Abstract
The influence of the PvuII polymorphism (intron 6) of the lipoprotein lipase (LPL) gene on cord plasma lipid traits was studied in 252 ethnic Chinese and 240 ethnic Indian newborns of Singapore. The allelic frequencies of P+ (presence of the restriction site) were 0.67 and 0.56 in the Chinese and Indian newborns, respectively, similar to their respective adult populations. The genotype distributions at the Pvu II site were at Hardy Weinberg equilibrium in both ethnic Chinese(χ2 = 2.0) and ethnic Indians (χ2 = 3.6). Cord blood HDL-cholesterol (HDL-C) levels are higher in newborn Chinese than newborn Indians. In addition, cord blood LDL-cholesterol (LDL-C), apoB, and lipoprotein(a) levels are lower in newborn Chinese than newborn Indians. Both newborn Chinese and Indian male homozygotes for P- allele have higher cord blood LDL-C levels than newborns with the more common P+P+ or P-P+ genotypes. In Chinese male newborns, the LDL-C levels were 0.76 ± 0.61 mmol/L, 0.53 ± 0.29 mmol/L and 0.46 ± 0.25 mmol/L, respectively (p = 0.01). In Indian male newborns, the LDL-C levels were 0.88 ± 0.35 mmol/L for the P-P- genotype and 0.65 ± 0.24 mmol/L for the P+P+ genotype (p = 0.003). In addition, the influence of the P- allele on LDL-C levels is remarkably similar in both ethnic groups, accounting for 8.48% of the population variance in the Chinese newborns and 8.09% in the Indian newborns. In contrast, no obvious effect of genotype is seen in this lipid parameter in the newborn females of either ethnic groups. There is presence of significant genotype specific influence on the LDL-C levels in cord plasma in male newborns, suggesting an early expression of the LPL gene locus.
Similar content being viewed by others
Main
In Singapore, ethnic Indians (South Asians) have a prevalence and mortality rate of CAD about three times that of ethnic Chinese(1, 2). Clinical studies have demonstrated a close association between lipoprotein abnormalities and susceptibility to CAD. The types of lipoprotein abnormalities encountered in patients with CAD include increased LDL-C levels, decreased HDL-C levels along with increased TG or VLDL levels and increased Lp(a) levels(3–8). Family studies have revealed that individual variations in the levels of lipoprotein in the blood are due largely to hereditary influences and have been shown to account for 50-65% of individual lipid variation of T-C and LDL-C(9–11).
The cord blood lipid profiles in newborns were investigated in the different ethnic groups of Singapore and revealed significant ethnic differences concordant with coronary risks(12, 13). In these studies, ethnic Indian newborns were shown to have significantly higher plasma lipid levels of TG, LDL-C, apoB, and Lp(a), and significantly lower levels of HDL-C and apoA-I when compared with ethnic Chinese newborns.
LPL plays a critical role in the determination of plasma lipid and lipoprotein profile. With its pivotal role in lipoprotein metabolism, it is a rate-limiting enzyme that affects the clearance of TG-rich lipoproteins, including VLDL and chylomicrons from the circulation. The LPL gene has been found to be located at the short arm of chromosome 8 (8p22)(14), and the gene structure and cDNA sequence have been elucidated. Several RFLPs in the LPL gene have been reported, including the Pvu II. RFLP in intron 6 have been studied for associations with plasma lipids. Associations of the PvuII polymorphism with plasma lipid levels in adults have, however, been inconsistent. Some of these studies reported raised TGs levels in individuals with P+ allele(15, 16), whereas others reported a lack of influence(17–22).
It is expected that the fetal-environmental variations are minimal in utero. In view of the above inconsistent results in the adults, we report in this communication the influence of the PvuII polymorphism of the LPL gene on cord plasma lipid traits in the two ethnic groups (viz., Chinese and Indians) whose newborn lipid profiles have been shown to differ significantly in the same gradient as their ethnic CAD risk levels.
METHODS
Study subjects. A subsample of 492 normal newborns of pure Chinese and Indian heritage from whom DNA samples were available was studied. Verbal informed consent was obtained from parents of the newborns. The cord blood samples of these newborns were studied. The details of the samples and methods of plasma lipid and apo level estimation of the present study have been reported earlier(12, 13).
