Abstract
Previous genome-wide association studies have discovered significant association at ITGAX-ITGAM on 16p11.2 for IgA nephropathy (IgAN). In this study, we performed a two-stage association study that enrolled 1700 IgAN cases and 2400 controls to further investigate the relationship of ITGAX and ITGAM gene polymorphisms with IgAN. Seven single-nucleotide polymorphisms (SNPs) were selected for genotyping in 1000 IgAN cases and 1000 healthy controls in the discovery stage, and the significant SNP was further validated in additional 700 IgAN cases and 1400 healthy controls. We found that four SNPs (rs11150619, rs11150614, rs7190997, and rs4597342) showed potential associations with IgAN susceptibility in the discovery stage, but only SNP rs11150619 was further genotyped in the validation stage after multiple testing. The results indicated that rs11150619 was significantly associated with IgAN in the combined samples (OR = 0.81, 95%CI = 0.71–0.91, and dominant P = 6.68 × 10−4). Moreover, patients with TT genotype of rs11150619 exhibited increased estimated glomerular filtration rate levels and a reduced proportion of global sclerosis compared with those with TC and CC genotypes. Our results suggested that ITGAX and ITGAM gene polymorphisms were associated with IgAN in a Chinese Han population, and the rs11150619-T allele showed a potential protective role for IgAN.
Similar content being viewed by others
Introduction
IgA nephropathy (IgAN) is one of the most type of primary glomerulonephritis in the Chinese population [1]. Currently, IgAN can only be diagnosed by renal biopsy, which is featured by dominant IgA-containing immune complexes deposition in the glomerular mesangial region [2]. The activation of mesangial cells is amplified by the pathogenetic complexes to activate the complement system, and complement component C3 is often seen in the renal biopsies of IgAN [3].
IgAN shows a slow but persistent clinical course and 30–40% of cases will progress to end-stage renal disease within 20 years [4]. Numerous lines of evidences indicated that IgAN is a complex polygenic disease, and multiple genetic susceptibility genes with minor to moderate effects likely contribute to the development and progression of the disease [5]. Since 2011, five genome-wide association studies (GWAS) have discovered a large number of susceptibility loci of IgAN [6,7,8,9,10], and the ITGAX-ITGAM locus on 16p11.2 is significantly associated with IgAN in European and Chinese populations [9, 10].
ITGAM (CD11b) and ITGAX (CD11c) encode integrins alphaM and alphaX, which integrate with the integrin β2 chain to form leukocyte-specific complement receptors 3 (CR3) and 4 (CR4), respectively [11]. The receptors (CR3 and CR4) are expressed on most white blood cells (WBCs) and play a role in the adherence of neutrophils and monocytes to stimulate endothelium cells and phagocytosis of complement-coated particles [12]. Both CR3 and CR4 have binding sites for inactive complement component 3b (iC3b) and fibrinogen [13], and are involved in the glomerular infiltration of immune cells in IgAN as complement receptors [14]. These observations indicate that CD11b and CD11c may act as an important player in complement activation and the etiology of IgAN.
ITGAM and ITGAX were also reported as candidate genes for several autoimmune diseases, including systemic lupus erythematosus (SLE) [15], systemic sclerosis [16], and Behcet’s disease [17]. Recently, an IgAN GWAS conducted in European populations has identified two significant signals at rs11574637 and rs11150612 within the ITGAX-ITGAM region [9]. Interestingly, the frequency of rs11574637-C allele in European population is 0.18, while absent in Asian populations, suggesting the contribution of genetic susceptibility to disease prevalence in different ethnic populations. However, the association analysis of ITGAX and ITGAM variants with IgAN susceptibility and clinical significance were not performed till now. Therefore, we conducted this two-stage association study to further explore the potential role of ITGAX and ITGAM gene polymorphisms with IgAN in a Chinese Han population.
Materials and methods
Study subjects
A total of 1700 sporadic IgAN patients (1000 cases for the discovery stage and 700 cases for the validation stage) were recruited from the Department of Nephrology, The First Affiliated Hospital of Sun Yat-sen University. All patients were confirmed by histopathological examination of renal biopsy. Patients with secondary IgAN such as chronic liver cirrhosis, SLE, and Henoch–Schonlein purpura were removed. Demographic and clinical information of IgAN patients were collected at the time of diagnosis.
