Abstract
Circulating tumor DNA (ctDNA) has emerged as a tumor-specific biomarker for the early detection of various cancers. To date, several techniques have been devised to enrich the extremely small amounts of ctDNA present in plasma, but they are still insufficient for cancer diagnosis, especially at the early stage. Here, we developed a novel method, CUT (CRISPR-mediated, Ultrasensitive detection of Target DNA)-PCR, which uses CRISPR endonucleases to enrich and detect the extremely small amounts of tumor DNA fragments among the much more abundant wild-type DNA fragments by specifically eliminating the wild-type sequences. We computed that by using various orthologonal CRISPR endonucleases such as SpCas9 and FnCpf1, the CUT-PCR method would be applicable to 80% of known cancer-linked substitution mutations registered in the COSMIC database. We further verified that CUT-PCR together with targeted deep sequencing enables detection of a broad range of oncogenes with high sensitivity (<0.01%) and accuracy, which is superior to conventional targeted deep sequencing. In the end, we successfully applied CUT-PCR to detect sequences with oncogenic mutations in the ctDNA of colorectal cancer patients’ blood, suggesting that our technique could be adopted for diagnosing various types of cancer at early stages.
Similar content being viewed by others
Introduction
On the basis of advances in sequencing technology, it is well accepted that cancer is generally driven by oncogenic mutations.1, 2 Several studies have provided evidence that tumorigenesis strongly correlates with the prevalence of somatic mutations in certain types of cancer.3, 4 The COSMIC (Catalogue of Somatic Mutations in Cancer) database includes hundreds of thousands of human cancer-associated somatic mutations that are classified by tumor type and disease.5 Recently, circulating tumor DNA (ctDNA), which is released into the bloodstream from tumor cells as cell-free DNA (cfDNA) fragments, has been proposed as a tumor-specific biomarker candidate.6, 7, 8, 9, 10, 11, 12 Thus, diagnosing tumors at early stages might be possible by simply detecting tumor-specific somatic mutations in the ctDNA from a patient’s blood.10, 13 However, cfDNAs in the blood plasma generally contain extremely small amounts of tumor DNA, which are reportedly dependent on the tumor burden or cancer stage.13, 14 Therefore, especially in the early stages of cancer, a highly sensitive and specific method would be required to diagnose a tumor by detecting ctDNA.15, 16, 17, 18, 19, 20, 21, 22
The clustered regularly interspaced short palindromic repeats (CRISPR) and CRISPR-associated (Cas) protein system, an adaptive immune response in prokaryotes, are well known for their specific DNA target recognition and cleavage.23, 24 The versatility of the CRISPR system has allowed us and other groups to broadly utilize it, not only for genome editing in various organisms,25, 26, 27, 28, 29, 30 but also to cleave DNA in vitro.31 CRISPR endonucleases, such as type II Cas9 or type V Cpf1, specifically induce double-stranded breaks in target DNA by recognizing a protospacer-adjacent motif (PAM) downstream or upstream, respectively, of target DNA sequences corresponding to that of a guide RNA (gRNA).32, 33, 34, 35, 36, 37
To use ctDNA for cancer diagnosis, it is necessary to detect, in a highly accurate manner, the small amounts of ctDNAs with missense mutations among the relatively very large amounts of wild-type cfDNAs. By adapting the accurate and specific cleavage ability of CRISPR endonucleases in vitro, it is possible to enrich tumor-specific versus wild-type alleles by specifically cleaving the wild-type DNAs. Here, we devised a new method employing the CRISPR system, termed CUT (CRISPR-mediated, Ultrasensitive detection of Target DNA)-PCR (polymerase chain reaction), that efficiently enriches oncogenic mutant DNAs by eliminating wild-type DNAs before PCR amplification. We note that by altering gRNAs corresponding to various wild-type DNAs, one can easily and precisely reduce different background DNA signals in an unbiased manner. After the wild-type DNAs are cleaved by CRISPR endonucleases, the mutant target regions are amplified by PCR and then exquisitely identified by targeted deep sequencing using next-generation sequencing facilities. Because PCR amplification is performed after background wild-type sequences are reduced, the CUT-PCR process minimizes polymerase-generated errors and maximizes target cleavage specificity.
