Abstract
Structural and numerical alterations of chromosome 21 are extremely common in hematological malignancies. While the functional impact of chimeric transcripts from fused chromosome 21 genes such as TEL-AML1, AML1-ETO, or FUS-ERG have been extensively studied, the role of gain of chromosome 21 remains largely unknown. Gain of chromosome 21 is a frequently occurring aberration in several types of acute leukemia and can be found in up to 35% of cases. Children with Down syndrome (DS), who harbor constitutive trisomy 21, highlight the link between gain of chromosome 21 and leukemogenesis, with an increased risk of developing acute leukemia compared with other children. Clinical outcomes for DS-associated leukemia have improved over the years through the development of uniform treatment protocols facilitated by international cooperative groups. The genetic landscape has also recently been characterized, providing an insight into the molecular pathogenesis underlying DS-associated leukemia. These studies emphasize the key role of trisomy 21 in priming a developmental stage and cellular context susceptible to transformation, and have unveiled its cooperative function with additional genetic events that occur during leukemia progression. Here, using DS-leukemia as a paradigm, we aim to integrate our current understanding of the role of trisomy 21, of critical dosage-sensitive chromosome 21 genes, and of associated mechanisms underlying the development of hematological malignancies. This review will pave the way for future investigations on the broad impact of gain of chromosome 21 in hematological cancer, with a view to discovering new vulnerabilities and develop novel targeted therapies to improve long term outcomes for DS and non-DS patients.
Similar content being viewed by others
Introduction
Copy Number Variation (CNV), i.e. gains or losses of entire chromosomes or of specific genomic regions, are hallmarks of cancer. Understanding the impact of CNVs in tumor development is challenging since they can sometimes alter the dosage of hundreds or even thousands of genes simultaneously, thereby modifying mRNA and protein abundance, ultimately impacting cellular fitness. In hematological cancer, numerous CNVs are found and they are markedly different to those found in solid tumors. Gain of chromosome 21 (+21) is one of the most frequent CNVs observed in hematological malignancies [1, 2]. Using large cohorts, it has been shown that +21 is rarely seen in solid tumors and that trisomy 21, the most common type of +21, is found in nearly all subtypes of hematological malignancies, ranging from 2.2% in chronic lymphocytic leukemia (CLL) to nearly 15% in acute lymphoblastic leukemia (ALL) (Mitelman database, available at https://mitelmandatabase.isb-cgc.org). It is rarely observed as a sole cytogenetic abnormality and its prognostic value varies depending on the cohort analyzed and the type of hematological malignancy [1, 3]. Children with acute megakaryoblastic leukemia (AMKL) and B-cell precursor ALL (B-ALL) harbor +21 most frequently, occurring in approximately one-third of cases [4, 5]. To date, reasons for such an association remain elusive, but strongly suggest that somatic +21 is clonally selected during leukemia development and that megakaryocytic and B-cell progenitor/precursors are profoundly susceptible to the increased dosage of chromosome 21 genes.
Individuals with Down syndrome (DS) harbor constitutive trisomy 21 [6] and are predisposed to childhood acute leukemia [7]. Constitutive trisomy 21 is the most common cytogenetic abnormality seen at birth (1 in 700–1000 newborns), and predominantly results from nondisjunction of chromosome 21 during meiosis (95%) [8,9,10]. Although this mechanism is different to somatic gain of chromosome 21, studying the predisposing and leukemia promoting role of trisomy 21 in DS human specimens, cell lines and murine models has been instrumental in undertanding the role of +21 alone and in cooperation with other secondary genetic alterations. Evidence indicates that trisomy 21 (or gain of chromosome 21), regardless of whether constitutive or acquired, is a promoting event in hematological malignancies.
This review will present a snapshot of our knowledge on DS-associated leukemia, integrating studies addressing clinical features, therapy and molecular mechanisms of leukemogenesis, and will consider them in parallel to other hematological malignancies harboring somatic +21 with the view to emphasize similarities and differences at the clinical and biological level.
Clinical features, therapy, and outcome of DS-associated leukemia
Clinical features
Common clinical features seen in individuals with DS are intellectual disability, congenital heart defects, Alzheimer’s disease and immunodeficiency, among many others [8]. Incidence and severity of these phenotypes can vary due to trisomy 21 itself (complete or segmental trisomy) and/or potential modifier genes that remain elusive to date. Individuals with DS have a unique pattern of malignancies compared with the general population, characterized by a decreased incidence of solid tumors in adults and predisposition to leukemia during childhood, predominantly myeloid leukemia (ML–DS, >100-fold increased risk), which has a high prevalence of AMKL, and ALL (DS–ALL) [7].
Indicative of an intrinsic effect of trisomy 21 on hematopoiesis, almost all neonates with DS have quantitative and/or qualitative disorders of the myeloid compartment such as macrocytosis, dysplastic platelets, leukocytosis, and on average 4% blasts in the peripheral blood [11]. Approximately 10% develop transient myeloproliferative disorder (TMD), which is classically defined by the presence of megakaryoblasts in the peripheral blood, liver and bone marrow [12]. TMD spontaneously resolves within the first months of life, suggesting that mechanisms regulated by the fetal microenvironment maintain perturbed hematopoiesis. Up to 30% of children with DS who have classically defined TMD develop ML–DS before 5 years of age. This occurs through stepwise pathogenesis with an incremental acquisition of genetic alterations (including GATA1 mutations, see below). However, ~20% of neonates with DS have been identified as having ‘silent TMD,’ defined by a peripheral blast count of ≤10% and detection of a GATA1 gene mutation by next generation sequencing [11]. This discovery has implications for the population at risk of transforming to ML–DS. Children with ML–DS have remarkably good prognosis compared with non-DS children with acute myeloid leukemia (AML), with 5-years overall and event-free survival of 89–93% and 87–90% respectively [13, 14].
Children with DS have a 27-fold increased risk of developing ALL [7]. A recent study has emphasized that these are almost exclusively B-cell phenotype, with T-cell ALL identified in only 5 of 653 children with DS [15]. In contrast to ML–DS, children with DS–ALL have an inferior outcome compared with non-DS children with ALL due to higher relapse rates, increased risk of infection, treatment-related mortality and induction failure [15, 16].
Therapy and outcome
Significant progress has been made in the treatment of children with DS and leukemia, with outcomes summarized in Table 1. Several studies have prospectively collected and published data on TMD in neonates with DS. These studies have been instrumental in identifying that the majority of patients undergo spontaneous remission, demonstrated a clear benefit for treating babies with high-risk features with low-dose cytarabine (which reduces TMD-related mortality but does not prevent progression to ML–DS) and indicated that persistence of minimal residual disease (MRD) can be used to predict risk for developing ML–DS [17,18,19,20,21]. The next generation of trials should seek to uniformly define high-risk criteria for therapy, identify the optimal dose and schedule for cytarabine treatment and identify the molecular mechanisms that underpin progression to ML–DS. Given the rarity of the disease, consideration should be given to a unified international protocol, which will provide uniformity of outcomes and permit a greater number of questions to be answered.
Prior to the 1980s, children with ML–DS were undertreated resulting in a high rate of treatment failure [22]. ML–DS patients were subsequently registered on protocols used for non-DS AML. Increased survival was evident, with lower rates of induction failure and relapse, however treatment-related mortality was more frequent resulting in protocol adaptation to dose-reduce therapy or prolong the interval between chemotherapy courses [23]. Over the last 20 years, children have been enrolled onto uniform ML–DS specific protocols (Table 1) [13, 14, 24,25,26,27]. These studies have been instrumental in highlighting the benefit for reduced-intensity ML–DS specific protocols, with outcomes for children with ML–DS significantly better than non-DS AML. Due to concerns regarding increased treatment-related toxicity in children with ML–DS [28,29,30], sequential protocols successfully reduced cumulative exposure to several agents including daunorubicin, etoposide, and intrathecal cytarabine without impacting on overall outcome [13, 14, 26, 27]. High-dose cytarabine was established as an important component of therapy, with early administration leading to improved outcomes [13] and subsequent attempts to omit high-dose cytarabine in standard-risk patients resulting in significantly lower event-free survival [31]. Several studies have identified older age as an unfavorable independent prognostic feature [26, 27, 32]. ML–DS diagnosed in children over 4 years of age has been shown to lack GATA1 mutations and has a cytogenetic profile more akin to children with sporadic AML [33]. Given that biologically the disease appears to represent sporadic AML occurring in children with DS rather than ML–DS, undertreatment on the less intensive ML–DS protocols may account for the adverse prognosis in this age group. As such, it has been suggested that children within this older age group may benefit from more intensive therapy corresponding to that given to children with sporadic AML [33]. Monosomy 7 [24], gain of chromosome 8 [14], normal karyotype and high white blood cell count ≥20 × 109/l at presentation [32] have also been identified in individual reports as independent variables associated with an inferior outcome. Detection of MRD after induction therapy using both deep sequencing of GATA1 and flow cytometric methods has been identified as a significant prognostic factor for predicting relapse [34], with current trials implementing risk-adapted therapy according to MRD response (NCT02521493; jRCTs041190047). Future studies should prospectively establish whether additional clinical and biological features can be utilized in addition to MRD response assessment for risk-stratification, which may help further reduce therapy in low-risk patients without compromising outcome and enable treatment intensification for high-risk patients to prevent relapse.
