks) had Otsu thresholding redone on the greyscale image without the Laplacian filter, followed by morphological closing. Connected components on the binary mask were identified, and objects and holes smaller than 100,000 μm were removed and filled, respectively. Finally, the image was converted to hue, saturation, value (HSV) space and objects with mean hue <0.68, corresponding to any remaining pen marks, were removed.
Image patches of size 512 × 512 pixels were extracted using a sliding window approach, and only images that contained a minimum amount of tissue or annotated region were saved (90% and 75%, respectively). Images were extracted at 40x objective magnification using strides of 512 pixels (no overlap). Training on tumor slides used only annotated regions, whereas testing on tumor slides used the entire slide image with only tissue segmentation applied.
Stain normalization was implemented prior to training using the Vahadane method [11] through an open-source python implementation (https://github.com/Peter554/StainTools), in reference to a single image taken from a Vancouver General Hospital surgical specimen. Color jitter was applied during CNN training using random shifts in the HSV space with maximum values as follows: brightness ± 0.1, saturation ± 0.1, hue ± 0.05.
Convolutional neural network (CNN) training and prediction
Model training was performed using five-fold cross-validation, which split patients into non-overlapping training and testing sets, with approximately one-fifth of patients allocated to testing for each split. All slides and images from the same patient were allocated to either testing or training. From the training set, 10% of the images were allocated to internal validation for monitoring training progression.
Convolutional neural network training was implemented in Pytorch [12] on a single GPU of the BC Genome Sciences Centre GPU cluster (an Nvidia Tesla V100, 2080Ti, or RTX5000). A ResNet18 [13] model was selected due to its high classification accuracy in initial experiments and because it had relatively few layers and parameters, reducing both training time and the risk of overfitting. The model was slightly modified, with the final fully connected layer having two features followed by a softmax layer. A stochastic gradient descent optimizer was used with learning rate = 0.001 and momentum = 0.9, and cross entropy loss was used without class weighting. To balance the training data, for each epoch, 1500 images were randomly selected from each case in the training set, for a total of approximately 150,000 images. Color jitter, as well as random horizontal/vertical flips and rotations (90, 180, or 270 degrees), were applied during training. The models were trained for 15 epochs, which took approximately 13-14 hours. The weights from the epoch with the best internal validation set performance were used for subsequent prediction using the network. Using the trained network, the probability of malignancy was predicted and saved for all patches in the benign and unannotated tumor image sets.
Patient level prediction
Patient level prediction used a multiple instance learning (MIL) paradigm [14] in which it was assumed that patients labeled ‘negative’ had only negative training instances (i.e. images), and patients labeled ‘positive’ had at least one positive instance. For our pooled implementation of MIL (MIL-pool), all image patches from a given patient were ranked by the softmax value, corresponding to probability of malignancy. The top 0.5% of patches were selected, with a minimum of 10 images used per patient. These values were then averaged, giving the predicted probability that a patient has malignant mesothelioma.
Our recurrent neural network (RNN) implementation of MIL (MIL-RNN) was based on a previously described model, with adjustments to account for the substantially smaller size of our training set [15]. WSIs were separated into regions of 5120 × 5120 pixels (corresponding to 100 image patches), each labeled as benign or malignant. The top 10 image patches (ranked by softmax value) in each region were used as input to train the RNN. Following RNN training, the network was used to predict the probability of malignancy for each region in the test set. The top 0.5% of regions were selected and averaged to give the probability of a patient having malignancy, with a minimum of 3 regions used per patient. The RNN used contained three linear layers, with a 64 dimensional state vector, and was trained with stochastic gradient descent (weight decay = 0.1) for 5 epochs.
For deployment on the referral and external test sets, patient level predictions were obtained from each of the five models trained in cross validation, and these values were averaged to produce the final probability that the specimen represented a malignancy. For cross-validation, only the model trained in a given split was used to predict the corresponding test split.