Results and discussion
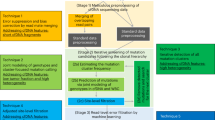
By using the specific recognition property of CRISPR endonucleases for DNA PAM sequences, it is possible to cleave PAM-containing wild-type DNA sequences selectively, reducing background DNA signals and enriching cancer-specific mutant DNA signals (Figure 1a). The representative type II CRISPR endonuclease Cas9, derived from Streptococcus pyogenes (SpCas9), and type V Cpf1, from Francisella novicida (FnCpf1), respectively, recognize the PAM sequences 5′-NGG-3′, located downstream of the target DNA,33 and 5′-TTN-3′, located upstream of the target DNA.37 Therefore, if oncogenic mutant sequences are generated by single-base substitutions, such as these (NGG>NGH or NGG>NHG, H is A, C or T; TTN>TVN or TTN>VTN, V is A, C or G, in wild-type DNA sequences) wild-type DNAs can be selectively and precisely eliminated by SpCas9 or FnCpf1, respectively.23, 37 After wild-type DNA cleavage, pooled DNAs can be identified directly with PCR amplification followed by targeted deep sequencing.38
CUT-PCR method and its applicable targets in the COSMIC database. (a) Schematic of the CUT-PCR enrichment process. To cleave wild-type DNA specifically, single-guide RNAs were designed to target PAM sites that are destroyed by oncogenic mutations. Such mutant alleles are not recognized by the CRISPR endonucleases and largely avoid cleavage. After cleavage of wild-type DNA, the DNA in the pooled solution was amplified with PCR. (b) The classification of human cancer-associated somatic mutations registered in the COSMIC database. (c) The ratio of CUT-PCR applicable targets among the mutations of indel (blue bar) and substitution (red bar) registered in the COSMIC database.
We inspected all possible target sites registered in the COSMIC database in silico to determine whether various orthologonal Cas9 or Cpf1 proteins would be applicable for the specific destruction of the corresponding wild-type sequences. As shown in Figure 1b, among 325 856 mutations registered in the COSMIC database (version 77), 90.4% are single- or multiple-nucleotide substitutions. Insertions and deletions account for 2.9% and 6.6% of the entries, respectively, and unknown patterns represent 0.2%. For each indel, we searched for an adjacent PAM sequence to design sgRNA because CRISPR endonucleases barely cleave mutant DNA that contains indels in the gRNA target region.31, 37 In the case of the substitutions, however, CRISPR endonucleases can typically cleave mutant DNAs as well as wild-type DNAs in vitro because of their ability to recognize sites that vary by one or a few nucleotides from the gRNA sequence (off-target effects).39 However, PAM recognition by CRISPR endonucleases is much stricter; they rarely cleave target DNA that lacks a PAM sequence even if the target DNA is exactly complementary to the gRNA. Thus, we determined whether each substitution mutation might have destroyed a PAM sequence in the corresponding wild-type DNA, which would mean that the wild-type DNA would be cleaved much more readily than the oncogenic DNA that lacked a proper PAM. Further analyses resultantly showed that 98.9% of the indels (Supplementary Table 1) and 80.5% of the substitutions (Supplementary Table 2) represent DNA targets that can be selectively cleaved by the various orthologonal Cas9 or Cpf1 proteins reported to date, suggesting that our CUT-PCR method would be useful for detecting about 80% of the oncogenic mutations in the COSMIC database (Figure 1c).