Children with DS–ALL are treated on standard ALL chemotherapeutic protocols [35,36,37,38]. Similar to ML–DS, sequential treatment protocols have identified that children with DS–ALL are more prone to treatment-related toxicity due to heightened sensitivity to chemotherapeutic agents, particularly methotrexate, and infectious complications, further contributing to their inferior outcome [16]. Consequently, this has resulted in modification of treatment to reduce intensity and implementation of intensified supportive care measures for children with DS–ALL [39, 40]. We are rapidly approaching a therapeutic plateau to which we can intensify conventional chemotherapeutic agents in order to balance the equilibrium between relapse and treatment-related toxicity, indicating the need to investigate novel agents in children with DS–ALL. Integration of the bispecific T-cell engager, blinatumomab, into the treatment backbone is being investigated in the current upfront COG (NCT03914625) and AIEOP-BFM (NCT03643276) studies and children with high-risk DS–ALL who are MRD positive at the end of consolidation are eligible for the single arm, phase 2 study of tisagenlecleucel (NCT03876769).
Outcomes for relapsed/refractory leukemia in children with DS are extremely poor [13, 41, 42]. Stem cell transplantation has been associated with high rates of relapse and treatment-related mortality in children with DS [43,44,45]. CAR T-cell therapy may provide an alternative option to stem cell transplantation in this setting. Children with relapsed/refractory DS–ALL have been included in studies using tisagenlecleucel, with preliminary results identifying comparable safety and efficacy to children without DS [46]. Such findings highlight the importance of including children with DS in studies of new agents for the treatment of leukemia to identify innovative approaches to reduce rates of relapse and treat relapsed disease [47]. Novel therapeutic strategies targeting the somatic events found in DS leukemia or the mechanisms altered by trisomy 21 as the initiating event in DS leukemogenesis should also be explored, which ultimately may be applicable to a broad spectrum of hematological malignancies that present with a similar genetic background.
Genetic landscape of DS-associated leukemia
Several studies have recently reported on the genetic landscape of DS-associated leukemia, providing new insights into leukemia development. Ongoing functional characterization of secondary alterations that have been identified may shed light on novel actionable targets that could be therapeutically exploited.
The multi-step pathogenesis of ML–DS development
Development of ML–DS, from predisposing trisomy 21 to pre-leukemia TMD to frank ML–DS, has been considered a model of sequential acquisition of secondary alterations for many years.
Discovery of GATA1 mutations in nearly all patients with TMD and ML–DS was the first major breakthrough in our understanding of ML–DS [48,49,50]. The transcription factor GATA1 is a master regulator of erythroid and megakaryocytic lineages. In TMD/ML–DS, GATA1 mutations are almost exclusively localized in exon 2 (97% of cases) and are predominantly insertions, deletions or duplications [51]. All GATA1 mutations lead to the appearance of a premature stop codon. Consequently, a ‘short’ GATA1 isoform (GATA1s), truncated from its amino-terminal transactivation domain is expressed. Identical GATA1 mutations are seen in TMD and at progression to ML–DS, regardless of whether they are present in the major clone [51,52,53,54,55]. The majority of these clones have leukemia-initiating and self-renewal potential [56, 57]. To date, the type of GATA1 mutation, the level of GATA1s expression, the TMD blast karyotype and the size of dominant GATA1-bearing clones have not been shown to predict progression from TMD to ML–DS [51, 54, 55]. However, features identified as being predictive of progression include the persistence of immunophenotypic TMD blasts (>0.1%) or detection of GATA1 mutations by quantitative PCR using patient-specific oligonucleotides at week 12 and the presence of pleural effusions at diagnosis of TMD [19, 21].
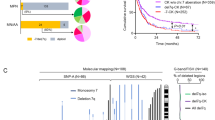
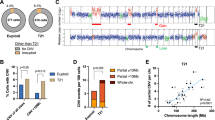
Clonal evolution from TMD to ML–DS has been linked to acquisition of several secondary chromosomal and genetic alterations. There is a lower incidence of CNVs and chromosomal translocations in ML–DS compared with non-DS children with AML; the most common cytogenetic alterations being +8, dup(1q), and a fourth chromosome 21 [4, 5]. Next generation sequencing experiments have uncovered the mutational spectrum of TMD/ML–DS [54, 55, 58], revealing GATA1 mutations as the only somatic mutations seen in TMD in most cases. During progression to ML–DS, two to five additional mutations are found. Among them, the most frequently altered genes encode signaling effectors (JAK1/2/3, MPL, RAS) in 56% of cases, members of the cohesin complex or associated components (STAG2, RAD21, SMC1A) in 48%, and epigenetic regulators (EZH2, SUZ12, and BCOR) in 38% (Fig. 1a). A novel clonal gain of function mutation affecting the CSF2RB gene, encoding the common beta chain of the IL3, IL5, and CSF cytokine receptors, was recently identified in almost 5% of patients with ML–DS and was mutually exclusive with JAK1-3, MPL or RAS mutations [55]. In ML–DS, somatic variants in signaling effectors and cohesin complex components are likely clonal and frequently co-occur, indicative of a potential cooperative effect in trisomic GATA1s-expressing hematopoietic progenitors [54, 55]. To date, nearly 75–80% of patients with ML–DS have been shown to harbor secondary alterations, emphasizing the model of progression from the pre-leukemic TMD stage. The molecular bases of clonal evolution for the remainder remains obscure to date.
Numerous in vitro and in vivo assays have reported on the role of somatic alterations found in TMD/ML–DS. GATA1s expression has been shown to promote megakaryoblastic progenitor expansion during fetal life but does not to lead to ML–DS [59]. Most of the somatic alterations found in ML–DS are shared with non-DS leukemia, and have been shown to promote self-renewal, differentiation blockade, proliferation and survival of hematopoietic stem cells (HSC) and/or myeloid progenitor populations [60, 61]. To assess leukemia progression from GATA1s-expressing cells without trisomy 21, models of oncogenic cooperation have been developed (retroviral insertional mutagenesis, ectopic expression and ‘loss of function screening’ using a CRISPR/Cas9 strategy and transgenic models) [55, 62, 63]. On average, 2.7 alterations on top of Gata1s expression was sufficient for the development of an erythro-megakaryoblastic leukemia in vivo, resembling ML–DS but strongly biased toward the erythroid lineage (CD117+Ter119+ population) [55]. The most common alterations found in these recipients were variants found in signaling effectors (73.7%) and epigenetic regulators (79%). The loss of function mutations in genes encoding components of the cohesin complex were underrepresented (16% in murine recipients vs 48% in patients with ML–DS). Whether this is due to human versus murine differences, or indicative of the lack of trisomy 21 in this model remains unknown.
In summary, observations in humans and mice have emphasized the essential role of trisomy 21, the fetal hematopoietic context of GATA1s expression and the requirement of additional genetic events predominantly affecting signaling effectors, epigenetic regulators and cohesin complex components to drive progression toward ML–DS.
Somatic alterations found in DS–ALL
DS–ALL is a heterogeneous subtype of B-ALL that has a distinct cytogenetic profile compared with other types of childhood B-ALL (non-DS–ALL). Approximately 40% of DS–ALL cases have a normal karyotype other than the constitutional trisomy 21, 8–10% express the ETV6-RUNX1 fusion transcript, while 9–11% have high hyperdiploidy (HeH) [4, 15]. The most frequent CNV in DS–ALL is gain of chromosome X (38% vs 21% in non-DS–ALL); a twofold increase of del(9p) is also found in DS–ALL [4]. DS–ALL also has a higher proportion of rearrangements affecting the CRLF2 locus (~50% in DS–ALL vs 4–5% in non-DS–ALL), with interstitial deletion of the pseudoautosomal region PAR1 (that fuses CRLF2 to the first non-coding exon of P2RY8) or chromosomal translocation to the immunoglobulin heavy chain (IgH) locus [64,65,66]. These rearrangements lead to overexpression of the CRLF2 gene, encoding a protein that heterodimerizes with IL7RA to form the thymic stromal lymphopoietin receptor. To date, there are no fusion proteins uniquely found in DS–ALL [67, 68].
At the gene level, JAK2 activating mutations are present in 20–40% of patients with DS–ALL and predominantly affect the arginine 683 residue, located in the pseudokinase domain of JAK2 [69] (Fig. 1b). These JAK2 mutations are virtually always found in CRLF2-overexpressing cases, indicative of a mechanism of oncogenic cooperation, as shown in experimental models [65, 66, 70]. In non-JAK2 mutated or non-CRLF2-overexpressing cases, gain of function mutations are found in CRLF2 or IL7RA genes, strongly implicating cytokine signaling as a major transforming process in DS–ALL [66, 70, 71]. Activating mutations affecting other signaling effectors, such as NRAS, KRAS, KIT, FLT3, and PTPN11, are also frequently found and are often mutually exclusive with JAK2 mutations, highlighting the high incidence (>60%) of constitutively active signaling pathways in DS–ALL [67, 72]. In line with these findings, a recent study revealed enrichment of a Philadelphia-like transcriptional signature in DS–ALL [68]. Other common genetic abnormalities found in DS–ALL affect genes encoding cell cycle regulators (CDKN2A/B, RB1), transcription factors (PAX5, IKZF1, and ETV6) and epigenetic modifiers (EZH2, SETD2, and CREBBP) (Fig. 1b). Of interest, a recent genome-wide association meta-analysis explored inherited genetic susceptibility to ALL in children with DS, highlighting increased germline penetrance of the rs3731249 CDKN2A risk locus (9p21.3) [73].
To our knowledge, there is only one in vivo model of DS–ALL [74] and several other models assessing the cooperation between secondary mutations commonly found in DS–ALL. One study hypothesized that loss of USP9X (seen in 4/17 CRLF2-rearranged cases), which encodes a deubiquitinase known to stabilize activated JAK2, may naturally buffer the toxic effect of JAK/STAT signaling hyperactivation in DS–ALL [67]. In addition, activating mutations in signaling effectors have been shown to functionally cooperate with loss of Ink4a/Arf and Pax5 alterations to drive B-cell leukemia development in vivo [75, 76].