Isolation of DNA and genotyping of LPL gene PvuII polymorphism. Genomic DNA was extracted from leukocytes obtained from the cord blood samples by sarcosine method(23). DNA samples were amplified by polymerase chain reaction in a Perkin-Elmer Co. thermal cycler 480, in 25μL of reaction mixture containing commercial buffer (Life Technologies, Inc.), 1.5 mM magnesium chloride, 200 μM dNTPs, 0.3 μM of each primer, 0.2-0.5 μg of genomic DNA, and 1.0 U of Taq DNA polymerase. The primer sequences for PvuII (intron 6) site were as follows(18): forward primer, 5′-TAG AGG TTG AGG CAC CTG TGC-3′; reverse primer, 5′-GTG GGT GAA TCA CCT GAG GTC-3′.
Amplification was carried out for 25 cycles at 95°C for 1 min, at 70°C for 2 min, and 72°C for 2 min according to Ahn et al.(18). Ten microliters of the amplified product were digested with 10 U of PvuII overnight at 37°C, size-fractionated in 2% agarose gels with incorporated ethidium bromide, and photographed on a UV transilluminator. The presence of the PvuII site (P+) yielded fragments of 592 and 266 bp.
Statistical analysis. Statistical analysis was performed using the Version 6 of the Statistical Package for the Social Sciences (SPSS) for Windows. The significance of differences for the means of plasma T-C, TG, HDL-C, LDL-C, apoA-I, apoB, and Lp(a) levels between races and gender were determined by the t test. Allelic frequencies of the PvuII RFLP of the LPL gene were estimated by gene counting. Hardy-Weinberg equilibrium for the distribution of genotypes was estimated by χ2 analysis. The significance of difference of proportions of genotypes and alleles was tested by χ2 test. ANOVA was performed to determine the effect of the PvuII polymorphism of the LPL gene on different lipid traits in the different ethnic-gender groups after adjustment for significant covariates. The percentage of variance (r2 × 100) was calculated from the sum of squares. The significance of the sample variance was tested by F and p values. The raw data for TG and Lp(a) were transformed by natural logarithm [ln TG and ln Lp(a)] before comparison by t test and by ANOVA. Because the values of TGs in mmol/L in cord blood specimens were small with values below 1, and as the natural logarithm of <1 is negative, logarithmic transformation for these parameters was done after addition of 1 to the raw values(24–26). Statistical significance was taken at the 0.05 level.
RESULTS
Four hundred ninety-two newborns comprising 245 male (113 Chinese and 132 Indians) and 247 female (139 Chinese and 108 Indians) subjects were available for final analysis. Table 1 shows the means and SD of plasma lipoprotein, apo, and Lp(a) levels in the male and female newborns of the two ethnic groups. The mean birth weight and gestation age of the babies of the same sex were not significantly different between the two ethnic groups. In Indians, female babies had a significantly higher gestational period than male babies (p < 0.01). Female Chinese newborns had significantly higher LDL-C levels than their male counterparts (p< 0.05), whereas Indian females newborns had significantly higher HDL-C(p < 0.05) than the Indian male newborns. Between the two ethnic groups, Chinese female newborns were found to have significantly higher HDL-C levels (p < 0.001) and apoA-I (p < 0.01); lower levels of LDL-C (p < 0.01); apoB (p < 0.001) and ln[Lp(a) + 1] (p < 0.001] than ethnic Indian female newborns. Similarly, between the male subjects of the two ethnic groups, HDL-C and TG were found to be significantly higher in ethnic Chinese (p < 0.001 and < 0.05, respectively). The other adverse risk factors of LDL-C, apoB, and Lp(a) were also significantly higher and unfavorable in Indian male newborns with statistical significance of p < 0.001, < 0.001, and < 0.01, respectively.
The allelic frequencies and genotype distributions of the PvuII polymorphism of the LPL gene in the Chinese and Indian newborns are shown in Table 2. Frequencies for presence of the restriction site(P+) were 0.67 and 0.56 in Chinese and Indians, respectively. No deviations from from Hardy Weinberg equilibrium could be detected in ethnic Chinese (χ2 = 2.0) and in ethnic Indians (χ2 = 3.6).
