Abstract
In this review, we discuss the ways in which recent studies of low-complexity (LC) domains have challenged our understanding of the mechanisms underlying cellular organization. LC sequences, long believed to function in the absence of a molecular structure, are abundant in the proteomes of all eukaryotic organisms. Over the past decade, the phase separation of LC domains has emerged as a fundamental mechanism driving dynamic multivalent interactions of many cellular processes. We review the key evidence showing the role of phase separation of individual proteins in organizing cellular assemblies and facilitating biological function while implicating the dynamics of phase separation as a key to biological validity and functional utility. We also highlight the evidence showing that pathogenic LC proteins alter various phase separation-dependent interactions to elicit debilitating human diseases, including cancer and neurodegenerative diseases. Progress in understanding the biology of phase separation may offer useful hints toward possible therapeutic interventions to combat the toxicity of pathogenic proteins.
Similar content being viewed by others
Introduction
Over 20% of eukaryotic proteomes consist of polypeptide sequences that are of low complexity (LC)1,2,3,4,5. Instead of using a balanced distribution of the 20 amino acids typically deployed to facilitate the folding of a protein into a distinct, three-dimensional shape, LC domains are often composed of a limited number of amino acids6. A major group of proteins carrying LC domains includes the RNA- and DNA-binding proteins used in gene regulation7. Many of these proteins function as dynamic ribonucleoprotein (RNP) complexes in membraneless organelles, and it has long been recognized that LC domains are present in the major constituents and promoters of membraneless organelle assembly8,9. Other examples of LC domain-containing proteins include intermediate filaments, proteins of the central channel of nuclear pores, and the membrane-bound proteins of the Golgi apparatus and mitochondria10,11,12,13,14. Simply put, LC domains are deployed liberally in all aspects of cell biology.
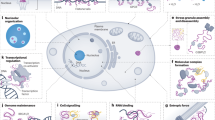
LC domains have long been thought to function in the absence of a molecular order and thus are termed intrinsically disordered. A decade ago, however, it was observed that these LC domains can undergo phase separation out of aqueous solution to form either liquid-like droplets (LLDs) or amyloid-like reversible polymers upon in vitro incubation15,16 (Fig. 1). A large body of research has revealed that phase separation provides a general strategy for the formation of membraneless organelles such as nucleoli, stress granules, and Cajal bodies by mediating reversible and dynamic protein–protein or protein–RNA interactions8,9. Moreover, phase separation-based interactions can contribute to the formation of a dynamic complex of proteins and RNA in a variety of cellular processes, including gene regulation, DNA repair, cell fate decisions, and even immune responses17,18,19. Consistently, disturbances in the phase separation process can cause disease. Cancer-related proteins can drive aberrant gene transcription and tumorigenesis through unbridled phase separation of the LC domains20,21. Amyotrophic lateral sclerosis (ALS)- or frontotemporal dementia (FTD)-causing mutations impair the dynamics and reversibility of phase separation, leading to disturbances in the dynamics and function of the RNP complex22,23.
Many LC domain-containing proteins consist of structured domains, such as the RBD (RNA binding domain, rectangles) and LC domain (dark blue lines). While previously thought to function in the absence of structural order, LC domains undergo reversible phase separation into liquid-like droplets and/or amyloid-like polymers. By mediating homotypic or heterotypic protein‒protein interactions, phase separation can regulate the functions of LC domain-containing proteins in diverse biological processes. The dynamic and reversible nature of phase separation is key to its biological validity and functional utility.
Phase separation is now recognized as a fundamental principle in cellular organization. It provides the molecular basis of the dynamics of intracellular spatial organization that cannot be readily explained by traditional lock-and-key models of protein‒protein interactions. If we are to properly understand the role of phase separation in disease, we must learn how phase separation-mediated interactions actually work in biological reactions and processes. In this review, we discuss phase separation, highlight the regulatory mechanisms of phase separation, and describe how studies of model systems have revealed a role for controlled and reversible phase separation in cellular function and diseases.
Phase separation of the LC domains
LC domains known to undergo phase separation
How can we demonstrate that a specific protein is capable of phase separation? The most common way to initially detect phase separation is light microscopy24,25. In this technique, a solution of protein samples is first observed under conditions in which there are no droplets. Then, by making a change in conditions to favor phase separation (e.g., by adding or diluting salt, changing the pH or temperature, or adding RNA) followed by incubation of the sample for a fixed amount of time (minutes or hours), the solution can be imaged microscopically, and the presence of droplets can be identified. Upon prolonged incubation, the LC domains can also transition into hydrogels composed of amyloid-like polymers. The gelation of LC domain polymers has been adapted into confocal microscopic assays, wherein small hydrogel droplets are formed on the wells of chamber slides26. The polymeric fibers can be further imaged using fluorescence or electron microscopy24,26,27.
Many LC domains that undergo phase separation contain repeats of the aromatic amino acids tyrosine or phenylalanine10. Tyrosine residues are usually flanked by either glycine or serine residues (G/S-Y-G/S), while phenylalanine residues have a glycine residue on their amino or carboxyl sides (FG motifs). Mutational analysis has revealed that repetitive tyrosine or phenylalanine residues are critical to the phase separation and function of LC domains. For example, tyrosine-to-serine (Y-to-S) substitutions in a triplet sequence (G/S-Y-G/S) within the LC domain of FUS or TAF15 effectively impede phase transition into the hydrogel-like state composed of polymers21 (Fig. 2a). In the case of the key nucleolar protein fibrillarin, we found that phenylalanine-to-serine (F-to-S) substitutions of the FG repeats are sufficient to interrupt the phase transition of fibrillarin into LLDs or hydrogel droplets and to prevent the incorporation of fibrillarin into hydrogels or LLDs composed of wild-type LC domains25 (Fig. 2b). Similarly, an F-to-S substitution in the LC domain of heterogeneous nuclear ribonucleoprotein A2 (hnRNPA2) can interrupt the incorporation of the LC domain into preexisting LLDs or hydrogel droplets28. In addition, studies of the LAF-1 P-granule protein revealed that the arginine/glycine-rich (RGG) domain plays a key role in driving phase separation29. Despite years of research, however, the molecular code that drives phase separation continues to be enigmatic30. Cross-β, π:π, and cation:π interactions have been proposed to be important drivers of reversible phase separation10,31, but many key questions regarding the chemical basis of phase separation remain unanswered. Further research using mutations in the LC domains will eventually yield knowledge to enable prediction of the properties of phase separation based on amino acid sequences.
a Mutation of tyrosine residues in the FUS LC domain impairs both hydrogel binding and stress granule association. The [G/S]-Y-[G/S] triplet repeats in the FUS LC domain are shown at the top. When the GFP-linked WT or mutant LC domains harboring 5 or 15 Y-to-S mutations are assayed for binding to mCherry hydrogel droplets composed of the FUS LC domain (mCherry:FUS-LC), moderate retention is observed for the 5 Y-to-S mutant, and little retention is observed for the 15 Y-to-S mutant. As FUS is a major constituent of stress granules, WT FUS localizes to stress granules in human U2OS cells upon application of sodium arsenite. Predictably, the 5 Y-to-S mutant localizes to stress granules, but the 15 Y-to-S mutant does not. Cell nuclei were visualized with DAPI. b Mutation of phenylalanine residues in the fibrillarin LC domain impaired both LLD formation and nucleolar localization. The 6 FG repeats in the fibrillarin LC domain are shown at the top. In the LLD formation assay, droplets formed by the F18/40 S LC domain are smaller in size than those made by the WT LC domain, and no droplets are assembled by the FallS LC domain, in which all 6 phenylalanine residues are substituted with serine. These mutations have correlative effects on the ability of fibrillarin to localize to the nucleolus in living cells. WT fibrillarin is evenly localized to the nucleolar dense fibrillar component region, but F18/40 S and FallS fibrillarin show aberrant, dot-like expression patterns similar to those observed with fibrillarin without the LC domain (ΔLC) (red signal). Nucleoli and nuclei are visualized with anti-nucleophosmin (green) or DAPI (blue), respectively. Panel a reproduced from Kato et al.16; Panel b reproduced from Kim et al.25.