Two independent groups of 2400 age- and gender-matched healthy controls (1000 healthy controls for discovery stage and 1400 healthy controls for validation stage) with normal urinalysis and no history of kidney diseases were collected from the Health Examination Center, The First Affiliated Hospital of Sun Yat-sen University. All subjects were unrelated and self-described as Han Chinese. The study was approved by the Research Ethics Committee, The First Affiliated Hospital of Sun Yat-sen University and conformed to the principles outlined in the Helsinki declaration. Informed consent was obtained from all the study participants.
SNP selection
We used the HapMap (http://www.HapMap.org) to download the single-nucleotide polymorphism (SNP) genotyping data for ITGAX and ITGAM (Release 27, Phase I + II + III, CHB populations). Haploview software (https://www.broadinstitute.org/haploview/haploview) was used to analyze the HapMap data. Five tag SNPs (rs7199585, rs11150614, rs11150619, rs2929, and rs9929832) were chosen for the ITGAX gene, and only one tag SNP (rs4597342) was chosen for ITGAM owing to the high linkage disequilibrium (LD) among the common SNPs of this gene. The selection of these SNPs was based on the criteria of the minor allele frequency ≥5% and r2 threshold of 0.80. Another functional SNP of ITGAX (rs1140195) was also selected after searching the SNPinfo (http://snpinfo.niehs.nih.gov/snpinfo/snpfunc.htm) website for its potential role as microRNA (miRNA) binding site. Then, the LD analysis was conducted for these six SNPs of ITGAX by using Haploview software. We found that rs7199585 and rs1140195 were in high LD (r2 = 0.86), and the potentially functional SNP rs1140195 was left for further analysis. The GWAS-reported SNP rs7190997 was also included in this study. Finally, a total of seven SNPs (rs11150614, rs11150619, rs2929, rs9929832, rs4597342, rs1140195, and rs7190997) were chosen for genotyping.
Genotyping assay
DNA was isolated from 2 ml whole blood of 4100 samples by using a commercial DNA extraction Kit (Qiagen, Hilden, Germany) according to the manufacturer’s instructions. The Sequenom MassARRAY system (Sequenom, San Diego, USA) was used to genotype the seven candidate SNPs in the discovery stage, and Kompetitive Alelle Specific PCR technology with SNPLine platform (LGC Genomics, the United Kingdom) was used to genotype the potentially significant SNP in the validation stage. The corresponding polymerase chain reaction (PCR) primers used for each SNP are listed in Supplementary Tables 1 and 2.
Measurement of ITGAX and ITGAM mRNA levels
To evaluate ITGAX and ITGAM gene expression levels in WBCs, we enrolled 74 IgAN patients and 54 gender- and age-matched healthy controls. Total RNA from WBCs was isolated by using TRIzol reagent (Invitrogen, Carlsbad, CA, USA). The mRNA levels were evaluated by TaqMan quantitative real-time polymerase chain reaction (qRT-PCR) experiments using the ABI 7900 (ABI prism 7900HT Sequence Detection System, Life Technology, USA). TaqMan probes and primers for the target ITGAX gene (Hs00174217-m1), ITGAM gene (Hs00167304-m1), and the internal reference gene GAPDH (Hs03929097-gl) were obtained from Applied Biosystems (Carlsbad, CA, USA). ΔCt values were calculated (Ct of the target ITGAX and ITGAM gene minus Ct of the GAPDH gene) in each sample that was evaluated in duplicate. The relative mRNA expression levels were assessed as the expression of the target ITGAX and ITGAM gene normalized to the expression of GAPDH by using the 2−ΔCt method.
Bioinformatics analysis of the significant SNP
We used HaploReg 4.1 (http://compbio.mit.edu/HaploReg) to explore the tagged variants of rs11150619 according to the LD information from the ASN population (CHB, Chinese individuals in Beijing; +JPT, Japanese individuals in Tokyo) with r2 ≥ 0.8, and RegulomeDB online website (http://www.regulomedb.org/) was used to calculate the regulome database score.
