Abstract
Programmed cell death protein 1/programmed cell death protein ligand1 (PD-1/PD-L1) interaction is an important immune checkpoint targeted by anti-PD-1/PD-L1 immunotherapies. However, the observed prognostic significance of PD-1/PD-L1 expression in diffuse large B-cell lymphoma treated with the standard of care has been inconsistent and even contradictory. To clarify the prognostic role of PD-1/PD-L1 expression and interaction in diffuse large B-cell lymphoma, in this study we used 3-marker fluorescent multiplex immunohistochemistry and Automated Quantitative Analysis Technology to assess the CD3+, PD-L1+, and PD-1+CD3+ expression in diagnostic samples and PD-1/PD-L1 interaction as indicated by presence of PD-1+CD3+ cells in the vicinity of PD-L1+ cells, analyzed their prognostic effects in 414 patients with de novo diffuse large B-cell lymphoma, and examined whether PD-1/PD-L1 interaction is required for the prognostic role of PD-1+/PD-L1+ expression. We found that low T-cell tissue cellularity, tissue PD-L1+ expression (irrespective of cell types), PD-1+CD3+ expression, and PD-1/PD-L1 interaction showed hierarchical adverse prognostic effects in the study cohort. PD-1/PD-L1 interaction showed higher sensitivity and specificity than PD-1+ and PD-L1+ expression in predicting inferior prognosis in patients with high CD3+ tissue cellularity (“hot”/inflammatory tumors). However, both PD-1+ and PD-L1+ expression showed adverse prognostic effects independent of PD-1/PD-L1 interaction, and PD-1/PD-L1 interaction showed favorable prognostic effect in PD-L1+ patients without high CD3+ tissue cellularity. Macrophage function and tumor-cell MYC expression may contribute to the PD-1-independent adverse prognostic effect of PD-L1+ expression. In summary, low T-cell tissue cellularity has unfavorable prognostic impact in diffuse large B-cell lymphoma, and tissue PD-L1+ expression and T-cell-derived PD-1+ expression have significant adverse impact only in patients with high T-cell infiltration. PD-1/PD-L1 interaction in tissue is essential but not always responsible for the inhibitory effect of PD-L1+/PD-1+ expression. These results suggest the benefit of PD-1/PD-L1 blockade therapies only in patients with sufficient T-cell infiltration, and the potential of immunofluorescent assays and Automated Quantitative Analysis in the clinical assessment of PD-1/PD-L1 expression and interaction.
Similar content being viewed by others
Introduction
Programmed cell death protein 1 (PD-1) [1] and its ligands PD-L1 [2] and PD-L2 [3] are important immune checkpoint proteins that limit the antitumor response of T cells. Immunotherapies targeting the PD-1 pathway have shown impressive efficacy in many types of advanced cancers as well as promising clinical activity in refractory or relapsed diffuse large B-cell lymphoma, the most common type of non-Hodgkin lymphoma [4]. However, the differential efficacy of anti-PD-1 immunotherapy across different cancer entities is incompletely understood [5, 6]. Predictive biomarkers are being intensively pursued to guide selection of patients who are more likely to respond to PD-1/PD-L1 blockade, including PD-1/PD-L1 expression (the targets of PD-1/PD-L1 blockade therapy) and T-cell infiltration which were inconsistently associated with higher response rates to anti-PD-1/PD-L1 immunotherapy. There are no known predictive biomarkers for PD-1 blockade in diffuse large B-cell lymphoma currently [5, 6].
Investigating the baseline T-cell infiltration levels and PD-1/PD-L1 expression in a large number of patients with diffuse large B-cell lymphoma can gain insight into the potential of anti-PD-1/PD-L1 immunotherapy in diffuse large B-cell lymphoma. PD-1 can be expressed by both T cells and lymphoma B cells [7,8,9]. The primary PD-1 ligand PD-L1 can be expressed on a wide variety of cells. Several study groups have assessed expression of PD-1 [10,11,12,13] or PD-L1 [11, 12, 14,15,16,17] by single or double-staining immunohistochemistry in patients with diffuse large B-cell lymphoma. However, most studies have only subjectively assessed PD-1+ lymphocytes overall or PD-L1 expression on lymphoma cells, but not PD-1+ T cells or PD-L1 expression in the tumor microenvironment, which can also ligate with PD-1 on T cells mediating immune suppression. In addition, PD-1 and PD-L1 expression may be not directly linked to their active interaction and suppressive function. Active PD-1–PD-L1 interaction has been evaluated by proximity between PD-1 and PD-L1, which showed significant association with the efficacy of anti-PD-1 immunotherapy [18] and the survival of patients treated with standard therapy in solid cancer [19]. In Hodgkin lymphoma, recent topological analysis revealed the enrichment of PD-L1+ (versus PD-L1−) macrophages and CD4+ (versus CD8+) T cells in the vicinity of PD-L1+ Hodgkin Reed-Sternberg cells, the spatial enrichment of PD-1+ or PD-1− CD4+ T cells in direct contact (versus without contact) with PD-L1+ Hodgkin Reed-Sternberg cells and tumor-associated macrophages, the spatial enrichment of PD-1+ or PD-1− CD8+ T cells in direct contact (versus without contact) with PD-L1+ tumor-associated macrophages, and the characteristics of the tumor microenvironment having high PD-L1 expression from abundant macrophages and low PD-1 expression in T cells [20]. However, whether macrophages and CD4+ T cells that are more abundant locally are more functionally relevant for clinical outcome, whether only PD-1+ cells in direct contact with (but not those distant to) PD-L1+ cells are functionally suppressed, and whether only the topologically relevant PD-1+/PD-L1+ cells under microscopy have clinical impact and therapeutic implications in Hodgkin lymphoma have not been studied.
To investigate the prognostic significance of PD-1/PD-L1 expression and interaction in diffuse large B-cell lymphoma, in this study, we employed 3-marker fluorescent multiplex immunohistochemistry to simultaneously analyze tumor-infiltrating CD3+ T cells, PD-1 expression in tumor-infiltrating T cells, overall tissue PD-L1 expression, and PD-1/PD-L1 spatial proximity in a large cohort of diffuse large B-cell lymphoma patients. To minimize variation and subjectivity of manual scoring, Automated Quantitative Analysis (AQUA) Technology [21, 22] were used to analyze the local /PD-L1+/PD-1+/CD3+ tissue expression and interaction.