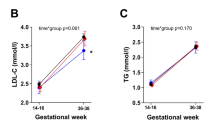
The levels of the various plasma lipid traits were adjusted for the significant covariates of gestation and birth weight before comparison between the PvuII genotypes by ANOVA test in Chinese male and female newborns separately (Table 3) and in Indian male and female newborns separately (Table 4). In Chinese male newborns, the adjusted LDL-C levels were highest and significantly so in the P- homozygotes (0.76 ± 0.61 mmol/L; p = 0.014) with the PvuII genotype accounting for 8.5% of the variance. No significant effects of the LPL gene was observed in ethnic Chinese female newborns. In ethnic Indian male newborns, the adjusted plasma levels of T-C, LDL-C, and apoA-I were significantly higher in P-P- genotype compared with the P+P+ genotype (p = 0.003; 0.003, and 0.016, respectively). Individuals with P- allele also showed a nonsignificant trend toward higher levels of HDL-C (p= 0.063). In ethnic Indian female newborns, the only lipid trait that showed significant genotype-specific variation was TG, which was significantly lower in the P- homozygotes (p = 0.018).
DISCUSSION
The ethnic and gender differences of cord plasma levels of lipids, apoA-I, apoB, and Lp(a) in the present series are similar to that of the earlier reports(12, 13). The P+ allele frequencies for the PvuII RFLP in this study (0.67 in Chinese, 0.56 in Indians) are higher to those reported in adult Caucasian populations(15, 17, 27) and in healthy Japanese(15). They are, however, similar to the adult Chinese and Indian populations of Singapore, the P+ allele frequency being 0.61 and 0.50 in Chinese and Indians, respectively (N. Saha, unpublished results).
Studies on the relationship between genetic variation at the LPL locus and lipid profiles have produced rather equivocal results, in particular with regard to the PvuII RFLP (intron 6). The precise mechanisms whereby the PvuII polymorphism LPL gene could act on plasma lipid levels remain unclear. Although preceding studies suggested mainly no or only very weak associations of this polymorphism with lipid levels in hyperlipidemic persons, the P- allele has been found to be associated with variation in levels of TGs, HDL-C, LDL-C, and T-C in normolipidemic populations(15, 19, 20). The P+ allele of the PvuII RFLP has been found, although inconsistently, to be associated with higher TG(15) and lower HDL-C levels(19). In another study in adult Caucasians, the mean plasma fasting TG levels for normal individuals with genotypes P+P+ were found to be 30% higher than those with genotypes P-P-(15). Yet another study on healthy adult Mediterranean migrants in Australia had revealed no significant association of the PvuII polymorphism with any lipid, although there was some evidence of an independent effect of this polymorphism on both LDL-C and T-C levels in normolipidemic individuals(20).
In this investigation on the cord plasma of newborns, we have found that the adjusted LDL-C levels of the male newborns of both ethnic groups were significantly influenced by the PvuII polymorphism genotype which contributed 8.1-8.5% to the variance. In ethnic Indian male newborns, the LDL-C and apoA-I levels were highest in the P- homozygotes, intermediate in the heterozygotes, and lowest in the P+ homozygotes, suggesting a gene-dosage effect. Mattu et al.(19) also reported higher serum LDL-C levels in healthy adult male carriers of the P- allele but not in female carriers(19). Additionally, in ethnic Indian male newborns, the adjusted plasma levels of T-C and apoA-I were also significantly higher in the P-P- genotype. Coupled with the genotype-related variation of apoA-I, the corresponding trend of HDL-C levels is higher in the Indian male newborn P- homozygotes (p = 0.063). In all the four gender-ethnic groups, TG levels tended to be higher in the P+ homozygotes, but reaching significance only in female Indian newborns, which is in conformance with the observations in Caucasian populations(15). The absence of gene dosage effect on LDL-C in male Chinese could also be a spurious finding due to the low numbers of the rare homozygotes.
The findings in these normal newborns again suggest the influence of the P- allele in normolipidemic states even before any environment factors could operate. Genotypic differences produced by variation in the LPL gene may need augmentation by interaction with postnatal environmental factors such as high fat, high cholesterol diet, and this could explain the relative low level of influence of the LPL gene PvuII polymorphism on lipid variation at birth when interaction with dietary fat has not yet taken place. In newborns, the VLDL-cholesterol levels at birth could also account for a weaker association of LPL gene polymorphism with LDL-C levels.
As these newborns have yet to be exposed to environmental influence, the presence of these significant PvuII genotype-specific influences on the lipid profile in cord plasma suggests an early expression of the LPL gene locus on plasma lipid traits. Further, the influence of the LPL gene in the present study is predominantly limited to male subjects, and the gender-specific influence suggests some hormonal modulation of genotype specific effects on plasma lipids.
