Abstract
Background
Leptin regulates satiety and energy homoeostasis, and plays a key role in placentation in pregnancy. Previous studies have demonstrated regulation of leptin gene (LEP) expression and/or methylation in placenta and cord blood in association with early life exposures, but most have been small and have not considered the influence of genetic variation. Here, we investigated the relationship between maternal factors in pregnancy, infant anthropometry and LEP genetic variation with LEP promoter methylation at birth and 12 months of age.
Methods
LEP methylation was measured in cord (n = 877) and 12-month (n = 734) blood in the Barwon Infant Study, a population-based pre-birth cohort. Infant adiposity at birth and 12-months was measured as triceps and subscapular skinfold thickness. Cross-sectional regression tested associations of methylation with pregnancy and anthropometry measures, while longitudinal regression tested if birth anthropometry predicted 12-month LEP methylation levels.
Results
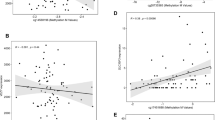
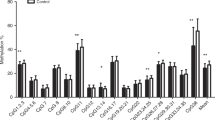
Male infants had lower LEP methylation in cord blood (−2.07% average methylation, 95% CI (−2.92, −1.22), p < 0.001). Genetic variation strongly influenced DNA methylation at a single CpG site, which was also negatively associated with birth weight (r = −0.10, p = 0.003). Pre-eclampsia was associated with lower cord blood methylation at another CpG site (−6.06%, 95% CI (−10.70, −1.42), p = 0.01). Gestational diabetes was more modestly associated with methylation at two other CpG units. Adiposity at birth was associated with 12-month LEP methylation, modified by rs41457646 genotype. There was no association of LEP methylation with 12-month anthropometric measures.
Conclusions
Infant sex, weight, genetic variation, and exposure to pre-eclampsia and gestational diabetes, are associated with LEP methylation in cord blood. Infant adiposity at birth predicts 12-month blood LEP methylation in a genotype-dependent manner. These findings are consistent with genetics and anthropometry driving altered LEP epigenetic profile and expression in infancy. Further work is required to confirm this and to determine the long-term impact of altered LEP methylation on health.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Data and code availability
The data and code used in this analysis are available upon reasonable request.
References
Drake AJ, Reynolds RM. Impact of maternal obesity on offspring obesity and cardiometabolic disease risk. Reproduction. 2010;140:387–98.
West NA, Crume TL, Maligie MA, Dabelea D. Cardiovascular risk factors in children exposed to maternal diabetes in utero. Diabetologia. 2011;54:504–7.
Davis EF, Lazdam M, Lewandowski AJ, Worton SA, Kelly B, Kenworthy Y, et al. Cardiovascular risk factors in children and young adults born to preeclamptic pregnancies: a systematic review. Pediatrics. 2012;129:e1552–e61.
Smith CJ, Ryckman KK. Epigenetic and developmental influences on the risk of obesity, diabetes, and metabolic syndrome. Diabetes Metab Syndr Obes. 2015;8:295–302.
Friedman JM, Halaas JL. Leptin and the regulation of body weight in mammals. Nature. 1998;395:763.
Grattan DR, Ladyman SR, Augustine RA. Hormonal induction of leptin resistance during pregnancy. Physiol Behav. 2007;91:366–74.
Jansson N, Greenwood S, Johansson B, Powell T, Jansson T. Leptin stimulates the activity of the system A amino acid transporter in human placental villous fragments. J Clin Endocrinol Metab. 2003;88:1205–11.
Myers MG,Jr. Leibel RL, Seeley RJ, Schwartz MW. Obesity and leptin resistance: distinguishing cause from effect. Trends Endocrinol Metab. 2010;21:643–51.
Briffa JF, McAinch AJ, Romano T, Wlodek ME, Hryciw DH. Leptin in pregnancy and development: a contributor to adulthood disease? Am J Physiol Endocrinol Metab. 2015;308:E335–50.
Green ED, Maffei M, Braden VV, Proenca R, DeSilva U, Zhang Y, et al. The human obese (OB) gene: RNA expression pattern and mapping on the physical, cytogenetic, and genetic maps of chromosome 7. Genome Res. 1995;5:5–12.
Bado A, Levasseur S, Attoub S, Kermorgant S, Laigneau J-P, Bortoluzzi M-N, et al. The stomach is a source of leptin. Nature. 1998;394:790.
Hogg K, Robinson WP, Beristain AG. Activation of endocrine-related gene expression in placental choriocarcinoma cell lines following DNA methylation knock-down. Mol Hum Reprod. 2014;20:677–89.