Materials and methods
Patients
The study cohort included 414 patients with de novo diffuse large B-cell lymphoma with available diagnostic biopsy specimens who were treated with standard immunochemotherapy, consisting of rituximab plus cyclophosphamide, doxorubicin, vincristine, and prednisone (R-CHOP), as part of the International DLBCL Rituximab-CHOP Consortium Program Study. Patients with primary mediastinal large B-cell lymphoma, primary cutaneous diffuse large B-cell lymphoma, primary diffuse large B-cell lymphoma of the central nervous system, HIV-associated diffuse large B-cell lymphoma, or diffuse large B-cell lymphoma transformed from low-grade B-cell lymphoma have been excluded [23]. Gene expression profiling data (GSE#31312) by the Affymetrix GeneChip Human Genome U133 Plus 2.0 array [24] were available for 361 patients, which classified 328 cases as germinal center B-cell-like or activated B-cell-like subtype. For other cases, cell of origin was determined for 80 cases according to immunohistochemistry analysis using the Visco-Young and/or Choi algorithms [24]. The study was conducted in accordance with the Declaration of Helsinki, and the protocol was approved as being of minimal to no risk or as exempt by the institutional review board of each participating institution.
Quantitative immunofluorescence imaging and analysis
Tissue microarrays with core size of 1.0 mm were constructed for diagnostic formalin-fixed, paraffin-embedded tumor samples. Fluorescent staining with CD3 (EP41, Biocare Medical), PD-1 (NAT105, Biocare Medical), and PD-L1 (E1L3N, Cell Signaling Technology) antibodies and 4′,6-diamidino-2-phenylindole (DAPI) (cell nuclei) was performed using an automated stainer (Intellipath, Biocare Medical). Fluorescent images were all acquired on the Vectra imaging system (PerkinElmer) at × 20 using various channels including DAPI (blue), Cy5 (red), Cy3.5 (orange), and Opal 520 (green).
Vectra images were analyzed by proprietary analysis algorithms AQUA® (Navigate BioPharma Services, Inc) [21]. Cell population analysis was performed by establishing cell masks that represent all cells and individual cell types in the tissue microarray cores. All cell compartments were derived by converting DAPI signals to binary images and then dilated to approximate size of an entire cell. CD3, PD-1, and PD-L1 cells were identified by creating binary masks without dilation, as membrane staining outlines actual cell size. CD3/PD-1 double-positive cells were established by identifying overlap areas between CD3 and PD-1 binary cell compartments. In colored multiplexed images, overlapping regions show overlaid color. For example, colocalization of green and red colors produces a degree of yellow color. To evaluate PD-1/PD-L1 interaction, PD-L1 binary masks were first dilated to create an interaction mask encompassing the nearest neighbor cells. Interaction scores were then generated by calculating overlap areas between the PD-1+/CD3+ compartment and the dilated PD-L1 mask multiplied by 10,000 to create whole numbers representing an interaction score for each specimen.
Statistical analysis
Clinicopathologic parameters were compared between groups using the Fisher’s exact test. Overall survival was calculated from the date of diagnosis to the date of last follow-up or death. Progression-free survival was calculated from the date of diagnosis to the date of disease progression or death. Survival of patients was compared by Kaplan–Meier curves and the log-rank (Mantel–Cox) test. P values of ≤ 0.05 were considered statistically significant. As multiple comparisons were performed, the significance was adjusted by both the Benjamini–Hochberg procedure and the Bonferroni correction methods. Gene expression profiling analysis was performed by the R software. Differential expressed genes were identified by multiple t tests and the obtained P values were corrected for multiple testing controlling the false discovery rate as described previously [24, 25].
Results
T-cell infiltration and tissue PD-L1/PD-1 expression and interaction in diffuse large B-cell lymphoma
Figure 1 shows immunofluorescent staining images for two representative samples of diffuse large B-cell lymphoma. Numbers of CD3+, PD-1+, PD-L1+, and PD-1+CD3+ (i.e., PD-1+ T) cells in the entire tissue area were simultaneously quantified. PD-1+ T cells within the vicinity of PD-L1 compartment were identified and evaluated by a PD-1/PD-L1 interaction score. CD3/PD-1/PD-L1 expression levels were assessed by total tissue cellularity, dividing the positive cell numbers by total nucleated cells (DAPI +) and rounding to the nearest 0.1%. Most (90%) lymphoma cases had T-cell infiltration with a mean tissue cellularity of 13.6%. PD-L1 was expressed in 53% of the cohort, and in these cases, PD-L1 was expressed by a mean tissue cellularity of 6.7%. PD-1 was expressed in 30% of lymphoma cases, and in these cases, PD-1 was expressed by a mean tissue cellularity of 1.46%. T-cell-specific PD-1 expression was detected in 18% of lymphoma cases with a mean tissue cellularity of 1.1%. In 20% of the cohort, PD-1 was also expressed on CD3− cells with a mean tissue cellularity of 1.24%. Interaction scores were in the range of 0–3910. About 23% of cases had a positive interaction score, with a mean score of 190. The mean tissue cellularity of CD3+, PD-L1+, PD-1+, and PD-1+CD3+ cells in the entire cohort were 12.1%, 3.6%, 0.44%, and 0.19%, respectively. Activated B-cell-like compared with germinal center B-cell-like diffuse large B-cell lymphoma had a significantly higher mean tissue cellularity of PD-L1+ cells (P = 0.0013; Supplementary Figure 1a), but not CD3+ cells or PD-1+CD3+ cells.
Representative images of CD3+, PD-1+, and PD-L1+ expression in diffuse large B-cell lymphoma samples by fluorescent multiplex immunohistochemistry (counter-stained with DAPI). a A representative case with PD-1+ T cells proximal to PD-L1+ cells in the tissue specimen. Cells with PD-1+ (red fluorescence) and CD3+ (green fluorescence; T-cell marker) co-expression show yellow color in the color imaging overlay. b A representative case having PD-L1+ cells but no PD-1+ T cells in the tissue
Hierarchical unfavorable prognostic effects associated with tissue cellularity of CD3+, PD-1+CD3+, and PD-L1+ cells
CD3/PD-1/PD-L1 expression and PD-1/PD-L1 interaction scores were correlated with patient survival. In the overall diffuse large B-cell lymphoma cohort, patients with a low percentage of CD3+ T cells in the tumor tissue (CD3−/lo, 0% to 3.5% of all cells were CD3+) had a poorer overall survival rate compared with patients with a high CD3+ tissue cellularity (CD3hi, > 3.5% of all cells were CD3+) (P = 0.02; Fig. 2a). Patients with a positive tissue cellularity of PD-L1+ cells or PD-1+ T cells had a non-significantly poorer progression-free survival rate (P = 0.057 and P = 0.079, respectively) but a similar overall survival rate compared with cases with a negative tissue cellularity. The distribution of cases with CD3hi, PD-L1+, or PD-1+CD3+ tissue cellularity or with PD-1/PD-L1 interaction in the cohort is shown in Fig. 2b.
