Abstract
Background
Deterioration of the adipogenic potential of preadipocytes may contribute to adipose tissue dysfunction in obesity and type 2 diabetes (T2D). Here, we hypothesized that extracellular factors in obesity epigenetically reprogram adipogenesis potential and metabolic function of preadipocytes.
Methods
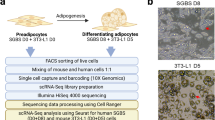
The transcriptomic profile of visceral adipose tissue preadipocytes collected from Lean, Obese and Obese with T2D was assessed throughout in vitro differentiation using RNA sequencing. Reduced Representation Bisulfite Sequencing was used to establish the genome-wide DNA methylation profile of human preadipocytes and 3T3-L1 preadipocytes treated by the inflammatory cytokine Tumour Necrosis Factor-α (TNF-α) or palmitate.
Results
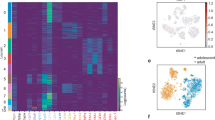
While preadipocytes from all obese subjects (Obese+Obese T2D), compared to those of Lean, were transcriptionally different in response to differentiation in culture, preadipocytes from Obese T2D showed impaired insulin signalling and a further transcriptomic shift towards altered adipocyte function. Cultures with a lower expression magnitude of adipogenic genes throughout differentiation (PLIN1, CIDEC, FABP4, ADIPOQ, LPL, PDK4, APOE, LIPE, FABP3, LEP, RBP4 and CD36) were associated with DNA methylation remodelling at genes controlling insulin sensitivity and adipocytokine signalling pathways. Prior incubation of 3T3-L1 preadipocytes with TNF-α or palmitate markedly altered insulin responsiveness and metabolic function in the differentiated adipocytes, and remodelled DNA methylation and gene expression at specific genes, notably related to PPAR signalling.
Conclusions
Our findings that preadipocytes retain the memory of the donor in culture and can be reprogrammed by extracellular factors support a mechanism by which adipocyte precursors are epigenetically reprogrammed in vivo. Epigenetic reprogramming of preadipocytes represents a mechanism by which metabolic function of visceral adipose tissue may be affected in the long term by past exposure to obesity- or T2D-specific factors.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Hotamisligil GS, Arner P, Caro JF, Atkinson RL, Spiegelman BM. Increased adipose tissue expression of tumor necrosis factor-a in human obesity and insulin resistance. J Clin Invest. 1995;95:2409–15.
Hajer GR, Van Haeften TW, Visseren FLJ. Adipose tissue dysfunction in obesity, diabetes, and vascular diseases. Eur Hear J. 2008;29:2959–71.
Rønningen T, Shah A, Reiner AH, Collas P, Moskaug JØ. Epigenetic priming of inflammatory response genes by high glucose in adipose progenitor cells. Biochem Biophys Res Commun. 2015;467:979–86.
Isakson P, Hammarstedt A, Gustafson B, Smith U. Impaired preadipocyte differentiation in human abdominal obesity: role of Wnt, tumor necrosis factor-alpha, and inflammation. Diabetes. 2009;58:1550–7.
Permana PA, Nair S, Lee Y, Luczy-bachman G, De Courten BV, Tataranni PA, et al. Subcutaneous abdominal preadipocyte differentiation in vitro inversely correlates with central obesity. Am J Physiol Endocrinol Metab. 2004;286:958–62.
Lessard J, Laforest S, Pelletier M, Leboeuf M, Blackburn L, Tchernof A. Low abdominal subcutaneous preadipocyte adipogenesis is associated with visceral obesity, visceral adipocyte hypertrophy, and a dysmetabolic state. Adipocyte. 2014;3:197–205.
Rossmeislová L, Mališová L, Kračmerová J, Tencerová M, Kováčová Z, Koc M, et al. Weight loss improves the adipogenic capacity of human preadipocytes and modulates their secretory profile. Diabetes. 2013;62:1990–5.
Siersbæk MS, Loft A, Aagaard MM, Nielsen R, Schmidt SF, Petrovic N, et al. Genome-wide profiling of peroxisome proliferator-activated receptor γ in primary epididymal, inguinal, and brown adipocytes reveals depot-selective binding correlated with gene expression. Mol Cell Biol. 2012;32:3452–63.
Mikkelsen TS, Xu Z, Zhang X, Wang L, Gimble JM, Lander ES, et al. Comparative epigenomic analysis of murine and human adipogenesis. Cell. 2010;143:156–69.
Benton MC, Johnstone A, Eccles D, Harmon B, Hayes MT, Lea RA, et al. An analysis of DNA methylation in human adipose tissue reveals differential modification of obesity genes before and after gastric bypass and weight loss. Genome Biol. 2015;16:8:1–21.