To investigate whether rs11150619 had a functional effect on gene expression in Epstein-Barr virus-transformed lymphoblastoid cell lines, we conducted genotype-phenotype analysis using the publicly available SNPexp online tool (http://app3.titan.uio.no/biotools/tool.php?app=snpexp). The genotype data were obtained from the HapMap (phase II release 23 data set) database including 270 individuals with different ethnicities, among which 45 were unrelated CHB individuals.
Statistical analyses
Mean ± standard deviation (SD) was applied to present the normally distributed variables. Median (interquartile ranges) and percentages were used for non-normally distributed variables and categorical data, respectively. Student’s t-test, nonparametric Mann–Whitney U-test, one-way analysis of variance and Kruskal–Wallis test were performed for comparison of continuous variables.
We evaluated the Hardy-Weinberg equilibrium (HWE) by chi-square (χ2) test for each candidate SNP in healthy controls. The associations between SNPs and IgAN susceptibility were evaluated by odds ratios (ORs) and 95% confidence intervals (CIs) using multivariate logistic regression analysis with adjustment for age and gender under additive, dominant and recessive genetic models as previously described [18]. We used Haploview 4.2 software (Broad Institute, Cambridge, MA, USA) to detect the LD of SNPs within ITGAX and ITGAM. Bonferroni correction for multiple tests was applied by lowering the significance level of a P value < 0.0083 for the six SNPs.
The Mann–Whitney U-test and student’s t-test were used to compare ITGAX and ITGAM mRNA levels in WBCs between IgAN patients and healthy controls, respectively. Statistical comparison of ITGAX and ITGAM mRNA levels among SNP genotypes was assessed by using the Mann–Whitney U-test and Student’s t-test. The correlations between ITGAX and ITGAM mRNA levels and clinical parameters of IgAN patients were tested using Spearman and Pearson correlation analysis. P ≤ 0.05 was considered to be statistical significance. All analyses were assessed by using PLINK 1.07 [19], SPSS 19.0 (SPSS, Inc., Chicago, IL, USA), and GraphPad Prism 5.0 (GraphPad Software Inc., San Diego, CA, USA).
Results
General characteristics of the study subjects
The basic characteristics for the discovery stage (1000 IgAN cases and 1000 controls) and the validation stage (700 IgAN cases and 1400 controls) are summarized in Table 1. The mean (SD) age of 1700 cases and 2400 controls in the combined samples was 35.54 ± 10.81 and 34.92 ± 11.06 years, respectively. No significant differences in the age (P = 0.076) and gender distribution (P = 0.260) were noted between the two groups (Table 1).
Association of ITGAX and ITGAM gene polymorphisms with IgAN susceptibility
Seven candidate SNPs were all successfully genotyped and all the SNPs had call rates >95%. However, SNP rs2929 was excluded for further analysis due to deviation from HWE in the control samples (P < 0.05). Four SNPs (rs11150619, rs11150614, rs7190997, and rs4597342) showed potential associations with the susceptibility of IgAN (P < 0.05), but only rs11150619 achieved significance after Bonferroni’s correction under the additive model (OR = 0.83, 95%CI = 0.72–0.95, P = 0.006) (Table 2).
Then, we recruited 700 IgAN cases and 1400 controls to further validate the association of rs11150619 with IgAN. A marginal association was noted between rs11150619 and IgAN risk in the validation stage under the dominant model (OR = 0.83, 95% CI = 0.69–1.00, P = 0.051) (Table 3). However, when we further analyzed the association in the combined samples, the significant effect of rs11150619 became more evident for IgAN susceptibility (OR = 0.86, 95%CI = 0.78–0.94, additive P = 1.56 × 10−3; OR = 0.81, 95%CI = 0.71–0.91, dominant P = 6.68 × 10−4) (Table 3).