To validate that CRISPR endonucleases could selectively cleave target DNA as specified in the CUT-PCR protocol, we performed in vitro cleavage assays with T-vector cloned sequences containing various missense mutations (Supplementary Table 3). We expected that CRISPR endonucleases with gRNAs specific to the wild-type sequence would specifically deplete the wild-type DNA. We chose five recurrent cancer-associated mutations in the KRAS gene (KRAS c.35G>A, c.35G>T, c.34G>T, c.35G>C and c.34G>C) for testing type II SpCas9 and one in the GNAQ gene (GNAQ c.626A>T) for testing type V FnCpf1. Both oncogenes are well known for their tumorigenicity.40, 41
For testing the SpCas9, we constructed one plasmid containing the wild-type KRAS sequence and five plasmids containing patient-mimic sequences in which the PAM sequence was changed, as shown in Figure 2a. We validated that each plasmid can be linearized with NcoI restriction enzyme and then treated each linearized plasmid in vitro with the SpCas9 complex containing a single-guide RNA (sgRNA) specific to the wild-type sequence. As a result, it showed that the SpCas9 complex selectively cleaved wild-type DNA resulting in shorter DNA fragments but generally did not cleave the other mutant sequences that lacked a functional PAM (Figure 2b). We note that one mutant plasmid containing KRAS (c.35G>A), which has a 5′-NGA-3′ PAM sequence, is marginally targeted by SpCas9 specific for the wild-type sequence, a result already reported in a previous study.42 For testing the FnCpf1 nuclease, we constructed two different plasmids, one containing the wild-type GNAQ sequence and the other the patient-mimic mutated sequence, as shown in Figure 2c. Both plasmids can be also linearized with NcoI. In this case, we designed one CRISPR RNA specific to the wild-type sequence and treated each plasmid in vitro with the FnCpf1 complex. Results showed that the FnCpf1 complex selectively cleaved wild-type plasmid DNA but not the mutant sequence that lacked a functional PAM (Figure 2d), suggesting that FnCpf1 would also be useful for mutant sequence enrichment in the CUT-PCR process.
In vitro cleavage assay with plasmids containing sequences with PAM mutations. (a) Top: schematic of the plasmids containing sequences with wild-type and oncogenic mutations. PCR amplicons of relevant wild-type proto-oncogene cDNAs were subcloned into the commercial T-blunt cloning vector (T-Blunt PCR cloning Kit, SolGent, Seoul, South Korea) using the manufacturer’s protocol. Single-base-pair-substituted mutations (KRAS: c.35G>A, c.35G>T, c.34G>T, c.35G>C, c.34G>C, GNAQ: c.626A>T) were constructed with site-directed mutagenesis using appropriate primer sets (Supplementary Table 3). Plasmids can be linearized by the restriction enzyme NcoI. Bottom: the sequences of wild-type and recurrent KRAS mutations in the COSMIC database. The PAM sequence (5′-TGG-3′) for SpCas9 is underlined in blue. Missense mutations are shown in red. (b) In vitro cleavage assay using SpCas9 with linearized plasmids containing wild-type and mutant KRAS sequences. Target plasmids were first linearized with the restriction enzyme NcoI (New England Biolabs, Ipswich, MA, USA) for 37 °C for 1 h (10 μl reaction in NEB buffer 3.1). The linearized product was further cleaved by treatment with a wild-type sequence-specific CRISPR nuclease (Cas9 100 ng, sgRNA 70 ng, 10 μl reaction in NEB buffer 3.1 at 37 °C, 1 h). CF, cleaved fragment; LF, linearized fragment. (c) The sequences of wild-type and recurrent GNAQ mutation in the COSMIC database. The PAM sequence (5′-TTG-3′) for FnCpf1 is underlined in blue. (d) In vitro cleavage assay using FnCpf1 with linearized plasmids containing wild-type and mutant GNAQ sequences.
To investigate whether CUT-PCR could be used to detect rare oncogene-specific mutations, we next prepared mixtures in which the plasmid containing a mutant sequence was serially diluted with the plasmid containing the wild-type sequence. We then treated each plasmid mixture in vitro with a CRISPR endonuclease and a gRNA specific to the wild-type sequence, after which we amplified the target region using PCR (Figure 3a). We expected that plasmids containing the wild-type sequence would be selectively cleaved by CRISPR complexes, resulting in relatively less amplification, whereas plasmids containing the mutant sequences would not be cleaved and would therefore be amplified more. When a mixture of plasmids containing either wild-type or mutant KRAS (c.35G>T) sequences was treated with wild-type-specific SpCas9 complexes in vitro, the total amount of PCR amplicons gradually decreased as the abundance of the KRAS mutant plasmid decreased (Figure 3b, lanes 1–5), in contrast to untreated samples (Figure 3b, lanes 6–10). This result indicates that most wild-type sequences were eliminated by SpCas9, suggesting that mutant sequences were enriched relative to the wild-type sequences in the mixture of PCR amplicons. As a quantitative control for PCR amplification in each reaction, we added pairs of internal control primers to each mixture and determined the PCR outcomes relative to these control PCR products.