Nylander V, Ingerslev LR, Andersen E, Fabre O, Garde C, Rasmussen M, et al. Ionizing radiation potentiates high-fat diet–induced insulin resistance and reprograms skeletal muscle and adipose progenitor cells. Diabetes. 2016;65:3573–84.
Nilsson E, Jansson PA, Perfilyev A, Volkov P, Pedersen M, Svensson MK, et al. Altered DNA methylation and differential expression of genes influencing metabolism and inflammation in adipose tissue from subjects with type 2 diabetes. Diabetes. 2014;63:2962–276.
Rönn T, Volkov P, Davegårdh C, Dayeh T, Hall E, Olsson AH, et al. A six months exercise intervention influences the genome-wide DNA methylation pattern in human adipose tissue. PLoS Genet. 2013;9:e1003572.
Mikkelsen TS, Ku M, Jaffe DB, Issac B, Lieberman E, Giannoukos G, et al. Genome-wide maps of chromatin state in pluripotent and lineage-committed cells. Nature. 2007;448:553–60.
Reik W. Stability and flexibility of epigenetic gene regulation in mammalian development. Nature. 2007;447:425–32.
Arner P, Arner E, Hammarstedt A, Smith U. Genetic predisposition for type 2 diabetes, but not for overweight/obesity, is associated with a restricted adipogenesis. PLoS ONE. 2011;6:e18284.
Pattamaprapanont P, Garde C, Fabre O, Barrès R. Muscle contraction induces acute hydroxymethylation of the exercise-responsive gene Nr4a3. Front Endocrinol (Lausanne). 2016;7:1–9.
Yates A, Akanni W, Amode MR, Barrell D, Billis K, Carvalho-Silva D, et al. Ensembl 2016. Nucleic Acids Res. 2016;44:D710–6.
Bray NL, Pimentel H, Melsted P, Pachter L. Near-optimal probabilistic RNA-seq quantification. Nat Biotechnol. 2016;34:525–7.
Pimentel H, Bray NL, Puente S, Melsted P, Pachter L. Differential analysis of RNA-seq incorporating quantification uncertainty. Nat Methods. 2017;14:687–90.
Busser H, Najar M, Raicevic G, Pieters K, Velez Pombo R, Philippart P, et al. Isolation and characterization of human mesenchymal stromal cell subpopulations: comparison of bone marrow and adipose tissue. Stem Cells Dev. 2015;24:2142–57.
Boyle P, Clement K, Gu H, Smith ZD, Ziller M, Fostel JL, et al. Gel-free multiplexed reduced representation bisulfite sequencing for large-scale DNA methylation profiling. Genome Biol. 2012;13:R92.
Krueger F, Andrews SR. Bismark: a flexible aligner and methylation caller for Bisulfite-Seq applications. Bioinformatics. 2011;27:1571–2.
Hebestreit K, Dugas M, Klein H-U. Detection of significantly differentially methylated regions in targeted bisulfite sequencing data. Bioinformatics. 2013;29:1647–53.
Bjornsson HT, Sigurdsson MI, Fallin MD, Irizarry RA, Aspelund T, Cui H, et al. Intra-individual change over time in DNA methylation with familial clustering. JAMA. 2008;299:2877–83.
Broholm C, Olsson AH, Perfilyev A, Gillberg L, Hansen NS, Ali A, et al. Human adipogenesis is associated with genome-wide DNA methylation and gene-expression changes. Epigenomics. 2016;8:1601–17.
Van Harmelen V, Röhrig K, Hauner H. Comparison of proliferation and differentiation capacity of human adipocyte precursor cells from the omental and subcutaneous adipose tissue depot of obese subjects. Metabolism. 2004;53:632–7.
Pinnick KE, Nicholson G, Manolopoulos KN, McQuaid SE, Valet P, Frayn KN, et al. Distinct developmental profile of lower-body adipose tissue defines resistance against obesity-associated metabolic complications. Diabetes. 2014;63:3785–97.
Baglioni S, Cantini G, Poli G, Francalanci M, Squecco R, Franco A, et al. Functional differences in visceral and subcutaneous fat pads originate from differences in the adipose stem cell. PLoS ONE. 2012;7:e36569.
Böhm A, Halama A, Meile T, Zdichavsky M, Lehmann R, Weigert C, et al. Metabolic signatures of cultured human adipocytes from metabolically healthy versus unhealthy obese individuals. PLoS ONE. 2014;9:e93148.