Associations between rs11150619 and the clinical phenotypes of IgAN
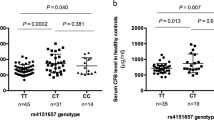
We collected the clinical data of IgAN patients in the discovery stage and further investigated the associations between the significant SNP rs11150619 and the clinical phenotypes. As shown in Table 4, our results indicated that the protective allele T of rs11150619 was associated with younger age (P = 0.019), increased eGFR levels (P = 0.034), increased serum C3 levels (P = 0.051) and a reduced proportion of global glomerulosclerosis (P = 0.031).
Comparison of ITGAX and ITGAM mRNA levels among rs11150619 genotypes
The ITGAX mRNA expression levels in WBCs of IgAN patients were increased compared with those of normal controls [0.55 (0.37–0.85) versus 0.44 (0.36–0.50), P = 0.004] (Fig. 1a), but no significant difference was found between IgAN patients and controls for ITGAM mRNA expression levels (Fig. 1b). Moreover, there was no significant correlation between ITGAX and ITGAM mRNA levels and clinical variables in IgAN patients (Supplementary Table 3).
ITGAX mRNA expression levels in healthy controls and IgAN patients [0.55 (0.37–0.85) versus 0.44 (0.36–0.50), P = 0.004] (a). ITGAM mRNA expression levels in healthy controls and IgAN patients (0.016 ± 0.008 versus 0.016 ± 0.009, P = 0.833) (b). Bars represent median and mean values of groups in a and b, respectively
To evaluate the influence of rs11150619 on ITGAX and ITGAM expression, we compared ITGAX and ITGAM mRNA levels among different genotypes in IgAN patients and healthy controls. As shown in Fig. 2, no significant differences of ITGAX and ITGAM mRNA levels were found among individuals with different rs11150619 genotypes (P > 0.05), which was consistent with the observation regarding the rs11150619 genotypes and ITGAX expression in the CHB population according to the SNPexp database (P = 0.168, Supplementary Table 4).
ITGAX and ITGAM mRNA levels in individuals with different genotypes of rs11150619 in IgAN patients and healthy controls. There was no statistical difference between different genotypes of rs11150619 of ITGAX and ITGAM mRNA levels in IgAN patients (a, c) and healthy controls (b, d). Bars represent median (a, b) and mean (c, d) values of groups
Functional annotation of SNP rs11150619
SNP rs11150619 was located in the intron of ITGAX, and 12 variants (r2 ≥ 0.8) were tagged by rs11150619 using the HaploReg v4.1 database. All these 12 SNPs are located within the ITGAX gene, and we examined their potential biological significance in RegulomeDB (Supplementary Table 5). SNP rs11150619 was annotated as regulatory with “score 3a: TF binding + any motif + DNase peak”. Among the 12 proxy variants, rs11150616 and rs13332545 had the highest evidence for potential function (score 1f: eQTL + TF binding/DNase peak), and a missense variant rs2230429 exhibited high LD (r2 = 0.95) with rs11150619.
Discussion
In current study, we conducted a two-stage association study with a total of 1700 sporadic IgAN patients and 2400 healthy controls in a Chinese Han population to explore the genetic association of ITGAX and ITGAM SNPs with the susceptibility of IgAN. Our results showed that the rs11150619-T allele was significantly associated with reduced risk and disease severity of IgAN. Thus, it may serve as an important protective role for IgAN.
The ITGAX-ITGAM region was associated with numerous autoimmune diseases [15,16,17], and SNP rs7190997 was identified to be associated with IgAN in the Chinese population [10]. In the current study, our data showed that rs7190997 was nominally associated with IgAN susceptibility, but we found another SNP rs11150619 which was in high LD (r2 = 0.78, Fig. 3) with rs7190997 showed a strong association with IgAN. After functional annotation analysis, RegulomeDB [20] annotated that rs11150619 (scored 3a) was less likely to affect binding and had no eQTL data. When we investigated whether rs11150619 have a functional influence on ITGAX and ITGAM gene expression, no statistical significance was noted among different genotypes of rs11150619, which was consistent with the results from SNPexp [21]. Using HaploReg [22], 12 proxy SNPs to rs11150619 were identified and five of these SNPs had a high degree of potential regulatory function (RegulomeDB scored 2b-1f), suggesting that ITGAX variants may influence disease by regulating gene expression. Moreover, of the 12 SNPs, the missense variant rs2230429 was predicted to have a probable damaging effect on the protein according to PolyPhen-2 database [23] and may influence the formation of CR4 protein. Although we observed the association of rs11150619 polymorphism with IgAN susceptibility, we couldn’t speculate whether it is due to the SNP itself, or other causal SNPs having LD with rs11150619. Further comprehensive analysis of ITGAX-ITGAM locus should be performed to identify the causal variants within this region.
