Abstract
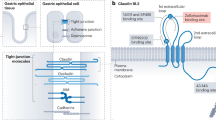
TJs are large intercellular adhesion complexes that maintain cell polarity in normal epithelia and endothelia. During the metastatic process, TJs must be ‘loosened’ or dismantled in cancer cells to enable migration and dissemination. Diminished TJ integrity must also occur within endothelial cells to allow intravasation and extravasation of cancer cells across endothelial barriers. Claudins are critical components of TJs, forming homo- and heteromeric interactions between the adjacent cells, which have been implicated as key modulators of carcinogenesis and metastasis. Numerous epithelial-derived cancers display altered claudin expression patterns and certain claudins can now be used as biomarkers to predict patient prognosis. Moreover, claudins have been functionally implicated in numerous steps of the metastatic cascade. The distinct roles played by claudins during the cancer progression to metastatic disease are just starting to be elucidated. A more complete understanding of the mechanisms through which claudins augment cancer metastasis is required to develop new therapeutic agents against this family of proteins. In this review, we will summarize the relationship between the claudin expression and clinical outcomes in diverse cancers, discuss tumor intrinisic roles through which claudins regulate metastasis and explore claudin-mediated functions within stromal cells that influence the metastatic process. Finally, we will consider possible strategies for targeting claudins that have the potential to improve the management of metastatic cancer.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Krause G, Winkler L, Mueller SL, Haseloff RF, Piontek J, Blasig IE . Structure and function of claudins. Biochim Biophys Acta 2008; 1778: 631–645.
Furuse M, Sasaki H, Tsukita S . Manner of interaction of heterogeneous claudin species within and between tight junction strands. J Cell Biol 1999; 147: 891–903.
Piontek J, Winkler L, Wolburg H, Müller SL, Zuleger N, Piehl C et al. Formation of tight junction: determinants of homophilic interaction between classic claudins. FASEB J 2008; 22: 146–158.
Angelow S, Ahlstrom R, Yu AS . Biology of claudins. Am J Physiol Renal Physiol 2008; 295: F867–F876.
Lal-Nag M, Morin PJ . The claudins. Genome Biol 2009; 10: 235.
Singh AB, Dhawan P . Claudins and cancer: Fall of the soldiers entrusted to protect the gate and keep the barrier intact. Semin Cell Dev Biol 2015; 42: 58–65.
Krause G, Winkler L, Mueller SL, Haseloff RF, Piontek J, Blasig IE . Structure and function of claudins. Biochim Biophys Acta 2008; 1778: 631–645.
Colegio OR, Van Itallie CM, McCrea HJ, Rahner C, Anderson JM . Claudins create charge-selective channels in the paracellular pathway between epithelial cells. Am J Physiol Cell Physiol 2002; 283: C142–C147.
Ruffer C, Gerke V . The C-terminal cytoplasmic tail of claudins 1 and 5 but not its PDZ-binding motif is required for apical localization at epithelial and endothelial tight junctions. Eur J Cell Biol 2004; 83: 135–144.
Piontek J, Winkler L, Wolburg H, Muller SL, Zuleger N, Piehl C et al. Formation of tight junction: determinants of homophilic interaction between classic claudins. Faseb J 2008; 22: 146–158.
Gonzalez-Mariscal L, Tapia R, Chamorro D . Crosstalk of tight junction components with signaling pathways. Biochim Biophys Acta 2008; 1778: 729–756.
Tsukita S, Furuse M, Itoh M . Multifunctional strands in tight junctions. Nat Rev Mol Cell Biol 2001; 2: 285–293.
Van Itallie CM, Gambling TM, Carson JL, Anderson JM . Palmitoylation of claudins is required for efficient tight-junction localization. J Cell Sci 2005; 118: 1427–1436.
Van Itallie CM, Mitic LL, Anderson JM . SUMOylation of claudin-2. Ann NY Acad Sci 2012; 1258: 60–64.
Van Itallie CM, Tietgens AJ, LoGrande K, Aponte A, Gucek M, Anderson JM . Phosphorylation of claudin-2 on serine 208 promotes membrane retention and reduces trafficking to lysosomes. J Cell Sci 2012; 125: 4902–4912.
Amasheh S, Meiri N, Gitter AH, Schoneberg T, Mankertz J, Schulzke JD et al. Claudin-2 expression induces cation-selective channels in tight junctions of epithelial cells. J Cell Sci 2002; 115: 4969–4976.
Yu AS, Cheng MH, Angelow S, Gunzel D, Kanzawa SA, Schneeberger EE et al. Molecular basis for cation selectivity in claudin-2-based paracellular pores: identification of an electrostatic interaction site. J Gen Physiol 2009; 133: 111–127.
