Abstract
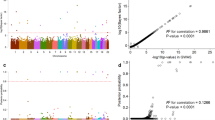
Genome-wide association study has limited to discover single-nucleotide polymorphisms (SNPs) in several ethnicities. Here, we investigated an initial GWAS to identify genetic modifiers predicting with adult moyamoya disease (MMD) in Koreans. GWAS was performed in 216 patients with MMD and 296 controls using the large-scale Asian-specific Axiom Precision Medicine Research Array. A subsequent fine-mapping analysis was conducted to assess the causal variants associated with adult MMD. A total of 489,966 out of 802,688 SNPs were subjected to quality control analysis. Twenty-one SNPs reached a genome-wide significance threshold (p = 5 × 10−8) after pruning linkage disequilibrium (r2 < 0.8) and mis-clustered SNPs. Among these variants, the 17q25.3 region including TBC1D16, CCDC40, GAA, RNF213, and ENDOV genes was broadly associated with MMD (p = 3.1 × 10−20 to 4.2 × 10−8). Mutations in RNF213 including rs8082521 (Q1133K), rs10782008 (V1195M), rs9913636 (E1272Q), rs8074015 (D1331G), and rs9674961 (S2334N) showed a genome-wide significance (1.9 × 10−8 < p < 4.3 × 10−12) and were also replicated in the East-Asian populations. In subsequent analysis, RNF213 mutations were validated in a fine-mapping outcome (log10BF > 7). Most of the loci associated with MMD including 17q25.3 regions were detected with a statistical power greater than 80%. This study identifies several novel and known variations predicting adult MMD in Koreans. These findings may good biomarkers to evaluate MMD susceptibility and its clinical outcomes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Roder C, Nayak NR, Khan N, Tatagiba M, Inoue I, Krischek B. Genetics of Moyamoya disease. J Hum Genet. 2010;55:711–16.
Ahn IM, Park DH, Hann HJ, Kim KH, Kim HJ, Ahn HS. Incidence, prevalence, and survival of moyamoya disease in Korea: A nationwide, population-based study. Stroke 2014;45:1090–95.
Im SH, Cho CB, Joo WI, Chough CK, Park HK, Lee KJ, et al. Prevalence and Epidemiological Features of Moyamoya Disease in Korea. J Cerebrovasc Endovasc Neurosurg. 2012;14:75–8.
Kuroda S, Houkin K. Moyamoya disease: Current concepts and future perspectives. Lancet Neurol. 2008;7:1056–66.
Nanba R, Kuroda S, Tada M, Ishikawa T, Houkin K, Iwasaki Y. Clinical features of familial moyamoya disease. Childs Nerv Syst. 2016;22:258–62.
Kamada F, Aoki Y, Narisawa A, Abe Y, Komatsuzaki S, Kikuchi A, et al. A genome-wide association study identifies RNF213 as the first Moyamoya disease gene. J Hum Genet. 2011;56:34–40.
Liu W, Morito D, Takashima S, Mineharu Y, Kobayashi H, Hitomi T, et al. A Identification of RNF213 as a susceptibility gene for moyamoya disease and its possible role in vascular development. PLoS One. 2011;6:e22542.
Duan L, Wei L, Tian Y, Zhang Z, Hu P, Wei Q, et al. Novel susceptibility loci for moyamoya disease revealed by a genome-wide association study. Stroke 2018;49:11–8.
Jang MA, Chung JW, Yeon JY, Kim JS, Hong SC, Bang OY, et al. Frequency and significance of rare RNF213 variants in patients with adult moyamoya disease. PLoS One. 2017;12:e0179689.
Kim EH, Yum MS, Ra YS, Park JB, Ahn JS, Kim GH, et al. Importance of RNF213 polymorphism on clinical features and long-term outcome in moyamoya disease. J Neurosurg. 2016;124:1221–27.
Benner C, Havulinna AS, Jarvelin MR, Salomaa V, Ripatti S, Pirinen M. Prospects of fine-mapping trait-associated genomic regions by using summary statistics from genome-wide association studies. Am J Hum Genet. 2017;101:539–51.
Freebern E, Santos DJA, Fang L, Jiang J, Parker Gaddis KL, Liu GE, et al. GWAS and fine-mapping of livability and six disease traits in Holstein cattle. BMC Genomics. 2020;21:41.