CUT-PCR-based enrichment of plasmid-borne sequences containing missense mutation. (a) Schematic of two sets of primers for target and internal control region. After treatment of wild-type-specific CRISPR endonucleases, each plasmid mixture containing sequences with wild-type and oncogenic mutation was amplified using PCR. (b) CUT-PCR experiment for various ratios of plasmid mixtures containing either wild-type or mutant KRAS (c.35G>T) sequence. DNA plasmids containing wild-type and mutant sequences were mixed in various ratios and subjected to CRISPR cleavage in vitro. The plasmid mixture was treated with a wild-type-specific CRISPR nuclease (11 ng total plasmid DNA, 100 ng Cas9, 70 ng sgRNA, 10 μl reaction in NEB buffer 3.1 at 37 °C for 1 h) to cleave wild-type DNA. After proteinase (Qiagen, Venlo, Netherlands) treatment, samples were purified using a PCR cleanup kit (DOCTOR PROTEIN, Seoul, South Korea, MD008) and each sample was amplified by PCR using targeted primer sets. To quantify target-specific cleavage, the fold increase in the target sequence was compared with that of an internal control product, which was amplified with internal control primers (Supplementary Table 3). The amount of the amplified KRAS target region relative to the internal control PCR product in each lane was calculated. (c) For the KRAS (c.35G>T) mutation, targeted deep sequencing after CUT-PCR was treated (red bars) or not (gray bars) were conducted for the plasmid mixtures in which mutant plasmids were originally mixed with wild-type plasmids at a ratio of from 100% to 0.01%. CUT-PCR-enriched plasmids were further amplified with adaptor primers (Supplementary Table 3) using Phusion polymerase (New England Biolabs). The resulting PCR amplicons were subjected to paired-end sequencing with the Illumina MiSeq system. Paired-end reads were then analyzed by comparing wild-type and mutant sequences using Cas-Analyzer (www.rgenome.net/cas-analyzer). For the mixture at a ratio of 0.01%, frequencies of mutant DNA fragments (d) were measured and the values of fold increase (e) were calculated after multiple rounds of CUT-PCR treatments. (f) For the four recurrent KRAS mutations (c.35G>A, c.34G>T, c.35G>C and c.34G>C), the frequencies of wild-type (blue bars) and mutant (red bars) fragments were measured using deep sequencing. In all cases, KRAS mutant plasmids were originally mixed with wild-type plasmids at a ratio of 0.1%. (g) Fold increase after CUT-PCR in each KRAS mutant DNA frequency was calculated from the data of f. (h) DNA frequencies of wild-type and GNAQ mutant (c.626A>T) fragments measured using deep sequencing after FnCpf1 mediated CUT-PCR. GNAQ mutant plasmids were originally mixed with wild-type plasmids at a ratio of 0.1%. (i) Fold increase in the mutant DNA fragment frequency for the recurrent GNAQ mutation calculated from h. Error bars mean s.e.m.; n=2 for c and 3 for d, f and h; *P<0.05; **P<0.01.
To examine the sensitivity of CUT-PCR method, we conducted targeted deep sequencing for each mixture with various ratios of mutant plasmids using Illumina (San Diego, CA, USA) MiSeq. Every sample was read at a sequencing depth of at least 10 000 ×. We sought to compare CUT-PCR-based deep sequencing against the conventional deep-sequencing data. For KRAS (c.35G>T) sample, mutant plasmids were originally mixed with wild-type plasmids at a ratio of 100 to 0.01%. Then, DNA target sites were amplified with PCR after being CUT-PCR-treated or not. As a result, conventional deep sequencing for missense mutations was limited to 0.1% as in a previous study.39 However, mutant DNA fragments were entirely enriched after SpCas9-based CUT-PCR treatments (Figure 3c). For the mixture that mutant plasmids were mixed at a ratio of 0.01%, CUT-PCR-treated samples showed a sixfold increase in the mutated DNA fragment frequency, relative to untreated samples. For the fold enrichment calculation, we used the value from the CUT-PCR-untreated sample as the background frequency. In addition, the mutated DNA fragment frequencies were more increased (Figure 3d) after multiple rounds of CUT-PCR, resulting in a greater fold increase (Figure 3e). These results indicate that the sensitivity of CUT-PCR-based deep sequencing is more than 0.01%. For the additional comparison with quantitative real-time PCR, it is hard to detect missense mutations among the mixtures at a ratio of 1% of mutant plasmids (Supplementary Figure 1).