Model evaluation
The primary metric of area under the receiver operating characteristic curve (AUC) was calculated using the metrics.roc_auc_score function in Scikit-Learn (v0.22.2). Additional metrics that were assessed included accuracy, sensitivity, and specificity. Confidence intervals were calculated through bootstrapping, using 1000 resamples with replacement. Associations between SpindleMesoNET scores and clinico-pathologic factors used Mann-Whitney U tests for two-category tests, Kruskal-Wallis tests for multi-category tests, and Spearman correlations for continuous variables, performed using the R Project for Statistical Computing version 3.5.2 through the RStudio (version 1.2.1335) package ‘stats’ (version 3.6.2).
Results
SpindleMesoNET was first trained and tested on the ‘training/cross-validation’ set, which consisted of in-house and consultation cases of benign spindle cell mesothelial proliferations and sarcomatoid mesotheliomas (81 and 58 cases, respectively; slide and tile numbers in Supplemental Table 1). Five-fold cross-validation was performed on this set, in which each of the cases was used for testing during one of the five runs. Concatenating the test set results across all five runs produced an overall AUC of 0.932 (Fig. 1A). There was a weak but significant positive correlation between the number of tiles that could be extracted from regions annotated as tumor and the SpindleMesoNET scores for malignant cases (Supplemental Fig. 1A), suggesting performance may be suboptimal on samples with very low tumor content. There was no significant association between SpindleMesoNET scores and the number of tiles from benign cases, which approximates specimen size (Supplemental Fig. 1B). SpindleMesoNET scores were not significantly associated with the anatomic site, morphologic subtype or source (in-house vs referred from other hospitals) of malignant cases (Supplemental Fig. 2A–C), indicating robust performance in relation to these factors. In addition, we noted that performance was similar regardless of whether cases had desmoplastic or sarcomatoid morphology (Supplemental Fig. 2B).
We then compared the location of highly predictive tiles from the ‘test’ cases of the five-fold cross validation to the location of tumor annotations. Tiles highly predictive of malignancy tended to overlap with annotated tumor areas (Fig. 2), despite SpindleMesoNET not being exposed to those annotations within the ‘fold’ of cross-validation that generated the scores. This finding supports the notion that SpindleMesoNET identifies areas similar to those recognized by pathologists as malignant.
(A, C) areas with tiles highly predictive of malignancy and (B, D) areas annotated as sarcomatoid malignancy (circled in red) on H&E stain. Representative benign and malignant areas from the H&E images are shown at 20X. For (A, C), tile scores closer to 1 (yellow) were most predictive of malignancy. Areas scoring 0.90 or less (low likelihood of malignancy) are shown in purple.
We next validated SpindleMesoNET on an independent set of referral cases not seen by the model during training or cross-validation. This ‘referral’ test set (Supplemental Table 1) consisted of 40 cases sent for expert opinion regarding whether they represented benign or malignant spindle cell mesothelial proliferations. These difficult cases were considered to represent those for which SpindleMesoNET predictions may provide the most clinical value and may therefore represent future ‘typical use’ scenarios. The prediction values from each of the five models trained on the training/cross-validation set were averaged to produce the final probability for each case in the referral test set, resulting in a referral test set AUC of 0.925 (Fig. 1B).
To provide a human reference, the accuracy of SpindleMesoNET was compared with that of three experienced pathologists (two with thoracic subspecialty training, one with other subspecialty training, and none involved with mesothelioma expert review panels) on the referral test set slides. The deep learning model outputs continuous values (the probability of malignancy), which were converted to binary ‘benign’ or ‘malignant’ predictions. When using the SpindleMesoNET score threshold that maximized accuracy (threshold 0.95), SpindleMesoNET accuracy (37/40, 92.5%) was similar to the average accuracy of the pathologists (91.7%, Table 1). The only cases incorrectly called by SpindleMesoNET were three malignant cases incorrectly called benign, of which one was also incorrectly called by a pathologist (Fig. 3). Representative images of cases incorrectly called by either a pathologist or SpindleMesoNET are shown in Fig. 4. SpindleMesoNET therefore had 100% specificity and 85.7% sensitivity for malignancy, similar to these metrics for pathologists (mean pathologist specificity 96.5% and mean sensitivity 87.3%; Table 1). Interestingly, 9/40 cases were incorrectly called by at least one pathologist (2 benign cases and 7 malignant cases), and SpindleMesoNET was able to provide a correct diagnosis for 8 of these 9 cases. The combined sensitivity of a malignant call by either SpindleMesoNET or a pathologist was 95–100% (depending on the pathologist), greater than the sensitivity of either alone.