Together, these studies unravel the high genetic complexity observed in DS–ALL, raising critical questions regarding the role of somatic alterations in the context of trisomy 21 in leukemia development. New models of DS–ALL will be required in the future to better understand the impact of trisomy 21, the mechanisms of cooperation and develop novel targeted therapies to improve outcome.
Constitutive trisomy 21 as a ‘priming’ and cooperating event
Constitutive trisomy 21 (T21) results in altered hematopoiesis affecting many lineages, is developmental stage selective, but is not sufficient to lead to leukemia. Studying embryonic, fetal and adult hematopoiesis in individuals with DS and from genetically engineered models has unraveled the cellular and molecular bases of DS-leukemogenesis, and allowed identification of chromosome 21 dosage-sensitive genes.
Insights from human DS fetal hematopoiesis
Compared with gestation-matched controls, analyses of second trimester human DS fetal livers revealed that trisomy 21 disturbs fetal hematopoiesis [77,78,79]. There is an increased proportion of HSC and of megakaryocyte-erythroid progenitors (MEP) in T21 fetal livers, at the expense of granulocyte-monocyte progenitors (GMP) and committed B progenitors. This bias toward the erythro-megakaryocytic lineage has been confirmed by clonogenic assays, liquid cultures, transplantation assays in immunodeficient mice and transcriptional analyses. A recent study showed that engineered GATA1s expression in human fetal liver HSC cooperates with trisomy 21 to promote blast and megakaryocyte expansion in xenotransplantation models [80]. The marked impairment of B lymphopoiesis in T21 fetal livers is also associated with a reduced proportion of both preproB (CD34+CD19+CD10−) and proB (CD34+CD19+CD10+) cells, and a decreased expression of early lymphoid or key lymphoid specific genes (such as IKZF1, FLT3, PAX5, and IL7RA) in early lymphoid progenitors [79]. Importantly, this reduced B-cell progenitor compartment is also seen in T21 fetal bone marrow, and is associated with impaired B-cell differentiation potential in vitro that may be linked to inflammatory signatures driven by the trisomic microenvironment [81].
This perturbed fetal hematopoiesis may provide insight into the cellular context of susceptibility for transformation of the megakaryocytic and B-cell lineages. However, several questions remain regarding the dynamic changes of these phenotypes during gestation before and after the second trimester, if they are maintained postnatally and throughout the life of individuals with DS, and how they may be affected by their microenvironment (yolk-sac, fetal liver and fetal and adult bone marrow).
Insights from human DS-induced pluripotent stem cells
Through the establishment of DS-derived induced pluripotent stem cells (iPSCs), independent studies have shown that trisomy 21 alone significantly accelerates the early stages of hematopoiesis [82,83,84,85]. Regardless of whether primitive (yolk-sac type) or definitive (fetal liver type) hematopoiesis was induced, all groups reported an increase in the clonogenic potential of erythroid progenitors and enhanced erythroid differentiation. However, in contrast to primitive-like hematopoietic progenitors [83], mimicking fetal hematopoiesis from trisomic iPSCs led to multi-lineage expansion compared with disomic cells, characterized by significant expansion of myeloid and megakaryocytic progenitors in colony-forming unit (CFU) assays [82, 84, 85]. These observations were confirmed by silencing one chromosome 21 in DS-derived iPSCs using an inducible XIST strategy [84]. Extensive analyses of trisomic/GATA1s iPSCs, established from primary patient TMD samples or engineered by genome editing, have shown that GATA1s expression correlates with defective embryonic erythropoiesis and confers a strong bias toward the myelo-megakaryocytic compartment, indicating that both events cooperate in the development of TMD during fetal life [85, 86].
A recent study reported that trisomic CD34+ hematopoietic progenitors derived from embryoid bodies also have decreased ability to generate CD19+ B cells compared with isogenic controls [87]. This phenotype has been linked to reduced expression of endothelin signaling, and provides potential explanation for the impaired B-cell differentiation in human trisomic fetal livers [79], and for the decreased number of circulating B cells in individuals with DS [88].
Overall, these findings suggest that trisomy 21 perturbs early hematopoiesis to create permissive cellular contexts, for the development of both ML–DS and DS–ALL, through the acquisition of additional genetic alterations.
Partially trisomic mice as models to study DS-leukemia
In the mouse genome, syngeneic regions of human chromosome 21 (Hsa21) are located on the three murine chromosomes (Mmu) 16, 17, and 10. Several partially trisomic murine models, containing some or all these syngeneic regions, have been used to study the impact of trisomy 21 on hematopoiesis, alone or in cooperation with GATA1s expression. The phenotypes of these models have been extensively reviewed [12, 89]. Here, we will briefly describe the phenotypes of murine transgenic DS models that have been used to define a minimal trisomic region and identify Hsa21 dosage-sensitive genes.
Ts65Dn is the most commonly used model to understand the phenotypes associated with DS [90]. This strain contains 104 trisomic genes, all located on Mmu16. Ts65Dn mice display an increased number of HSC and GMP, as well as decreased MEP during adulthood [91]. These mice develop progressive myeloproliferative disorder characterized by megakaryocytic hyperplasia, thrombocytosis, and myelofibrosis in the bone marrow and spleen. Ts65Dn mice also display a decreased proportion of common lymphoid progenitors associated with a lower level of IL7 receptor expression [92].
The Dp(16); Dp(17); Dp(10) strain is the only model trisomic for all syngeneic regions of human chromosome 21. These mice display macrocytic anemia and also develop myeloproliferative disorder; these phenotypes are conserved in the Dp(16) model alone [93], strongly suggesting that the minimal region implicated in DS-myeloid disorders is contained within Mmu16. In contrast to the Ts65Dn strain, 15-month-old Dp(16); Dp(17); Dp(10) mice have an increased percentage of MEP and a decreased percentage of GMP in the bone marrow. The reason for these differences are not known to date but may result from differences in strain and/or modifier genes.
The Cre/LoxP system has been used to develop trisomy of genes contained in the Down syndrome critical region (DSCR): creating the.... Ts1Rhr model [94]. The DSCR was triplicated on Mmu16 from the CBR1 to FAM3B genes and contains 31 protein coding genes and 2 antisense RNAs [94]. The Ts1Rhr model has been extensively used to assess the role of trisomy of the DSCR in leukemia predisposition and development. Triplication of these 33 regulatory elements alone has no major effect on fetal hematopoiesis, apart from a significant increase in phenotypic HSC. Adult Ts1Rhr mice develop a phenotype similar to the Ts65Dn strain, indicating that trisomy of the DSCR is the minimally required region associated with these myeloid phenotypes [63]. Expression of Gata1s in Ts1Rhr led to increased size of CFU-megakaryocyte colonies and transient thrombocytosis mimicking features of TMD. Reproducing the multi-step pathogenesis seen in patient samples by adding a third event in the Ts1Rhr/Gata1s model has provided insight on the role of trisomy 21 in TMD/ML–DS development. First, endogenous expression of JAK3 activating mutations enhance a TMD phenotype during fetal hematopoiesis alone, reinforcing the concept of developmental stage selectivity [95]. Moreover, bone marrow transplantation assays revealed that MPLW515L overexpression functionally cooperates with Gata1s and Ts1Rhr to drive megakaryocytic hyperplasia presenting with phenotypic features of DS–AMKL [63].
The Ts1Rhr strain has also been used to assess the impact of trisomy 21 on B-cell lineage. Compared with wild-type littermates, trisomic mice display a decreased proportion of bone marrow B220+CD43+ early B-cell progenitors, especially Hardy’s fractions B and C (proB cells), and increased clonogenic potential of CFU-preB colonies [74]. Similar to the megakaryocytic lineage, these mice will not spontaneously develop B-cell leukemia and four additional events found in DS–ALL samples (CRLF2 overexpression, Jak2R683G, Pax5 haploinsufficiency and expression of the dominant negative Ikaros isoform IK6) are required to drive a B-ALL phenotype, although not to full penetrance [74]. As a surrogate, Ts1Rhr was shown to cooperate with p210 BCR-ABL overexpression to develop B-ALL in vivo with shorter latency and complete penetrance, thus demonstrating the impact of trisomy of the DSCR in B-cell leukemogenesis.
In summary, the use of partially trisomic murine models clearly emphasize the role of trisomy 21 in leukemia predisposition and development. While these models can be limited by potential differences between species, they provide the relevant genetic background to study fetal, neonatal and adult hematopoiesis, to investigate the impact of the microenvironment on leukemia predisposition and progression, to identify specific chromosome 21 genes and assess their cooperation with secondary mutations found in human DS-leukemia samples.
Dosage-sensitive Hsa21 genes and mechanisms altered by trisomy 21 in DS-leukemia
The combination of genetically engineered iPSCs, partially trisomic animal models and characterization of cases with segmental trisomy 21 in patients with DS-leukemia, have been instrumental in identifying dosage-sensitive genes implicated in leukemia predisposition and development in children with DS (Fig. 2 and Table 2) [96, 97].
Transcription factors
The chromosome 21 ERG oncogene, which encodes a transcription factor from the E-twenty-six (ETS) family, is overexpressed in DS and de novo AMKL [98]. In mice, ERG overexpression promotes the expansion of fetal megakaryocytic progenitors, cooperates with Gata1s expression and leads to AMKL in vivo [99,100,101]. Moreover, the loss of one copy of Erg (ERGmld2 mice) reverts the myeloproliferative phenotype seen in the Ts65Dn model [102]. Possible mechanisms of cooperation between ERG and the secondary alterations found in DS-leukemia include increased chromatin accessibility for ERG (along with RUNX1; another transcription factor encoded by chromosome 21), through the alteration of cohesin complex components [103], and molecular interplay between ERG and the RAS/MAPK pathway, in which ERG induces the transcriptional signature of RAS/MAPK activation and RAS/MAPK regulates ERG activity [104, 105]. Whether this feed-forward loop participates in leukemia development and maintenance in both ML–DS and DS–ALL remains to be investigated. A study using genetically engineered DS-iPSCs suggests that trisomy of ERG, together with trisomy of ETS2 and RUNX1, enhances early hematopoiesis and cooperates with GATA1s expression [85].