Gunzel D, Stuiver M, Kausalya PJ, Haisch L, Krug SM, Rosenthal R et al. Claudin-10 exists in six alternatively spliced isoforms that exhibit distinct localization and function. J Cell Sci 2009; 122: 1507–1517.
Yu AS . Claudins and the kidney. J Am Soc Nephrol 2015; 26: 11–19.
Furuse M, Hata M, Furuse K, Yoshida Y, Haratake A, Sugitani Y et al. Claudin-based tight junctions are crucial for the mammalian epidermal barrier: a lesson from claudin-1-deficient mice. J Cell Biol 2002; 156: 1099–1111.
Nitta T, Hata M, Gotoh S, Seo Y, Sasaki H, Hashimoto N et al. Size-selective loosening of the blood-brain barrier in claudin-5-deficient mice. J Cell Biol 2003; 161: 653–660.
Tatum R, Zhang Y, Salleng K, Lu Z, Lin JJ, Lu Q et al. Renal salt wasting and chronic dehydration in claudin-7-deficient mice. Am J Physiol Renal Physiol 2010; 298: F24–F34.
Ding L, Lu Z, Foreman O, Tatum R, Lu Q, Renegar R et al. Inflammation and disruption of the mucosal architecture in claudin-7-deficient mice. Gastroenterology 2012; 142: 305–315.
Tanaka H, Takechi M, Kiyonari H, Shioi G, Tamura A, Tsukita S . Intestinal deletion of Claudin-7 enhances paracellular organic solute flux and initiates colonic inflammation in mice. Gut 2015; 64: 1529–1538.
Gow A, Southwood CM, Li JS, Pariali M, Riordan GP, Brodie SE et al. CNS myelin and sertoli cell tight junction strands are absent in Osp/claudin-11 null mice. Cell 1999; 99: 649–659.
Ben-Yosef T, Belyantseva IA, Saunders TL, Hughes ED, Kawamoto K, Van Itallie CM et al. Claudin 14 knockout mice, a model for autosomal recessive deafness DFNB29, are deaf due to cochlear hair cell degeneration. Hum Mol Genet 2003; 12: 2049–2061.
Kitajiri S, Miyamoto T, Mineharu A, Sonoda N, Furuse K, Hata M et al. Compartmentalization established by claudin-11-based tight junctions in stria vascularis is required for hearing through generation of endocochlear potential. J Cell Sci 2004; 117: 5087–5096.
Tamura A, Kitano Y, Hata M, Katsuno T, Moriwaki K, Sasaki H et al. Megaintestine in claudin-15-deficient mice. Gastroenterology 2008; 134: 523–534.
Tamura A, Hayashi H, Imasato M, Yamazaki Y, Hagiwara A, Wada M et al. Loss of claudin-15, but not claudin-2, causes Na+ deficiency and glucose malabsorption in mouse small intestine. Gastroenterology 2011; 140: 913–923.
Wada M, Tamura A, Takahashi N, Tsukita S . Loss of claudins 2 and 15 from mice causes defects in paracellular Na+ flow and nutrient transport in gut and leads to death from malnutrition. Gastroenterology 2013; 144: 369–380.
Miyamoto T, Morita K, Takemoto D, Takeuchi K, Kitano Y, Miyakawa T et al. Tight junctions in Schwann cells of peripheral myelinated axons: a lesson from claudin-19-deficient mice. J Cell Biol 2005; 169: 527–538.
Chao YC, Pan SH, Yang SC, Yu SL, Che TF, Lin CW et al. Claudin-1 is a metastasis suppressor and correlates with clinical outcome in lung adenocarcinoma. Am J Respir Crit Care Med 2009; 179: 123–133.
Che J, Yang Y, Xiao J, Zhao P, Yan B, Dong S et al. Decreased expression of claudin-3 is associated with a poor prognosis and EMT in completely resected squamous cell lung carcinoma. Tumour Biol 2015; 36: 6559–6568.
Yamamoto T, Oshima T, Yoshihara K, Yamanaka S, Nishii T, Arai H et al. Reduced expression of claudin-7 is associated with poor outcome in non-small cell lung cancer. Oncol Lett 2010; 1: 501–505.
Micke P, Mattsson JS, Edlund K, Lohr M, Jirstrom K, Berglund A et al. Aberrantly activated claudin 6 and 18.2 as potential therapy targets in non-small-cell lung cancer. Int J Cancer 2014; 135: 2206–2214.