Control of reversible phase separation
Phase separation into LLDs and polymers is both dynamic and reversible (Fig. 1). Although morphologically indistinguishable from the irreversible pathogenic amyloid fibers, phase separation-driven polymeric fibers are readily labile to disassembly upon dilution, as assayed by semidenaturing detergent agarose gel electrophoresis (SDD-AGE)6. Pathogenic amyloid polymers remain intact and migrate through electrophoresis gels as large polymers, but even upon simple dilution, phase separation-driven polymeric fibers dissolve into gel-loading buffer and migrate in the monomeric state into SDD-AGE gels. Another test involves the application of aliphatic alcohols to determine whether the phase separation of the LC domain is dynamic and reversible. The McKnight Laboratory demonstrated the capacity of 1,6-hexanediol (1,6-HD) to melt LLDs, polymers, and intracellular organelles known to be enriched in LC domain-containing proteins, including RNA granules, nuclear speckles, and Cajal bodies6. As discussed below, the dynamic nature of phase separation seems to be key to its biological relevance.
Substantial evidence suggests that the control of reversible phase separation occurs through posttranslational modifications (PTMs). A variety of PTMs can alter the charge, hydrophobicity, size, and structure of LC domains. Phosphorylation is the most common PTM32, and phosphorylation by kinases and dephosphorylation by phosphatases provide a major control mechanism for many fundamental processes in eukaryotic cells. The phosphoryl group is negatively charged, so its addition changes a polar, uncharged residue into a negatively charged one. Depending on the protein context, the phosphate modification of an amino acid can either favor or disfavor phase separation. FUS is an example of serine/threonine phosphorylation that disrupts phase separation. The McKnight and the Tycko groups mapped 14 sites of DNA-dependent protein kinase (DNAPK)-mediated phosphorylation in the LC domain of FUS proteins and mutated pairs of serine and threonine residues to alanine as a means of blocking phosphorylation at specific sites. They found that phosphorylation by DNAPK disrupts the hydrogel binding of FUS LC domains16 and melts droplets of the FUS LC domain33. One possible explanation for this is that the negative charges introduced by phosphorylation may exert repulsive forces that reduce polymer stability6. The inhibition of phase separation by phosphorylation has been further documented for the head domains of intermediate filaments14. All 73 human intermediate filament proteins share a common domain structure consisting of a central α-helical rod domain flanked by an N-terminal head and a C-terminal tail, which are both LC domains. The head domains of the neurofilament light (NFL) and desmin intermediate filaments undergo phase separation, and this phase separation is required for assembly into mature intermediate filaments13. The McKnight and the Tycko groups have also shown that the PKA-mediated phosphorylation of the head domains of the NFL and desmin intermediate filaments released the head domains from the hydrogels14. This is in line with the known role of phosphorylation in promoting filament disassembly34,35.
Phosphorylation can promote the phase separation potential of certain proteins. Fragile X mental retirement protein (FMRP) is an RNA-binding protein found in neuronal granules and is involved in many biological processes, such as pre-mRNA processing36, translational regulation37, neural granule transport38, and ion channel binding39. Forman-Kay and colleagues found that phosphorylation in the LC domain of FMRP by casein kinase II promotes phase separation in vitro40. Similar results were observed for the microtubule-binding protein tau. Phosphorylation displayed facilitative effects for the phase separation of both tau K18 segments41 and full-length tau42. This effect was more obvious as the number of phosphorylation sites increased. Phosphomimetic mutants (serine and threonine mutated into glutamate) can only simulate this change to a limited extent, suggesting that, in addition to introducing negative charges, phosphorylation could also change the conformation of tau42.
Methionine oxidation provides an additional means of regulating LC domain phase separation. Yeast ataxin-2, also known as Pbp1, senses the activity state of mitochondria and is critical for autophagy upon changes in the supply of growth nutrients43. Kato et al. showed that the methionines of ataxin-2 LC domains can be oxidized both in vitro and in vivo and that oxidation leads to the melting of ataxin-2 LLDs27. Conversely, the H2O2-mediated melting of LLDs is reversed through the re-reduction of oxidized methionines via the coupled reactions of two methionine sulfoxide reductases, thioredoxin, thioredoxin reductase, and NADPH. A follow-up study showed that phase separation of TDP43 is also regulated by methionine oxidation, which implies that methionine residues might endow ataxin-2 and TDP43 with the capacity to sense the cellular redox state43,44.
Related to the important role of RGG domains in phase separation, PTMs of arginine residues are also important regulators of phase separation. For example, the arginine methylation of FUS reduces phase separation and stress granule association45. Conversely, FUS hypomethylation, a molecular phenotype of FUS inclusions in FTD, drives FUS gelation to more stable cross-β structures46. In addition to phosphorylation, oxidation, and methylation, a number of other PTMs have been cataloged and implicated in tuning phase separation47,48,49,50,51,52. Future structural studies may allow us to understand how PTMs impact intermolecular interactions and hence tune the phase separation behavior.
Phase separation in cellular function
The biological functions of a protein depend on its physical interaction with other molecules. By mediating reversible and dynamic protein–protein interactions, phase separation can regulate the function of LC domain-containing proteins. Phase separation-dependent interactions can be achieved via two modes: homotypic interactions between the same LC domain and heterotypic interactions between different LC domains (Fig. 1). In the following, we provide two archetypes of the functional repertoire of phase separation; however, evidence for a variety of possible functions of phase separation in cells is still being acquired, and it is necessary to identify the full functions and roles of phase separation.
Phase separation is a mechanism for membraneless organelle assembly
Inside eukaryotic cells, macromolecules are partitioned into membrane-bound compartments, including the nucleus, lysosomes, endoplasmic reticulum, chloroplast, mitochondria, and Golgi apparatus. Their membranes create discrete chemical environments and achieve separation of constituents from the bulk cytoplasm. Enclosing membrane-bound compartments requires dedicated machinery to construct and maintain the lipid bilayer and transport substances across the membrane53,54. Many other well-known intracellular structures, including the nucleolus, Cajal bodies, nuclear speckles, paraspeckles, stress granules, and P granules, lack membranes55,56,57, introducing the potential for greater dynamics. These membraneless organelles rapidly exchange components with the cellular milieu, and their properties are readily altered in response to environmental cues, often implicating membraneless organelles in response to stress signaling. However, the mechanistic principles of their assembly and disassembly remain unclear.
Phase separation is an appealing answer. In 2009, a study of P granules (RNA and protein-containing bodies in nematode embryos) showed that they exhibit liquid-like behavior in vivo and that LAF-1, a DDX3 RNA helicase found in P granules, phase separates into P granule-like droplets in vitro. That study demonstrated that RNAi knockdown of LAF-1 results in the dissolution of P granules in the early embryo, suggesting that LAF-1 droplets are important for P granule assembly15.