Signaling effectors
Trisomy of DYRK1A has been shown to promote TMD/DS–AMKL development in human and murine models [63]. DYRK1A encodes the dual-specificity tyrosine phosphorylation regulated kinase 1A; a kinase that has multiple targets, thus regulating diverse functions in a cellular context-dependent manner [106]. In murine cells, increased dosage of Dyrk1a cooperates with Gata1s expression to increase megakaryocytic expansion through inhibition of the calcineurin/NFAT pathway [63]. Another chromosome 21 gene, RCAN1 (also known as DSCR1), encodes a negative regulator of the NFAT pathway, that also contributes to megakaryopoiesis [107]. Since increased dosage of both DYRK1A and RCAN1 cooperate to inhibit neo-angiogenesis in solid tumor development through calcineurin/NFAT pathway inhibition [108], it may be reasonable to assume that a similar additive effect contributes to ML–DS.
Interestingly, Dyrk1a has also been shown to regulate B lymphopoiesis [109]. Genetic disruption or pharmacological inhibition of Dyrk1a completely abolished CFU-preB colony formation. Dyrk1a also controls the transition between proliferative large preB to quiescent small preB, by triggering cyclin D3 degradation required to exit the cell cycle. Together, these phenotypes indicate that trisomy of DYRK1A may promote B-cell leukemia and warrants further investigation.
Over-activation and hypersensitivity to interferon (IFN) signaling, resulting from increased expression of IFN-related genes located on Hsa21 (including IFNAR1, IFNAR2, IFNGR2 and IL10RB) outside of the DSCR region, has been observed in multiple cell types in individuals with DS [110]. During adulthood, individuals with DS display a perturbed immune system, consistent with a state of chronic inflammation [111, 112]. IFN and inflammatory response transcriptional signatures have been recently observed in murine and human trisomic hematopoietic progenitors, as well as in DS–ALL samples [81] (Laurent A. and Malinge S., unpublished observations), and may be partly mediated by the microenvironment. These observations emphasize the link between IFN signaling, inflammation and immune deficiency in DS. Whether this is implicated in DS–ALL development and reflects the higher rate of B-cell leukemia in children with DS by increasing the risk of infections, which has been suggested as a causal factor for childhood ALL [113, 114], is a promising area for investigation. Moreover, since interferon α signaling has an anti-proliferative effect on DS-associated myeloid disorders in adult bone marrow but not during fetal life [115], this potential interplay between IFN signaling and DS-associated leukemia may also be time and spatially dependent.
Epigenetic regulators
HMGN1 encodes the high mobility group nucleosome-binding protein N1, which modulates accessibility of the histone H3 tail to other epigenetic regulators. Decreased expression or loss of one copy of Hmgn1 reverts the CFU-preB colonies seen in the Ts1Rhr model, indicative of its key role in leukemia predisposition [74, 116]. Mechanistically, increased dosage of Hmgn1 increases global H3K27ac and is associated with upregulation of B-cell specific transcriptional signatures [116], which are conserved in DS–ALL and normally associated with a H3K27me3 repressive mark in non-trisomic samples. Another chromatin associated protein encoded by chromosome 21 is CHAF1B, a component of the CAF-1 complex known to drive the first step of nucleosome formation after replication. CHAF1B has been shown to be overexpressed in AML, where it maintains the leukemic cells in an undifferentiated state by interfering with the occupancy of the regulator of myeloid differentiation CEBPA [117]. CHAF1B is also overexpressed in DS–AMKL [63], but whether it has a similar extra-canonical function in DS-associated leukemia remains elusive to date.
Other mechanisms
Chromosome 21 encodes microRNA (miR) and alters several other mechanisms that may impact leukemia development in children with DS. miR-125b-2 is overexpressed in several leukemia subtypes including DS–AMKL and B-ALL, but not in DS–ALL [118, 119]. miR-125b-2 overexpression has been shown to enhance proliferation and self-renewal of megakaryocytic progenitors and synergize with Gata1s expression to enhance the DS–AMKL phenotype [118].
Trisomy 21 has also been associated with DNA hypomethylation in TMD/ML–DS, with downregulation of endothelin signaling and over-activity of insulin-like growth factor (IGF) signaling in DS-leukemia primary patient samples, iPSCs and murine models [62, 84, 87, 120], although the chromosome 21 genes and the underlying mechanisms associated with these molecular features remain largely unknown.
Gain of chromosome 21 in non-DS leukemia
As a somatic event, +21 is one of the most common alterations in hematological cancer (Table 3). Whether findings associated with +21 in DS-leukemia can be extrapolated to other subtypes of leukemia with +21 remains to be seen. Here, we present several subtypes of non-DS leukemia or blood cancer harboring +21, to consider whether studying leukemogenesis in children with DS can have broader application.
Gain of chromosome 21 in pediatric non-DS AMKL
Analyses of large cohorts have revealed that +21 is often seen in AMKL, occurring most frequently in children with AMKL [5]. In de novo AMKL, trisomy 21 has been found in all cytogenetically defined subtypes: ETO2-GLIS2, OTT-MAL, NUP98-KDM5A, HOX rearranged and Other [121]. Strikingly, 9.2% of childhood AMKL has a genetic background similar to DS–AMKL (DS–AMKL-like: i.e. alterations in genes encoding GATA1, cohesin complex components and signaling effectors together with acquired trisomy 21), with excellent outcome also seen for this subgroup of non-DS children [121]. DS–AMKL-like development has been reported in a patient with Cornelia de Lange syndrome, characterized by a mutation in NIPBL (encoding a cohesin complex component), with acquired trisomy 21, a GATA1 mutation, along with EZH2 and JAK/RAS alterations [122]. Moreover, underlying the impact of +21 in TMD development, rare TMD-like cases (occurrence of TMD in children without DS) have been described [123,124,125]. These studies on TMD/DS–AMKL-like disorders not only emphasize the cooperative role of trisomy 21, but also raise the question regarding the order of acquisition of these somatic events. Whether trisomy 21 is a founder alteration that is required to ‘prime’ the cellular context susceptible for GATA1s expression as highlighted in DS–TMD, or whether +21 cooperates with other genetic alterations in non-DS leukemia, regardless of whether it is an early or late event, remains unknown.
Gain of chromosome 21 in pediatric B-ALL
B-ALL is the most common type of childhood cancer, and accounts for approximately two-thirds of all pediatric acute leukemia. Complete or partial gain of chromosome 21 is one of the most frequent chromosomal alterations in childhood B-ALL, found in nearly 30% of cases compared with ~11% in adults. It is predominantly seen in HeH, which comprises 25–30% of all pediatric B-ALL, where more than 90% of cases harbor between one and three additional copies of whole chromosome 21, with the majority being tetrasomic [126]. Intra-chromosomal amplification of chromosome 21 (iAMP21) occurs in 2% of pediatric B-ALL. iAMP21 is characterized by a rearranged chromosome 21 patterned with amplified and deleted genomic regions [127, 128]. As in DS–ALL, there is a high incidence of genetic alterations affecting signaling effectors (NRAS, KRAS, FLT3, and SH2B3) in HeH and iAMP21 subtypes [129,130,131]. However, the molecular bases of this possible oncogenic cooperation are currently unknown. The observation that a minimal region of amplification on chromosome 21, found in both HeH and iAMP21 and overlapping with the DSCR [126, 128, 132] (Fig. 2), indicates that increased dosage of specific chromosome 21 genes located in this region, regardless of whether they are amplified by a constitutive or somatic alteration, may play a role in B-cell leukemogenesis.
Gain of chromosome 21 in other hematological malignancies
Trisomy 21 is found in 4–5% of AML overall (8.5% in children and 3.9% in adults) [133, 134] (Table 3). This incidence may be slightly underestimated since focal amplifications of specific regions of chromosome 21, such as band 21q22 that contains the ERG oncogene, are also found in AML [135,136,137]. In human and animal models, ERG overexpression leads to the development of lymphoid and myeloid leukemia, and promotes a stem cell and progenitor signature [138, 139]. This may provide a susceptible context for oncogenic cooperation with additional alterations that have been found in primary patient samples (cohesin complex components and RAS/MAPK signaling). Overexpression of the chromosome 21 gene CHAF1B has been shown to maintain a stem cell signature in murine AML models [117]. Whether other genes located in the 21q22 region are involved in AML development or maintenance remains unknown.
Complete or partial gain of chromosome 21 is also frequently seen in adult hematological cancers such as lymphoma of B or T-cell origin (Table 3). Trisomy 21 has been shown to be an independent risk factor in follicular lymphoma [140]. Gain of chromosome 21 is seen in 8.5% of cutaneous T-cell lymphoma (CTCL) [141, 142], and its role in CTCL development has been assessed in vivo in cooperation with JAK3 activating mutations, through establishment of a trisomic murine model of CTCL [95]. However, the chromosome 21 genes involved in these disorders and the associated molecular mechanisms require further investigation.