Wang Q, Zhang Y, Zhang T, Han ZG, Shan L . Low claudin-6 expression correlates with poor prognosis in patients with non-small cell lung cancer. Onco Targets Ther 2015; 8: 1971–1977.
Sheehan GM, Kallakury BV, Sheehan CE, Fisher HA, Kaufman Jr RP, Ross JS . Loss of claudins-1 and -7 and expression of claudins-3 and -4 correlate with prognostic variables in prostatic adenocarcinomas. Hum Pathol 2007; 38: 564–569.
Landers KA, Samaratunga H, Teng L, Buck M, Burger MJ, Scells B et al. Identification of claudin-4 as a marker highly overexpressed in both primary and metastatic prostate cancer. Br J Cancer 2008; 99: 491–501.
Szasz AM, Nyirady P, Majoros A, Szendroi A, Szucs M, Szekely E et al. beta-catenin expression and claudin expression pattern as prognostic factors of prostatic cancer progression. BJU Int 2010; 105: 716–722.
Szasz AM, Majoros A, Rosen P, Srivastava S, Dobi A, Szendroi A et al. Prognostic potential of ERG (ETS-related gene) expression in prostatic adenocarcinoma. Int Urol Nephrol 2013; 45: 727–733.
Hibbs K, Skubitz KM, Pambuccian SE, Casey RC, Burleson KM, Oegema TR Jr. et al. Differential gene expression in ovarian carcinoma: identification of potential biomarkers. Am J Pathol 2004; 165: 397–414.
Hough CD, Sherman-Baust CA, Pizer ES, Montz FJ, Im DD, Rosenshein NB et al. Large-scale serial analysis of gene expression reveals genes differentially expressed in ovarian cancer. Cancer Res 2000; 60: 6281–6287.
Rangel LB, Agarwal R, D'Souza T, Pizer ES, Alo PL, Lancaster WD et al. Tight junction proteins claudin-3 and claudin-4 are frequently overexpressed in ovarian cancer but not in ovarian cystadenomas. Clin Cancer Res 2003; 9: 2567–2575.
Choi YL, Kim J, Kwon MJ, Choi JS, Kim TJ, Bae DS et al. Expression profile of tight junction protein claudin 3 and claudin 4 in ovarian serous adenocarcinoma with prognostic correlation. Histol Histopathol 2007; 22: 1185–1195.
Kleinberg L, Holth A, Trope CG, Reich R, Davidson B . Claudin upregulation in ovarian carcinoma effusions is associated with poor survival. Hum Pathol 2008; 39: 747–757.
Dahiya N, Becker KG, Wood WH 3rd, Zhang Y, Morin PJ . Claudin-7 is frequently overexpressed in ovarian cancer and promotes invasion. PLoS One 2011; 6: e22119.
Kim CJ, Lee JW, Choi JJ, Choi HY, Park YA, Jeon HK et al. High claudin-7 expression is associated with a poor response to platinum-based chemotherapy in epithelial ovarian carcinoma. Eur J Cancer 2011; 47: 918–925.
Wang H, Yang X . The expression patterns of tight junction protein claudin-1, -3, and -4 in human gastric neoplasms and adjacent non-neoplastic tissues. Int J Clin Exp Pathol 2015; 8: 881–887.
Huang J, Li J, Qu Y, Zhang J, Zhang L, Chen X et al. The expression of claudin 1 correlates with beta-catenin and is a prognostic factor of poor outcome in gastric cancer. Int J Oncol 2014; 44: 1293–1301.
Zhu JL, Gao P, Wang ZN, Song YX, Li AL, Xu YY et al. Clinicopathological significance of claudin-4 in gastric carcinoma. World J Surg Oncol 2013; 11: 150.
Japanese Gastric Cancer A. Japanese classification of gastric carcinoma: 3rd English edition. Gastric Cancer 2011; 14: 101–112.
Ming SC . Gastric carcinoma. A pathobiological classification. Cancer 1977; 39: 2475–2485.
Hwang TL, Lee LY, Wang CC, Liang Y, Huang SF, Wu CM . Claudin-4 expression is associated with tumor invasion, MMP-2 and MMP-9 expression in gastric cancer. Exp Ther Med 2010; 1: 789–797.
Liu JX, Wei ZY, Chen JS, Lu HC, Hao L, Li WJ . Prognostic and clinical significance of claudin-4 in gastric cancer: a meta-analysis. World J Surg Oncol 2015; 13: 207.
Ohtani S, Terashima M, Satoh J, Soeta N, Saze Z, Kashimura S et al. Expression of tight-junction-associated proteins in human gastric cancer: downregulation of claudin-4 correlates with tumor aggressiveness and survival. Gastric Cancer 2009; 12: 43–51.