Prognostic analysis for CD3, PD-L1, and T-cell-derived PD-1 expression in the study cohort. a High tissue cellularity of CD3+ T cells was associated with significantly better overall survival in patients with diffuse large B-cell lymphoma. b A plot illustrating the case distribution of CD3hi, PD-L1+, and PD-1+CD3+ tissue cellularity and PD-1/PD-L1 proximity (PD-1:PD-L1 interaction). Each column represents one patient. Positive cases were indicated by corresponding colors. Abbreviations: ABC, activated B-cell-like; GCB, germinal center B-cell-like. c In CD3hi cases, PD-1 expression on T cells was associated with significantly poorer progression-free survival. d–e In CD3hi cases, PD-L1 expression and positive PD-1/PD-L1 interaction scores were associated with significantly poorer overall survival
Although the CD3hi group was prognostically favorable compared with the CD3−/lo group, it had higher mean cellularity of PD-L1+ cells and PD-1+CD3+ cells (P < 0.0001 and P = 0.0002, respectively; Supplementary Figure 1b), which remained significant after exclusion of CD3− cases (0% tissue cellularity) from the CD3−/lo group (that is, CD3lo subset) (P < 0.0001 and P = 0.0007, respectively). Moreover, in the CD3hi group of patients, the presence of PD-1+ tumor-infiltrating T cells was associated with significantly poorer overall survival (P = 0.028; Supplementary Figure 1c) and progression-free survival (P = 0.0099, Fig. 2c), although PD-1+CD3+ cases had a higher T-cell tissue cellularity (P < 0.0001). PD-L1+ cellularity was also associated with significantly poorer overall survival (P = 0.0063; Fig. 2d) and progression-free survival (P = 0.0058; Supplementary Figure 1c) in the CD3hi group, but it was not associated with higher CD3+ tissue cellularity (P = 0.3). Similarly, positive PD-1/PD-L1 interaction scores, suggesting interaction between PD-1+ T cells and PD-L1+ cells, were associated with significantly poorer overall survival (P = 0.0006; Fig. 2e) and progression-free survival (P = 0.0003; Supplementary Figure 1c) in the CD3hi patients, despite the association with a higher CD3+ tissue cellularity (P = 0.0012). In contrast, the presence of PD-1+CD3− cells was not prognostic in CD3hi patients (P = 0.99 for overall survival and P = 0.95 for progression-free survival) or the entire cohort.
When CD3hi cases were divided into germinal center B-cell-like and activated B-cell-like subtypes, the adverse prognostic effects associated with PD-1+/PD-L1+ expression in patients with CD3hi tissue cellularity were only significant in the activated B-cell-like diffuse large B-cell lymphoma subset, whereas PD-1/PD-L1 interaction was associated with significantly poorer overall survival in both germinal center B-cell-like (P = 0.024) and activated B-cell-like subtypes (P = 0.01) as well as poorer progression-free survival in the activated B-cell-like subtype (P = 0.0007; Supplementary Figure 1d).
Clinical features were compared between groups of patients with different prognosis. Patients with CD3−/lo cellularity more often had > 1 extranodal site of involvement and an International Prognostic Index score of > 2 (Table 1). PD-L1 expression was associated with primary nodal and an advanced stage of disease and elevated serum lactate dehydrogenase level in the overall cohort (Table 1) and with advanced stage, B-symptom, and an International Prognostic Index score of > 2 in the CD3hi subcohort. PD-1+CD3+ expression in CD3hi patients was associated with a performance status score > 1 (Table 1). PD-1/PD-L1 interaction in CD3hi patients was associated with an International Prognostic Index score of > 2, advanced stage, a performance status score of > 1, and > 1 extranodal site of involvement (Supplementary Table 1).
Independent adverse prognostic effects of PD-1+CD3+ and PD-L1+ expression and PD-1/PD-L1 interaction in CD3hi patients
Because the presence of PD-1+ T cells was associated with a higher mean PD-L1+ tissue cellularity in the overall diffuse large B-cell lymphoma cohort (P < 0.0001; Supplementary Figure 1e) and the CD3hi subset (P = 0.019), we examined the dependency of the prognostic effects associated with PD-1+CD3+ expression and PD-L1+ expression in patients with CD3hi cellularity. We found that compared with patients having neither PD-1 nor PD-L1 expression, patients with PD-1+ T cells had significantly poorer overall/progression-free survival regardless of positive/negative PD-L1 expression (PD-1+CD3+/PD-L1+ patients: P = 0.0002 for overall survival and P < 0.0001 for progression-free survival; PD-1+CD3+/PD-L1─ patients: P = 0.012 for overall survival and P = 0.016 for progression-free survival; Fig. 3a and Supplementary Figure 2a). PD-1+CD3+ expression also had significant adverse prognostic effect regardless of presence/absence of PD-1/PD-L1 interaction (Fig. 3b and Supplementary Figure 2b).
Independent prognostic effects of PD-1 and PD-L1 expression and the role of PD-1/PD-L1 interaction in patients with high CD3+ tissue cellularity (CD3hi cases). a In CD3hi cases, PD-1 expression in T cells was associated with significantly poorer overall survival with or without PD-L1 expression. b In CD3hi cases, PD-1 expression were associated with significantly poorer overall survival with or without a positive PD-1/PD-L1 interaction score. c In CD3hi cases without PD-1+ expression (PD-1−CD3hi), PD-L1 expression were associated with significantly poorer overall survival. d–e Positive PD-1/PD-L1 interaction scores were associated with significantly poorer overall survival in all PD-1−CD3hi cases and in PD-1−CD3hi cases with PD-L1+ expression. f After exclusion of cases with a positive PD-1/PD-L1 interaction score, PD-L1+ expression remained to be associated with significantly poorer overall survival in PD-1−CD3hi cases
Conversely, in CD3hi patients with PD-1+CD3+ expression, PD-L1 expression and PD-1/PD-L1 interaction were not associated with a significant prognostic effect. However, in CD3hi patients without PD-1+CD3+ expression (PD-1−CD3hi), PD-L1+ expression was associated with significantly poorer overall survival (P = 0.0034; Fig. 3c) and progression-free survival (P = 0.0059; Supplementary Figure S2a), independent of whether the tumors were of germinal center B-cell-like or activated B-cell-like subtype. Although this suggests that PD-L1 had inhibitory function independent of PD-1 expression on T cells, PD-L1 positivity was not associated with higher PD-1+ expression on non-T (CD3−) cells (P = 0.23 for decrease) or CD3+ T cellularity (P = 0.81). The inferior survival may not result from interaction with PD-1 expressed on non-T cells also because the presence of PD-1+CD3− cells was not prognostic in PD-1−CD3hi patients. On the other hand, we surprisingly found that a small number of PD-L1+/PD-1−CD3hi cases (n = 23; Fig. 2b) had a low but positive PD-1/PD-L1 interaction score (1–53), suggesting presence of low numbers of PD-1+ T cells (< 0.1% tissue cellularity) having active interaction with PD-L1+ cells. Presence of (albeit low) PD-1/PD-L1 interaction had significantly poorer survival in the PD-1−CD3hi subcohort (P < 0.0001 for OS, P = 0.0001 for progression-free survival; Fig. 3d and Supplementary Figure 2b), independent of the germinal center B-cell-like (P = 0.0005 for overall survival and P = 0.0058 for progression-free survival) and activated B-cell-like subtypes (P = 0.033 for overall survival and P = 0.014 for progression-free survival). Furthermore, PD-1/PD-L1 interaction significantly contributed to the adverse prognostic effect of PD-L1+ expression in PD-1−CD3hi patients, as it was associated with significantly poorer survival in PD-1−CD3hi cases with PD-L1+ expression (P = 0.0083 for both overall survival and progression-free survival; Fig. 3e). Nonetheless, after cases with positive PD-1/PD-L1 interaction scores were excluded, PD-L1 expression remained to be associated with significantly poorer survival (P = 0.032 for OS; Fig. 3f and P = 0.049 for progression-free survival).