In 2012, McKnight and colleagues demonstrated that the components of stress granules—namely, the LC domains of FUS and hnRNPA2 RNA-binding proteins—can reversibly phase separate into polymeric and amyloid-like fibers, and this reversible transition can mediate their dynamic movement in and out of stress granules16. A stress granule is a cytoplasmic membraneless organelle that forms in response to a variety of cellular stressors and signaling and promotes cell survival by condensing translationally stalled mRNAs, ribosomal components, translation initiation factors, and RNA-binding proteins. The researchers showed that mCherry:FUS and mCherry:hnRNPA2 hydrogel droplets are capable of trapping the LC domains of heterologous RNA-binding proteins found in stress granules. Y-to-S mutations in G/S-Y-G/S triplet motifs of the FUS LC domain that abolish phase separation have correlative effects on the ability of FUS to be incorporated into stress granules in living cells (Fig. 2a). Furthermore, phosphorylation by DNAPK interferes with the phase separation of the FUS LC domain, explaining the dynamic translocation of FUS upon DNAPK signaling33. Together, these findings suggest that reversible phase separation can drive the inclusion and exclusion of RNA-binding proteins in stress granules in a way that can be regulated by the local concentration of RNA-binding proteins and PTMs, such as phosphorylation. Subsequent studies demonstrated that the phase separation of LC domains in a variety of different RNA-binding proteins—including TDP-43, TIA1, Lsm, RBM14, nucleophosmin, and fibrillarin25,58,59,60,61,62,63,64,65—can contribute to the assembly of stress granules, P bodies, paraspeckles, Cajal bodies, and nucleoli. For example, phase separation of the N-terminal LC domain in fibrillarin regulates its binding to RNA-binding proteins and proper nucleolus localization25. The nucleolus has a multilayer organization, which has been proposed to underlie the sequential assembly of ribosomal subunits66. Notably, LC hydrogels of fibrillarin can trap not only the same LC domains via homotypic protein–protein interactions but also heterotypic LC domains derived from RNA-binding proteins other than fibrillarin. Mutational analysis demonstrates that F-to-S mutations in FG repeats of the fibrillarin LC domain that abolish phase separation prevent both the interaction of fibrillarin with RNA-binding proteins and the normal localization of fibrillarin into dense fibrillar components within the nucleolus (Fig. 2b).
The notion that phase separation drives the dynamic assembly of membraneless organelles is further supported by the observation that the disease-causing mutations in the LC domain of RNA-binding proteins not only reduce the dynamics of the respective LC domain polymers but also reduce the RNP granule dynamics and functions in cells. For the RNA-binding proteins hnRNPA1, hnRNPA2, and hnRNPDL, D-to-V mutations in the LC domain have been identified in patients with ALS and limb-girdle muscular dystrophy67,68. The Taylor group found that the D-to-V mutations in hnRNPA1 and hnRNPA2 alter the dynamics of stress granule assembly in cells58,59,68. The McKnight Laboratory discovered a possible reason for this: the D-to-V mutations in all three proteins, hnRNPA1, hnRNPA2, and hnRNPDL, cause the respective LC domains to phase-separate into labile polymers with enhanced stability, as measured using SDD-AGE13. Other studies of TDP-4362, TIA163, and FUS60,61 have consistently shown that disease mutations result in a propensity for more stable polymers, affecting phase separation-based interactions and slowing the assembly and disassembly of the membraneless organelles where they reside. These observations may indicate that phase separation represents the underlying biological utility of LC domains, allowing proteins to dynamically move into and out of subcellular compartments that are not membrane bound.
Phase separation as a regulatory mechanism of gene expression
Phase separation also contributes to gene regulation, possibly by promoting the dynamic assembly of transcription factors and RNA-binding proteins. Studies of the RNA-binding proteins FUS, EWS and TAF15, offered an early example of the association between phase separation and transcription initiation machinery. These three RNA-binding proteins are referred to as FET (FUS/EWS/TAF15) proteins69. The translocation of the N-terminal LC domains of FET proteins to any of a number of different DNA-binding domains (DBDs) represents an oncogenic event leading to many forms of cancer70,71,72,73,74. DBDs direct the cancer-causing fusion proteins to the appropriate genes to facilitate cell growth or survival, while the N-terminal LC domains of FET proteins function as transcriptional activation domains. Despite a concrete understanding of how DBDs function, however, the mechanisms behind the function of the activation domains remain unknown. McKnight and colleagues have provided compelling evidence that the LC domains of FET proteins directly recruit the C-terminal domain (CTD) of RNA polymerase II and that the molecular determinant for this interaction is a phase separation of the N-terminal LC domains of FET proteins21 (Fig. 3). The CTD of mammalian RNA polymerase II contains 52 repeats of the heptad sequence YSPTSPS75,76. The CTD, which is 350 residues in length, is composed almost exclusively of just four amino acids—Y, S, P, and T—corresponding to the LC sequence. In vitro binding assays revealed that the CTD of RNA polymerase II is trapped by hydrogel droplets formed from the LC domains of FUS, EWS and TAF15. Y-to-S mutations in the G/S-Y-G/S triplet repeats in the N-terminal LC domain of TAF15, which abrogate phase separation capacity, correlatively reduce both CTD interaction and the transcriptional activation capacity in cells. Furthermore, CTD binding to FET protein hydrogels is reversed upon phosphorylation of the CTD by cyclin-dependent kinases (CDK7 or CDK9), which are known to phosphorylate the CTD in living cells21 (Fig. 3). These observations not only suggest how FET fusion drives oncogenic gene expression but also answer the question of how RNA polymerase II is recruited to the transcription initiation complex of gene promoters. However, the role of intact FUS or TAF15 proteins under normal conditions has yet to be established.
Upper panel, schematic of RNA polymerase II recruitment by LC domain polymers. During transcription initiation, the DNA-binding domain (DBD) of FET fusion proteins binds to their cognate genes, and their LC domains form reversible polymers that can recruit RNA polymerase II (Pol II) via direct interaction with the C-terminal domain (CTD). Phosphorylation of the CTD heptad repeats by CDK7 or CDK9 facilitates release of the LC domain polymer-bound Pol II so that it can escape the preinitiation complex and enter the process of transcriptional initiation and elongation. Lower panel, hydrogel binding assay showing phosphorylation-regulated binding of GFP-linked heptad repeats 27–52 of Pol II CTD (GFP-CTDC26) to mCherry-TAF15 hydrogel droplets. Bound protein is exposed to both CDK7 protein kinase and ATP (top), CDK7 alone (middle), or ATP alone (bottom). Bound GFP-CTDC26 is released from mCherry-TAF15 hydrogel droplets in a time-dependent manner only in the presence of both CDK7 and ATP. Figure adapted from Kwon et al.21.
In addition to transcription initiation, a phase separation model can explain the mechanism underlying the transition of RNA polymerase II from an initiation complex to an elongation complex (Fig. 3)21,77,78. During the transcription cycle (initiation, elongation, and termination), RNA polymerase II is recruited to active genes in its unphosphorylated state and released for elongation following the phosphorylation of the CTD; this model assumes that both the transcription-initiation machinery and the splicing machinery can form phase-separated condensates. The transcription-initiation machinery consists of mediators, transcription factors, coactivators, and nonphosphorylated RNA polymerase II; the splicing machinery is a transient elongation condensate 50–100 bp downstream of the initiation site and consists of phosphorylated RNA polymerase II, nascent RNA, elongation factors, RNA processing factors, and specific elongation coactivators. The hypophosphorylated CTD of RNA polymerase II is preferentially incorporated into phase-separated initiation condensate, while the hyperphosphorylated CTD is preferentially incorporated into condensates that are formed by splicing factors. When RNA polymerase II reaches the end of the gene, hypophosphorylated CTD is released and transferred back to the initiation condensate.