We repeated the CUT-PCR procedure with other KRAS mutant plasmid mixtures as described above (Supplementary Figure 2). For the mixture that mutant plasmids were originally mixed at a ratio of 0.1%, mutant DNA fragments were significantly enriched by SpCas9-based CUT-PCR relative to untreated samples (Figure 3f). Moreover, the values of a fold increase in the mutated DNA fragment frequency were from 29.6 to 76.3 (Figure 3g). We noted that, in the case of the KRAS (c.35G>A) mutant sequence, some of the plasmids were cleaved by wild-type-specific SpCas9 as shown in Figure 2b, but the relative amount of mutant DNA fragments was strongly increased after CUT-PCR, which might indicate that wild-type DNA fragments were preferentially eliminated.
We further tested whether CUT-PCR enrich mutant DNA for different target sites and different CRISPR types. For the GNAQ (c.626A>T) mutant and wild-type plasmid mixture, we verified that FnCpf1 would cleave the wild-type DNA fragment selectively and sufficiently (Supplementary Figure 3). In line with SpCas9-based CUT-PCR, we determined that a mixture treated with FnCpf1-based CUT-PCR showed a 27-fold increase in the mutant fragment frequency as compared with untreated samples when GNAQ mutant and wild-type plasmids were originally mixed at a ratio of 0.1% (Figures 3h and i). To test whether CUT-PCR could be used more generally, we applied the process to other oncogenes. We used FnCpf1-based CUT-PCR with CTNNB1 containing a substitution mutation (Supplementary Figure 4) and SpCas9-based CUT-PCR with EGFR containing a deletion (Supplementary Figure 5). We also found that one cycle of CUT-PCR efficiently enriched mutant DNA fragments as in the above results.
We ultimately applied the CUT-PCR technique to detect cell-free ctDNA extracted from the blood plasma of eight colorectal cancer patients at various stages of the disease. As a control, we used plasma from four healthy donors. KRAS mutations are frequently found in colon cancer. To enrich mutant KRAS ctDNAs, we prepared sgRNA for SpCas9 specific to the wild-type KRAS sequence as described above. As shown in Figure 3, KRAS sequences containing five different oncogenic mutations (KRAS c.35G>A, c.35G>T, c.34G>T, c.35G>C and c.34G>C) can be enriched using one common sgRNA because of PAM sequence substitution. Because total amounts of cfDNAs in plasma are low and mutant ctDNA fragments are present at very low abundance, especially at early stages of disease, we conducted multiple rounds of CUT-PCR. After each round of CUT-PCR, we measured mutant and wild-type KRAS allele frequencies (AFs) by targeted deep sequencing (Figure 4a and Supplementary Table 4). After the third round of CUT-PCR, we measured the enriched mutant AF and calculated its fold increase relative to the wild-type AF (Figure 4b).
CUT-PCR-based enrichment of sequences containing recurrent KRAS mutations in cfDNAs from colorectal cancer (CRC) patients and healthy donors. (a) The AFs of recurrent KRAS mutation candidates (c.35G>A, c.35G>T, c.34G>T, c.35G>C and c.34G>C) were analyzed from cfDNAs in plasma of eight CRC patients (pink boxes) and four healthy donors (blue boxes). Peripheral blood samples from patients were obtained from the Pusan National University Hospital (Busan, Korea). This study was reviewed and approved by the Institutional Review Board (IRB) of PNUH(H-1412-011-024) and UNIST(UNISTIRB-13-002-A). To get cell-free DNA from CRC patients and healthy volunteers, plasma was obtained from blood sample by using Ficoll-Paque PLUS (GE Healthcare, Chicago, IL, USA) and cell-free DNA was purified from 1 ml of the plasma with a QIAamp Circulating Nucleic Acid Kit (Qiagen) according to the manufacturer’s protocol. After the multiple rounds of CUT-PCR treatment using the wild-type KRAS-specific SpCas9 nucleases, each AF of recurrent KRAS mutation sequence was measured using targeted deep sequencing for all samples, respectively. (b) Fold increase in each KRAS mutant sequence calculated from a after the third round of CUT-PCR. The cutoff baseline for KRAS-mutated ctDNA observation was determined by averaging the mutant AFs of CRISPR-untreated sample in healthy controls. Error bars mean s.e.m.; n=3; **P<0.01, ***P<0.001.