Nakagawa S, Miyoshi N, Ishii H, Mimori K, Tanaka F, Sekimoto M et al. Expression of CLDN1 in colorectal cancer: a novel marker for prognosis. Int J Oncol 2011; 39: 791–796.
Shibutani M, Noda E, Maeda K, Nagahara H, Ohtani H, Hirakawa K . Low expression of claudin-1 and presence of poorly-differentiated tumor clusters correlate with poor prognosis in colorectal cancer. Anticancer Res 2013; 33: 3301–3306.
Yoshida T, Kinugasa T, Akagi Y, Kawahara A, Romeo K, Shiratsuchi I et al. Decreased expression of claudin-1 in rectal cancer: a factor for recurrence and poor prognosis. Anticancer Res 2011; 31: 2517–2525.
Dhawan P, Singh AB, Deane NG, No Y, Shiou SR, Schmidt C et al. Claudin-1 regulates cellular transformation and metastatic behavior in colon cancer. J Clin Invest 2005; 115: 1765–1776.
Huo Q, Kinugasa T, Wang L, Huang J, Zhao J, Shibaguchi H et al. Claudin-1 protein is a major factor involved in the tumorigenesis of colorectal cancer. Anticancer Res 2009; 29: 851–857.
Kinugasa T, Huo Q, Higashi D, Shibaguchi H, Kuroki M, Tanaka T et al. Selective up-regulation of claudin-1 and claudin-2 in colorectal cancer. Anticancer Res 2007; 27: 3729–3734.
Bornholdt J, Friis S, Godiksen S, Poulsen SS, Santoni-Rugiu E, Bisgaard HC et al. The level of claudin-7 is reduced as an early event in colorectal carcinogenesis. BMC Cancer 2011; 11: 65.
Bhat AA, Pope JL, Smith JJ, Ahmad R, Chen X, Washington MK et al. Claudin-7 expression induces mesenchymal to epithelial transformation (MET) to inhibit colon tumorigenesis. Oncogene 2015; 34: 4570–4580.
Oshima T, Kunisaki C, Yoshihara K, Yamada R, Yamamoto N, Sato T et al. Reduced expression of the claudin-7 gene correlates with venous invasion and liver metastasis in colorectal cancer. Oncol Rep 2008; 19: 953–959.
Darido C, Buchert M, Pannequin J, Bastide P, Zalzali H, Mantamadiotis T et al. Defective claudin-7 regulation by Tcf-4 and Sox-9 disrupts the polarity and increases the tumorigenicity of colorectal cancer cells. Cancer Res 2008; 68: 4258–4268.
Kuhn S, Koch M, Nubel T, Ladwein M, Antolovic D, Klingbeil P et al. A complex of EpCAM, claudin-7, CD44 variant isoforms, and tetraspanins promotes colorectal cancer progression. Mol Cancer Res 2007; 5: 553–567.
Nakayama F, Semba S, Usami Y, Chiba H, Sawada N, Yokozaki H . Hypermethylation-modulated downregulation of claudin-7 expression promotes the progression of colorectal carcinoma. Pathobiology 2008; 75: 177–185.
Pitule P, Vycital O, Bruha J, Novak P, Hosek P, Treska V et al. Differential expression and prognostic role of selected genes in colorectal cancer patients. Anticancer Res 2013; 33: 4855–4865.
Ma F, Ding X, Fan Y, Ying J, Zheng S, Lu N et al. A CLDN1-negative phenotype predicts poor prognosis in triple-negative breast cancer. PLoS One 2014; 9: e112765.
Morohashi S, Kusumi T, Sato F, Odagiri H, Chiba H, Yoshihara S et al. Decreased expression of claudin-1 correlates with recurrence status in breast cancer. Int J Mol Med 2007; 20: 139–143.
Szasz AM, Tokes AM, Micsinai M, Krenacs T, Jakab C, Lukacs L et al. Prognostic significance of claudin expression changes in breast cancer with regional lymph node metastasis. Clin Exp Metastasis 2011; 28: 55–63.
Soini Y . Claudins 2, 3, 4, and 5 in Paget's disease and breast carcinoma. Hum Pathol 2004; 35: 1531–1536.
Kim TH, Huh JH, Lee S, Kang H, Kim GI, An HJ . Down-regulation of claudin-2 in breast carcinomas is associated with advanced disease. Histopathology 2008; 53: 48–55.
Kimbung S, Kovacs A, Bendahl PO, Malmstrom P, Ferno M, Hatschek T et al. Claudin-2 is an independent negative prognostic factor in breast cancer and specifically predicts early liver recurrences. Mol Oncol 2014; 8: 119–128.