We looked for prognostic biomarkers differentially associated with PD-L1 expression according to PD-1 expression status in diffuse large B-cell lymphoma. We found that in the PD-1−CD3hi subcohort, but not in the CD3hi subcohort with PD-1+CD3+ expression, PD-L1+ expression was significantly associated with a higher mean level of tumor Myc expression (P = 0.015; Fig. 4a), a higher frequency (19% vs. 2%, P = 0.011) of Myc/Bcl-2 dual expression (MYC+BCL2+, an adverse prognostic factor [23, 25, 26]), and a lower frequency of CD30 positivity [27] (14% vs. 29%, P = 0.05) evaluated by standard immunohistochemistry. CD30− was also associated with PD-1+CD3+ expression and PD-1/PD-L1 interaction in CD3hi cases (Table 1 and Supplementary Table 1). After exclusion of cases with positive PD-1/PD-L1 interaction scores from the PD-1−CD3hi cases, the association of PD-L1+ with Myc overexpression remained significant (P = 0.041; Fig. 4b). PD-L1 expression in these patients and in the overall PD-1─CD3hi cases was also associated with activated B-cell-like subtype and female sex (Supplementary Table 1).
Tumor-cell Myc expression and survival analysis in cases with PD-1−CD3hi tissue cellularity (PD-1−CD3hi cases). a–b In PD-1−CD3hi cases, PD-L1+ expression was associated with a higher mean level of Myc expression by traditional immunohistochemistry. c In PD-1−CD3hi cases with PD-L1+ expression, MYC+BCL2+ double-expression was associated with significantly poorer overall survival. d After exclusion of MYC+BCL2+ double-expressors from PD-1−CD3hi cases, PD-L1 expression remained to be associated with significantly poorer progression-free survival. e In cases with PD-L1+/CD3−/lo tissue cellularity, PD-1/PD-L1 interaction was associated with significantly better overall survival. f After exclusions of cases with PD-1/PD-L1 interaction and cases with MYC+BCL2+ expression from the CD3−/lo subcohort, PD-L1+ expression was associated with significantly poorer overall survival
The association of MYC+BCL2+ cases significantly contributed to the adverse prognostic effect of PD-L1 expression in the PD-1−CD3hi group, in that MYC+BCL2+ expression was associated with significantly poorer overall survival and progression-free survival both in PD-1−CD3hi patients overall and in patients with PD-L1+/PD-1−CD3hi cellularity (Fig. 4c) (all P < 0.0001). Nonetheless, after exclusion of cases with MYC+BCL2+ expression from the PD-1−CD3hi cases, PD-L1 expression remained to be associated with significantly poorer progression-free survival (P = 0.013; Fig. 4d), and a trend toward poorer overall survival also was seen (P = 0.074). After exclusion of both MYC+BCL2+ cases and cases with a positive PD-1/PD-L1 interaction score from the PD-1−CD3hi subcohort, PD-L1 expression was not associated with overall survival or progression-free survival in PD-1−CD3hi cases (P = 0.18 and P = 0.29 for poorer overall survival and progression-free survival, respectively).
Unfavorable prognostic effect of PD-L1+ expression and favorable prognostic effect of PD-1/PD-L1 interaction in CD3−/lo patients
In patients with CD3−/lo tissue cellularity, presence of PD-1/PD-L1 interaction (positive score 1–3366) was associated with a higher mean CD3+ and PD-1+CD3+ tissue cellularity (all P < 0.0001), and non-significantly better overall survival (P = 0.065; Supplementary Figure 2c) and progression-free survival (P = 0.08). In the subset of CD3−/lo patients with PD-L1+ tissue cellularity, PD-1/PD-L1 interaction was associated with significantly better overall survival (P = 0.014, Fig. 4e) and progression-free survival (P = 0.011; Supplementary Figure 2d), especially in the activated B-cell-like subtype of diffuse large B-cell lymphoma (P = 0.038 for overall survival and P = 0.0088 for progression-free survival; Supplementary Figure 3a).
Like PD-1/PD-L1 interaction, PD-1+CD3+ expression in the CD3lo patients was associated with a higher mean CD3+ cellularity (P = 0.014), but the expression was not prognostic either in the CD3−/lo or CD3lo subset. PD-L1 expression was also significantly associated with a higher mean CD3+ tissue cellularity in both the CD3−/lo and CD3lo groups (P < 0.0001 and P = 0.0006, respectively), as well as activated B-cell-like subtype (P = 0.036), elevated serum lactate dehydrogenase level (P = 0.0011), primary nodal disease (P = 0.0041), and CD30 positivity (25% vs. 13%, P = 0.043). After patients with PD-1/PD-L1 interaction (favorable) and patients with MYC+BCL2+ expression (unfavorable, P < 0.0001) were excluded from the CD3−/lo subcohort, PD-L1 expression was associated with significantly poorer overall survival (P = 0.02, Fig. 4f) and progression-free survival (P = 0.018; Supplementary Figure 3b), even though PD-L1+ expression was still associated with higher CD3+ tissue cellularity (P = 0.0003).