Perspective
A significant number of studies have been reported over the last decade. From the clinical aspect, we are now reaching a chemotherapeutic plateau and new therapies are required to further improve outcome for DS–ALL and relapsed ML–DS. Studying the molecular basis of leukemia predisposition and development in children with DS has been instrumental in dissecting the role of trisomy 21 on hematopoiesis, both alone and in cooperation with other genetic events, and led to the identification of several key dosage-sensitive chromosome 21 genes. Integration of fundamental research will provide new actionable targets to improve the outcome for children with DS. However, several important questions remain: Are the same chromosome 21 genes/mechanisms implicated in leukemia predisposition, development and maintenance? Does +21 have a role in response to treatment? Could we therapeutically target chromosome 21 proteins or the mechanisms of cooperation between +21 and somatic alterations? As +21 is frequently seen in non-DS-associated blood cancer, pursuing study of +21 in DS-leukemia may have high translational potential to ultimately provide clinical benefit for many patients with hematological malignancies.
References
Mitelman F, Heim S, Mandahl N. Trisomy 21 in neoplastic cells. Am J Med Genet. 1990;7(Suppl.):262–6.
Duijf PH, Schultz N, Benezra R. Cancer cells preferentially lose small chromosomes. Int J Cancer. 2013;132:2316–26.
Raimondi SC, Chang MN, Ravindranath Y, Behm FG, Gresik MV, Steuber CP, et al. Chromosomal abnormalities in 478 children with acute myeloid leukemia: clinical characteristics and treatment outcome in a cooperative pediatric oncology group study-POG 8821. Blood. 1999;94:3707–16.
Forestier E, Izraeli S, Beverloo B, Haas O, Pession A, Michalova K, et al. Cytogenetic features of acute lymphoblastic and myeloid leukemias in pediatric patients with Down syndrome: an iBFM-SG study. Blood. 2008;111:1575–83.
Hama A, Muramatsu H, Makishima H, Sugimoto Y, Szpurka H, Jasek M, et al. Molecular lesions in childhood and adult acute megakaryoblastic leukaemia. Br J Haematol. 2012;156:316–25.
Lejeune J, Turpin R, Gautier M. Mongolism; a chromosomal disease (trisomy). Bull Acad Natl Med. 1959;143:256–65.
Hasle H, Friedman JM, Olsen JH, Rasmussen SA. Low risk of solid tumors in persons with Down syndrome. Genet Med. 2016;18:1151–7.
Roizen NJ, Patterson D. Down’s syndrome. Lancet. 2003;361:1281–9.
Mai CT, Isenburg JL, Canfield MA, Meyer RE, Correa A, Alverson CJ, et al. National population-based estimates for major birth defects, 2010-2014. Birth Defects Res. 2019;111:1420–35.
Sherman SL, Freeman SB, Allen EG, Lamb NE. Risk factors for nondisjunction of trisomy 21. Cytogenet Genome Res. 2005;111:273–80.
Roberts I, Alford K, Hall G, Juban G, Richmond H, Norton A, et al. GATA1-mutant clones are frequent and often unsuspected in babies with Down syndrome: identification of a population at risk of leukemia. Blood. 2013;122:3908–17.
Mateos MK, Barbaric D, Byatt SA, Sutton R, Marshall GM. Down syndrome and leukemia: insights into leukemogenesis and translational targets. Transl Pediatr. 2015;4:76–92.
Taub JW, Berman JN, Hitzler JK, Sorrell AD, Lacayo NJ, Mast K, et al. Improved outcomes for myeloid leukemia of Down syndrome: a report from the Children’s Oncology Group AAML0431 trial. Blood. 2017;129:3304–13.
Uffmann M, Rasche M, Zimmermann M, von Neuhoff C, Creutzig U, Dworzak M, et al. Therapy reduction in patients with Down syndrome and myeloid leukemia: the international ML-DS 2006 trial. Blood. 2017;129:3314–21.
Buitenkamp TD, Izraeli S, Zimmermann M, Forestier E, Heerema NA, van den Heuvel-Eibrink MM, et al. Acute lymphoblastic leukemia in children with Down syndrome: a retrospective analysis from the Ponte di Legno study group. Blood. 2014;123:70–7.
Salazar EG, Li Y, Fisher BT, Rheingold SR, Fitzgerald J, Seif AE, et al. Supportive care utilization and treatment toxicity in children with Down syndrome and acute lymphoid leukaemia at free-standing paediatric hospitals in the United States. Br J Haematol. 2016;174:591–9.
Yamato G, Muramatsu H, Watanabe T, Deguchi T, Iwamoto S, Hasegawa D, et al. Predictive factors of the development of leukemia in patients with transient abnormal myelopoiesis and Down syndrome: The Jccg Study JPLSG TAM-10. Blood. 2019;134(Supplement_1):3833.
Massey GV, Zipursky A, Chang MN, Doyle JJ, Nasim S, Taub JW, et al. A prospective study of the natural history of transient leukemia (TL) in neonates with Down syndrome (DS): Children’s Oncology Group (COG) study POG-9481. Blood. 2006;107:4606–13.
Klusmann JH, Creutzig U, Zimmermann M, Dworzak M, Jorch N, Langebrake C, et al. Treatment and prognostic impact of transient leukemia in neonates with Down syndrome. Blood. 2008;111:2991–8.
Gamis AS, Alonzo TA, Gerbing RB, Hilden JM, Sorrell AD, Sharma M, et al. Natural history of transient myeloproliferative disorder clinically diagnosed in Down syndrome neonates: a report from the Children’s Oncology Group Study A2971. Blood. 2011;118:6752–9.
Flasinski M, Scheibke K, Zimmermann M, Creutzig U, Reinhardt K, Verwer F, et al. Low-dose cytarabine to prevent myeloid leukemia in children with Down syndrome: TMD Prevention 2007 study. Blood Adv. 2018;2:1532–40.
Shah A, Stiller C, Lancaster D, Vincent T, Coleman MP. Leukaemia survival trends in children with Down’s syndrome in Great Britain, 1971-2000: a population-based study. J Epidemiol Community Health. 2010;64:604–9.
Caldwell JT, Ge Y, Taub JW. Prognosis and management of acute myeloid leukemia in patients with Down syndrome. Expert Rev Hematol. 2014;7:831–40.
Kudo K, Kojima S, Tabuchi K, Yabe H, Tawa A, Imaizumi M, et al. Prospective study of a pirarubicin, intermediate-dose cytarabine, and etoposide regimen in children with Down syndrome and acute myeloid leukemia: the Japanese Childhood AML Cooperative Study Group. J Clin Oncol. 2007;25:5442–7.
Taga T, Shimomura Y, Horikoshi Y, Ogawa A, Itoh M, Okada M, et al. Continuous and high-dose cytarabine combined chemotherapy in children with down syndrome and acute myeloid leukemia: report from the Japanese children’s cancer and leukemia study group (JCCLSG) AML 9805 down study. Pediatr Blood Cancer. 2011;57:36–40.
Sorrell AD, Alonzo TA, Hilden JM, Gerbing RB, Loew TW, Hathaway L, et al. Favorable survival maintained in children who have myeloid leukemia associated with Down syndrome using reduced-dose chemotherapy on Children’s Oncology Group trial A2971: a report from the Children’s Oncology Group. Cancer. 2012;118:4806–14.
Taga T, Watanabe T, Tomizawa D, Kudo K, Terui K, Moritake H, et al. Preserved high probability of overall survival with significant reduction of chemotherapy for myeloid leukemia in Down syndrome: a nationwide prospective study in Japan. Pediatr Blood Cancer. 2016;63:248–54.
Krischer JP, Epstein S, Cuthbertson DD, Goorin AM, Epstein ML, Lipshultz SE. Clinical cardiotoxicity following anthracycline treatment for childhood cancer: the Pediatric Oncology Group experience. J Clin Oncol. 1997;15:1544–52.
O’Brien MM, Taub JW, Chang MN, Massey GV, Stine KC, Raimondi SC, et al. Cardiomyopathy in children with Down syndrome treated for acute myeloid leukemia: a report from the Children’s Oncology Group Study POG 9421. J Clin Oncol. 2008;26:414–20.
Hassler A, Bochennek K, Gilfert J, Perner C, Schoning S, Creutzig U, et al. Infectious complications in children with acute myeloid leukemia and Down syndrome: analysis of the prospective multicenter trial AML-BFM 2004. Pediatr Blood Cancer. 2016;63:1070–4.
Hitzler JK, Berman J, Gerbing RB, Beckman A, Hirsch BA, Raimondi SC, et al. High-dose cytarabine is indispensable for the survival of children with myeloid leukemia in Down syndrome Despite negative minimal residual disease Post-induction. Blood. 2019;134(Supplement_1):118.
Blink M, Zimmermann M, von Neuhoff C, Reinhardt D, de Haas V, Hasle H, et al. Normal karyotype is a poor prognostic factor in myeloid leukemia of Down syndrome: a retrospective, international study. Haematologica. 2014;99:299–307.
Hasle H, Abrahamsson J, Arola M, Karow A, O’Marcaigh A, Reinhardt D, et al. Myeloid leukemia in children 4 years or older with Down syndrome often lacks GATA1 mutation and cytogenetics and risk of relapse are more akin to sporadic AML. Leukemia. 2008;22:1428–30.
Taga T, Tanaka S, Terui K, Iwamoto S, Hiramatsu H, Miyamura T, et al. Post-induction minimal residual disease measured by flow cytometry and deep sequencing of mutant GATA1 are both significant prognostic factors for children with myeloid leukemia and Down syndrome: a nationwide prospective study of the Japanese Pediatric Leukemia/Lymphoma Study Group. Blood. 2019;134(Supplement_1):3848.
Lee P, Bhansali R, Izraeli S, Hijiya N, Crispino JD. The biology, pathogenesis and clinical aspects of acute lymphoblastic leukemia in children with Down syndrome. Leukemia. 2016;30:1816–23.
Zawitkowska J, Odoj T, Drabko K, Zaucha-Prazmo A, Rudnicka J, Romiszewski M, et al. Outcome of acute lymphoblastic leukemia in children with down syndrome-Polish pediatric leukemia and lymphoma study group report. Pediatr Hematol Oncol. 2017;34:199–205.