Park JJ, Kim BJ, Youn DH, Choi HJ, Jeon JP. A Preliminary Study of the Association between SOX17 Gene Variants and Intracranial Aneurysms Using Exome Sequencing. J Korean Neurosurg Soc. 2020;63:559–65.
Hong EP, Kim BJ, Cho SS, Yang JS, Choi HJ, Kang SH, et al. Genomic Variations in Susceptibility to Intracranial Aneurysm in the Korean Population. J Clin Med. 2019;8:275.
Kim JE, Kim KM, Kim JG, Kang HS, Bang JS, et al. Clinical features of adult moyamoya disease with special reference to the diagnosis. Neurol Med Chir (Tokyo). 2012;52:311–7.
Jeon JS, Ahn JH, Moon YJ, Cho WS, Son YJ, Kim SK, et al. Expression of cellular retinoic acid-binding protein-I (CRABP-I) in the cerebrospinal fluid of adult onset moyamoya disease and its association with clinical presentation and postoperative haemodynamic change. J Neurol Neurosurg Psychiatry. 2014;85:726–31.
Chang CC, Chow CC, Tellier LC, Vattikuti S, Purcell SM, Lee JJ. Second-generation PLINK: rising to the challenge of larger and richer datasets. Gigascience 2015;4:7.
Pruim RJ, Welch RP, Sanna S, Teslovich TM, Chines PS, Gliedt TP, et al. Locuszoom: Regional visualization of genome-wide association scan results. Bioinformatics 2010;26:2336–7.
Barrett JC, Fry B, Maller J, Daly MJ. Haploview: Analysis and visualization of ld and haplotype maps. Bioinformatics 2005;21:263–5.
Wang K, Li M, Hakonarson H. ANNOVAR. functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 2010;38:e164.
Moteki Y, Onda H, Kasuya H, Yoneyama T, Okada Y, Hirota K, et al. Systematic Validation of RNF213 Coding Variants in Japanese Patients With Moyamoya Disease. J Am Heart Assoc. 2015;4:e001862.
Machiela MJ, Chanock SJ. LDlink: a web-based application for exploring population-specific haplotype structure and linking correlated alleles of possible functional variants. Bioinformatics 2015;31:3555–7.
Benner C, Spencer CC, Havulinna AS, Salomaa V, Ripatti S, Pirinen M. FINEMAP: efficient variable selection using summary data from genome-wide association studies. Bioinformatics 2016;32:1493–501.
van Rooij FJA, Qayyum R, Smith AV, Zhou Y, Trompet S, Tanaka T, et al. Genome-wide Trans-ethnic Meta-analysis Identifies Seven Genetic Loci Influencing Erythrocyte Traits and a Role for RBPMS in Erythropoiesis. Am J Hum Genet. 2017;100:51–63.
Hong EP, Youn DH, Kim BJ, Ahn JH, Park JJ, Rhim JK, et al. Fine-mapping of intracranial aneurysm susceptibility based on a genome-wide association study. Sci Rep. 2022;12:2717.
Purcell S, Cherny SS, Sham PC. Genetic Power Calculator: Design of linkage and association genetic mapping studies of complex traits. Bioinformatics 2003;19:149–50.
Hong EP, Park JW. Sample size and statistical power calculation in genetic association studies. Genomics Inf. 2012;10:117–22.
Kim JS. Moyamoya disease: Epidemiology, clinical features, and diagnosis. J Stroke. 2016;18:2–11.
Zhu B, Liu X, Zhen X, Li X, Wu M, Zhang Y, et al. RNF213 gene polymorphism rs9916351 and rs8074015 significantly associated with moyamoya disease in Chinese population. Ann Transl Med. 2020;8:851.
Yamauchi T, Tada M, Houkin K, Tanaka T, Nakamura Y, Kuroda S, et al. Linkage of familial moyamoya disease (spontaneous occlusion of the circle of Willis) to chromosome 17q25. Stroke 2000;31:930–5.
Mineharu Y, Liu W, Inoue K, Matsuura N, Inoue S, Takenaka K, et al. Autosomal dominant moyamoya disease maps to chromosome 17q25.3. Neurology 2008;70:2357–63.
Koizumi A, Kobayashi H, Hitomi T, Harada KH, Habu T, Youssefian S. A new horizon of moyamoya disease and associated health risks explored through RNF213. Environ Health Prev Med. 2016;21:55–70.