For the (KRAS c.35G>A) mutation shown in the upper panel of Figures 4a and b, no significant increase in mutant AFs was observed after multiple rounds of CUT-PCR in the case of control samples (Figure 4a, blue region). However, we found that mutant AFs from patients 2, 3, 4, 5 and 7 increased considerably (Figure 4a, pink region) compared with the average value from the healthy controls, resulting in fold increases from 164 to 640 as shown in the upper panel in Figure 4b. We calculated the fold increase of mutant AFs in the same way for the other mutations and summarized the results in Supplementary Table 5. As a result, our CUT-PCR data from cfDNAs provide as much information as the pyrosequencing data from tissue samples even in the patients with stage I cancer.
In conclusion, the CUT-PCR method enriches and thus enables the sensitive and precise detection of extremely small amounts of circulating mutant DNA sequences derived from tumor cells via the removal of background signals through the specific cleavage of wild-type sequences by CRISPR endonucleases in vitro. We emphasize that cleaving target genomic DNA in vitro before PCR amplification increases the fidelity of mutant DNA enrichment by eliminating the chance of enriching false mutations, which may be generated by DNA polymerase during PCR amplification. Furthermore, if researchers engineer existing CRISPR endonucleases to have altered PAM specificities43 or discover new CRISPR endonucleases that recognize different PAM sequences, CUT-PCR-targetable sites would be further extended, which enlarges the utility of this method.
References
Park JY, Kricka LJ, Fortina P . Next-generation sequencing in the clinic. Nat Biotechnol 2013; 31: 990–992.
Kruglyak KM, Lin E, Ong FS . Next-generation sequencing in precision oncology: challenges and opportunities. Expert Rev Mol Diagn 2014; 14: 635–637.
Vogelstein B, Papadopoulos N, Velculescu VE, Zhou SB, Diaz LA, Kinzler KW . Cancer genome landscapes. Science 2013; 339: 1546–1558.
Wood LD, Parsons DW, Jones S, Lin J, Sjoblom T, Leary RJ et al. The genomic landscapes of human breast and colorectal cancers. Science 2007; 318: 1108–1113.
Forbes SA, Tang G, Bindal N, Bamford S, Dawson E, Cole C et al. COSMIC (the Catalogue of Somatic Mutations in Cancer): a resource to investigate acquired mutations in human cancer. Nucleic Acids Res 2010; 38: D652–D657.
De Mattos-Arruda L, Caldas C . Cell-free circulating tumour DNA as a liquid biopsy in breast cancer. Mol Oncol 2016; 10: 464–474.
Sorber L, Zwaenepoel K, Deschoolmeester V, Van Schil PEY, Van Meerbeeck J, Lardon F et al. Circulating cell-free nucleic acids and platelets as a liquid biopsy in the provision of personalized therapy for lung cancer patients. Lung Cancer 2017; 107: 100–107.
Heitzer E, Ulz P, Geigl JB . Circulating tumor DNA as a liquid biopsy for cancer. Clin Chem 2015; 61: 112–123.
Cheng FF, Su L, Qian C . Circulating tumor DNA: a promising biomarker in the liquid biopsy of cancer. Oncotarget 2016; 7: 48832–48841.
Schwarzenbach H, Hoon DSB, Pantel K . Cell-free nucleic acids as biomarkers in cancer patients. Nat Rev Cancer 2011; 11: 426–437.
Alix-Panabieres C, Schwarzenbach H, Pantel K . Circulating tumor cells and circulating tumor DNA. Annu Rev Med 2012; 63: 199–215.
Freidin MB, Freydina DV, Leung M, Fernandez AM, Nicholson AG, Lim E . Circulating tumor DNA outperforms circulating tumor cells for KRAS mutation detection in thoracic malignancies. Clin Chem 2015; 61: 1299–1304.