Athale UH, Puligandla M, Stevenson KE, Asselin B, Clavell LA, Cole PD, et al. Outcome of children and adolescents with Down syndrome treated on Dana-Farber Cancer Institute Acute Lymphoblastic Leukemia Consortium protocols 00-001 and 05-001. Pediatr Blood Cancer. 2018;65:e27256.
Matloub Y, Rabin KR, Ji L, Devidas M, Hitzler J, Xu X, et al. Excellent long-term survival of children with Down syndrome and standard-risk ALL: a report from the Children’s Oncology Group. Blood Adv. 2019;3:1647–56.
Kroll M, Kaupat-Bleckmann K, Moricke A, Alten J, Schewe DM, Stanulla M, et al. Methotrexate-associated toxicity in children with Down syndrome and acute lymphoblastic leukemia during consolidation therapy with high dose methotrexate according to ALL-BFM treatment regimen. Haematologica. 2020;105:1013–20.
Maloney KW. Acute lymphoblastic leukaemia in children with Down syndrome: an updated review. Br J Haematol 2011;155:420–5.
Meyr F, Escherich G, Mann G, Klingebiel T, Kulozik A, Rossig C, et al. Outcomes of treatment for relapsed acute lymphoblastic leukaemia in children with Down syndrome. Br J Haematol. 2013;162:98–106.
Taga T, Saito AM, Kudo K, Tomizawa D, Terui K, Moritake H, et al. Clinical characteristics and outcome of refractory/relapsed myeloid leukemia in children with Down syndrome. Blood. 2012;120:1810–5.
Rubin CM, Mick R, Johnson FL. Bone marrow transplantation for the treatment of haematological disorders in Down’s syndrome: toxicity and outcome. Bone Marrow Transplant 1996;18:533–40.
Hitzler JK, He W, Doyle J, Cairo M, Camitta BM, Chan KW, et al. Outcome of transplantation for acute myelogenous leukemia in children with Down syndrome. Biol Blood Marrow Transplant. 2013;19:893–7.
Hitzler JK, He W, Doyle J, Cairo M, Camitta BM, Chan KW, et al. Outcome of transplantation for acute lymphoblastic leukemia in children with Down syndrome. Pediatr Blood Cancer. 2014;61:1126–8.
Laetsch TW, Maude SL, Grupp SA, Boyer MW, Harris AC, Qayed M, et al. CTL019 therapy appears safe and effective in pediatric patients with Down syndrome with relapsed/refractory (r/r) acute lymphoblastic leukemia. Blood. 2017;130(Supplement 1):1280.
Rabin K, Izraeli S, Hijiya N, Hitzler J. Need for new thinking: treatment of relapsed leukemia in children with Down syndrome. Pediatr Blood Cancer. 2019;66:e27644.
Wechsler J, Greene M, McDevitt MA, Anastasi J, Karp JE, Le Beau MM, et al. Acquired mutations in GATA1 in the megakaryoblastic leukemia of Down syndrome. Nat Genet. 2002;32:148–52.
Rainis L, Bercovich D, Strehl S, Teigler-Schlegel A, Stark B, Trka J, et al. Mutations in exon 2 of GATA1 are early events in megakaryocytic malignancies associated with trisomy 21. Blood. 2003;102:981–6.
Groet J, McElwaine S, Spinelli M, Rinaldi A, Burtscher I, Mulligan C, et al. Acquired mutations in GATA1 in neonates with Down’s syndrome with transient myeloid disorder. Lancet. 2003;361:1617–20.
Alford KA, Reinhardt K, Garnett C, Norton A, Bohmer K, von Neuhoff C, et al. Analysis of GATA1 mutations in Down syndrome transient myeloproliferative disorder and myeloid leukemia. Blood. 2011;118:2222–38.
Hitzler JK, Cheung J, Li Y, Scherer SW, Zipursky A. GATA1 mutations in transient leukemia and acute megakaryoblastic leukemia of Down syndrome. Blood. 2003;101:4301–4.
Ahmed M, Sternberg A, Hall G, Thomas A, Smith O, O’Marcaigh A, et al. Natural history of GATA1 mutations in Down syndrome. Blood. 2004;103:2480–9.
Yoshida K, Toki T, Okuno Y, Kanezaki R, Shiraishi Y, Sato-Otsubo A, et al. The landscape of somatic mutations in Down syndrome-related myeloid disorders. Nat Genet. 2013;45:1293–9.
Labuhn M, Perkins K, Matzk S, Varghese L, Garnett C, Papaemmanuil E, et al. Mechanisms of progression of myeloid preleukemia to transformed myeloid leukemia in children with Down syndrome. Cancer Cell. 2019;36:123–38e10.
Chen J, Li Y, Doedens M, Wang P, Shago M, Dick JE, et al. Functional differences between myeloid leukemia-initiating and transient leukemia cells in Down’s syndrome. Leukemia. 2010;24:1012–7.
Saida S, Watanabe K, Sato-Otsubo A, Terui K, Yoshida K, Okuno Y, et al. Clonal selection in xenografted TAM recapitulates the evolutionary process of myeloid leukemia in Down syndrome. Blood. 2013;121:4377–87.
Nikolaev SI, Santoni F, Vannier A, Falconnet E, Giarin E, Basso G, et al. Exome sequencing identifies putative drivers of progression of transient myeloproliferative disorder to AMKL in infants with Down syndrome. Blood. 2013;122:554–61.
Li Z, Godinho FJ, Klusmann JH, Garriga-Canut M, Yu C, Orkin SH. Developmental stage-selective effect of somatically mutated leukemogenic transcription factor GATA1. Nat Genet. 2005;37:613–9.
Gruber TA, Downing JR. The biology of pediatric acute megakaryoblastic leukemia. Blood. 2015;126:943–9.
Cuartero S, Innes AJ, Merkenschlager M. Towards a better understanding of cohesin mutations in AML. Front Oncol. 2019;9:867.
Klusmann JH, Godinho FJ, Heitmann K, Maroz A, Koch ML, Reinhardt D, et al. Developmental stage-specific interplay of GATA1 and IGF signaling in fetal megakaryopoiesis and leukemogenesis. Genes Dev. 2010;24:1659–72.
Malinge S, Bliss-Moreau M, Kirsammer G, Diebold L, Chlon T, Gurbuxani S, et al. Increased dosage of the chromosome 21 ortholog Dyrk1a promotes megakaryoblastic leukemia in a murine model of Down syndrome. J Clin Investig. 2012;122:948–62.
Russell LJ, Capasso M, Vater I, Akasaka T, Bernard OA, Calasanz MJ, et al. Deregulated expression of cytokine receptor gene, CRLF2, is involved in lymphoid transformation in B-cell precursor acute lymphoblastic leukemia. Blood. 2009;114:2688–98.
Mullighan CG, Collins-Underwood JR, Phillips LA, Loudin MG, Liu W, Zhang J, et al. Rearrangement of CRLF2 in B-progenitor- and Down syndrome-associated acute lymphoblastic leukemia. Nat Genet. 2009;41:1243–6.
Hertzberg L, Vendramini E, Ganmore I, Cazzaniga G, Schmitz M, Chalker J, et al. Down syndrome acute lymphoblastic leukemia, a highly heterogeneous disease in which aberrant expression of CRLF2 is associated with mutated JAK2: a report from the International BFM Study Group. Blood. 2010;115:1006–17.
Schwartzman O, Savino AM, Gombert M, Palmi C, Cario G, Schrappe M, et al. Suppressors and activators of JAK-STAT signaling at diagnosis and relapse of acute lymphoblastic leukemia in Down syndrome. Proc Natl Acad Sci USA. 2017;114:E4030–9.
Kubota Y, Uryu K, Ito T, Seki M, Kawai T, Isobe T, et al. Integrated genetic and epigenetic analysis revealed heterogeneity of acute lymphoblastic leukemia in Down syndrome. Cancer Sci. 2019;110:3358–67.
Bercovich D, Ganmore I, Scott LM, Wainreb G, Birger Y, Elimelech A, et al. Mutations of JAK2 in acute lymphoblastic leukaemias associated with Down’s syndrome. Lancet. 2008;372:1484–92.
Yoda A, Yoda Y, Chiaretti S, Bar-Natan M, Mani K, Rodig SJ, et al. Functional screening identifies CRLF2 in precursor B-cell acute lymphoblastic leukemia. Proc Natl Acad Sci USA. 2010;107:252–7.
Shochat C, Tal N, Bandapalli OR, Palmi C, Ganmore I, te Kronnie G, et al. Gain-of-function mutations in interleukin-7 receptor-alpha (IL7R) in childhood acute lymphoblastic leukemias. J Exp Med. 2011;208:901–8.
Nikolaev SI, Garieri M, Santoni F, Falconnet E, Ribaux P, Guipponi M, et al. Frequent cases of RAS-mutated Down syndrome acute lymphoblastic leukaemia lack JAK2 mutations. Nat Commun. 2014;5:4654.
Brown AL, de Smith AJ, Gant VU, Yang W, Scheurer ME, Walsh KM, et al. Inherited genetic susceptibility to acute lymphoblastic leukemia in Down syndrome. Blood. 2019;134:1227–37.
Lane AA, Chapuy B, Lin CY, Tivey T, Li H, Townsend EC, et al. Triplication of a 21q22 region contributes to B cell transformation through HMGN1 overexpression and loss of histone H3 Lys27 trimethylation. Nat Genet. 2014;46:618–23.
Sewastianik T, Jiang M, Sukhdeo K, Patel SS, Roberts K, Kang Y, et al. Constitutive Ras signaling and Ink4a/Arf inactivation cooperate during the development of B-ALL in mice. Blood Adv. 2017;1:2361–74.