Joo SP, Kim TS, Lee IK, Kim JT, Park MS, Cho KH. A genome-wide study of moyamoya-type cerebrovascular disease in the korean population. J Korean Neurosurg Soc. 2011;50:486–91.
Kanoke A, Fujimura M, Niizuma K, Ito A, Sakata H, Sato-Maeda M, et al. Temporal profile of the vascular anatomy evaluated by 9.4-tesla magnetic resonance angiography and histological analysis in mice with the R4859K mutation of RNF213, the susceptibility gene for moyamoya disease. Brain Res. 2015;1624:497–505.
Sugihara M, Morito D, Ainuki S, Hirano Y, Ogino K, Kitamura A, et al. The AAA+ ATPase/ubiquitin ligase mysterin stabilizes cytoplasmic lipid droplets. J Cell Biol. 2019;218:949–60.
Mineharu Y, Miyamoto S. RNF213 and GUCY1A3 in Moyamoya Disease: Key regulators of metabolism, inflammation, and vascular stability. Front Neurol. 2021;12:687088.
Hsu SH, Shyu HW, Hsieh-Li HM, Li H. Spz1, a novel bhlh-zip protein, is specifically expressed in testis. Mech Dev. 2001;100:177–87.
Liu X, Han X, Wan X, He C, Wang Y, Mao A, et al. Spz1 is critical for chemoresistance and aggressiveness in drug-resistant breast cancer cells. Biochem Pharm. 2018;156:43–51.
Wang LT, Chiou SS, Chai CY, Hsi E, Chiang CM, Huang SK, et al. Transcription factor SPZ1 promotes TWIST-mediated epithelial-mesenchymal transition and oncogenesis in human liver cancer. Oncogene 2017;36:4405–14.
Bhak Y, Jeon Y, Jeon S, Yoon C, Kim M, Blazyte A, et al. Polygenic risk score validation using Korean genomes of 265 early-onset acute myocardial infarction patients and 636 healthy controls. PLoS One. 2021;16:e0246538.
Sekar A, Bialas AR, de Rivera H, Davis A, Hammond TR, Kamitaki N, et al. Schizophrenia risk from complex variation of complement component 4. Nature 2016;530:177–83.
Akan G, Kisenge P, Sanga TS, Mbugi E, Adolf I, Turkcan MK, et al. Common SNP-based haplotype analysis of the 9p21.3 gene locus as predictor coronary artery disease in Tanzanian population. Cell Mol Biol (Noisy-le-Gd). 2019;65:33–43.
Kuriyama S, Kusaka Y, Fujimura M, Wakai K, Tamakoshi A, Hashimoto S, et al. Prevalence and clinicoepidemiological features of moyamoya disease in Japan: findings from a nationwide epidemiological survey. Stroke 2008;39:42–7.
Miyatake S, Miyake N, Touho H, Nishimura-Tadaki A, Kondo Y, et al. Homozygous c.14576G>A variant of RNF213 predicts early-onset and severe form of moyamoya disease. Neurology 2012;78:803–10.
Acknowledgements
This research was supported by a grant of the Seoul National University Hospital (grant number: 0320180090) and Korea Health Technology R&D Project through the Korea Health Industry Development Institute (KHIDI), funded by the Ministry of Health & Welfare, Republic of Korea (grant number: HR21C0198). In addition, all authors thank all members manufacturing “The First Korean Stroke Genetics Association Research (The FirstKSGAR, https://1ksgh.org/)” consortium, who correct biospecimen and clinical datasets.
Author information
Authors and Affiliations
Contributions
JEK designed and managed this study. All of analysis and result interpretation were equally done by JPJ and EPH Sample preparation and data collection were done by EJH, BJK, DHY, SL, HCL, KMK, SHL, WSC, and HSK. Drafting and reviewing of manuscript were done by JPJ, EPH, EJH, BJK, DHY, SL, HCL, KMK, SHL, WSC, HSK, and JEK.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Jeon, J.P., Hong, E.P., Ha, E.J. et al. Genome-wide association study identifies novel susceptibilities to adult moyamoya disease. J Hum Genet 68, 713–720 (2023). https://doi.org/10.1038/s10038-023-01167-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s10038-023-01167-9