Of the two other nominally associated SNPs (rs11150614 and rs4597342), SNP rs11150614 is located in the intron of ITGAX with little evidence of predicted functional effects (RegulomeDB scored 4). However, SNP rs4597342 was predicted to be located in the 3′ UTR of ITGAM and affect binding between the miRNA and ITGAM based on the SNPinfo website [24], which may influence the translation of ITGAM mRNA and disease susceptibility. Identification of multiple variants in the ITGAX-ITGAM region indicated a potential effect of ITGAX-ITGAM SNPs on the susceptibility for IgAN. The investigators have demonstrated that variants in ITGAX-ITGAM region have functional effect on the ligand-binding and phagocytosis of iC3b-coated targets [25]. The CR4-mediated inflammatory responses were enhanced by increased CR4 expression and ligand binding [26], and iC3b/CR4 interaction can amplify persistent glomerular infiltration of immune cells in IgAN [14]. Moreover, CD11b (ITGAM) participates in the regulation of IgA-producing cells in the intestine [27], and CR3 (CD11b/CD18) is also required for the interaction between CD89 (FcαR) and secretory IgA [28]. In IgAN, IgA1 can bind with CD89 to form an IgA1-CD89 complex, and then interact with CD71 (TfR1) on mesangial cells and accumulate in the kidney [29]. Thus, we speculated that ITGAX and ITGAM gene polymorphisms may affect the gene expression, structures and ligand binding activity of CR3 and CR4, eventually leading to the deposition of IgA1-CD89 complex in the kidney, aberrant inflammatory and complement response during the development of IgAN.
Given that IgAN patients exhibited different clinical features, we further investigated the correlation between ITGAX rs11150619 and various clinical phenotypes of IgAN. The protective allele T of rs11150619 was found to be associated with less disease severity including increased eGFR levels, serum C3 levels and reduced proportion of global glomerulosclerosis in IgAN patients. However, the patients with TT genotypes were younger, and these patients with higher eGFR levels and less global glomerulosclerosis may partially due to the age. In addition, it was noteworthy that increased serum C3 levels indicated better prognosis of IgAN [30], suggesting that rs11150619 may have functional relevance in complement activation by influencing the serum C3 levels. Taken together, our findings indicated that the SNP rs11150619 may not only influence disease susceptibility but can also serve as an important factor to predict IgAN disease severity.
ITGAX and ITGAM play a pivotal role in cellular signaling and immunoregulation of multiple inflammatory diseases by T cell activation and cytokine production [31, 32]. In addition, ITGAX was demonstrated to be a treatment biomarker to predict response to a TNF inhibitor drug in rheumatoid arthritis patients [33]. Previous studies showed that ITGAX and ITGAM were highly expressed in the kidney of IgAN and positively associated with proteinuria [34]. It is likely that ITGAX and ITGAM may act as an important player on the process of binding with the ligands, eventually influencing the inflammatory response during the development of IgAN. We observed that IgAN patients had higher ITGAX mRNA levels compared with healthy controls. However, we did not identify a correlation between ITGAX mRNA expression levels and clinical variables. Whether ITGAX expression levels can be applied as a biomarker for predicting disease severity and progression require further elucidation.
In brief, the current study suggested that the variants of ITGAX and ITGAM are involved in the susceptibility of IgAN, and the rs11150619-T allele showed a protective role for IgAN. More experimental data are needed to elucidate the biologic mechanisms of the associations of ITGAX and ITGAM with IgAN.
References
Liu ZH. Nephrology in china. Nat Rev Nephrol. 2013;9:523–8.
Lai KN, Tang SC, Schena FP, Novak J, Tomino Y, Fogo AB, et al. IgA nephropathy. Nat Rev Dis Primers. 2016;2:16001.