Jamrog L, Chemin G, Fregona V, Coster L, Pasquet M, Oudinet C, et al. PAX5-ELN oncoprotein promotes multistep B-cell acute lymphoblastic leukemia in mice. Proc Natl Acad Sci USA. 2018;115:10357–62.
Chou ST, Opalinska JB, Yao Y, Fernandes MA, Kalota A, Brooks JS, et al. Trisomy 21 enhances human fetal erythro-megakaryocytic development. Blood. 2008;112:4503–6.
Tunstall-Pedoe O, Roy A, Karadimitris A, de la Fuente J, Fisk NM, Bennett P, et al. Abnormalities in the myeloid progenitor compartment in Down syndrome fetal liver precede acquisition of GATA1 mutations. Blood. 2008;112:4507–11.
Roy A, Cowan G, Mead AJ, Filippi S, Bohn G, Chaidos A, et al. Perturbation of fetal liver hematopoietic stem and progenitor cell development by trisomy 21. Proc Natl Acad Sci USA. 2012;109:17579–84.
Wagenblast E, Gan OI, Azkanaz M, Smith SA, Araújo J, Shakib L, et al. Understanding pre-leukemia in trisomy 21 human HSC and modeling progression towards Down syndrome associated leukemia Using CRISPR/Cas9 at single cell resolution. Blood. 2019;134(Supplement_1):2531.
O’Byrne SI, Elliott N, Buck G, Rice S, O’Connor D, Oswald J, et al. Trisomy 21 driven pro-inflammatory signalling in fetal bone marrow may play a role in perturbed B-Lymphopoiesis and acute lymphoblastic leukemia of Down syndrome. Blood. 2019;134(Supplement_1):1206.
Maclean GA, Menne TF, Guo G, Sanchez DJ, Park IH, Daley GQ, et al. Altered hematopoiesis in trisomy 21 as revealed through in vitro differentiation of isogenic human pluripotent cells. Proc Natl Acad Sci USA. 2012;109:17567–72.
Chou ST, Byrska-Bishop M, Tober JM, Yao Y, Vandorn D, Opalinska JB, et al. Trisomy 21-associated defects in human primitive hematopoiesis revealed through induced pluripotent stem cells. Proc Natl Acad Sci USA. 2012;109:17573–8.
Chiang JC, Jiang J, Newburger PE, Lawrence JB. Trisomy silencing by XIST normalizes Down syndrome cell pathogenesis demonstrated for hematopoietic defects in vitro. Nat Commun. 2018;9:5180.
Banno K, Omori S, Hirata K, Nawa N, Nakagawa N, Nishimura K, et al. Systematic cellular disease models reveal synergistic interaction of trisomy 21 and GATA1 mutations in hematopoietic abnormalities. Cell Rep. 2016;15:1228–41.
Byrska-Bishop M, VanDorn D, Campbell AE, Betensky M, Arca PR, Yao Y, et al. Pluripotent stem cells reveal erythroid-specific activities of the GATA1 N-terminus. J Clin Investig. 2015;125:993–1005.
MacLean GA, McEldoon J, Huang J, Allred J, Canver MC, Orkin SH. Downregulation of endothelin receptor B contributes to defective B cell lymphopoiesis in trisomy 21 pluripotent stem cells. Sci Rep. 2018;8:8001.
Carsetti R, Valentini D, Marcellini V, Scarsella M, Marasco E, Giustini F, et al. Reduced numbers of switched memory B cells with high terminal differentiation potential in Down syndrome. Eur J Immunol. 2015;45:903–14.
Malinge S, Izraeli S, Crispino JD. Insights into the manifestations, outcomes, and mechanisms of leukemogenesis in Down syndrome. Blood. 2009;113:2619–28.
Davisson MT, Schmidt C, Akeson EC. Segmental trisomy of murine chromosome 16: a new model system for studying Down syndrome. Prog Clin Biol Res. 1990;360:263–80.
Kirsammer G, Jilani S, Liu H, Davis E, Gurbuxani S, Le Beau MM, et al. Highly penetrant myeloproliferative disease in the Ts65Dn mouse model of Down syndrome. Blood. 2008;111:767–75.
Lorenzo LP, Chen H, Shatynski KE, Clark S, Yuan R, Harrison DE, et al. Defective hematopoietic stem cell and lymphoid progenitor development in the Ts65Dn mouse model of Down syndrome: potential role of oxidative stress. Antioxid Redox Signal. 2011;15:2083–94.
Liu C, Yu T, Xing Z, Jiang X, Li Y, Pao A, et al. Triplications of human chromosome 21 orthologous regions in mice result in expansion of megakaryocyte-erythroid progenitors and reduction of granulocyte-macrophage progenitors. Oncotarget. 2018;9:4773–86.
Olson LE, Richtsmeier JT, Leszl J, Reeves RH. A chromosome 21 critical region does not cause specific Down syndrome phenotypes. Science. 2004;306:687–90.
Rivera-Munoz P, Laurent AP, Siret A, Lopez CK, Ignacimouttou C, Cornejo MG, et al. Partial trisomy 21 contributes to T-cell malignancies induced by JAK3-activating mutations in murine models. Blood Adv. 2018;2:1616–27.
Korbel JO, Tirosh-Wagner T, Urban AE, Chen XN, Kasowski M, Dai L, et al. The genetic architecture of Down syndrome phenotypes revealed by high-resolution analysis of human segmental trisomies. Proc Natl Acad Sci USA. 2009;106:12031–6.
Takahashi T, Inoue A, Yoshimoto J, Kanamitsu K, Taki T, Imada M, et al. Transient myeloproliferative disorder with partial trisomy 21. Pediatr Blood Cancer. 2015;62:2021–4.
Rainis L, Toki T, Pimanda JE, Rosenthal E, Machol K, Strehl S, et al. The proto-oncogene ERG in megakaryoblastic leukemias. Cancer Res. 2005;65:7596–602.
Salek-Ardakani S, Smooha G, de Boer J, Sebire NJ, Morrow M, Rainis L, et al. ERG is a megakaryocytic oncogene. Cancer Res. 2009;69:4665–73.
Stankiewicz MJ, Crispino JD. ETS2 and ERG promote megakaryopoiesis and synergize with alterations in GATA-1 to immortalize hematopoietic progenitor cells. Blood. 2009;113:3337–47.
Birger Y, Goldberg L, Chlon TM, Goldenson B, Muler I, Schiby G, et al. Perturbation of fetal hematopoiesis in a mouse model of Down syndrome’s transient myeloproliferative disorder. Blood. 2013;122:988–98.
Ng AP, Hyland CD, Metcalf D, Carmichael CL, Loughran SJ, Di Rago L, et al. Trisomy of Erg is required for myeloproliferation in a mouse model of Down syndrome. Blood. 2010;115:3966–9.
Mazumdar C, Shen Y, Xavy S, Zhao F, Reinisch A, Li R, et al. Leukemia-associated cohesin mutants dominantly enforce stem cell programs and impair human hematopoietic progenitor differentiation. Cell Stem Cell. 2015;17:675–88.
Goldberg L, Tijssen MR, Birger Y, Hannah RL, Kinston SJ, Schutte J, et al. Genome-scale expression and transcription factor binding profiles reveal therapeutic targets in transgenic ERG myeloid leukemia. Blood. 2013;122:2694–703.
Huang Y, Thoms JA, Tursky ML, Knezevic K, Beck D, Chandrakanthan V, et al. MAPK/ERK2 phosphorylates ERG at serine 283 in leukemic cells and promotes stem cell signatures and cell proliferation. Leukemia. 2016;30:1552–61.
Kay LJ, Smulders-Srinivasan TK, Soundararajan M. Understanding the multifaceted role of human Down syndrome kinase DYRK1A. Adv Protein Chem Struct Biol. 2016;105:127–71.
Zaslavsky A, Chou ST, Schadler K, Lieberman A, Pimkin M, Kim YJ, et al. The calcineurin-NFAT pathway negatively regulates megakaryopoiesis. Blood. 2013;121:3205–15.
Baek KH, Zaslavsky A, Lynch RC, Britt C, Okada Y, Siarey RJ, et al. Down’s syndrome suppression of tumour growth and the role of the calcineurin inhibitor DSCR1. Nature. 2009;459:1126–30.
Thompson BJ, Bhansali R, Diebold L, Cook DE, Stolzenburg L, Casagrande AS, et al. DYRK1A controls the transition from proliferation to quiescence during lymphoid development by destabilizing Cyclin D3. J Exp Med. 2015;212:953–70.
Sullivan KD, Lewis HC, Hill AA, Pandey A, Jackson LP, Cabral JM, et al. Trisomy 21 consistently activates the interferon response. eLife. 2016;5:e16220.
Sullivan KD, Evans D, Pandey A, Hraha TH, Smith KP, Markham N, et al. Trisomy 21 causes changes in the circulating proteome indicative of chronic autoinflammation. Sci Rep. 2017;7:14818.
Waugh KA, Araya P, Pandey A, Jordan KR, Smith KP, Granrath RE, et al. Mass cytometry reveals global immune remodeling with multi-lineage hypersensitivity to Type I interferon in Down Syndrome. Cell Rep. 2019;29:1893–908 e4.
Martin-Lorenzo A, Hauer J, Vicente-Duenas C, Auer F, Gonzalez-Herrero I, Garcia-Ramirez I, et al. Infection exposure is a causal factor in B-cell precursor acute lymphoblastic leukemia as a result of Pax5-inherited susceptibility. Cancer Discov. 2015;5:1328–43.
Greaves M. A causal mechanism for childhood acute lymphoblastic leukaemia. Nat Rev Cancer. 2018;18:471–84.