Blanchard AA, Skliris GP, Watson PH, Murphy LC, Penner C, Tomes L et al. Claudins 1, 3, and 4 protein expression in ER negative breast cancer correlates with markers of the basal phenotype. Virchows Arch 2009; 454: 647–656.
Kulka J, Szasz AM, Nemeth Z, Madaras L, Schaff Z, Molnar IA et al. Expression of tight junction protein claudin-4 in basal-like breast carcinomas. Pathol Oncol Res 2009; 15: 59–64.
Lanigan F, McKiernan E, Brennan DJ, Hegarty S, Millikan RC, McBryan J et al. Increased claudin-4 expression is associated with poor prognosis and high tumour grade in breast cancer. Int J Cancer 2009; 124: 2088–2097.
Kominsky SL, Argani P, Korz D, Evron E, Raman V, Garrett E et al. Loss of the tight junction protein claudin-7 correlates with histological grade in both ductal carcinoma in situ and invasive ductal carcinoma of the breast. Oncogene 2003; 22: 2021–2033.
Sauer T, Pedersen MK, Ebeltoft K, Naess O . Reduced expression of Claudin-7 in fine needle aspirates from breast carcinomas correlate with grading and metastatic disease. Cytopathology 2005; 16: 193–198.
Martin TA, Harrison GM, Watkins G, Jiang WG . Claudin-16 reduces the aggressive behavior of human breast cancer cells. J Cell Biochem 2008; 105: 41–52.
Martin TA, Lane J, Ozupek H, Jiang WG . Claudin-20 promotes an aggressive phenotype in human breast cancer cells. Tissue Barriers 2013; 1: e26518.
Mina LA, Sledge GW Jr . Rethinking the metastatic cascade as a therapeutic target. Nat Rev Clin Oncol 2011; 8: 325–332.
Huang J, Zhang L, He C, Qu Y, Li J, Zhang J et al. Claudin-1 enhances tumor proliferation and metastasis by regulating cell anoikis in gastric cancer. Oncotarget 2015; 6: 1652–1665.
Yoda S, Soejima K, Hamamoto J, Yasuda H, Nakayama S, Satomi R et al. Claudin-1 is a novel target of miR-375 in non-small-cell lung cancer. Lung Cancer 2014; 85: 366–372.
Qin W, Ren Q, Liu T, Huang Y, Wang J . MicroRNA-155 is a novel suppressor of ovarian cancer-initiating cells that targets CLDN1. FEBS Lett 2013; 587: 1434–1439.
Zhang GJ, Xiao HX, Tian HP, Liu ZL, Xia SS, Zhou T . Upregulation of microRNA-155 promotes the migration and invasion of colorectal cancer cells through the regulation of claudin-1 expression. Int J Mol Med 2013; 31: 1375–1380.
Bhat AA, Sharma A, Pope J, Krishnan M, Washington MK, Singh AB et al. Caudal homeobox protein Cdx-2 cooperates with Wnt pathway to regulate claudin-1 expression in colon cancer cells. PLoS One 2012; 7: e37174.
Singh AB, Sharma A, Dhawan P . Claudin-1 expression confers resistance to anoikis in colon cancer cells in a Src-dependent manner. Carcinogenesis 2012; 33: 2538–2547.
Pope JL, Ahmad R, Bhat AA, Washington MK, Singh AB, Dhawan P . Claudin-1 overexpression in intestinal epithelial cells enhances susceptibility to adenamatous polyposis coli-mediated colon tumorigenesis. Mole Cancer 2014; 13: 167.
Bos PD, Zhang XH, Nadal C, Shu W, Gomis RR, Nguyen DX et al. Genes that mediate breast cancer metastasis to the brain. Nature 2009; 459: 1005–1009.
Kang Y, Siegel PM, Shu W, Drobnjak M, Kakonen SM, Cordon-Cardo C et al. A multigenic program mediating breast cancer metastasis to bone. Cancer Cell 2003; 3: 537–549.
Minn AJ, Gupta GP, Siegel PM, Bos PD, Shu W, Giri DD et al. Genes that mediate breast cancer metastasis to lung. Nature 2005; 436: 518–524.
Tabariès S, Dong Z, Annis MG, Omeroglu A, Pepin F, Ouellet V et al. Claudin-2 is selectively enriched in and promotes the formation of breast cancer liver metastases through engagement of integrin complexes. Oncogene 2011; 30: 1318–1328.
Herschkowitz JI, Simin K, Weigman VJ, Mikaelian I, Usary J, Hu Z et al. Identification of conserved gene expression features between murine mammary carcinoma models and human breast tumors. Genome Biol 2007; 8: R76.