Melzner I, Scott V, Dorsch K, Fischer P, Wabitsch M, Brüderlein S, et al. Leptin gene expression in human preadipocytes is switched on by maturation-induced demethylation of distinct CpGs in its proximal promoter. J Biol Chem. 2002;277:45420–7.
Iliopoulos D, Malizos KN, Tsezou A. Epigenetic regulation of leptin affects MMP-13 expression in osteoarthritic chondrocytes: possible molecular target for osteoarthritis therapeutic intervention. Ann Rheum Dis. 2007;66:1616–21.
Lesseur C, Armstrong DA, Paquette AG, Koestler DC, Padbury JF, Marsit CJ. Tissue-specific Leptin promoter DNA methylation is associated with maternal and infant perinatal factors. Mol Cell Endocrinol. 2013;381:160–7.
Kadakia R, Zheng Y, Zhang Z, Zhang W, Hou L, Josefson JL. Maternal pre-pregnancy BMI downregulates neonatal cord blood LEP methylation. Pediatr Obes. 2017;12(S1):57–64.
Allard C, Desgagne V, Patenaude J, Lacroix M, Guillemette L, Battista MC, et al. Mendelian randomization supports causality between maternal hyperglycemia and epigenetic regulation of leptin gene in newborns. Epigenetics. 2015;10:342–51.
Lesseur C, Armstrong DA, Paquette AG, Li Z, Padbury JF, Marsit CJ. Maternal obesity and gestational diabetes are associated with placental leptin DNA methylation. Am JObstet Gynecol. 2014;211:654.e1–9.
Bouchard L, Thibault S, Guay SP, Santure M, Monpetit A, St-Pierre J, et al. Leptin gene epigenetic adaptation to impaired glucose metabolism during pregnancy. Diabetes Care. 2010;33:2436–41.
Hogg K, Blair JD, von Dadelszen P, Robinson WP. Hypomethylation of the LEP gene in placenta and elevated maternal leptin concentration in early onset pre-eclampsia. Mol Cell Endocrinol. 2013;367:64–73.
Xiang Y, Cheng Y, Li X, Li Q, Xu J, Zhang J, et al. Up-regulated expression and aberrant DNA methylation of LEP and SH3PXD2A in pre-eclampsia. PloS ONE. 2013;8:e59753.
Laivuori H, Gallaher M, Collura L, Crombleholme W, Markovic N, Rajakumar A, et al. Relationships between maternal plasma leptin, placental leptin mRNA and protein in normal pregnancy, pre-eclampsia and intrauterine growth restriction without pre-eclampsia. Mol Hum reprod. 2006;12:551–6.
Ødegård RA, Vatten LJ, Nilsen ST, Salvesen KÅ, Austgulen R. Umbilical cord plasma leptin is increased in preeclampsia. Am J Obstet Gynecol. 2002;186:427–32.
Ng EK, Leung TN, Tsui NB, Lau TK, Panesar NS, Chiu RW, et al. The concentration of circulating corticotropin-releasing hormone mRNA in maternal plasma is increased in preeclampsia. Clini Chem. 2003;49:727–31.
Tian F-Y, Rifas-Shiman SL, Cardenas A, Baccarelli AA, DeMeo DL, Litonjua AA, et al. Maternal corticotropin-releasing hormone is associated with LEP DNA methylation at birth and in childhood: an epigenome-wide study in Project Viva. Int J Obes. 2018:1.
Wang Y-H, Xu X-X, Sun H, Han Y, Lei Z-F, Wang Y-C, et al. Cord blood leptin DNA methylation levels are associated with macrosomia during normal pregnancy. Pediatr Res. 2019;86:305–10.
Xu X, Yang X, Liu Z, Wu K, Liu Z, Lin C, et al. Placental leptin gene methylation and macrosomia during normal pregnancy. Mol Med Rep. 2014;9:1013–8.
Tobi EW, Lumey LH, Talens RP, Kremer D, Putter H, Stein AD, et al. DNA methylation differences after exposure to prenatal famine are common and timing- and sex-specific. Hum Mol Genet. 2009;18:4046–53.
Schultz NS, Broholm C, Gillberg L, Mortensen B, Jorgensen SW, Schultz HS, et al. Impaired leptin gene expression and release in cultured preadipocytes isolated from individuals born with low birth weight. Diabetes. 2014;63:111–21.
Souren NY, Paulussen AD, Steyls A, Loos RJ, Stassen AP, Gielen M, et al. Common SNPs in LEP and LEPR associated with birth weight and type 2 diabetes-related metabolic risk factors in twins. Int J Obes. 2008;32:1233.
Jiang Y, Wilk J, Borecki I, Williamson S, DeStefano A, Xu G, et al. Common variants in the 5′ region of the leptin gene are associated with body mass index in men from the National Heart, Lung, and Blood Institute Family Heart Study. Am J Hum Genetics. 2004;75:220–30.