The role of phase separation in RNA splicing is further supported by studies of hnRNPs. hnRNPs are a large family of RNA-binding proteins that control key events in RNA biogenesis under both normal and diseased cellular conditions. Blencowe and colleagues showed that phase separation-dependent interactions control the assembly of hnRNPA/D family members, which, in turn, function to regulate splicing79. hnRNP complexes also contain different kinds of RNAs and RNA-binding proteins other than hnRNPs, and their assembly and disassembly occur rapidly80. hnRNPH1 is a prototypical hnRNP containing two distinctive LC domains on its C-terminus (LC1 and LC2). Using mutagenesis of these LC domains of hnRNPH1, we showed that a triple tyrosine substitution reduced the ability of the hnRNPH1 LC1 domain to phase separate and simultaneously reduced the capacity of hnRNPH1 to interact with RNA-binding proteins and regulate RNA splicing in living cells (Fig. 4)24. These results suggest that phase separation of the LC1 domain can promote the higher-order assembly of hnRNPH1 and other RNA-binding proteins that are required for the splicing activity of hnRNPH1. Thus, we speculate that phase separation-mediated assemblies relay the passage of genetic information from one site to another within a cell, ensuring that the process is of extreme fidelity.
The domain architecture of hnRNPH1 shows that the N-terminal half consists of two RNA-recognition motifs (RRM1 and RRM2) and the C-terminal half consists of two LC domains (LC1 and LC2) separated by RRM3. RRMs are in yellow, and LC domains are in blue. The oncogenic MEF2D-hnRNPH1 fusion protein is shown on the top, in which hNRNPH1 maintained two LC domains and one RRM at the C-terminus and MEF2D maintained the critical MADS-box domain at the N-terminus. Out of the two LC domains of hnRNPH1, LC1 can undergo phase separation into LLDs or hydrogel droplets composed of reversible polymers. Y-to-S mutations impairing the LLD formation of hnRNPH1 LC1 domains significantly reduced interaction with different kinds of RNA-binding proteins and the alternative-splicing activity of hnRNPH1, providing insight into the role of LC1 phase separation in the control of hnRNPH1 splicing activity (bottom left). While the LC2 domain does not undergo phase separation by itself, it is required for transcriptional activation in the context of a fusion gene with the GAL4 DNA binding domain. A single tyrosine-to-serine substitution (Y408S) in the LC2 domain is sufficient to interrupt transcriptional activity. Unlike the mCherry hydrogel droplets composed of the WT C-terminal half of hnRNPH1, which were melted substantially by 1,6-HD, hydrogel droplets composed of the C-terminus containing the Y408S mutation were resistant to 1,6-HD even after overnight exposure at 37 °C, implying the importance of phase separation dynamics in hnRNPH1 functions (bottom right). Images were reproduced from Kim et al.24.
Phase separation in disease
Evidence accumulated over many years in studies of genetics, cell biology, and pathology has revealed that phase separation is relevant to numerous human pathological conditions, including cancer and neurodegenerative diseases. Therapeutic strategies to regulate phase separation dynamics in cells could help treat diseases related to aberrant phase separation.
Role of phase separation in cancer
Cancer is a disease where cells reproduce uncontrollably. It is governed by biochemical pathways that have escaped the regulatory bounds of normal homeostatic balance. This balance is maintained through precise spatiotemporal regulation of these pathways. Phase separation is increasingly implicated as a previously hidden driver of aberrant spatiotemporal organization and protein dynamics involved in oncogenic activity. As discussed above, FET fusion oncoproteins provide a striking example of how the phase separation of oncoproteins acts to assemble transcription machinery and causes aberrant transcription21 (Fig. 3). When appended to a DBD, as is the case in oncogenic FET fusion proteins, the LC domains of FET proteins act to directly recruit the CTD of RNA polymerase II. To achieve this task, the LC domains of FET proteins must be capable of phase separation. Together with the fact that FET proteins in their intact form are endowed with an RNA binding domain and are not associated with cancer development, McKnight and colleagues proposed that the binding of multiple copies of the DBDs from fusion oncoproteins to their cognate genes may concentrate the FET LC domain to a level sufficient for phase separation, driving aberrant transcription6. This is in accordance with previous studies showing that portions of EWS, FUS, and TAF15 are functionally interchangeable in FET fusion proteins, while the transcription factor DNA-binding moiety determines the tumor phenotype81.
Consistent with the notion that the dynamic aspect of phase separation is the key to biological validity and the functional utility of phase separation, changes in phase separation dynamics functionally impact gene transcription and biochemical outcomes in cancer. One prominent example of this is the correlative effects of Y-to-S mutation of the LC domain in hnRNPH1 on phase separation dynamics and transcriptional activity in the context of fusion oncoproteins (Fig. 4). The hnRNPH1-myocyte-specific enhancer factor 2D (hnRNPH1-MEF2D) fusion gene has been identified in acute lymphoblastic leukemia. The resulting gene product is a fusion protein in which a DBD of MEF2D is connected to the C-terminal region of hnRNPH1, retaining the LC1 and LC2 domains. We have demonstrated that the LC2 domain in the C-terminal region functions as a transcriptional activation domain25. Remarkably, the Y408S mutation in the LC2 domain abolished its transcriptional activity. The same LC2 mutation induced the phase separation of the hnRNPH1 C-terminus into irreversible hydrogel droplets, suggesting that the Y408S mutation may decrease the transcriptional activity of hnRNPH1 by restricting the movement and interactions of other macromolecules, such as transcription factors and coactivators. This hypothesis was supported by the observation that the Y408S mutation of the LC2 domain enhanced homotypic and heterotypic binding to the LC domains of various RNA-binding proteins, including FUS, TAF15, DHX9, hnRNPA1, hnRNPA2, and hnRNPF25. Similar observations have been made regarding AKAP95, which is frequently overexpressed in breast and ovarian cancers82,83. Mutation of all six tyrosine residues to phenylalanine (Y-to-F) in the LC domain of AKAP95 enhances phase separation propensity, renders condensates into a more solid-like state, and impairs the ability of AKAP95 to regulate RNA splicing of cancer-related targets and tumorigenesis83. Given that AKAP95 is dispensable in normal cell growth, its overexpression might increase the concentration of the AKAP95 LC domain to a level that is sufficient for phase separation, which in turn might mediate the oncogenic interactions of AKAP95 with other macromolecules, such as splicing modulators and RNA substrates. Collectively, it is plausible that overexpression or the translocation product causative of cancer might recruit the macromolecules necessary for the initiation of oncogenic gene expression in a phase separation-dependent manner. Accordingly, therapies that target the phase separation dynamics of oncoproteins are currently in development for cancer treatment20.
Pathogenic LC domain proteins in neurodegenerative diseases
Protein aggregation is a pathological hallmark of neurodegenerative diseases, including the β-amyloid (Aβ) and tau proteins of Alzheimer’s disease, the huntingtin of Huntington’s disease, and the α-synuclein of Parkinson’s diseases84. Many proteins found in pathological aggregates contain intrinsic disorder/LC domains85,86. Clear examples of a link between neurodegenerative diseases and LC proteins have been found in studies of several ALS-associated proteins. ALS is a progressive adult-onset neurodegenerative disease characterized by the selective death of motor neurons in the brain and spinal cord87. Different RNA-binding proteins harboring LC domains (TDP-4388,89, FUS90,91, hnRNPA168,92, hnRNPA268, matrin 393, TIA163, hnRNPDL67, and annexin A1194,95) are associated with ALS. ALS-causing mutations in the LC domains of these RNA-binding proteins accelerate phase separation into less dynamic or irreversible fibrils that can produce the fibrillar pathology observed in patient cells13,59,60,61,62,63,68. As mentioned above, these mutations can cause disturbances in the dynamics of membraneless organelle assembly, which can impair functions with adverse consequences for multiple aspects of RNA metabolism96.