Prat A, Parker JS, Karginova O, Fan C, Livasy C, Herschkowitz JI et al. Phenotypic and molecular characterization of the claudin-low intrinsic subtype of breast cancer. Breast Cancer Res 2010; 12: R68.
Prat A, Perou CM . Deconstructing the molecular portraits of breast cancer. Mol Oncol 2011; 5: 5–23.
Tabariès S, Dupuy F, Dong Z, Monast A, Annis MG, Spicer J et al. Claudin-2 promotes breast cancer liver metastasis by facilitating tumor cell interactions with hepatocytes. Mol Cell Biol 2012; 32: 2979–2991.
Tabariès S, Annis MG, Hsu BE, Tam CE, Savage P, Park M et al. Lyn modulates Claudin-2 expression and is a therapeutic target for breast cancer liver metastasis. Oncotarget 2015; 6: 9476–9487.
Dhawan P, Ahmad R, Chaturvedi R, Smith JJ, Midha R, Mittal MK et al. Claudin-2 expression increases tumorigenicity of colon cancer cells: role of epidermal growth factor receptor activation. Oncogene 2011; 30: 3234–3247.
Mima S, Takehara M, Takada H, Nishimura T, Hoshino T, Mizushima T . NSAIDs suppress the expression of claudin-2 to promote invasion activity of cancer cells. Carcinogenesis 2008; 29: 1994–2000.
Agarwal R, D'Souza T, Morin PJ . Claudin-3 and claudin-4 expression in ovarian epithelial cells enhances invasion and is associated with increased matrix metalloproteinase-2 activity. Cancer Res 2005; 65: 7378–7385.
Huang YH, Bao Y, Peng W, Goldberg M, Love K, Bumcrot DA et al. Claudin-3 gene silencing with siRNA suppresses ovarian tumor growth and metastasis. Proc Natl Acad Sci USA 2009; 106: 3426–3430.
Hwang TL, Changchien TT, Wang CC, Wu CM . Claudin-4 expression in gastric cancer cells enhances the invasion and is associated with the increased level of matrix metalloproteinase-2 and -9 expression. Oncol Lett 2014; 8: 1367–1371.
Ren Y, Wu Q, Liu Y, Xu X, Quan C . Gene silencing of claudin6 enhances cell proliferation and migration accompanied with increased MMP2 activity via p38 MAPK signaling pathway in human breast epithelium cell line HBL100. Mol Med Rep 2013; 8: 1505–1510.
Augeron C, Laboisse CL . Emergence of permanently differentiated cell clones in a human colonic cancer cell line in culture after treatment with sodium butyrate. Cancer Res 1984; 44: 3961–3969.
Philip R, Heiler S, Mu W, Buchler MW, Zoller M, Thuma F . Claudin-7 promotes the epithelial-mesenchymal transition in human colorectal cancer. Oncotarget 2015; 6: 2046–2063.
Ladwein M, Pape UF, Schmidt DS, Schnolzer M, Fiedler S, Langbein L et al. The cell-cell adhesion molecule EpCAM interacts directly with the tight junction protein claudin-7. Exp Cell Res 2005; 309: 345–357.
Hemler ME . Tetraspanin functions and associated microdomains. Nat Rev Mol Cell Biol 2005; 6: 801–811.
Lazo PA . Functional implications of tetraspanin proteins in cancer biology. Cancer Sci 2007; 98: 1666–1677.
Sharma RK, Chheda ZS, Das Purkayastha BP, Gomez-Gutierrez JG, Jala VR, Haribabu B . A spontaneous metastasis model reveals the significance of claudin-9 overexpression in lung cancer metastasis. Clin Exp Metastasis 2016; 33: 263–275.
Gril B, Evans L, Palmieri D, Steeg PS . Translational research in brain metastasis is identifying molecular pathways that may lead to the development of new therapeutic strategies. Eur J Cancer 2010; 46: 1204–1210.
Jia W, Martin TA, Zhang G, Jiang WG . Junctional adhesion molecules in cerebral endothelial tight junction and brain metastasis. Anticancer Res 2013; 33: 2353–2359.
Wilhelm I, Molnar J, Fazakas C, Hasko J, Krizbai IA . Role of the blood-brain barrier in the formation of brain metastases. Int J Mol Sci 2013; 14: 1383–1411.
Abbott NJ, Ronnback L, Hansson E . Astrocyte-endothelial interactions at the blood-brain barrier. Nat Rev Neurosci 2006; 7: 41–53.