In view of the present findings and those of other studies in normolipidemic adult populations, it appears that the LPL gene plays an important role in lipid metabolism. As the PvuII polymorphism is in the noncoding region of the gene, it cannot have a direct effect on plasma lipids. Rather, the effects on normal population levels of plasma lipids may be mediated by another linked functional mutation elsewhere in the gene. The Hin dIII and the PvuII sites have been shown to be in strong linkage disequilibrium in Caucasian as well as in Japanese populations(15, 21, 27). It has also been proposed that the PvuII site predisposes individuals to raised plasma TGs but requires coinheritance of a variant such as the H- allele for full expression. Recently, the PvuII polymorphism has been shown to be associated with the severity of CAD in Australian white subjects(27). Ser447 terminal mutation in exon 9 is in strong linkage with P- (PvuII site, intron 6)(28, 29). However, the truncated protein has normal lipolytic activity, although its kinetic properties and substrate specificity could be different from the normal protein(28, 30). This or some other functional mutation in the coding region of the LPL gene may be linked to the PvuII site of intron 6. Further studies are needed to elucidate the underlying functional mechanisms of the LPL gene and to identify its functional variants.
Abbreviations
- T-C:
-
total cholesterol
- HDL-C:
-
HDL cholesterol
- LDL-C:
-
LDL cholesterol
- TG:
-
triglyceride
- Lp(a):
-
lipoprotein(a)
- LPL:
-
lipoprotein lipase
- CAD:
-
coronary artery disease
- RFLP:
-
restriction fragment length polymorphism
References
Danaraj TJ, Acker MS, Danaraj W, Wong HO, Tan BY 1959 Ethnic group differences in coronary heart disease in Singapore-an analysis of necropsy records. Am Heart J 58: 516–526.
Hughes K, Lun KC, Yeo PPB 1990 Cardiovascular diseases in Chinese, Malays and Indians in Singapore. I. Differences in mortality. J Epidemiol Community Health 44: 24–28.
Miller NE 1987 Associations of high-density lipoprotein subclasses and apolipoproteins with ischemic heart disease and coronary atherosclerosis. Am Heart J 113: 589–597.
Castelli WP 1986 The triglyceride issue: A view from Framingham. Am Heart J 112: 432–437.
Austin MA 1989 Plasma triglyceride as a risk factor for coronary heart disease: The epidemiologic evidence and beyond. Am J Epidemiol 129: 249–259.
Rose G, Tunstall-Pedoe HD, Heller RF 1983 UK Heart Disease Prevention Project-incidence and mortality results. Lancet 1: 1062–1066.
Rosengren A, Wilhelmsen L, Eriksson E, Risberg B, Wedel H 1990 Lipoprotein(a) and coronary heart disease: a prospective case-control study in a general population sample of middle aged men. BMJ 301: 1248–1251.
Utermann G 1989 The mysteries of lipoprotein(a). Science 246: 904–910.
Sing CF, Orr JD 1978 Analysis of genetic and environmental sources of variation in serum cholesterol in Tecumseh, Michigan. IV. Separation of polygene from common environmental effects. Am J Hum Genet 30: 491–504.
Moll PP, Powsner R, Sing CF 1979 Analysis of genetic and environmental sources of variation in serum cholesterol in Tecumseh, Michigan. V. Variance component estimated from pedigrees. Ann Hum Genet 42: 343–354.
Namboodiri KK, Kaplan EB, Heach I, Elston RC, Green PP, Rao DC, Laskarzewski P, Glueck CJ, Rifkind BM 1985 The Collaborative Lipid Research Clinics Family Study: biological and cultural determinants of familial resemblance for plasma lipids and lipoproteins. Genet Epidemiol 2: 227–254.
Low PS, Saha N, Tay JSH, Hong S 1996 Ethnic variation of cord plasma apolipoprotein levels in relation to coronary risk level: a study in three ethnic groups of Singapore. Acta Paediatr 85: 1476–1482.
Low PS, Heng CK, Saha N, Tay JSH 1996 Racial variation of cord plasma lipoprotein(a) levels in relation to coronary risk level: a study in three ethnic groups of Singapore. Pediatr Res 40: 718–722.
Sparkes RS, Zollman S, Klisak I, Kirchgessner TG, Komaromy MC, Mohandas T, Schotz MC, Lusis AJ 1987 Human genes involved in lipolysis of plasma lipoproteins: mapping of loci for lipoprotein lipase to 8p22 and hepatic lipase to 15q21. Genomic 1: 138–144.