Woo AJ, Wieland K, Huang H, Akie TE, Piers T, Kim J, et al. Developmental differences in IFN signaling affect GATA1s-induced megakaryocyte hyperproliferation. J Clin Invest. 2013;123:3292–304.
Mowery CT, Reyes JM, Cabal-Hierro L, Higby KJ, Karlin KL, Wang JH, et al. Trisomy of a Down syndrome critical region globally amplifies transcription via HMGN1 overexpression. Cell Rep. 2018;25:1898–911 e5.
Volk A, Liang K, Suraneni P, Li X, Zhao J, Bulic M, et al. A CHAF1B-dependent molecular switch in hematopoiesis and leukemia pathogenesis. Cancer Cell. 2018;34:707–23 e7.
Klusmann JH, Li Z, Bohmer K, Maroz A, Koch ML, Emmrich S, et al. miR-125b-2 is a potential oncomiR on human chromosome 21 in megakaryoblastic leukemia. Genes Dev. 2010;24:478–90.
Gefen N, Binder V, Zaliova M, Linka Y, Morrow M, Novosel A, et al. Hsa-mir-125b-2 is highly expressed in childhood ETV6/RUNX1 (TEL/AML1) leukemias and confers survival advantage to growth inhibitory signals independent of p53. Leukemia. 2010;24:89–96.
Malinge S, Chlon T, Dore LC, Ketterling RP, Tallman MS, Paietta E, et al. Development of acute megakaryoblastic leukemia in Down syndrome is associated with sequential epigenetic changes. Blood. 2013;122:e33–43.
de Rooij JD, Branstetter C, Ma J, Li Y, Walsh MP, Cheng J, et al. Pediatric non-Down syndrome acute megakaryoblastic leukemia is characterized by distinct genomic subsets with varying outcomes. Nat Genet. 2017;49:451–6.
Vial Y, Lachenaud J, Verloes A, Besnard M, Fenneteau O, Lainey E, et al. Down syndrome-like acute megakaryoblastic leukemia in a patient with Cornelia de Lange syndrome. Haematologica. 2018;103:e274–6.
Apollonsky N, Shende A, Ouansafi I, Brody J, Atlas M, Aygun B. Transient myeloproliferative disorder in neonates with and without Down syndrome: a tale of 2 syndromes. J Pediatr Hematol Oncol. 2008;30:860–4.
Tsai MH, Hou JW, Yang CP, Yang PH, Chu SM, Hsu JF, et al. Transient myeloproliferative disorder and GATA1 mutation in neonates with and without Down syndrome. Indian J Pediatr. 2011;78:826–32.
Ono R, Hasegawa D, Hirabayashi S, Kamiya T, Yoshida K, Yonekawa S, et al. Acute megakaryoblastic leukemia with acquired trisomy 21 and GATA1 mutations in phenotypically normal children. Eur J Pediatr. 2015;174:525–31.
Paulsson K, Forestier E, Lilljebjorn H, Heldrup J, Behrendtz M, Young BD, et al. Genetic landscape of high hyperdiploid childhood acute lymphoblastic leukemia. Proc Natl Acad Sci USA. 2010;107:21719–24.
Harrison CJ, Moorman AV, Schwab C, Carroll AJ, Raetz EA, Devidas M, et al. An international study of intrachromosomal amplification of chromosome 21 (iAMP21): cytogenetic characterization and outcome. Leukemia. 2014;28:1015–21.
Li Y, Schwab C, Ryan S, Papaemmanuil E, Robinson HM, Jacobs P, et al. Constitutional and somatic rearrangement of chromosome 21 in acute lymphoblastic leukaemia. Nature. 2014;508:98–102.
Malinowska-Ozdowy K, Frech C, Schonegger A, Eckert C, Cazzaniga G, Stanulla M, et al. KRAS and CREBBP mutations: a relapse-linked malicious liaison in childhood high hyperdiploid acute lymphoblastic leukemia. Leukemia. 2015;29:1656–67.
Paulsson K. High hyperdiploid childhood acute lymphoblastic leukemia: chromosomal gains as the main driver event. Mol Cell Oncol. 2016;3:e1064555.
Sinclair PB, Ryan S, Bashton M, Hollern S, Hanna R, Case M, et al. SH2B3 inactivation through CN-LOH 12q is uniquely associated with B-cell precursor ALL with iAMP21 or other chromosome 21 gain. Leukemia. 2019;33:1881–94.
Rand V, Parker H, Russell LJ, Schwab C, Ensor H, Irving J, et al. Genomic characterization implicates iAMP21 as a likely primary genetic event in childhood B-cell precursor acute lymphoblastic leukemia. Blood. 2011;117:6848–55.
Harrison CJ, Hills RK, Moorman AV, Grimwade DJ, Hann I, Webb DK, et al. Cytogenetics of childhood acute myeloid leukemia: United Kingdom Medical Research Council Treatment trials AML 10 and 12. J Clin Oncol. 2010;28:2674–81.
Grimwade D, Hills RK, Moorman AV, Walker H, Chatters S, Goldstone AH, et al. Refinement of cytogenetic classification in acute myeloid leukemia: determination of prognostic significance of rare recurring chromosomal abnormalities among 5876 younger adult patients treated in the United Kingdom Medical Research Council trials. Blood. 2010;116:354–65.
Baldus CD, Liyanarachchi S, Mrozek K, Auer H, Tanner SM, Guimond M, et al. Acute myeloid leukemia with complex karyotypes and abnormal chromosome 21: Amplification discloses overexpression of APP, ETS2, and ERG genes. Proc Natl Acad Sci USA. 2004;101:3915–20.
Weber S, Haferlach C, Jeromin S, Nadarajah N, Dicker F, Noel L, et al. Gain of chromosome 21 or amplification of chromosome arm 21q is one mechanism for increased ERG expression in acute myeloid leukemia. Genes Chromosomes Cancer. 2016;55:148–57.
Nibourel O, Guihard S, Roumier C, Pottier N, Terre C, Paquet A, et al. Copy-number analysis identified new prognostic marker in acute myeloid leukemia. Leukemia. 2017;31:555–64.
Carmichael CL, Metcalf D, Henley KJ, Kruse EA, Di Rago L, Mifsud S, et al. Hematopoietic overexpression of the transcription factor Erg induces lymphoid and erythro-megakaryocytic leukemia. Proc Natl Acad Sci USA. 2012;109:15437–42.
Tursky ML, Beck D, Thoms JA, Huang Y, Kumari A, Unnikrishnan A, et al. Overexpression of ERG in cord blood progenitors promotes expansion and recapitulates molecular signatures of high ERG leukemias. Leukemia. 2015;29:819–27.
Mitsui T, Yokohama A, Koiso H, Saito A, Toyama K, Shimizu H, et al. Prognostic impact of trisomy 21 in follicular lymphoma. Br J Haematol. 2019;184:570–7.
da Silva Almeida AC, Abate F, Khiabanian H, Martinez-Escala E, Guitart J, Tensen CP, et al. The mutational landscape of cutaneous T cell lymphoma and Sezary syndrome. Nat Genet. 2015;47:1465–70.
Choi J, Goh G, Walradt T, Hong BS, Bunick CG, Chen K, et al. Genomic landscape of cutaneous T cell lymphoma. Nat Genet. 2015;47:1011–9.
Derouet A, Petit A, Baruchel A, Clavel J, Brethon B, Bertrand Y, et al. Impact of therapy in a cohort of unselected children with Down syndrome associated Acute lymphoblastic leukaemia. Br J Haematol. 2016;174:983–5.
Klampfl T, Harutyunyan A, Berg T, Gisslinger B, Schalling M, Bagienski K, et al. Genome integrity of myeloproliferative neoplasms in chronic phase and during disease progression. Blood. 2011;118:167–76.
Gondek LP, Tiu R, O’Keefe CL, Sekeres MA, Theil KS, Maciejewski JP. Chromosomal lesions and uniparental disomy detected by SNP arrays in MDS, MDS/MPD, and MDS-derived AML. Blood. 2008;111:1534–42.
Schneider NR, Carroll AJ, Shuster JJ, Pullen DJ, Link MP, Borowitz MJ, et al. New recurring cytogenetic abnormalities and association of blast cell karyotypes with prognosis in childhood T-cell acute lymphoblastic leukemia: a pediatric oncology group report of 343 cases. Blood. 2000;96:2543–9.
Acknowledgements
APL was supported by Cancéropôle Ile-de-France and Fondation pour la Recherche Médicale. RSK is supported by a Fellowship from the National Health and Medical Research Council of Australia (NHMRC APP1142627) and the Children’s Leukaemia and Cancer Research Foundation (CLCRF, Australia). SM is supported by the Children’s Leukaemia and Cancer Research Foundation (CLCRF, Australia), Fondation Jérôme Lejeune (#1806, France), and by a Fellowship from Cancer Council Western Australia (CCWA).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Laurent, A.P., Kotecha, R.S. & Malinge, S. Gain of chromosome 21 in hematological malignancies: lessons from studying leukemia in children with Down syndrome. Leukemia 34, 1984–1999 (2020). https://doi.org/10.1038/s41375-020-0854-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41375-020-0854-5
This article is cited by
-
Inherent genome instability underlies trisomy 21-associated myeloid malignancies
Leukemia (2024)
-
Effects of aneuploidy on cell behaviour and function
Nature Reviews Molecular Cell Biology (2022)
-
Trib1 promotes the development of acute myeloid leukemia in a Ts1Cje mouse model of Down syndrome
Leukemia (2022)
-
Cytogenetics analysis as the central point of genetic testing in acute myeloid leukemia (AML): a laboratory perspective for clinical applications
Clinical and Experimental Medicine (2022)
-
Clinical and biological aspects of myeloid leukemia in Down syndrome
Leukemia (2021)