Fanning AS, Jameson BJ, Jesaitis LA, Anderson JM . The tight junction protein ZO-1 establishes a link between the transmembrane protein occludin and the actin cytoskeleton. J Biol Chem 1998; 273: 29745–29753.
Jia W, Lu R, Martin TA, Jiang WG . The role of claudin-5 in blood-brain barrier (BBB) and brain metastases (review). Mol Med Rep 2014; 9: 779–785.
Rodriguez PL, Jiang S, Fu Y, Avraham S, Avraham HK . The proinflammatory peptide substance P promotes blood-brain barrier breaching by breast cancer cells through changes in microvascular endothelial cell tight junctions. Int J Cancer 2014; 134: 1034–1044.
Avraham HK, Jiang S, Fu Y, Nakshatri H, Ovadia H, Avraham S . Angiopoietin-2 mediates blood-brain barrier impairment and colonization of triple-negative breast cancer cells in brain. J Pathol 2014; 232: 369–381.
Feng S, Cen J, Huang Y, Shen H, Yao L, Wang Y et al. Matrix metalloproteinase-2 and -9 secreted by leukemic cells increase the permeability of blood-brain barrier by disrupting tight junction proteins. PLoS One 2011; 6: e20599.
Krizbai IA, Gasparics A, Nagyoszi P, Fazakas C, Molnar J, Wilhelm I et al. Endothelial-mesenchymal transition of brain endothelial cells: possible role during metastatic extravasation. PLoS One 2015; 10: e0123845.
Kominsky SL . Claudins: emerging targets for cancer therapy. Expert Rev Mol Med 2006; 8: 1–11.
Suzuki H, Kondoh M, Takahashi A, Yagi K . Proof of concept for claudin-targeted drug development. Ann NY Acad Sci 2012; 1258: 65–70.
Takahashi A, Kondoh M, Suzuki H, Yagi K . Claudin as a target for drug development. Curr Med Chem 2011; 18: 1861–1865.
Cocco E, Casagrande F, Bellone S, Richter CE, Bellone M, Todeschini P et al. Clostridium perfringens enterotoxin carboxy-terminal fragment is a novel tumor-homing peptide for human ovarian cancer. BMC Cancer 2010; 10: 349.
Fujita K, Katahira J, Horiguchi Y, Sonoda N, Furuse M, Tsukita S . Clostridium perfringens enterotoxin binds to the second extracellular loop of claudin-3, a tight junction integral membrane protein. FEBS Lett 2000; 476: 258–261.
Gao Z, McClane BA . Use of Clostridium perfringens enterotoxin and the enterotoxin receptor-binding domain (C-CPE) for cancer treatment: opportunities and challenges. J Toxicol 2012; 2012: 981626.
McClane BA, Wnek AP, Hulkower KI, Hanna PC . Divalent cation involvement in the action of Clostridium perfringens type A enterotoxin. Early events in enterotoxin action are divalent cation-independent. J Biol Chem 1988; 263: 2423–2435.
Katahira J, Inoue N, Horiguchi Y, Matsuda M, Sugimoto N . Molecular cloning and functional characterization of the receptor for Clostridium perfringens enterotoxin. J Cell Biol 1997; 136: 1239–1247.
Katahira J, Sugiyama H, Inoue N, Horiguchi Y, Matsuda M, Sugimoto N . Clostridium perfringens enterotoxin utilizes two structurally related membrane proteins as functional receptors in vivo. J Biol Chem 1997; 272: 26652–26658.
Escudero-Esparza A, Jiang WG, Martin TA . The Claudin family and its role in cancer and metastasis. Front Biosci 2011; 16: 1069–1083.
Lal-Nag M, Battis M, Santin AD, Morin PJ . Claudin-6: a novel receptor for CPE-mediated cytotoxicity in ovarian cancer. Oncogenesis 2012; 1: e33.
Black JD, Lopez S, Cocco E, Schwab CL, English DP, Santin AD . Clostridium perfringens enterotoxin (CPE) and CPE-binding domain (c-CPE) for the detection and treatment of gynecologic cancers. Toxins (Basel) 2015; 7: 1116–1125.
Casagrande F, Cocco E, Bellone S, Richter CE, Bellone M, Todeschini P et al. Eradication of chemotherapy-resistant CD44+ human ovarian cancer stem cells in mice by intraperitoneal administration of Clostridium perfringens enterotoxin. Cancer 2011; 117: 5519–5528.
Cocco E, Shapiro EM, Gasparrini S, Lopez S, Schwab CL, Bellone S et al. Clostridium perfringens enterotoxin C-terminal domain labeled to fluorescent dyes for in vivo visualization of micrometastatic chemotherapy-resistant ovarian cancer. Int J Cancer 2015; 137: 2618–2629.