By far, the most common genetic cause of ALS and FTD is the expansion of the GGGGCC hexanucleotide repeat (HRE) in the first intron of the chromosome 9 open reading frame 72 (C9orf72) gene97,98,99. From the HRE, five different poly-dipeptides are produced, depending on the reading frames, namely, glycine-alanine, glycine-proline, glycine-arginine (GR), proline-alanine, and proline-arginine (PR) poly-dipeptides (Fig. 5a)100,101,102,103,104,105,106. These poly-dipeptides are obviously LC domain proteins composed of only two amino acids. Of the five poly-dipeptides derived from C9orf72 HRE, McKnight and colleagues recognized that PR and GR poly-dipeptides are reminiscent of the serine-arginine (SR) domains commonly found in SR splicing factors (SRSFs) that bind to hydrogels made up of the LC domain of the RNA-binding protein hnRNPA2 in vitro23,107,108. This SR binding to hydrogels is reversed upon the phosphorylation of serine residues by cyclin-like kinases (CLKs) known to regulate SRSF function in living cells (Fig. 5b)109. They thus proposed the intriguing hypothesis that soluble PR and/or GR poly-dipeptides might bind LC-domain hydrogel droplets but are impervious to CLK-mediated phosphorylation and thus irreversibly bind cellular targets such as the RNP complex in membraneless organelles in vivo. Indeed, they found that, unlike native SRSF2 protein, which is distributed across nuclear speckles, the two poly-dipeptides are irreversibly trapped in nucleoli, where they impair ribosomal RNA processing (Fig. 5a). McKnight and colleagues confirmed that the application of one of the dipeptides (PR) to cultured human astrocytes resulted in the mis-splicing of a number of gene transcripts, including the same mis-splicing of the glutamate transporter Eaat2 transcript found in C9orf72 ALS patients23,110. After this work, a number of studies produced confirmatory findings. PR and/or GR poly-dipeptides disrupt the phase separation of the nucleolar protein nucleophosmin65, impede nucleocytoplasmic transport111,112 and are themselves capable of phase separation113,114. Unbiased studies of the direct intracellular targets of PR poly-dipeptides showed that they bind to the polymeric forms of LC domains in a wide variety of cellular proteins, including RNA-binding proteins, nucleoporins, and intermediate filaments13,115. From these results, we speculate that the soluble forms of PR and GR poly-dipeptides, via irreversible binding to LC proteins, cause widespread disturbances to the integrity and dynamics of cellular structures13,23,65,114,115,116,117,118,119, resulting in broad impediments to the information flow from gene to mRNA to protein that eventually lead to cell death (Fig. 5a).
a Cellular processes impaired by PR and GR poly-dipeptides. The C9orf72 gene consists of 11 exons. Coding exons are indicated in green, and noncoding exons are indicated in blue (not to scale). Unaffected people contain 10-20 GGGGCC hexanucleotide repeats, but the repeat number expands to hundreds or even thousands in C9orf72 ALS cases. A hexanucleotide repeat expansion (HRE), (GGGGCC)n, can produce five different poly-dipeptides, including GR and PR poly-dipeptides. Four pathogenicities enacted by GR and PR poly-dipeptides are shown: (1) PR poly-dipeptide bound to nucleoli results in nucleolar dysfunction. Nucleolar localization of synthetic peptide containing 20 repeats of the PR sequence (PR20) (green signal) is deduced by colocalization with the nucleolar marker fibrillin (red signal). (2) Impediments in splicing of the transcript encoding the glutamate transporter EAAT2. Human astrocytes exposed to PR20 show aberrantly spliced EAAT2 transcripts (black and gray arrowheads). The arrow indicates a normal EAAT2 transcript. (3) Alterations in cell morphology and death. Upon exposure to 30 µM of the PR20 peptide for 24 h, almost all cells were detached from the culture substrate and dead. (4) Motor neuron hyperexcitability. PR20 peptide application (100 nM, 20 min) causes hyperexcitability in layer 5 (L5) pyramidal cells (PCs) of the primary motor cortex (cortical motor neurons). Reproduced from Kwon et al.23 and Jo et al.123. b Cartoon depicting the irreversible binding of PR and GR poly-dipeptides to intracellular targets. The serine-arginine (SR) domain of SR splicing factors binds LC proteins, which is dynamically reversible by cyclin-like kinases (CLKs). PR and GR poly-dipeptides bind to LC proteins but cannot be liberated by CLK enzymes, thereby impeding the flow of information from gene to mRNA to protein.
Our more recent study showed that PR and GR poly-dipeptides are drivers of cortical hyperexcitability, an early and key feature of ALS patients (Fig. 5a)120,121,122,123. Despite much research on the subject, its underlying mechanisms remain elusive. We demonstrated that PR and GR poly-dipeptides selectively induce hyperexcitability in cortical motor neurons, possibly by binding to and hyperactivating a voltage-gated sodium channel Nav1.2-β1-β4 complex. As discussed above, PR poly-dipeptides can impede splicing of the Eaat2 transcript in astrocytes23. We thus speculate that this impediment might decrease the reuptake of excess glutamate that is released by hyperactivated motor neurons, favoring the accumulation of glutamate in the synaptic cleft, which, in turn, leads to glutamate excitotoxicity and eventually to the selective neurodegeneration of lower motor neurons in C9orf72 ALS. Knowing that Nav1.2 and auxiliary β4 subunits have intrinsically disordered domains, we suggest that phase separation of these LC domains might be important for the assembly of the macromolecular sodium channel complex and interaction with other LC domain proteins, such as neuronal intermediate filaments124,125,126,127. If this is the case, we further hypothesize that PR and GR poly-dipeptides might bind to phase-separated LC domains otherwise adhering Nav channels to multiprotein complexes. We anticipate that the combination of both molecular and electrophysiological studies may help identify the role of intrinsically disordered domains associated with ion channel proteins in neurological and neurodegenerative disease pathophysiology, as well as in normal physiological conditions.
Conclusions and comments
The examples of phase-separating LC domains discussed within this review illustrate how phase separation is beneficial for cells to form dynamic, multivalent protein–protein interactions. These examples also highlight the importance of regulatory mechanisms to maintain the reversibility of phase separation. The functional relevance of reversibility and the dynamics of phase separation are particularly evident when considering that these properties are directly impacted by many disease-causing mutations. Together, phase separation dynamics serve as a biological framework to explore new therapeutic approaches to devastating human diseases, including cancer and neurodegeneration.
References
van der Lee, R. et al. Classification of intrinsically disordered regions and proteins. Chem. Rev. 114, 6589–6631 (2014).
Marcotte, E. M., Pellegrini, M., Yeates, T. O. & Eisenberg, D. A census of protein repeats. J. Mol. Biol. 293, 151–160 (1999).
Golding, G. B. Simple sequence is abundant in eukaryotic proteins. Protein Sci. 8, 1358–1361 (1999).
Alba, M. M. & Guigo, R. Comparative analysis of amino acid repeats in rodents and humans. Genome Res. 14, 549–554 (2004).
Karlin, S., Brocchieri, L., Bergman, A., Mrazek, J. & Gentles, A. J. Amino acid runs in eukaryotic proteomes and disease associations. Proc. Natl Acad. Sci. USA 99, 333–338 (2002).
Kato, M. & McKnight, S. L. A solid-state conceptualization of information transfer from gene to message to protein. Annu. Rev. Biochem. 87, 351–390 (2018).
King, O. D., Gitler, A. D. & Shorter, J. The tip of the iceberg: RNA-binding proteins with prion-like domains in neurodegenerative disease. Brain Res. 1462, 61–80 (2012).