Naukkarinen J, Heinonen S, Hakkarainen A, Lundbom J, Vuolteenaho K. Characterising metabolically healthy obesity in weight-discordant monozygotic twins. Diabetologia. 2014;57:167–76.
van Tienen FHJ, van der Kallen CJH, Lindsey PJ, Wanders RJ, van Greevenbroek MM, Smeets HJM. Preadipocytes of type 2 diabetes subjects display an intrinsic gene expression profile of decreased differentiation capacity. Int J Obes. 2011;35:1154–64.
Henninger AMJ, Eliasson B, Jenndahl LE, Hammarstedt A. Adipocyte hypertrophy, inflammation and fibrosis characterize subcutaneous adipose tissue of healthy, non-obese subjects predisposed to type 2 diabetes. PLoS ONE. 2014;9:1–8.
Oliva-olivera W, Gea AL, Lhamyani S, Coín-aragüez L, Torres JA, Bernal-lópez MR, et al. Differences in the osteogenic differentiation capacity of omental adipose-derived stem cells in syndrome. Endocrinology. 2015;156:4492–501.
Arner P, Sinha I, Thorell A, Rydén M, Dahlman-Wright K, Dahlman I. The epigenetic signature of subcutaneous fat cells is linked to altered expression of genes implicated in lipid metabolism in obese women. Clin Epigenet. 2015;7:1–13.
Broholm C, Olsson AH, Perfilyev A, Hansen NS, Schrölkamp M, Strasko KS, et al. Epigenetic programming of adipose-derived stem cells in low birthweight individuals. Diabetologia. 2016;59:2664–73.
Guénard F, Tchernof A, Deshaies Y, Pérusse L, Biron S, Lescelleur O, et al. Differential methylation in visceral adipose tissue of obese men discordant for metabolic disturbances. Physiol Genom. 2014;46:216–22.
Collas P. Programming differentiation potential in mesenchymal stem cells. Epigenetics. 2010;5:476–82.
Barrès R, Osler ME, Yan J, Rune A, Fritz T, Caidahl K, et al. Non-CpG Methylation of the PGC-1α promoter through DNMT3B controls mitochondrial density. Cell Metab. 2009;10:189–98.
Sharples AP, Polydorou I, Hughes DC, Owens DJ, Hughes TM, Stewart CE. Skeletal muscle cells possess a ‘memory’ of acute early life TNF-a exposure: role of epigenetic adaptation. Biogerontology. 2016;17:603–17.
Asterholm IW, Tao C, Morley TS, Wang QA, Delgado-Lopez F, Wang ZV, et al. Adipocyte inflammation is essential for healthy adipose tissue expansion and remodeling. Cell Metab. 2014;20:103–18.
Acknowledgements
OF was recipient of a research grant from the Danish Diabetes Academy supported by the Novo Nordisk Foundation. We would like to acknowledge The Danish National High-Throughput DNA Sequencing Centre, University of Copenhagen, for sequencing services. The Novo Nordisk Foundation Centre for Basic Metabolic Research is an independent research centre at the University of Copenhagen partially funded by an unrestricted donation from the Novo Nordisk Foundation.
Author contribution:
EA collected human samples, performed experiments, analysed the data and wrote the manuscript; LRI and AA performed bioinformatics analysis and generated figures; OF performed experiments, analysed the data and edited the manuscript; ID, SV, TB and VBK collected human samples, contributed to study design and edited the manuscript; DS provided expert advice and edited the manuscript; RB designed the study, analysed the data and wrote the manuscript. All authors approved the final version of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest:
The authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Andersen, E., Ingerslev, L.R., Fabre, O. et al. Preadipocytes from obese humans with type 2 diabetes are epigenetically reprogrammed at genes controlling adipose tissue function. Int J Obes 43, 306–318 (2019). https://doi.org/10.1038/s41366-018-0031-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41366-018-0031-3
This article is cited by
-
Age-dependent genes in adipose stem and precursor cells affect regulation of fat cell differentiation and link aging to obesity via cellular and genetic interactions
Genome Medicine (2024)
-
Identification and characterization of the long non-coding RNA NFIA-AS2 as a novel locus for body mass index in American Indians
International Journal of Obesity (2023)
-
Role of secretomes in cell-free therapeutic strategies in regenerative medicine
Cell and Tissue Banking (2023)
-
Insulin resistance rewires the metabolic gene program and glucose utilization in human white adipocytes
International Journal of Obesity (2022)
-
Epigenetics of type 2 diabetes mellitus and weight change — a tool for precision medicine?
Nature Reviews Endocrinology (2022)