The case in (A) was correctly called benign by SpindleMesoNET, but incorrectly called by one pathologist. Subtle zonation and layering support a benign diagnosis. The cases in (B) and (C) were correctly called malignant by SpindleMesoNET, but were incorrectly called by a pathologist. A diagnosis of malignancy is supported by fat invasion in (B) and short storiform architecture in (C). The cases in (D–F) were incorrectly called benign by SpindleMesoNET and either correctly called malignant by pathologists (D, E) or incorrectly called by one pathologist (F). A diagnosis of malignancy is supported by nodularity in (D) and (F) and by necrosis in (E).
The ten highest scoring tiles in all malignant referral set cases showed features compatible with or suggestive of sarcomatoid mesothelioma (described in Fig. 5A–H), indicating that SpindleMesoNET may utilize morphologic features similar to those used by pathologists. Of particular interest, review of the high scoring malignant tiles suggested that SpindleMesoNet put weight on high cellularity (a feature often used by pathologists) and nuclear atypia, a feature often difficult for pathologists to evaluate in this type of tumor. SpindleMesoNet also was able to recognize necrosis as a feature of malignancy (Fig. 5A). The ten lowest scoring tiles from each benign referral set case included recognizable benign tissue types and areas of paucicellular fibrous tissue (Fig. 5I–P).
Among the top ten highest scoring (i.e. most predictive of malignancy) tiles from each malignant cases were tiles showing A necrosis, B short storiform architecture, C possible fat invasion, D ropey collagen, E pleomorphism, F prominent nucleoli, and G, H high cellularity. Among the ten lowest scoring tiles (i.e. least likely to be malignant) from each case were tiles showing (I–L) dense hypocellular collagenous tissue, M capillary channels, N adipose tissue, O organizing blood clot, and P fibrin.
To assess the performance of SpindleMesoNET on slides with different staining characteristics, we also validated SpindleMesoNET on an externally stained test set consisting of benign and malignant cases stained at outside institutions (Supplemental Table 1). These externally stained cases were selected as clear representations of benign and malignant processes. As with the referral set, each of the five trained models produced by the training/cross-validation was used to classify each of the externally stained test set cases, and the prediction values were averaged to provide the final probability for a given case. This produced an AUC for the externally stained test set of 0.989 (Fig. 1C), supporting the notion that SpindleMesoNET is robust to inter-institutional differences in H&E staining.
The above implementations of SpindleMesoNET used both color jitter and stain normalization (see methods) for color adjustment, and used a recurrent neural network for patient level prediction. Retraining the neural network using only color jitter or only stain normalization produced similar or inferior results on the referral and externally-stained test sets compared to the use of both methods (Table 2, Supplemental Table 2). Interestingly, color jitter alone still performed well for the referral test set but not for the externally stained test set, whereas stain normalization alone performed well for the externally stained test set but not the referral test set. This reinforces the critical need for stain normalization when deploying a pathology deep learning model on a data set that was stained in a different laboratory than the training data. We also investigated an alternate ‘average pooled’ method of patient level data summarization (see Methods) but found worse performance than the recurrent neural network method on the referral and externally-stained test sets (Table 2, Supplemental Table 2).