Courchaine, E. M., Lu, A. & Neugebauer, K. M. Droplet organelles? EMBO J. 35, 1603–1612 (2016).
Boeynaems, S. et al. Protein phase separation: a new phase in cell biology. Trends Cell Biol. 28, 420–435 (2018).
Kato, M., Zhou, X. & McKnight, S. L. How do protein domains of low sequence complexity work? RNA 28, 3–15 (2022).
Frey, S. & Gorlich, D. A saturated FG-repeat hydrogel can reproduce the permeability properties of nuclear pore complexes. Cell 130, 512–523 (2007).
Frey, S., Richter, R. P. & Gorlich, D. FG-rich repeats of nuclear pore proteins form a three-dimensional meshwork with hydrogel-like properties. Science 314, 815–817 (2006).
Lin, Y. et al. Toxic PR poly-dipeptides encoded by the C9orf72 repeat expansion target LC domain polymers. Cell 167, 789–802.e712 (2016).
Zhou, X. et al. Transiently structured head domains control intermediate filament assembly. Proc. Natl Acad. Sci. USA 118, e2022121118 (2021).
Brangwynne, C. P. et al. Germline P granules are liquid droplets that localize by controlled dissolution/condensation. Science 324, 1729–1732 (2009).
Kato, M. et al. Cell-free formation of RNA granules: low complexity sequence domains form dynamic fibers within hydrogels. Cell 149, 753–767 (2012).
Liu, X. et al. Phase separation drives decision making in cell division. J. Biol. Chem. 295, 13419–13431 (2020).
Harami, G. M. et al. Phase separation by ssDNA binding protein controlled via protein-protein and protein-DNA interactions. Proc. Natl Acad. Sci. USA 117, 26206–26217 (2020).
Su, Q., Mehta, S. & Zhang, J. Liquid-liquid phase separation: orchestrating cell signaling through time and space. Mol. Cell 81, 4137–4146 (2021).
Taniue, K. & Akimitsu, N. Aberrant phase separation and cancer. FEBS J. 289, 17–39 (2022).
Kwon, I. et al. Phosphorylation-regulated binding of RNA polymerase II to fibrous polymers of low-complexity domains. Cell 155, 1049–1060 (2013).
Nedelsky, N. B. & Taylor, J. P. Pathological phase transitions in ALS-FTD impair dynamic RNA-protein granules. RNA 28, 97–113 (2022).
Kwon, I. et al. Poly-dipeptides encoded by the C9orf72 repeats bind nucleoli, impede RNA biogenesis, and kill cells. Science 345, 1139–1145 (2014).
Kim, G. H. & Kwon, I. Distinct roles of hnRNPH1 low-complexity domains in splicing and transcription. Proc. Natl Acad. Sci. USA 118, e2109668118 (2021).
Kim, E. & Kwon, I. Phase transition of fibrillarin LC domain regulates localization and protein interaction of fibrillarin. Biochem. J. 478, 799–810 (2021).
Kato, M., Lin, Y. & McKnight, S. L. Cross-beta polymerization and hydrogel formation by low-complexity sequence proteins. Methods 126, 3–11 (2017).
Kato, M. et al. Redox state controls phase separation of the yeast Ataxin-2 protein via reversible oxidation of its methionine-rich low-complexity domain. Cell 177, 711–721.e718 (2019).
Xiang, S. et al. The LC domain of hnRNPA2 adopts similar conformations in hydrogel polymers, liquid-like droplets, and nuclei. Cell 163, 829–839 (2015).
Elbaum-Garfinkle, S. et al. The disordered P granule protein LAF-1 drives phase separation into droplets with tunable viscosity and dynamics. Proc. Natl Acad. Sci. USA 112, 7189–7194 (2015).
Harrison, A. F. & Shorter, J. RNA-binding proteins with prion-like domains in health and disease. Biochem. J. 474, 1417–1438 (2017).
Dignon, G. L., Best, R. B. & Mittal, J. Biomolecular phase separation: from molecular driving forces to macroscopic properties. Annu. Rev. Phys. Chem. 71, 53–75 (2020).
Khoury, G. A., Baliban, R. C. & Floudas, C. A. Proteome-wide post-translational modification statistics: frequency analysis and curation of the swiss-prot database. Sci. Rep. 1, 90 (2011).
Murray, D. T. et al. Structure of FUS protein fibrils and its relevance to self-assembly and phase separation of low-complexity domains. Cell 171, 615–627.e616 (2017).
Snider, N. T. & Omary, M. B. Post-translational modifications of intermediate filament proteins: mechanisms and functions. Nat. Rev. Mol. Cell Biol. 15, 163–177 (2014).
Geisler, N. & Weber, K. Phosphorylation of desmin in vitro inhibits formation of intermediate filaments; identification of three kinase A sites in the aminoterminal head domain. EMBO J. 7, 15–20 (1988).
Zhou, L. T. et al. A novel role of fragile X mental retardation protein in pre-mRNA alternative splicing through RNA-binding protein 14. Neuroscience 349, 64–75 (2017).
Darnell, J. C. & Klann, E. The translation of translational control by FMRP: therapeutic targets for FXS. Nat. Neurosci. 16, 1530–1536 (2013).
Kao, D. I., Aldridge, G. M., Weiler, I. J. & Greenough, W. T. Altered mRNA transport, docking, and protein translation in neurons lacking fragile X mental retardation protein. Proc. Natl Acad. Sci. USA 107, 15601–15606 (2010).
Ferron, L. Fragile X mental retardation protein controls ion channel expression and activity. J. Physiol. 594, 5861–5867 (2016).
Tsang, B. et al. Phosphoregulated FMRP phase separation models activity-dependent translation through bidirectional control of mRNA granule formation. Proc. Natl Acad. Sci. USA 116, 4218–4227 (2019).
Ambadipudi, S., Biernat, J., Riedel, D., Mandelkow, E. & Zweckstetter, M. Liquid-liquid phase separation of the microtubule-binding repeats of the Alzheimer-related protein Tau. Nat. Commun. 8, 275 (2017).
Wegmann, S. et al. Tau protein liquid-liquid phase separation can initiate tau aggregation. EMBO J. 37, e98049 (2018).
Kato, M., Tu, B. P. & McKnight, S. L. Redox-mediated regulation of low complexity domain self-association. Curr. Opin. Genet. Dev. 67, 111–118 (2021).
Lin, Y. et al. Redox-mediated regulation of an evolutionarily conserved cross-beta structure formed by the TDP43 low complexity domain. Proc. Natl Acad. Sci. USA 117, 28727–28734 (2020).
Hofweber, M. et al. Phase separation of FUS is suppressed by its nuclear import receptor and arginine methylation. Cell 173, 706–719 e713 (2018).
Qamar, S. et al. FUS phase separation is modulated by a molecular chaperone and methylation of arginine cation-pi interactions. Cell 173, 720–734.e715 (2018).
Yasuda, S. et al. Stress- and ubiquitylation-dependent phase separation of the proteasome. Nature 578, 296–300 (2020).
Bock, A. S. et al. N-terminal acetylation modestly enhances phase separation and reduces aggregation of the low-complexity domain of RNA-binding protein fused in sarcoma. Protein Sci. 30, 1337–1349 (2021).
Ukmar-Godec, T. et al. Lysine/RNA-interactions drive and regulate biomolecular condensation. Nat. Commun. 10, 2909 (2019).
Saito, M. et al. Acetylation of intrinsically disordered regions regulates phase separation. Nat. Chem. Biol. 15, 51–61 (2019).