Multiple testing correction and multivariate survival analysis
Significance of multiple comparisons in the study cohort was adjusted using both the BenjaminiHochberg procedure and Bonferroni correction (Supplementary Table 2). By the Benjamini–Hochberg procedure with a false discovery rate of 0.10, the adverse prognostic effect of CD3−/lo cellularity in overall cohort, the adverse prognostic effects of PD-L1 expression, PD-1 expression, and PD-1/PD-L1 interaction in CD3hi cases, the adverse prognostic effect of PD-L1 expression in the CD3−/lo subset without PD-1 interaction and MYC+BCL2+ expression, and the favorable prognostic effect of PD-1/PD-L1 interaction in cases with PD-L1+/CD3−/lo tissue cellularity all remained significant. By the more stringent Bonferroni criterion for six tests, only the adverse prognostic effects of PD-1/PD-L1 interaction and PD-L1 expression in CD3hi cases were significant.
Multivariate survival analyses were performed to adjust for clinical parameters using the Cox regression model. In the overall cohort, the favorable effect of CD3hi cellularity on overall survival was not significant (P = 0.11, hazard ratio (HR) 0.74, 95% confidence interval (CI) 0.52–1.07); marginally significant P values of poorer progression-free survival were seen for PD-L1+ expression (P = 0.068, HR 1.39, 95% CI 0.98–1.97) and PD-1/PD-L1 interaction (P = 0.072, HR 1.42, 95% CI 0.97–2.09). However, in the CD3hi (but not CD3−/lo) subcohort, PD-1/PD-L1 interaction and PD-L1+ expression were identified as significant adverse prognostic factors independent of clinical factors (Table 2). Although PD-1+ expression was not an independent prognostic factor in the multivariate analysis, the adverse prognostic effects of PD-1/PD-L1 interaction and PD-L1+ expression in CD3hi patients were significant only in patients with PD-1−CD3hi tissue cellularity (Table 2) but not in CD3hi patients with PD-1+CD3+ expression.
After further adding the activated B-cell-like risk factor into the Cox regression models, only PD-1/PD-L1 interaction remained as an independent prognostic factor in CD3hi patients (P = 0.009, HR 2.01, 95% CI 1.19–3.41 for progression-free survival and P = 0.046, HR 1.79, 95% CI 1.01–3.16 for OS). PD-L1 expression only showed a marginal P value for being an independent factor for poorer progression-free survival with the additional adjustment of cell of origin (P = 0.08, data not shown).
Gene signatures associated with T-cell infiltration and PD-1/PD-L1 expression
Gene expression profiling data were analyzed to identify signatures associated with CD3hi cellularity in the overall cohort and with PD-1+CD3+ expression and PD-L1+ expression in the CD3hi and CD3−/lo subcohorts.
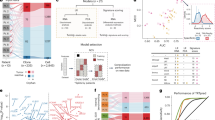
Patients with CD3hi tissue cellularity was associated with a distinct gene signature, including upregulation of genes involved in the T-cell receptor complex (TRBC1, CD3E, and TRAT1), adhesion (ITM2A, SIRPG, and CD2), chemotaxis (CXCL9 and CXCL10), T-cell cytolytic function (GZMK, GZMA, GBP1, and GBP2), cell survival (BCL11B and GIMAP1), SLAMF7 (a receptor regulating a wide variety of immune cells), and complement component C1QB (Fig. 5a, Table 3; false discovery rate < 0.01, fold change > 1.74)).
Gene expression profiling analysis and a schematic summary for this study. a Heatmap for differentially expressed genes between CD3−/lo and CD3hi cases (false discovery rate threshold: 0.01; fold change cutoff: > 1.74). b Heatmap for differentially expressed genes between PD-L1+ and PD-L1─ patients in the CD3hi cases overall (false discovery rate threshold: 0.01; fold change cutoff: > 1.7) and in the PD-1−CD3hi subset without PD-1/PD-L1 interaction (false discovery rate threshold: 0.30). c A schematic diagram illustrating the relevant regulation, interaction, and prognostic effects of PD-L1/CD3/PD-1 expression in this study or according to the literature. Green arrows indicate upregulation of PD-1/PD-L1 expression; red arrows indicate transmission of inhibitory signals by receptor/ligand interaction; dashed red or black arrow indicates uncertain inhibitory or unknown effects; red bar-headed line indicates inhibition; blue boxes highlight the detection of expression by fluorescent multiplex immunohistochemistry and AQUA in diffuse large B-cell lymphoma specimens; and black arrows indicate prognostic effects. Abbreviation: APC, antigen-presenting cells; FIHC, fluorescent multiplex immunohistochemistry; AQUA, Automated Quantitative Analysis
For PD-1+ expression in T cells, gene signatures were identified in the CD3+, CD3hi, and PD-L1+/CD3hi subcohorts, but not in the PD-L1─/CD3hi subcohort. NFATC2, a transcription factor regulating PDCD1 expression, was upregulated in PD-L1+/CD3hi patients with PD-1+CD3+ expression (Supplementary Figure 3c, Table 3). Presence of PD-1/PD-L1 interaction showed a gene signature in the activated B-cell-like subtype but not in the germinal center B-cell-like subtype of CD3hi cases (Supplementary Figure 3d, Table 3), which included upregulation of ubiquitin ligases (FBXO36 and RBCK1) and genes involved in biosynthesis and metabolism, and downregulation of PTCH1 (receptor for Hedgehog) and genes involved in biosynthesis (ST6GALNAC5), actin organization (RAB3IP), membrane trafficking (ZFYVE16), and protein folding (RICS).
For PD-L1+ expression, the gene signature identified in CD3hi patients included some genes shared with the CD3hi gene signature (such as CXCL9/10, GZMK/A, GBP1/2, SLAMF7, and C1QB) and some unique genes including CXCL11, CCL8, PRF1, KLRK1, FCGR1B, FCER1G, STAT1, and LILRB2 (Fig. 5b, false discovery rate < 0.01). In the PD-1−CD3hi subset, PD-L1+ expression was associated with upregulation of genes with T-cell cytolytic function (PRF1), complement (C1QB, FCGR1B, and C1QA), and MTDH (which activates NF-ĸB), and downregulation of the transcription factor TEAD1 with tumor suppression function and adhesion genes (FLRT2 and ITGA6) (Table 3). After exclusion of cases with a positive PD-1/PD-L1 interaction score from the PD-1−CD3hi subset, PD-L1+ expression was associated with a gene signature featuring upregulating of the macrophage markers CD163 and CD68 and monocyte chemoattractant genes CCL8, CXCL10, and CXCL11 (Fig. 5b, Table 3).
No gene signature was identified for PD-L1+/PD-1+ expression or interaction in CD3−/lo patients overall, but a distinct gene signature was identified for PD-L1+ expression in the CD3− activated B-cell-like subset, including upregulation of MYO1G, CDRT1, IL8RB, adhesion genes, and metabolism genes (Supplementary Figure 3e, Table 3).