Discussion
The accurate distinction of benign and malignant spindle cell mesothelial proliferations can be a significant diagnostic challenge for pathologists [5, 16, 17]. Morphology is sometimes diagnostic by itself, and specific immunohistochemical or FISH testing resolves the problem in some instances, but a significant number of cases cannot be classified with this approach and end up being referred for expert consultation. There is little room for error in the separation of this extremely aggressive malignancy from benign processes, as this distinction is critical for patient management and for any further subtyping of malignant cases. Here we develop the first neural network trained to distinguish sarcomatoid mesothelioma from benign spindle cell mesothelial proliferations, demonstrating excellent performance on a training/cross-validation set (AUC 0.932), a referral test set (AUC 0.925), and an externally stained test set (AUC 0.989).
Neural networks for mesothelioma have previously been trained to predict prognosis or the presence of a transitional morphologic subtype in cases already diagnosed as malignant mesothelioma [2, 6], but have not been trained to separate benign from malignant mesothelial processes. Neural networks that distinguish benign and malignant histology in other tissue types (e.g. lung [18, 19], lymph nodes with breast cancer metastases [20], prostate [7], brain [21], colon [22], and thyroid [23]) have also shown promise, but few studies have specifically tested on problem cases referred for expert opinion, as was done here. Our study is also unusual in focusing on an uncommon malignancy. The infrequency of sarcomatoid mesothelioma contributes to the difficulties that most pathologists have with its definitive diagnosis, and consequently to the need for ancillary test development. However, this rarity limits the number of samples available for test development.
We demonstrate an approach to neural network development that can provide highly accurate classification using only a limited number of training cases with careful annotation of malignant areas. As all malignant cases included areas of benign tissue, annotations were necessary to ensure that all individual tiles extracted from malignant cases contained malignant tissue and were, therefore, representations of ‘malignant’ morphology. However, the labor-intensive process of annotation restricts the practical size of the training set, and is limited by access to appropriate expertise. Moreover, using annotated tumor slides limits training of the neural network to areas that are already possible for a pathologist to recognise as malignant (which may exclude particularly bland tumor regions) and may introduce bias from the individual pathologists involved in annotation (as not all experts may agree on the ‘malignancy’ of a given area). Methods of aggregating multiple expert opinions into final annotations may prove useful for model refinement.
We envision a refined SpindleMesoNET model having future use as an ancillary test of value to both expert and non-expert pathologists for supporting diagnoses and informing on the need for referral. We demonstrate proof-of-concept that predictions with sufficient accuracy to be clinically valuable can be made by a neural network for this application. Indeed, SpindleMesoNET had non-inferior accuracy to that of subspecialty pathologists on referral set cases (SpindleMesoNET accuracy 92.5%, mean pathologist accuracy 91.7%). Importantly, by virtue of having been sent for expert opinion, the referral set consists entirely of highly challenging cases for which the initial pathologist felt additional support was needed in order to make a definitive diagnosis. The combination of either a pathologist or SpindleMesoNET predicting malignancy had greater sensitivity for malignancy (95–100%) than either method alone, highlighting how SpindleMesoNET predictions, interpreted in the context of a pathologist’s impression, may add clinical value.
Prior to clinical implementation, SpindleMesoNET would benefit from training and testing on additional externally stained cases from multiple outside institutions. We found that the use of color processing techniques allowed SpindleMesoNET to perform well on both the in-house and externally stained slides used in our study. Stain normalization (which maps features of H&E staining to a single reference image) was applied to all training and testing images to address baseline differences in staining between image sets, while color jitter (small random shifts in the color values) was applied during CNN training to increase variability and help prevent overfitting. These results support the notion that SpindleMesoNET may also produce robust results on other externally stained slide sets. Important caveats are that SpindleMesoNET is not intended to distinguish sarcomatoid mesothelioma from other malignant spindle cell neoplasms in the differential diagnosis, and that SpindleMesoNET development used predominantly pleural cases, where sarcomatoid morphology is most common. Application to non-pleural samples would likely benefit from further training and testing.