Bettegowda C, Sausen M, Leary RJ, Kinde I, Wang Y, Agrawal N et al. Detection of circulating tumor DNA in early- and late-stage human malignancies. Sci Transl Med 2014; 6: 224ra24.
Dawson SJ, Tsui DW, Murtaza M, Biggs H, Rueda OM, Chin SF et al. Analysis of circulating tumor DNA to monitor metastatic breast cancer. N Engl J Med 2013; 368: 1199–1209.
Li M, Diehl F, Dressman D, Vogelstein B, Kinzler KW . BEAMing up for detection and quantification of rare sequence variants. Nat Methods 2006; 3: 95–97.
Newman AM, Bratman SV, To J, Wynne JF, Eclov NC, Modlin LA et al. An ultrasensitive method for quantitating circulating tumor DNA with broad patient coverage. Nat Med 2014; 20: 548–554.
Forshew T, Murtaza M, Parkinson C, Gale D, Tsui DW, Kaper F et al. Noninvasive identification and monitoring of cancer mutations by targeted deep sequencing of plasma DNA. Sci Transl Med 2012; 4: 136ra168.
Chang-Hao Tsao S, Weiss J, Hudson C, Christophi C, Cebon J, Behren A et al. Monitoring response to therapy in melanoma by quantifying circulating tumour DNA with droplet digital PCR for BRAF and NRAS mutations. Sci Rep 2015; 5: 11198.
Kinde I, Wu J, Papadopoulos N, Kinzler KW, Vogelstein B . Detection and quantification of rare mutations with massively parallel sequencing. Proc Natl Acad Sci USA 2011; 108: 9530–9535.
Narayan A, Carriero NJ, Gettinger SN, Kluytenaar J, Kozak KR, Yock TI et al. Ultrasensitive measurement of hotspot mutations in tumor DNA in blood using error-suppressed multiplexed deep sequencing. Cancer Res 2012; 72: 3492–3498.
Song C, Liu Y, Fontana R, Makrigiorgos A, Mamon H, Kulke MH et al. Elimination of unaltered DNA in mixed clinical samples via nuclease-assisted minor-allele enrichment. Nucleic Acids Res 2016; 44: e146.
Chan KC, Jiang P, Zheng YW, Liao GJ, Sun H, Wong J et al. Cancer genome scanning in plasma: detection of tumor-associated copy number aberrations, single-nucleotide variants, and tumoral heterogeneity by massively parallel sequencing. Clin Chem 2013; 59: 211–224.
Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, Charpentier E . A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 2012; 337: 816–821.
Kim H, Kim JS . A guide to genome engineering with programmable nucleases. Nat Rev Genet 2014; 15: 321–334.
Hsu PD, Lander ES, Zhang F . Development and applications of CRISPR-Cas9 for genome engineering. Cell 2014; 157: 1262–1278.
Wu Y, Liang D, Wang Y, Bai M, Tang W, Bao S et al. Correction of a genetic disease in mouse via use of CRISPR-Cas9. Cell Stem Cell 2013; 13: 659–662.
Friedland AE, Tzur YB, Esvelt KM, Colaiacovo MP, Church GM, Calarco JA . Heritable genome editing in C. elegans via a CRISPR-Cas9 system. Nat Methods 2013; 10: 741–743.
Baek K, Kim DH, Jeong J, Sim SJ, Melis A, Kim JS et al. DNA-free two-gene knockout in Chlamydomonas reinhardtii via CRISPR-Cas9 ribonucleoproteins. Sci Rep 2016; 6: 30620.
Cho SW, Kim S, Kim JM, Kim JS . Targeted genome engineering in human cells with the Cas9 RNA-guided endonuclease. Nat Biotechnol 2013; 31: 230–232.
Chang N, Sun C, Gao L, Zhu D, Xu X, Zhu X et al. Genome editing with RNA-guided Cas9 nuclease in zebrafish embryos. Cell Res 2013; 23: 465–472.
Kim JM, Kim D, Kim S, Kim JS . Genotyping with CRISPR-Cas-derived RNA-guided endonucleases. Nat Commun 2014; 5: 3157.