Discussion
Unlike previous studies that separately quantified PD-1+ lymphocytes of any cell type and PD-L1+ percentage of tumor cells with traditional chromogen-based immunohistochemistry, this study simultaneously quantified CD3+ cells, PD-1+CD3+ T cells, and total PD-L1+ cells of any cell type as well as the PD-1/PD-L1 interaction score in the same tissue microarray cores using a 3-marker 4-color fluorescent multiplex immunohistochemistry platform. PD-1/PD-L1 interaction was also assessed with an interaction score, which increase with the proportion of PD-1+ T cells proximal to PD-L1+ cells. Computerized AQUA was used to quantitate expression and interaction, in order to provide more continuous and reproducible measures of PD-1/PD-L1 expression in terms of biomarker intensity, percent positivity, and receptor-ligand interactions. Gene expression profiling data, which was not available in earlier studies, were analyzed for gene signatures associated with CD3/PD-1/PD-L1 expression and PD-1/L1 interaction. The results demonstrated the capability of a fully automated process to stratify patients with diffuse large B-cell lymphoma treated with R-CHOP, and a significant hierarchical association of low intratumoral T cells, PD-1+ expression in T cells, overall PD-L1+ expression, and PD-1/PD-L1 interaction in the tissue with poorer survival. First, absence to a low tumor-infiltrating CD3+ T-cell tissue cellularity was associated with worse overall survival and a gene expression profiling signature indicating lower antitumor T-cell response. Second, high CD3+ T-cell tissue cellularity was associated with increased PD-1/PD-L1 expression, and the expression and interaction of PD-L1/PD-1 had adverse prognostic effects in patients with CD3hi tissue cellularity. Third, both PD-1+ and PD-L1+ expression had adverse prognostic impact independent of PD-1/PD-L1 interaction, likely attributable to the presence of other inhibitory interactions (such as with PD-L2 and CD80) [28,29,30] and other mechanisms (Fig. 5c).
In earlier studies, presence of PD-1+ lymphocyte was paradoxically associated with favorable or no differential survival in diffuse large B-cell lymphoma overall [6]. In a more recent study, Kiyasu et al. [14] showed that PD-L1hi percentage of tumor cells but not PD-1+ lymphocyte numbers had significant adverse effect on OS, resembling the no effect of T-cell PD-1+ expression on overall survival in overall patients identified through fluorescent co-expression in this study. However, PD-1+CD3+ expression showed a nonsignificant trend of poorer progression-free survival in the overall cohort. Moreover, by separating the prognostic effect of T-cell infiltration from that of PD-1+ expression, this study clearly demonstrated the unfavorable clinical effect of T-cell PD-1+ expression in CD3hi patients with diffuse large B-cell lymphoma, despite the association of PD-1+CD3+ expression with higher CD3+ tissue cellularity. Irrespective of cell resource of PD-1 expression, total PD-1 tissue cellularity with a cutoff of ≥ 0.4% of tissue cellularity was also associated with significantly poorer overall survival after exclusion of CD3−/lo patients (P = 0.036 for overall survival and P = 0.08 for progression-free survival). Although T-cell PD-1+ expression was associated with higher PD-L1+ tissue cellularity, the adverse prognostic effect of PD-1+CD3+ expression in CD3hi patients was independent of PD-L1 expression and PD-1/PD-L1 interaction. Our gene expression profiling analysis showed that PD-1+CD3+ expression in patients with PD-L1+/CD3hi tissue cellularity was associated with upregulation of NFATC2a, a transcription factor of the NFAT family that is known to regulate PD-1 expression and terminal differentiation of CD8+ T cells [5, 31, 32].
Prognostic analysis for the presence of PD-1/PD-L1 interaction (based on quantitative vicinity analysis for PD-1+CD3+ cells and PD-L1+ cells in the tissue) showed a greater adverse prognostic effect than that based on PD-1 or PD-L1 expression in the CD3hi patients overall and in the germinal center B-cell-like and activated B-cell-like subgroups. These greater prognostic effects and identification of cases with a positive PD-1/PD-L1 interaction score (albeit low) among cases with PD-1−CD3hi cellularity suggest that the PD-1/PD-L1 interaction score is a more sensitive and specific index for PD-1 inhibitory function than PD-1+CD3+ cellularity. However, only a weak gene expression profiling signature was identified for PD-1/PD-L1 interaction in the activated B-cell-like group, possibly because the major effect of PD-1/PD-L1 interaction is on signal transduction rather than transcriptional changes [33]. In addition, different from PD-1/PD-L1 expression, in patients with CD3−/lo/PD-L1+ tissue cellularity, presence of PD-1/PD-L1 interaction was associated with a significantly better survival, which could be attributable to the antitumor activity of T cells closely localized to tumors [19]. Also notably, an earlier study found that PD-L1 interacting with PD-1 or other unknown receptors on T cells promoted T-cell infiltration and cytokine production in a murine chronic colitis model [34]. However, the favorable effect of PD-1/PD-L1 interaction in CD3−/lo patients was not significant in multivariate analysis, and we did not identify a gene signature associated with PD-1/PD-L1 interaction in CD3−/lo patients.
Compared with earlier studies of tumor-cell PD-L1 expression in diffuse large B-cell lymphoma, this study extended the analysis to total PD-L1+ tissue cellularity, which can be conveniently quantified by fully automated methods. PD-L1 detected in this study could be expressed by tumor cells, antigen-presenting cells, T cells, or other cells in the tumor microenvironment. Tissue PD-L1+ expression was associated with higher CD3+ tissue cellularity in the overall cohort and CD3−/lo and CD3lo cohorts and with a gene signature indicating active T-cell responses in CD3hi patients. Furthermore, we specifically showed that the adverse prognostic effect by total PD-L1+ cells was in CD3hi patients primarily but also in CD3−/lo cases without MYC+BCL2+ expression. The total frequency of these unfavorable PD-L1+ cases was 27%. The adverse prognostic effect of tissue PD-L1 expression resembled that of tumor-cell PD-L1 expression by Kiyasu et al. [14] using a PD-L1/PAX5 double-staining technique, but the case numbers in this study was larger (414 vs. 273).