Daha MR, van Kooten C. Role of complement in IgA nephropathy. J Nephrol. 2016;29:1–4.
Reid S, Cawthon PM, Craig JC, Samuels JA, Molony DA, Strippoli GF. Non-immunosuppressive treatment for IgA nephropathy. Cochrane Database Syst Rev. 2011;3:CD003962.
Feehally J, Barratt J. The Genetics of IgA Nephropathy: an Overview from western countries. Kidney Dis. 2015;1:33–41.
Feehally J, Farrall M, Boland A, Gale DP, Gut I, Heath S, et al. HLA has strongest association with IgA nephropathy in genome-wide analysis. J Am Soc Nephrol. 2010;21:1791–7.
Gharavi AG, Kiryluk K, Choi M, Li Y, Hou P, Xie J, et al. Genome-wide association study identifies susceptibility loci for IgA nephropathy. Nat Genet. 2011;43:321–7.
Yu XQ, Li M, Zhang H, Low HQ, Wei X, Wang JQ, et al. A genome-wide association study in Han Chinese identifies multiple susceptibility loci for IgA nephropathy. Nat Genet. 2011;44:178–82.
Kiryluk K, Li Y, Scolari F, Sanna-Cherchi S, Choi M, Verbitsky M, et al. Discovery of new risk loci for IgA nephropathy implicates genes involved in immunity against intestinal pathogens. Nat Genet. 2014;46:1187–96.
Li M, Foo JN, Wang JQ, Low HQ, Tang XQ, Toh KY, et al. Identification of new susceptibility loci for IgA nephropathy in Han Chinese. Nat Commun. 2015;6:7270.
Fagerholm SC, MacPherson M, James MJ, Sevier-Guy C, Lau CS. The CD11b-integrin (ITGAM) and systemic lupus erythematosus. Lupus. 2013;22:657–63.
Erdei A, Lukacsi S, Macsik-Valent B, Nagy-Balo Z, Kurucz I, Bajtay Z. Non-identical twins: different faces of CR3 and CR4 in myeloid and lymphoid cells of mice and men. Semin Cell Dev Biol. 2019;85:110–21.
Myones BL, Dalzell JG, Hogg N, Ross GD. Neutrophil and monocyte cell surface p150,95 has iC3b-receptor (CR4) activity resembling CR3. J Clin Invest. 1988;82:640–51.
Ootaka T, Saito T, Soma J, Yusa A, Abe K. Intercellular adhesion molecule-1/leukocyte function associated antigen-1-mediated and complement receptor type 4-mediated infiltration and activation of glomerular immune cells in immunoglobulin A nephropathy. Am J Kidney Dis. 1996;28:40–6.
Hom G, Graham RR, Modrek B, Taylor KE, Ortmann W, Garnier S, et al. Association of systemic lupus erythematosus with C8orf13-BLK and ITGAM-ITGAX. N Engl J Med. 2008;358:900–9.
Anaya JM, Kim-Howard X, Prahalad S, Chernavsky A, Canas C, Rojas-Villarraga A, et al. Evaluation of genetic association between an ITGAM non-synonymous SNP (rs1143679) and multiple autoimmune diseases. Autoimmun Rev. 2012;11:276–80.
Park SR, Park KS, Park YJ, Bang D, Lee ES. CD11a, CD11c, and CD18 gene polymorphisms and susceptibility to Behcet’s disease in Koreans. Tissue Antigens. 2014;84:398–404.
Ma RC, Tam CH, Wang Y, Luk AO, Hu C, Yang X, et al. Genetic variants of the protein kinase C-beta 1 gene and development of end-stage renal disease in patients with type 2 diabetes. J Am Med Assoc. 2010;304:881–9.
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MA, Bender D, et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet. 2007;81:559–75.
Boyle AP, Hong EL, Hariharan M, Cheng Y, Schaub MA, Kasowski M, et al. Annotation of functional variation in personal genomes using RegulomeDB. Genome Res. 2012;22:1790–7.