Chamberlain JC, Thorn JA, Oka K, Galton DJ, Stocks J 1989 DNA polymorphisms at the lipoprotein lipase gene: associations in normal and hypertriglyceridaemic subjects. Atherosclerosis 79: 85–91.
Galton DJ, Mattu RK, Cavanna J 1994 Polymorphisms of the lipoprotein lipase gene and premature atherosclerosis. J Intern Med 236 ( suppl 736): 63–68.
Thorn JA, Chamberlain JC, Oka K, Chan L, Stocks J, Galton DJ 1990 Lipoprotein and hepatic lipase gene variants in coronary atherosclerosis. Atherosclerosis 85: 55–60.
Ahn YI, Kamboh MI, Hamman RF, Cole SA, Ferrell RE 1993 Two DNA polymorphisms in the lipoprotein lipase gene and their associations with factors related to cardiovascular diseases. J Lipid Res 34: 421–428.
Mattu RK, Needham EWA, Morgan R, Rees A, Hackshaw AK, Stocks J, Elwood PC, Galton DJ 1994 DNA variants at the LPL gene locus associate with angiographically defined severity of atherosclerosis and serum lipoprotein levels in a Welsh population. Arterioscler Thromb 14: 1090–1097.
Mitchell RJ, Earl L, Bray P, Fripp YJ, Williams J 1994 DNA polymorphisms at the lipoprotein lipase gene and their association with quantitative variation in plasma high-density lipoprotein and triacylglycerides. Hum Biol 66: 383–397.
Gerdes C, Gerdes LU, Hansen PS, Faergeman O 1995 Polymorphisms in the lipoprotein lipase gene and their associations with plasma lipid concentrations in 40-year-old Danish men. Circulation 92: 1765–1769.
Jemaa R, Fumeron F, Poirier O, Lecerf L, Evans A, Arveiler D, Luc G, Cambou J-P, Bard J-M, Fruchart J-C, Apfelbaum M, Cambien F, Tiret L 1995 Lipoprotein lipase gene polymorphisms: associations with myocardial infarction and lipoprotein levels, the ECTIM study. J Lipid Res 36: 2141–2146.
Parzer S Mannhalter C 1991 A rapid method for the isolation of genomic DNA from citrated whole blood. Biochem J 273: 229–231.
Gomez KA, Gomez AA 1984 Statistical Procedures for Agricultural Research, 2nd Ed. Wiley-Interscience, New York, pp 299–304.
Sokal RR, Rohlf JF 1969 Biometry. WH Freeman, San Francisco, p 384
Bartlett MS 1947 The use of transformations. Biometrics 3: 39–52.
Wang XL, McCredie RM, Wilcken DEL 1996 Common DNA polymorphisms at the lipoprotein lipase gene. Association with severity of coronary artery disease and diabetes. Circulation 63: 1339–1345.
Peacock RE, Hamsten A, Nilsson-Ehle P, Humphries SE 1992 Association between lipoprotein lipase gene polymorphisms and plasma correlations of lipids, lipoproteins and lipase activities in young myocardial infarction survivors and age-matched healthy individuals from Sweden. Atherosclerosis 97: 171–185.
Zhang Q, Cavanna J, Winkelman BR, Shine B, Gross W, Marz W, Galton DJ 1995 Common genetic variants of lipoprotein lipase that relate to lipid transport in patients with premature coronary artery disease. Clin Genet 48: 293–298.
Faustinella F, Chang A, Van Biervliet JP, Rosseneu M, Vinaimont N, Smith LC, Chen SH, Chan L 1991 Catalytic triad residue mutation(Asp156 → Gly) causing familial lipoprotein lipase deficiency. Co-inheritance with a nonsense mutation (Ser447 → stop codon) in a Turkish family. J Biol Chem 266: 14418–14424.
Acknowledgements
The authors gratefully acknowledge the technical assistance of Jumiah Bte Basair, Sally Hong, and the laboratory staff of SATA.
Author information
Authors and Affiliations
Additional information
Supported generously by the National University of Singapore and the Singapore Anti-Tuberculosis Association (SATA).
Rights and permissions
About this article
Cite this article
Low, PS., Saha, N., Tay, J. et al. Influence of PvuII (Intron 6) Polymorphism of the Lipoprotein Lipase Gene on Cord Plasma Lipid and Apoliporotein Levels in Indian and Chinese Newborns of Singapore. Pediatr Res 43, 240–244 (1998). https://doi.org/10.1203/00006450-199802000-00014
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1203/00006450-199802000-00014