Guidelines for how to interpret the continuous scale scores may also be refined. Thresholds for calling a specimen ‘benign’ or ‘malignant’ may be varied depending on the use case, without the classifier itself needing any re-training. A low threshold allows highly sensitive detection of potentially malignant cases, useful for capturing all cases with possible malignancy for expert consultation. Conversely, a high threshold enables highly specific diagnoses of malignancy, enhancing confidence that a specimen truly shows mesothelioma prior to signing it out. One other benefit of the way in which SpindleMesoNet is set up, with predictions based only on the values of the most predictive patches, is that it will not be fooled by a biopsy that has both benign areas (organizing pleuritis) and malignant areas but will make a diagnosis of malignancy.
In summary, we have developed the first neural network for distinguishing sarcomatoid mesothelioma from benign spindle cell mesothelial proliferations. We demonstrate robust performance on referral and externally stained test sets. SpindleMesoNET holds potential for future use as an ancillary test helping to inform diagnosis, and highlights key histologic features useful for identifying this aggressive yet diagnostically challenging malignancy.
Data availability
Source code will be made publicly available on Github following publication of this manuscript.
References
Galateau-Salle F, Churg A, Roggli V, Travis WD. The 2015 World Health Organization classification of tumors of the pleura: advances since the 2004 classification. J Thorac Oncol. 2016;11:142–54.
Galateau Salle F, Le Stang N, Tirode F, Courtiol P, Nicholson AG, Tsao M-S, et al. Comprehensive molecular and pathologic evaluation of transitional mesothelioma assisted by deep learning approach: a multi-institutional study of the international mesothelioma panel from the MESOPATH Reference Center. J Thorac Oncol. 2020;15:1037–53.
Nicholson AG, Sauter JL, Nowak AK, Kindler HL, Gill RR, Remy-Jardin M, et al. EURACAN/IASLC proposals for updating the histologic classification of pleural mesothelioma: towards a more multidisciplinary approach. J Thorac Oncol. 2020;15:29–49.
Churg A, Galateau-Salle F. The separation of benign and malignant mesothelial proliferations. Arch Pathol Lab Med. 2012;136:1217–26.
Churg A, Naso JR. The separation of benign and malignant mesothelial proliferations: new markers and how to use them. Am J Surg Pathol. 2020;44:e100–12.
Courtiol P, Maussion C, Moarii M, Pronier E, Pilcer S, Sefta M, et al. Deep learning-based classification of mesothelioma improves prediction of patient outcome. Nat Med. 2019;25:1519–25.
Ström P, Kartasalo K, Olsson H, Solorzano L, Delahunt B, Berney DM, et al. Artificial intelligence for diagnosis and grading of prostate cancer in biopsies: a population-based, diagnostic study. Lancet Oncol. 2020;21:222–32.
Goode A, Gilbert B, Harkes J, Jukic D, Satyanarayanan M. OpenSlide: a vendor-neutral software foundation for digital pathology. J Pathol Inf. 2013;4:27.
van der Walt S, Schönberger JL, Nunez-Iglesias J, Boulogne F, Warner JD, Yager N. scikit-image: image processing in Python. PeerJ. 2014;2:e453.
Otsu N. A Threshold selection method from gray-level histograms. IEEE Trans Syst, Man, Cyber. 1979;9:62–66.
Vahadane A, Peng T, Sethi A, Albarqouni S, Wang L, Baust M, et al. Structure-preserving color normalization and sparse stain separation for histological images. IEEE Trans Med Imaging. 2016;35:1962–71.
Paszke A, Gross S, Massa F, Lerer A, Bradbury J, Chanan G, et al. PyTorch: an imperative style, high-performance deep learning library. arXiv:191201703 [cs, stat] 2019.[cited 1 January 2021]. Available from: http://arxiv.org/abs/1912.01703.