Furthermore, our results showed that PD-L1/PD-1 interaction was not the only underlying mechanisms for the poorer prognosis associated with PD-L1 expression, as in PD-1−CD3hi cases without PD-L1 interaction with PD-1+ T cells, PD-1 expression on non-T cells was not prognostic, whereas PD-L1 expression continued to show an adverse prognostic effect, which potentially could be attributable to the PD-L1–CD80 axis or other PD-1-independent inhibitory functions. Notably, PD-L1+ expression in these PD-1−CD3hi cases having no PD-1/PD-L1 interaction was associated with a gene signature featuring macrophage-related genes. In addition, PD-L1+ expression in overall CD3hi cases was associated with upregulation of STAT1 (encoding a transcription factor which can activate PD-L1 gene expression) and LILRB2 (whose expression can suppress macrophage function and phagocytosis [35]), and PD-L1+ expression in CD3− patients was associated with upregulation of IL8RB (encoding a receptor for IL8, a chemokine produced by macrophages and other cells). Together, these results suggested that macrophage-related mechanisms may also be relevant to the PD-1-independent prognostic effect by PD-L1+ expression, and that traditional evaluation of PD-L1 expression on tumor cells only is insufficient.
In addition, intrinsic tumor mechanisms may also contribute to the adverse prognostic effect of PD-L1 expression independent of PD-1, as we found that PD-L1 expression in PD-1−CD3hi cases was associated with higher tumor Myc protein expression relevant for the poorer survival. Notably, others have shown that Myc transcriptionally regulates PD-L1 expression in a mouse model of MYC-induced T-cell acute lymphoblastic leukemia [36], whereas another study showed that BET/BRD4 (which regulates MYC transcription) upregulates PD-L1 expression independent of Myc in a Myc-driven B-cell lymphoma model [37]. Moreover, Myc also regulates expression of CD47, a critical “do not eat me” signals for macrophages [36]. Blocking anti-CD47 antibodies combined with rituximab have been shown to increase macrophage-mediated phagocytosis and antitumor T-cell responses in preclinical studies, and induced clinical response (objective response rate, 40%; complete response rate, 33%) in patients with relapsed or refractory diffuse large B-cell lymphoma in a recent phase Ib clinical trial [38,39,40].
In summary, tissue PD-L1 expression and T-cell-specific PD-1 expression showed adverse prognostic effects secondary to that of low T cells in the tumor microenvironment, and PD-1/PD-L1 interaction in tissue is essential but not always responsible for the inhibitory effect of PD-1 and PD-L1 expression in patients with CD3hi infiltrating diffuse large B-cell lymphoma. These results provide rationales for immunotherapies promoting T-cell activation/infiltration in CD3−/lo patients and PD-1/PD-L1 blockade immunotherapies in patients with CD3hi tissue cellularity. However, PD-L1+ expression does not always imply for PD-1 blockade immunotherapy, as in some PD-L1+ cases, PD-1-independent mechanisms are relevant for the poor prognosis. This study adds to our knowledge of the PD-1/PD-L1 axis in diffuse large B-cell lymphoma and has important implications for understanding the efficacy and biomarkers of PD-1/PD-L1 blockade therapies, as well as showed the clinical utility of automated immunofluorescent and quantitative analysis for prognostication and therapeutic decision. Notably, fluorescent multiplex immunohistochemistry has been implemented in clinical trials to characterize immune infiltrate in responders versus non-responders with solid cancers (for example, NCT03387761 and NCT03192202). However, further standardization and validation are warranted. In addition, the capacity of this simple three-antibody panel in prognostication in diffuse large B-cell lymphoma suggests that improvement of traditional chromogen-based immunohistochemistry and scoring methods in clinical practice in lieu of the availability of such specialized fluorescent multiplex immunohistochemistry and AQUA platform can also be valuable for patient stratification.
References
Ishida Y, Agata Y, Shibahara K, Honjo T. Induced expression of PD-1, a novel member of the immunoglobulin gene superfamily, upon programmed cell death. EMBO J. 1992;11:3887–95.
Dong H, Zhu G, Tamada K, Chen LB7-H1. a third member of the B7 family, co-stimulates T-cell proliferation and interleukin-10 secretion. Nat Med. 1999;5:1365–9.
Latchman Y, Wood CR, Chernova T, et al. PD-L2 is a second ligand for PD-1 and inhibits T cell activation. Nat Immunol. 2001;2:261–8.
Lesokhin AM, Ansell SM, Armand P, et al. Nivolumab in patients with relapsed or refractory hematologic alignancy: preliminary results of a phase Ib study. J Clin Oncol. 2016;34:2698–704.
Xu-Monette ZY, Zhang M, Li J, Young KH. PD-1/PD-L1 blockade: have we found the key to unleash the antitumor immuneresponse? Front Immunol. 2017;8:1597.
Xu-Monette ZY, Zhou J, Young KH. PD-1 expression and clinical PD-1 blockade in B-cell lymphomas. Blood. 2018;131:68–83.
Xerri L, Chetaille B, Serriari N, et al. Programmed death 1 is a marker of angioimmunoblastic T-cell lymphoma and B-cell small lymphocytic lymphoma/chronic lymphocytic leukemia. Hum Pathol. 2008;39:1050–8.
Muenst S, Hoeller S, Willi N, Dirnhofera S, Tzankov A. Diagnostic and prognostic utility of PD-1 in B cell lymphomas. Dis Markers. 2010;29:47–53.
Laurent C, Charmpi K, Gravelle P, et al. Several immune escape patterns in non-Hodgkin’s lymphomas. Oncoimmunology. 2015;4:e1026530.
Ahearne MJ, Bhuller K, Hew R, et al. Expression of PD-1 (CD279) and FoxP3 in diffuse large B-cell lymphoma. Virchows Arch. 2014;465:351–8.
Fang X, Xiu B, Yang Z, et al. The expression and clinical relevance of PD-1, PD-L1, and TP63 in patients with diffuse large B-cell lymphoma. Medicine (Baltim). 2017;96:e6398.
Kwon D, Kim S, Kim PJ, et al. Clinicopathological analysis of programmed cell death 1 and programmed cell death ligand 1 expression in the tumour microenvironments of diffuse large B cell lymphomas. Histopathology. 2016;68:1079–89.
Cohen M, Vistarop AG, Huaman F, et al. Cytotoxic response against Epstein Barr virus coexists with diffuse large B-cell lymphoma tolerogenic microenvironment: clinical features and survival impact. Sci Rep. 2017;7:10813.
Kiyasu J, Miyoshi H, Hirata A, et al. Expression of programmed cell death ligand 1 is associated with poor overall survival in patients with diffuse large B-cell lymphoma. Blood. 2015;126:2193–201.
Xing W, Dresser K, Zhang R, et al. PD-L1 expression in EBV-negative diffuse large B-cell lymphoma: clinicopathologic features and prognostic implications. Oncotarget. 2016;7:59976–86.
Menter T, Bodmer-Haecki A, Dirnhofer S, Tzankov A. Evaluation of the diagnostic and prognostic value of PDL1 expression in Hodgkin and B-cell lymphomas. Hum Pathol. 2016;54:17–24.