Hager J, Clement K, Francke S, Dina C, Raison J, Lahlou N, et al. A polymorphism in the 5′ untranslated region of the human ob gene is associated with low leptin levels. Int J Obes. 1998;22:200.
Meirhaeghe A, Cottel D, Amouyel P, Dallongeville J. Lack of association between certain candidate gene polymorphisms and the metabolic syndrome. Mol Genetics Metab. 2005;86:293–9.
Kilpeläinen TO, Carli JFM, Skowronski AA, Sun Q, Kriebel J, Feitosa MF, et al. Genome-wide meta-analysis uncovers novel loci influencing circulating leptin levels. Nat Commun. 2016;7:10494.
Vuillermin P, Saffery R, Allen KJ, Carlin JB, Tang ML, Ranganathan S, et al. Cohort Profile: The Barwon Infant Study. Int J Epidemiol. 2015;44:1148–60.
Nankervis A, McIntyre HD, Moses RG, Ross GP, Callaway LK. Testing for gestational diabetes mellitus in Australia. Diabetes Care. 2013;36:e64.
Tranquilli A, Dekker G, Magee L, Roberts J, Sibai B, Steyn W, et al. The classification, diagnosis and management of the hypertensive disorders of pregnancy: a revised statement from the ISSHP. Pregnancy Hypertens. 2014;4:97.
Schmelzle HR, Fusch C. Body fat in neonates and young infants: validation of skinfold thickness versus dual-energy X-ray absorptiometry. Am J Clin Nutr. 2002;76:1096–100.
de Onis M, Onyango AW, Van den Broeck J, Chumlea CW, Martorell R. Measurement and standardization protocols for anthropometry used in the construction of a new international growth reference. Food Nutr Bull. 2004;25(1_suppl1):S27–S36.
Cole TJ, Williams AF, Wright CM. Revised birth centiles for weight, length and head circumference in the UK-WHO growth charts. Ann Hum Biol. 2011;38:7–11.
WMGRS Group, Onis de, Growth MWHOChild. Standards based on length/height, weight and age. Acta Paediatr. 2006;95:76–85.
Mansell T, Novakovic B, Meyer B, Rzehak P, Vuillermin P, Ponsonby AL, et al. The effects of maternal anxiety during pregnancy on IGF2/H19 methylation in cord blood. Transl Psychiatry. 2016;6:e765.
Lam D, Ancelin ML, Ritchie K, Saffery R, Ryan J. DNA methylation and genetic variation of the angiotensin converting enzyme (ACE) in depression. Psychoneuroendocrinology. 2018;88:1–8.
Collier FM, Tang ML, Martino D, Saffery R, Carlin J, Jachno K, et al. The ontogeny of naïve and regulatory CD4+ T-cell subsets during the first postnatal year: a cohort study. ClinTransl Immunol. 2015;4:e34.
McCarthy S, Das S, Kretzschmar W, Delaneau O, Wood AR, Teumer A, et al. A reference panel of 64,976 haplotypes for genotype imputation. Nat Genet. 2016;48:1279–83.
Turner S, Armstrong LL, Bradford Y, Carlson CS, Crawford DC, Crenshaw AT, et al. Quality Control Procedures for Genome-Wide Association Studies. Curr Protoc Hum Genetics. 2011;68:1.19.1–1.8.
Martins MC, Trujillo J, Farias DR, GJNR Kac. Polymorphisms in the leptin (rs7799039) gene are associated with an increased risk of excessive gestational weight gain but not with leptin concentration during pregnancy. 2017;47:53–62.
Marcello MA, Calixto AR, de Almeida JFM, Martins MB, Cunha LL, Cavalari CAA, et al. Polymorphism in LEP and LEPR may modify leptin levels and represent risk factors for thyroid cancer. Int J Endocrinol. 2015;2015:1–8
Obermann-Borst SA, Eilers PH, Tobi EW, de Jong FH, Slagboom PE, Heijmans BT, et al. Duration of breastfeeding and gender are associated with methylation of the LEPTIN gene in very young children. Pediatr Res. 2013;74:344.
Muhlhausler BS, Duffield J, McMillen IC. Increased maternal nutrition stimulates peroxisome proliferator activated receptor-γ, adiponectin, and leptin messenger ribonucleic acid expression in adipose tissue before birth. Endocrinology. 2007;148:878–85.
Lesseur C, Armstrong DA, Murphy MA, Appleton AA, Koestler DC, Paquette AG, et al. Sex-specific associations between placental leptin promoter DNA methylation and infant neurobehavior. Psychoneuroendocrinology. 2014;40:1–9.