McGurk, L. et al. Poly(ADP-Ribose) prevents pathological phase separation of TDP-43 by promoting liquid demixing and stress granule localization. Mol. Cell 71, 703–717.e709 (2018).
Dao, T. P. et al. Ubiquitin modulates liquid–liquid phase separation of UBQLN2 via disruption of multivalent interactions. Mol. Cell 69, 965–978.e966 (2018).
Neupert, W. A perspective on transport of proteins into mitochondria: a myriad of open questions. J. Mol. Biol. 427, 1135–1158 (2015).
Grossman, E., Medalia, O. & Zwerger, M. Functional architecture of the nuclear pore complex. Annu. Rev. Biophys. 41, 557–584 (2012).
Mao, Y. S., Zhang, B. & Spector, D. L. Biogenesis and function of nuclear bodies. Trends Genet 27, 295–306 (2011).
Bond, C. S. & Fox, A. H. Paraspeckles: nuclear bodies built on long noncoding RNA. J. Cell Biol. 186, 637–644 (2009).
Anderson, P. & Kedersha, N. Stress granules: the Tao of RNA triage. Trends Biochem. Sci. 33, 141–150 (2008).
Lin, Y., Protter, D. S., Rosen, M. K. & Parker, R. Formation and maturation of phase-separated liquid droplets by RNA-binding proteins. Mol. Cell 60, 208–219 (2015).
Molliex, A. et al. Phase separation by low complexity domains promotes stress granule assembly and drives pathological fibrillization. Cell 163, 123–133 (2015).
Patel, A. et al. A liquid-to-solid phase transition of the ALS protein FUS accelerated by disease mutation. Cell 162, 1066–1077 (2015).
Murakami, T. et al. ALS/FTD mutation-induced phase transition of FUS liquid droplets and reversible hydrogels into irreversible hydrogels impairs RNP granule Function. Neuron 88, 678–690 (2015).
Conicella, A. E., Zerze, G. H., Mittal, J. & Fawzi, N. L. ALS mutations disrupt phase separation mediated by alpha-helical structure in the TDP-43 low-complexity C-terminal domain. Structure 24, 1537–1549 (2016).
Mackenzie, I. R. et al. TIA1 mutations in amyotrophic lateral sclerosis and frontotemporal dementia promote phase separation and alter stress granule dynamics. Neuron 95, 808–816.e809 (2017).
Feric, M. et al. Coexisting liquid phases underlie nucleolar subcompartments. Cell 165, 1686–1697 (2016).
White, M. R. et al. C9orf72 Poly(PR) dipeptide repeats disturb biomolecular phase separation and disrupt nucleolar function. Mol. Cell 74, 713–728.e716 (2019).
Fatica, A. & Tollervey, D. Making ribosomes. Curr. Opin. Cell Biol. 14, 313–318 (2002).
Vieira, N. M. et al. A defect in the RNA-processing protein HNRPDL causes limb-girdle muscular dystrophy 1G (LGMD1G). Hum. Mol. Genet 23, 4103–4110 (2014).
Kim, H. J. et al. Mutations in prion-like domains in hnRNPA2B1 and hnRNPA1 cause multisystem proteinopathy and ALS. Nature 495, 467–473 (2013).
Andersson, M. K. et al. The multifunctional FUS, EWS and TAF15 proto-oncoproteins show cell type-specific expression patterns and involvement in cell spreading and stress response. BMC Cell Biol. 9, 37 (2008).
Zinszner, H., Albalat, R. & Ron, D. A novel effector domain from the RNA-binding protein TLS or EWS is required for oncogenic transformation by CHOP. Genes Dev. 8, 2513–2526 (1994).
Rabbitts, T. H., Forster, A., Larson, R. & Nathan, P. Fusion of the dominant negative transcription regulator CHOP with a novel gene FUS by translocation t(12;16) in malignant liposarcoma. Nat. Genet. 4, 175–180 (1993).
Ichikawa, H., Shimizu, K., Katsu, R. & Ohki, M. Dual transforming activities of the FUS (TLS)-ERG leukemia fusion protein conferred by two N-terminal domains of FUS (TLS). Mol. Cell. Biol. 19, 7639–7650 (1999).
Crozat, A., Aman, P., Mandahl, N. & Ron, D. Fusion of CHOP to a novel RNA-binding protein in human myxoid liposarcoma. Nature 363, 640–644 (1993).
Bertolotti, A., Bell, B. & Tora, L. The N-terminal domain of human TAFII68 displays transactivation and oncogenic properties. Oncogene 18, 8000–8010 (1999).
Corden, J. L., Cadena, D. L., Ahearn, J. M. Jr & Dahmus, M. E. A unique structure at the carboxyl terminus of the largest subunit of eukaryotic RNA polymerase II. Proc. Natl Acad. Sci. USA 82, 7934–7938 (1985).
Lee, J. M. & Greenleaf, A. L. A protein kinase that phosphorylates the C-terminal repeat domain of the largest subunit of RNA polymerase II. Proc. Natl Acad. Sci. USA 86, 3624–3628 (1989).
Guo, Y. E. et al. Pol II phosphorylation regulates a switch between transcriptional and splicing condensates. Nature 572, 543–548 (2019).
Peng, L., Li, E. M. & Xu, L. Y. From start to end: phase separation and transcriptional regulation. Biochim. Biophys. Acta Gene Regul. Mech. 1863, 194641 (2020).
Gueroussov, S. et al. Regulatory expansion in mammals of multivalent hnRNP assemblies that globally control alternative splicing. Cell 170, 324–339 e323 (2017).
Chaudhury, A., Chander, P. & Howe, P. H. Heterogeneous nuclear ribonucleoproteins (hnRNPs) in cellular processes: focus on hnRNP E1’s multifunctional regulatory roles. RNA 16, 1449–1462 (2010).
Kovar, H. Dr. Jekyll and Mr. Hyde: the two faces of the FUS/EWS/TAF15 protein family. Sarcoma 2011, 837474 (2011).
Liu, W. et al. Roles of Cx43 and AKAP95 in ovarian cancer tissues in G1/S phase. Int. J. Clin. Exp. Pathol. 8, 14315–14324 (2015).
Li, W. et al. Biophysical properties of AKAP95 protein condensates regulate splicing and tumorigenesis. Nat. Cell Biol. 22, 960–972 (2020).
Mattson, M. P. & Magnus, T. Ageing and neuronal vulnerability. Nat. Rev. Neurosci. 7, 278–294 (2006).
Uversky, V. N. Intrinsic disorder in proteins associated with neurodegenerative diseases. Front. Biosci. 14, 5188–5238 (2009).
Zbinden, A., Perez-Berlanga, M., De Rossi, P. & Polymenidou, M. Phase separation and neurodegenerative diseases: a disturbance in the force. Dev. Cell 55, 45–68 (2020).
Brown, R. H. Jr & Al-Chalabi, A. Amyotrophic lateral sclerosis. N. Engl. J. Med. 377, 1602 (2017).
Sreedharan, J. et al. TDP-43 mutations in familial and sporadic amyotrophic lateral sclerosis. Science 319, 1668–1672 (2008).
Kabashi, E. et al. TARDBP mutations in individuals with sporadic and familial amyotrophic lateral sclerosis. Nat. Genet. 40, 572–574 (2008).
Kwiatkowski, T. J. Jr et al. Mutations in the FUS/TLS gene on chromosome 16 cause familial amyotrophic lateral sclerosis. Science 323, 1205–1208 (2009).
Vance, C. et al. Mutations in FUS, an RNA processing protein, cause familial amyotrophic lateral sclerosis type 6. Science 323, 1208–1211 (2009).