Rossille D, Azzaoui I, Feldman AL, et al. Soluble programmed death-ligand 1 as a prognostic biomarker for overall survival in patients with diffuse large B-cell lymphoma: a replication study and combined analysis of 508 patients. Leukemia. 2017;31:988–91.
Tumeh PC, Harview CL, Yearley JH, et al. PD-1 blockade induces responses by inhibiting adaptive immune resistance. Nature. 2014;515:568–71.
Carstens JL, Correa de Sampaio P, Yang D, et al. Spatial computation of intratumoral T cells correlates with survival of patients with pancreatic cancer. Nat Commun. 2017;8:15095.
Carey CD, Gusenleitner D, Lipschitz M, et al. Topological analysis reveals a PD-L1-associated microenvironmental niche for Reed-Sternberg cells in Hodgkin lymphoma. Blood. 2017;130:2420–30.
Dolled-Filhart M, Gustavson M, Camp RL, et al. Automated analysis of tissue microarrays. Methods Mol Biol. 2010;664:151–62.
Johnson DB, Bordeaux JM, Kim JY, et al. Quantitative spatial profiling of PD-1/PD-L1 interaction and HLA-DR/IDO-1 predicts improved outcomes of anti-PD-1 therapies in metastatic melanoma. Clin Cancer Res. 2018;24:5250–60.
Xu-Monette ZY, Li L, Byrd JC, et al. Assessment of CD37 B-cell antigen and cell of origin significantly improves risk prediction in diffuse large B-cell lymphoma. Blood. 2016;128:3083–3100.
Visco C, Li Y, Xu-Monette ZY, et al. Comprehensive gene expression profiling and immunohistochemical studies support application of immunophenotypic algorithm for molecular subtype classification in diffuse large B-cell lymphoma: a report from the International DLBCL Rituximab-CHOP Consortium Program Study. Leukemia. 2012;26:2103–13.
Xu-Monette ZY, Dabaja BS, Wang X, et al. Clinical features, tumor biology, and prognosis associated with MYC rearrangement and Myc overexpression in diffuse large B-cell lymphoma patients treated with rituximab-CHOP. Mod Pathol. 2015;28:1555–73.
Hu S, Xu-Monette ZY, Tzankov A, et al. MYC/BCL2 protein coexpression contributes to the inferior survival of activated B-cell subtype of diffuse large B-cell lymphoma and demonstrates high-risk gene expression signatures: a report from The International DLBCL Rituximab-CHOP Consortium Program. Blood. 2013;121:4021–31.
Hu S, Xu-Monette ZY, Balasubramanyam A, et al. CD30 expression defines a novel subgroup of diffuse large B-cell lymphoma with favorable prognosis and distinct gene expression signature: a report from the International DLBCL Rituximab-CHOP Consortium Program Study. Blood. 2013;121:2715–24.
Butte MJ, Keir ME, Phamduy TB, Sharpe AH, Freeman GJ. Programmed death-1 ligand 1 interacts specifically with the B7-1 costimulatory molecule to inhibit T cell responses. Immunity. 2007;27:111–22.
Paust S, Lu L, McCarty N, Cantor H. Engagement of B7 on effector T cells by regulatory T cells prevents autoimmune disease. Proc Natl Acad Sci USA. 2004;101:10398–403.
Park JJ, Omiya R, Matsumura Y, et al. B7-H1/CD80 interaction is required for the induction and maintenance of peripheral T-cell tolerance. Blood. 2010;116:1291–8.
Philip M, Fairchild L, Sun L, et al. Chromatin states define tumour-specific T cell dysfunction and reprogramming. Nature. 2017;545:452–6.
Pauken KE, Sammons MA, Odorizzi PM, et al. Epigenetic stability of exhausted T cells limits durability of reinvigoration by PD-1 blockade. Science. 2016;354:1160–5.
Mognol GP, Spreafico R, Wong V, et al. Exhaustion-associated regulatory regions in CD8+ tumor-infiltrating T cells. Proc Natl Acad Sci USA. 2017;114:E2776–E2785.
Kanai T, Totsuka T, Uraushihara K, et al. Blockade of B7-H1 suppresses the development of chronic intestinal inflammation. J Immunol. 2003;171:4156–63.
Barkal AA, Weiskopf K, Kao KS, et al. Engagement of MHC class I by the inhibitory receptor LILRB1 suppresses macrophages and is a target of cancer immunotherapy. Nat Immunol. 2017;19:76–84.
Casey SC, Tong L, Li Y, et al. MYC regulates the antitumor immune response through CD47 and PD-L1. Science. 2016;352:227–31.
Hogg SJ, Vervoort SJ, Deswal S, et al. BET-bromodomain inhibitors engage the host immune system and regulate expression of the immune checkpoint ligand PD-L1. Cell Rep. 2017;18:2162–74.
Chao MP, Alizadeh AA, Tang C, et al. Anti-CD47 antibody synergizes with rituximab to promote phagocytosis and eradicate non-Hodgkin lymphoma. Cell. 2010;142:699–713.
Tseng D, Volkmer JP, Willingham SB, et al. Anti-CD47 antibody-mediated phagocytosis of cancer by macrophages primes an effective antitumor T-cell response. Proc Natl Acad Sci USA. 2013;110:11103–8.
Advani R, Flinn I, Popplewell L, et al. CD47 blockade by Hu5F9-G4 and rituximab in non-Hodgkin’s lymphoma. N Engl J Med. 2018;379:1711–21.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Li, L., Sun, R., Miao, Y. et al. PD-1/PD-L1 expression and interaction by automated quantitative immunofluorescent analysis show adverse prognostic impact in patients with diffuse large B-cell lymphoma having T-cell infiltration: a study from the International DLBCL Consortium Program. Mod Pathol 32, 741–754 (2019). https://doi.org/10.1038/s41379-018-0193-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41379-018-0193-5
This article is cited by
-
The progress of novel strategies on immune-based therapy in relapsed or refractory diffuse large B-cell lymphoma
Experimental Hematology & Oncology (2023)
-
Immune priming with avelumab and rituximab prior to R-CHOP in diffuse large B-cell lymphoma: the phase II AvR-CHOP study
Leukemia (2023)
-
Immune evasion phenotype is common in Richter transformation diffuse large B-cell lymphoma variant
Virchows Archiv (2023)
-
Recent Advancements in Nanomedicine for ‘Cold’ Tumor Immunotherapy
Nano-Micro Letters (2021)
-
Epstein–Barr virus recruits PDL1-positive cells at the microenvironment in pediatric Hodgkin lymphoma
Cancer Immunology, Immunotherapy (2021)