He K, Zhang X, Ren S, Sun J. Deep residual learning for image recognition. In: 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR). Las Vegas, NV, USA: IEEE. p. 770-8 (2016).
Carbonneau M-A, Cheplygina V, Granger E, Gagnon G. Multiple instance learning: a survey of problem characteristics and applications. Pattern Recognit. 2018;77:329–53.
Campanella G, Hanna MG, Geneslaw L, Miraflor A, Werneck Krauss Silva V, Busam KJ. Clinical-grade computational pathology using weakly supervised deep learning on whole slide images. Nat Med. 2019;25:1301–9.
Mangano WE, Cagle PT, Churg A, Vollmer RT, Roggli VL. The diagnosis of desmoplastic malignant mesothelioma and its distinction from fibrous pleurisy: a histologic and immunohistochemical analysis of 31 cases including p53 immunostaining. Am J Clin Pathol. 1998;110:191–9.
Churg A, Colby TV, Cagle P, Corson J, Gibbs AR, Gilks B, et al. The separation of benign and malignant mesothelial proliferations. Am J Surg Pathol. 2000;24:1183–200.
Coudray N, Ocampo PS, Sakellaropoulos T, Narula N, Snuderl M, Fenyö D, et al. Classification and mutation prediction from non-small cell lung cancer histopathology images using deep learning. Nat Med. 2018;24:1559–67.
Kanavati F, Toyokawa G, Momosaki S, Rambeau M, Kozuma Y, Shoji F, et al. Weakly-supervised learning for lung carcinoma classification using deep learning. Sci Rep. 2020;10:9297.
Ehteshami Bejnordi B, Veta M, Johannes van Diest P, van Ginneken B, Karssemeijer N, Litjens G, et al. Diagnostic assessment of deep learning algorithms for detection of lymph node metastases in women with breast cancer. JAMA. 2017;318:2199–210.
Hollon TC, Pandian B, Adapa AR, Urias E, Save AV, Khalsa SSS, et al. Near real-time intraoperative brain tumor diagnosis using stimulated Raman histology and deep neural networks. Nat Med. 2020;26:52–58.
Rathore S, Hussain M, Aksam Iftikhar M, Jalil A. Novel structural descriptors for automated colon cancer detection and grading. Comput Methods Prog Biomed. 2015;121:92–108.
Dov D, Kovalsky SZ, Assaad S, Cohen J, Range DE, Pendse AA, et al. Weakly supervised instance learning for thyroid malignancy prediction from whole slide cytopathology images. Med Image Anal. 2021;67:101814.
Funding
This study was supported by the University of British Columbia Dept of Pathology Residency Training Program and the Dermatology Point-of-Care Intelligent Network, a Digital Technology Supercluster project.
Author information
Authors and Affiliations
Contributions
JRN and ABL drafted the manuscript and were involved in the analysis and interpretation of data. ABL performed neural network construction. JRN, AC, LRC, and SD acquired and reviewed specimens. JLW, CL, and H-MY provided histologic interpretations of referral set cases. AB, HF, and SJMJ contributed to the development of methodology, and provided technical and material support. SY and AC contributed to study conception and design, data interpretation, and review and revision of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
COMPETING INTERESTS
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Co-senior authors: Stephen Yip, Andrew Churg.
Supplementary information
Rights and permissions
About this article
Cite this article
Naso, J.R., Levine, A.B., Farahani, H. et al. Deep-learning based classification distinguishes sarcomatoid malignant mesotheliomas from benign spindle cell mesothelial proliferations. Mod Pathol 34, 2028–2035 (2021). https://doi.org/10.1038/s41379-021-00850-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41379-021-00850-6
This article is cited by
-
Deep learning model to differentiate Crohn’s disease from intestinal tuberculosis using histopathological whole slide images from intestinal specimens
Virchows Archiv (2024)
-
Early Malignant Mesothelioma Detection Using Ensemble of Naive Bayes Under Decorate Ensemble Framework
Journal of The Institution of Engineers (India): Series B (2024)