Krempler F, Breban D, Oberkofler H, Esterbauer H, Hell E, Paulweber B, et al. Leptin, peroxisome proliferator-activated receptor-γ, and CCAAT/enhancer binding protein-α mRNA expression in adipose tissue of humans and their relation to cardiovascular risk factors. Arterioscler Thromb Vas Biol. 2000;20:443–9.
Ong KKL, Ahmed ML, Sherriff A, Woods KA, Watts A, Golding J, et al. Cord blood leptin is associated with size at birth and predicts infancy weight gain in humans. J Clin Endocrinol Metab. 1999;84:1145–8.
Sayeed SK, Zhao J, Sathyanarayana BK, Golla JP, Vinson C. C/EBPβ (CEBPB) protein binding to the C/EBP| CRE DNA 8-mer TTGC| GTCA is inhibited by 5hmC and enhanced by 5mC, 5fC, and 5caC in the CG dinucleotide. Biochim et Biophys Acta. 2015;1849:583–9.
Mise H, Sagawa N, Matsumoto T, Yura S, Nanno H, Itoh H, et al. Augmented placental production of leptin in preeclampsia: possible involvement of placental hypoxia. J Clinl Endocrinol Metab. 1998;83:3225–9.
El Hajj N, Pliushch G, Schneider E, Dittrich M, Müller T, Korenkov M, et al. Metabolic programming of MEST DNA methylation by intrauterine exposure to gestational diabetes mellitus. Diabetes. 2013;62:1320–8.
Sherwood WB, Bion V, Lockett GA, Ziyab AH, Soto-Ramírez N, Mukherjee N, et al. Duration of breastfeeding is associated with leptin (LEP) DNA methylation profiles and BMI in 10-year-old children. Clin Epigenetics. 2019;11:1–10.
Saenen ND, Vrijens K, Janssen BG, Roels HA, Neven KY, Vanden Berghe W, et al. Lower placental leptin promoter methylation in association with fine particulate matter air pollution during pregnancy and placental nitrosative stress at birth in the ENVIRONAGE Cohort. Environmental health perspectives. 2017;125:262–8.
Wang Y, Eliot MN, Kuchel GA, Schwartz J, Coull BA, Mittleman MA, et al. Long-term exposure to ambient air pollution and serum leptin in older adults: results from the MOBILIZE Boston study. J Occup Environ Med. 2014;56:e73.
Pereira-Fernandes A, Dirinck E, Dirtu AC, Malarvannan G, Covaci A, Van Gaal L, et al. Expression of obesity markers and persistent organic pollutants levels in adipose tissue of obese patients: reinforcing the obesogen hypothesis? PloS ONE. 2014;9:e84816.
Kamstra JH, Hruba E, Blumberg B, Janesick A, Mandrup S, Hamers T, et al. Transcriptional and epigenetic mechanisms underlying enhanced in vitro adipocyte differentiation by the brominated flame retardant BDE-47. Environ Sci Technol. 2014;48:4110–9.
Acknowledgements
We thank QIMR Berghofer Medical Research Institute and the Erasmus MC University Medical Center for their role in coordinating and performing the genotyping of BIS samples. The establishment work and infrastructure for the BIS was provided by the Murdoch Children’s Research Institute, Deakin University and Barwon Health. Subsequent funding was secured from the National Health and Medical Research Council of Australia, The Jack Brockhoff Foundation, the Scobie Trust, the Shane O’Brien Memorial Asthma Foundation, the Our Women’s Our Children’s Fund Raising Committee Barwon Health, The Shepherd Foundation, the Rotary Club of Geelong, the Ilhan Food Allergy Foundation, GMHBA Limited and the Percy Baxter Charitable Trust, Perpetual Trustees. In-kind support was provided by the Cotton On Foundation and CreativeForce. The study sponsors were not involved in the collection, analysis, and interpretation of data; writing the report; or the decision to submit the report for publication. Research at Murdoch Children’s Research Institute is supported by the Victorian Government's Operational Infrastructure Support Program. This work was also supported by a Research Training Program Stipend through University of Melbourne [to TM], NHMRC Senior Research Fellowships [APP1008396 to ALP; APP1045161 to RS]; and an NHMRC Dementia Research Leader Fellowship [APP1135727 to JR].
Author information
Authors and Affiliations
Consortia
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Mansell, T., Ponsonby, AL., Collier, F. et al. Genetic variation, intrauterine growth, and adverse pregnancy conditions predict leptin gene DNA methylation in blood at birth and 12 months of age. Int J Obes 44, 45–56 (2020). https://doi.org/10.1038/s41366-019-0472-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41366-019-0472-3