Nishimasu H, Ran FA, Hsu PD, Konermann S, Shehata SI, Dohmae N et al. Crystal structure of Cas9 in complex with guide RNA and target DNA. Cell 2014; 156: 935–949.
Anders C, Niewoehner O, Duerst A, Jinek M . Structural basis of PAM-dependent target DNA recognition by the Cas9 endonuclease. Nature 2014; 513: 569–573.
Yamano T, Nishimasu H, Zetsche B, Hirano H, Slaymaker IM, Li Y et al. Crystal structure of Cpf1 in complex with guide RNA and target DNA. Cell 2016; 165: 949–962.
Lee SH, Bae S . Structural and dynamic views of the CRISPR-Cas system at the single-molecule level. BMB Rep 2016; 49: 201–207.
Kim D, Kim J, Hur JK, Been KW, Yoon SH, Kim JS . Genome-wide analysis reveals specificities of Cpf1 endonucleases in human cells. Nat Biotechnol 2016; 34: 863–868.
Zetsche B, Gootenberg JS, Abudayyeh OO, Slaymaker IM, Makarova KS, Essletzbichler P et al. Cpf1 is a single RNA-guided endonuclease of a class 2 CRISPR-Cas system. Cell 2015; 163: 759–771.
Park J, Lim K, Kim JS, Bae S . Cas-analyzer: an online tool for assessing genome editing results using NGS data. Bioinformatics 2017; 33: 286–288.
Cho SW, Kim S, Kim Y, Kweon J, Kim HS, Bae S et al. Analysis of off-target effects of CRISPR/Cas-derived RNA-guided endonucleases and nickases. Genome Res 2014; 24: 132–141.
Park JT, Johnson N, Liu S, Levesque M, Wang YJ, Ho H et al. Differential in vivo tumorigenicity of diverse KRAS mutations in vertebrate pancreas: a comprehensive survey. Oncogene 2015; 34: 2801–2806.
Xu X, Wei WB, Li B, Gao F, Zhang Z, Jonas JB . Oncogenic GNAQ and GNA11 mutations in uveal melanoma in Chinese. PLoS One 2014; 9: e109699.
Zhang Y, Ge X, Yang F, Zhang L, Zheng J, Tan X et al. Comparison of non-canonical PAMs for CRISPR/Cas9-mediated DNA cleavage in human cells. Sci Rep 2014; 4: 5405.
Kleinstiver BP, Prew MS, Tsai SQ, Topkar VV, Nguyen NT, Zheng Z et al. Engineered CRISPR-Cas9 nucleases with altered PAM specificities. Nature 2015; 523: 481–485.
Acknowledgements
We express our gratitude to all of the patients and healthy volunteers who contributed blood samples for this study. The biological specimens for this study were provided by the Pusan National University hospital, a member of the National Biobank of Korea, which is supported by the Ministry of Health and Welfare. All samples were obtained with informed consent according to the institutional review board-approved protocols. The deep-sequencing data are available at the NCBI Sequence Read Archive (http://www.ncbi.nlm.nih.gov/sra) under accession number SRX1970116. This work was supported by a grant from the Korea Healthcare technology R&D Project, Ministry for Health and Welfare Affairs (HI16C1012) to SB, (HI12C1845) to JP, DP and YC and Institute for Basic Science (IBS-R021-D1) to J-SK.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
SHL, SK and J-SK are co-inventors on a patent application covering the CUT-PCR method described in this manuscript. The remaining authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Oncogene website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Lee, S., Yu, J., Hwang, GH. et al. CUT-PCR: CRISPR-mediated, ultrasensitive detection of target DNA using PCR. Oncogene 36, 6823–6829 (2017). https://doi.org/10.1038/onc.2017.281
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2017.281
This article is cited by
-
Assessing and advancing the safety of CRISPR-Cas tools: from DNA to RNA editing
Nature Communications (2023)
-
The application of CRISPR-Cas in disease diagnosis and treatment
Science China Chemistry (2023)
-
CRISPR/Cas systems for the detection of nucleic acid and non-nucleic acid targets
Nano Research (2023)
-
Programmable Biosensors Based on RNA-Guided CRISPR/Cas Endonuclease
Biological Procedures Online (2022)
-
Massively targeted evaluation of therapeutic CRISPR off-targets in cells
Nature Communications (2022)