Liu, Q. et al. Whole-exome sequencing identifies a missense mutation in hnRNPA1 in a family with flail arm ALS. Neurology 87, 1763–1769 (2016).
Johnson, J. O. et al. Mutations in the Matrin 3 gene cause familial amyotrophic lateral sclerosis. Nat. Neurosci. 17, 664–666 (2014).
Leoni, T. B. et al. A novel multisystem proteinopathy caused by a missense ANXA11 variant. Ann. Neurol. 90, 239–252 (2021).
Smith, B. N. et al. Mutations in the vesicular trafficking protein annexin A11 are associated with amyotrophic lateral sclerosis. Sci. Transl. Med. 9, eaad9157 (2017).
Mathieu, C., Pappu, R. V. & Taylor, J. P. Beyond aggregation: Pathological phase transitions in neurodegenerative disease. Science 370, 56–60 (2020).
DeJesus-Hernandez, M. et al. Expanded GGGGCC hexanucleotide repeat in noncoding region of C9ORF72 causes chromosome 9p-linked FTD and ALS. Neuron 72, 245–256 (2011).
Renton, A. E. et al. A hexanucleotide repeat expansion in C9ORF72 is the cause of chromosome 9p21-linked ALS-FTD. Neuron 72, 257–268 (2011).
Taylor, J. P., Brown, R. H. Jr. & Cleveland, D. W. Decoding ALS: from genes to mechanism. Nature 539, 197–206 (2016).
Ash, P. E. et al. Unconventional translation of C9ORF72 GGGGCC expansion generates insoluble polypeptides specific to c9FTD/ALS. Neuron 77, 639–646 (2013).
Balendra, R. & Isaacs, A. M. C9orf72-mediated ALS and FTD: multiple pathways to disease. Nat. Rev. Neurol. 14, 544–558 (2018).
Donnelly, C. J. et al. RNA toxicity from the ALS/FTD C9ORF72 expansion is mitigated by antisense intervention. Neuron 80, 415–428 (2013).
Lagier-Tourenne, C. et al. Targeted degradation of sense and antisense C9orf72 RNA foci as therapy for ALS and frontotemporal degeneration. Proc. Natl Acad. Sci. USA 110, E4530–E4539 (2013).
Mizielinska, S. et al. C9orf72 frontotemporal lobar degeneration is characterised by frequent neuronal sense and antisense RNA foci. Acta Neuropathol. 126, 845–857 (2013).
Mori, K. et al. The C9orf72 GGGGCC repeat is translated into aggregating dipeptide-repeat proteins in FTLD/ALS. Science 339, 1335–1338 (2013).
Zu, T. et al. RAN proteins and RNA foci from antisense transcripts in C9ORF72 ALS and frontotemporal dementia. Proc. Natl Acad. Sci. USA 110, E4968–E4977 (2013).
Manley, J. L. & Tacke, R. SR proteins and splicing control. Genes Dev. 10, 1569–1579 (1996).
Roth, M. B., Murphy, C. & Gall, J. G. A monoclonal antibody that recognizes a phosphorylated epitope stains lampbrush chromosome loops and small granules in the amphibian germinal vesicle. J. Cell Biol. 111, 2217–2223 (1990).
Colwill, K. et al. The Clk/Sty protein kinase phosphorylates SR splicing factors and regulates their intranuclear distribution. EMBO J. 15, 265–275 (1996).
Lin, C. L. et al. Aberrant RNA processing in a neurodegenerative disease: the cause for absent EAAT2, a glutamate transporter, in amyotrophic lateral sclerosis. Neuron 20, 589–602 (1998).
Jovicic, A. et al. Modifiers of C9orf72 dipeptide repeat toxicity connect nucleocytoplasmic transport defects to FTD/ALS. Nat. Neurosci. 18, 1226–1229 (2015).
Freibaum, B. D. et al. GGGGCC repeat expansion in C9orf72 compromises nucleocytoplasmic transport. Nature 525, 129–133 (2015).
Chen, C. et al. Phase separation and toxicity of C9orf72 poly(PR) depends on alternate distribution of arginine. J. Cell Biol. 220, e202103160 (2021).
Boeynaems, S. et al. Phase separation of C9orf72 dipeptide repeats perturbs stress granule dynamics. Mol. Cell 65, 1044–1055.e1045 (2017).
Lee, K. H. et al. C9orf72 dipeptide repeats impair the assembly, dynamics, and function of membrane-less organelles. Cell 167, 774–788.e717 (2016).
Zhang, Y. J. et al. Poly(GR) impairs protein translation and stress granule dynamics in C9orf72-associated frontotemporal dementia and amyotrophic lateral sclerosis. Nat. Med. 24, 1136–1142 (2018).
Shi, K. Y. et al. Toxic PRn poly-dipeptides encoded by the C9orf72 repeat expansion block nuclear import and export. Proc. Natl Acad. Sci. USA. 114, E1111–E1117 (2017).
Wen, X. et al. Antisense proline-arginine RAN dipeptides linked to C9ORF72-ALS/FTD form toxic nuclear aggregates that initiate in vitro and in vivo neuronal death. Neuron 84, 1213–1225 (2014).
Haeusler, A. R. et al. C9orf72 nucleotide repeat structures initiate molecular cascades of disease. Nature 507, 195–200 (2014).
Selvaraj, B. T. et al. C9ORF72 repeat expansion causes vulnerability of motor neurons to Ca(2+)-permeable AMPA receptor-mediated excitotoxicity. Nat. Commun. 9, 347 (2018).
Schanz, O. et al. Cortical hyperexcitability in patients with C9ORF72 mutations: relationship to phenotype. Muscle Nerve 54, 264–269 (2016).
Perkins, E. M. et al. Altered network properties in C9ORF72 repeat expansion cortical neurons are due to synaptic dysfunction. Mol. Neurodegener. 16, 13 (2021).
Jo, Y., Lee, J., Lee, S. Y., Kwon, I. & Cho, H. Poly-dipeptides produced from C9orf72 hexanucleotide repeats cause selective motor neuron hyperexcitability in ALS. Proc. Natl Acad. Sci. USA. 119, e2113813119 (2022).
Shaw, J. E. & Koleske, A. J. Functional interactions of ion channels with the actin cytoskeleton: does coupling to dynamic actin regulate NMDA receptors? J. Physiol. 599, 431–441 (2021).
Morachevskaya, E. A. & Sudarikova, A. V. Actin dynamics as critical ion channel regulator: ENaC and Piezo in focus. Am. J. Physiol. Cell Physiol. 320, C696–C702 (2021).
Curran, J. & Mohler, P. J. Alternative paradigms for ion channelopathies: disorders of ion channel membrane trafficking and posttranslational modification. Annu. Rev. Physiol. 77, 505–524 (2015).
Steele, D. F. & Fedida, D. Cytoskeletal roles in cardiac ion channel expression. Biochim. Biophys. Acta 1838, 665–673 (2014).
Acknowledgements
This work was supported by the National Research Foundation of Korea grants NRF-2020R1A2C2012846 awarded to H.C. and NRF-2019R1A2C2003767 awarded to I.K.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lee, J., Cho, H. & Kwon, I. Phase separation of low-complexity domains in cellular function and disease. Exp Mol Med 54, 1412–1422 (2022). https://doi.org/10.1038/s12276-022-00857-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s12276-022-00857-2
This article is cited by
-
Clinical and genetic characteristics of Chinese patients with congenital fibrosis of the extraocular muscles
Orphanet Journal of Rare Diseases (2024)
-
FUS maintains TAZ fluidity and function
Nature Cell Biology (2024)
-
Impact of distinct FG nucleoporin repeats on Nup98 self-association
Nature Communications (2024)