Holm K, Melum E, Franke A, Karlsen TH. SNPexp - A web tool for calculating and visualizing correlation between HapMap genotypes and gene expression levels. BMC Bioinforma. 2010;11:600.
Ward LD, Kellis M. HaploReg v4: systematic mining of putative causal variants, cell types, regulators and target genes for human complex traits and disease. Nucleic Acids Res. 2016;44:D877–81.
Ramensky V, Bork P, Sunyaev S. Human non-synonymous SNPs: server and survey. Nucleic Acids Res. 2002;30:3894–900.
Xu Z, Taylor JA. SNPinfo: integrating GWAS and candidate gene information into functional SNP selection for genetic association studies. Nucleic Acids Res. 2009;37:W600–5.
Roberts AL, Thomas ER, Bhosle S, Game L, Obraztsova O, Aitman TJ, et al. Resequencing the susceptibility gene, ITGAM, identifies two functionally deleterious rare variants in systemic lupus erythematosus cases. Arthritis Res Ther. 2014;16:R114.
Walzog B, Weinmann P, Jeblonski F, Scharffetter-Kochanek K, Bommert K, Gaehtgens P. A role for beta(2) integrins (CD11/CD18) in the regulation of cytokine gene expression of polymorphonuclear neutrophils during the inflammatory response. FASEB J. 1999;13:1855–65.
Kunisawa J, Gohda M, Hashimoto E, Ishikawa I, Higuchi M, Suzuki Y, et al. Microbe-dependent CD11b+ IgA+ plasma cells mediate robust early-phase intestinal IgA responses in mice. Nat Commun. 2013;4:1772.
van Egmond M, van Vuuren AJ, Morton HC, van Spriel AB, Shen L, Hofhuis FM, et al. Human immunoglobulin A receptor (FcalphaRI, CD89) function in transgenic mice requires both FcR gamma chain and CR3 (CD11b/CD18). Blood. 1999;93:4387–94.
Daha MR, van Kooten C. Deposition of IgA in primary IgA nephropathy: it takes at least four to tango. Nephrol Dial Transplant. 2013;28:794–7.
Pan M, Zhang J, Li Z, Jin L, Zheng Y, Zhou Z, et al. Increased C4 and decreased C3 levels are associated with a poor prognosis in patients with immunoglobulin A nephropathy: a retrospective study. BMC Nephrol. 2017;18:231.
Ehirchiou D, Xiong Y, Xu G, Chen W, Shi Y, Zhang L. CD11b facilitates the development of peripheral tolerance by suppressing Th17 differentiation. J Exp Med. 2007;204:1519–24.
Vinay DS, Kwon BS. CD11c+CD8+ T cells: two-faced adaptive immune regulators. Cell Immunol. 2010;264:18–22.
Stuhlmuller B, Haupl T, Hernandez MM, Grutzkau A, Kuban RJ, Tandon N, et al. CD11c as a transcriptional biomarker to predict response to anti-TNF monotherapy with adalimumab in patients with rheumatoid arthritis. Clin Pharm Ther. 2010;87:311–21.
Ootaka T, Saito T, Soma J, Yusa A, Abe K. Mechanism of infiltration and activation of glomerular monocytes/macrophages in IgA nephropathy. Am J Nephrol. 1997;17:137–45.
Acknowledgements
This research was funded in part by the National Key Research and Development Project, China (No. 2016YFC0906101), the Operational Grant of Guangdong Provincial Key Laboratory, China (No. 2017B030314019), the Guangdong Provincial Program of Science and Technology, China (No. 2017A050503003; No. 2017B020227006), the National Natural Science Foundation of China (No. 81770661; No. 81570599; No. 81570750), the Guangzhou Municipal Program of Science and Technology, China (No. 2016201604030005; No. 201704020167; No. 201807010002) and the Young and Middle-aged Talents Program of The First Affiliated Hospital, Sun Yat-sen University.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Shi, D., Zhong, Z., Xu, R. et al. Association of ITGAX and ITGAM gene polymorphisms with susceptibility to IgA nephropathy. J Hum Genet 64, 927–935 (2019). https://doi.org/10.1038/s10038-019-0632-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s10038-019-0632-2