Neesse A, Hahnenkamp A, Griesmann H, Buchholz M, Hahn SA, Maghnouj A et al. Claudin-4-targeted optical imaging detects pancreatic cancer and its precursor lesions. Gut 2013; 62: 1034–1043.
Kominsky SL, Vali M, Korz D, Gabig TG, Weitzman SA, Argani P et al. Clostridium perfringens enterotoxin elicits rapid and specific cytolysis of breast carcinoma cells mediated through tight junction proteins claudin 3 and 4. Am J Pathol 2004; 164: 1627–1633.
Michl P, Buchholz M, Rolke M, Kunsch S, Lohr M, McClane B et al. Claudin-4: a new target for pancreatic cancer treatment using Clostridium perfringens enterotoxin. Gastroenterology 2001; 121: 678–684.
Neesse A, Griesmann H, Gress TM, Michl P . Claudin-4 as therapeutic target in cancer. Arch Biochem Biophys 2012; 524: 64–70.
Santin AD, Cane S, Bellone S, Palmieri M, Siegel ER, Thomas M et al. Treatment of chemotherapy-resistant human ovarian cancer xenografts in C.B-17/SCID mice by intraperitoneal administration of Clostridium perfringens enterotoxin. Cancer Res 2005; 65: 4334–4342.
Gao Z, Xu X, McClane B, Zeng Q, Litkouhi B, Welch WR et al. C terminus of Clostridium perfringens enterotoxin downregulates CLDN4 and sensitizes ovarian cancer cells to Taxol and Carboplatin. Clin Cancer Res 2011; 17: 1065–1074.
Yuan X, Lin X, Manorek G, Kanatani I, Cheung LH, Rosenblum MG et al. Recombinant CPE fused to tumor necrosis factor targets human ovarian cancer cells expressing the claudin-3 and claudin-4 receptors. Mol Cancer Ther 2009; 8: 1906–1915.
Yao Q, Cao S, Li C, Mengesha A, Low P, Kong B et al. Turn a diarrhoea toxin into a receptor-mediated therapy for a plethora of CLDN-4-overexpressing cancers. Biochem Biophys Res Commun 2010; 398: 413–419.
Saeki R, Kondoh M, Kakutani H, Matsuhisa K, Takahashi A, Suzuki H et al. A claudin-targeting molecule as an inhibitor of tumor metastasis. J Pharmacol Exp Ther 2010; 334: 576–582.
Kominsky SL, Tyler B, Sosnowski J, Brady K, Doucet M, Nell D et al. Clostridium perfringens enterotoxin as a novel-targeted therapeutic for brain metastasis. Cancer Res 2007; 67: 7977–7982.
Protze J, Eichner M, Piontek A, Dinter S, Rossa J, Blecharz KG et al. Directed structural modification of Clostridium perfringens enterotoxin to enhance binding to claudin-5. Cell Mol Life Sci 2015; 72: 1417–1432.
Kuwada M, Chihara Y, Luo Y, Li X, Nishiguchi Y, Fujiwara R et al. Pro-chemotherapeutic effects of antibody against extracellular domain of claudin-4 in bladder cancer. Cancer Lett 2015; 369: 212–221.
Dev KK . Making protein interactions druggable: targeting PDZ domains. Nat Rev Drug Discov 2004; 3: 1047–1056.
Nishimune A, Isaac JT, Molnar E, Noel J, Nash SR, Tagaya M et al. NSF binding to GluR2 regulates synaptic transmission. Neuron 1998; 21: 87–97.
Acknowledgements
We thank members of the Siegel laboratory for their thoughtful comments on the manuscript. Research conducted in the author’s laboratory was supported by grants from the CIHR (MOP-136907 to PMS). PMS is a William Dawson Scholar of McGill University.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Tabariès, S., Siegel, P. The role of claudins in cancer metastasis. Oncogene 36, 1176–1190 (2017). https://doi.org/10.1038/onc.2016.289
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2016.289
This article is cited by
-
Current and Future Biomarkers in Esophagogastric Adenocarcinoma
Journal of Gastrointestinal Cancer (2024)
-
Claudin-4-adhesion signaling drives breast cancer metabolism and progression via liver X receptor β
Breast Cancer Research (2023)
-
Downregulation of Claudin5 promotes malignant progression and radioresistance through Beclin1-mediated autophagy in esophageal squamous cell carcinoma
Journal of Translational Medicine (2023)
-
The role of claudins in homeostasis
Nature Reviews Nephrology (2023)
-
TMEM25 is a Par3-binding protein that attenuates claudin assembly during tight junction development
EMBO Reports (2023)