Abstract
Heat-shock transcription factor (HSF) family members function in stress protection and in human diseases including proteopathies, neurodegeneration and cancer. The mechanisms that drive distinct post-translational modifications, cofactor recruitment and target-gene activation for specific HSF paralogs are unknown. We present crystal structures of the human HSF2 DNA-binding domain (DBD) bound to DNA, revealing an unprecedented view of HSFs that provides insights into their unique biology. The HSF2 DBD structures resolve a new C-terminal helix that directs wrapping of the coiled-coil domain around DNA, thereby exposing paralog-specific sequences of the DBD surface for differential post-translational modifications and cofactor interactions. We further demonstrate a direct interaction between HSF1 and HSF2 through their coiled-coil domains. Together, these features provide a new model for HSF structure as the basis for differential and combinatorial regulation, which influences the transcriptional response to cellular stress.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Jiang, S. et al. Multifaceted roles of HSF1 in cancer. Tumour Biol. 36, 4923–4931 (2015).
Neef, D.W., Turski, M.L. & Thiele, D.J. Modulation of heat shock transcription factor 1 as a therapeutic target for small molecule intervention in neurodegenerative disease. PLoS Biol. 8, e1000291 (2010).
Akerfelt, M., Morimoto, R.I. & Sistonen, L. Heat shock factors: integrators of cell stress, development and lifespan. Nat. Rev. Mol. Cell Biol. 11, 545–555 (2010).
Jaeger, A.M., Makley, L.N., Gestwicki, J.E. & Thiele, D.J. Genomic heat shock element sequences drive cooperative human heat shock factor 1 DNA binding and selectivity. J. Biol. Chem. 289, 30459–30469 (2014).
Mendillo, M.L. et al. HSF1 drives a transcriptional program distinct from heat shock to support highly malignant human cancers. Cell 150, 549–562 (2012).
Whitesell, L. & Lindquist, S. Inhibiting the transcription factor HSF1 as an anticancer strategy. Expert Opin. Ther. Targets 13, 469–478 (2009).
Neef, D.W., Jaeger, A.M. & Thiele, D.J. Heat shock transcription factor 1 as a therapeutic target in neurodegenerative diseases. Nat. Rev. Drug Discov. 10, 930–944 (2011).
Littlefield, O. & Nelson, H.C. A new use for the 'wing' of the 'winged' helix-turn-helix motif in the HSF–DNA cocrystal. Nat. Struct. Biol. 6, 464–470 (1999).
Rabindran, S.K., Haroun, R.I., Clos, J., Wisniewski, J. & Wu, C. Regulation of heat shock factor trimer formation: role of a conserved leucine zipper. Science 259, 230–234 (1993).
Sorger, P.K. & Nelson, H.C. Trimerization of a yeast transcriptional activator via a coiled-coil motif. Cell 59, 807–813 (1989).
Farkas, T., Kutskova, Y.A. & Zimarino, V. Intramolecular repression of mouse heat shock factor 1. Mol. Cell. Biol. 18, 906–918 (1998).
Zuo, J., Baler, R., Dahl, G. & Voellmy, R. Activation of the DNA-binding ability of human heat shock transcription factor 1 may involve the transition from an intramolecular to an intermolecular triple-stranded coiled-coil structure. Mol. Cell. Biol. 14, 7557–7568 (1994).
Vihervaara, A. et al. Transcriptional response to stress in the dynamic chromatin environment of cycling and mitotic cells. Proc. Natl. Acad. Sci. USA 110, E3388–E3397 (2013).
Guertin, M.J., Martins, A.L., Siepel, A. & Lis, J.T. Accurate prediction of inducible transcription factor binding intensities in vivo. PLoS Genet. 8, e1002610 (2012).
Neef, D.W. et al. A direct regulatory interaction between chaperonin TRiC and stress-responsive transcription factor HSF1. Cell Reports 9, 955–966 (2014).
Hahn, J.-S., Neef, D.W. & Thiele, D.J. A stress regulatory network for co-ordinated activation of proteasome expression mediated by yeast heat shock transcription factor. Mol. Microbiol. 60, 240–251 (2006).
Scherz-Shouval, R. et al. The reprogramming of tumor stroma by HSF1 is a potent enabler of malignancy. Cell 158, 564–578 (2014).
Riva, L. et al. Poly-glutamine expanded huntingtin dramatically alters the genome wide binding of HSF1. J. Huntingtons Dis. 1, 33–45 (2012).
Shinkawa, T. et al. Heat shock factor 2 is required for maintaining proteostasis against febrile-range thermal stress and polyglutamine aggregation. Mol. Biol. Cell 22, 3571–3583 (2011).
Elsing, A.N. et al. Expression of HSF2 decreases in mitosis to enable stress-inducible transcription and cell survival. J. Cell Biol. 206, 735–749 (2014).
Ahlskog, J.K. et al. Anaphase-promoting complex/cyclosome participates in the acute response to protein-damaging stress. Mol. Cell. Biol. 30, 5608–5620 (2010).
El Fatimy, R. et al. Heat shock factor 2 is a stress-responsive mediator of neuronal migration defects in models of fetal alcohol syndrome. EMBO Mol. Med. 6, 1043–1061 (2014).
Mou, L. et al. A dominant-negative mutation of HSF2 associated with idiopathic azoospermia. Hum. Genet. 132, 159–165 (2013).
Akerfelt, M. et al. Promoter ChIP-chip analysis in mouse testis reveals Y chromosome occupancy by HSF2. Proc. Natl. Acad. Sci. USA 105, 11224–11229 (2008).
Wilkerson, D.C., Murphy, L.A. & Sarge, K.D. Interaction of HSF1 and HSF2 with the Hspa1b promoter in mouse epididymal spermatozoa. Biol. Reprod. 79, 283–288 (2008).
Rossi, A. et al. The proteasome inhibitor bortezomib is a potent inducer of zinc finger AN1-type domain 2a gene expression: role of heat shock factor 1 (HSF1)-heat shock factor 2 (HSF2) heterocomplexes. J. Biol. Chem. 289, 12705–12715 (2014).
Sandqvist, A. et al. Heterotrimerization of heat-shock factors 1 and 2 provides a transcriptional switch in response to distinct stimuli. Mol. Biol. Cell 20, 1340–1347 (2009).
Liu, J. et al. Structural basis of DNA recognition by PCG2 reveals a novel DNA binding mode for winged helix-turn-helix domains. Nucleic Acids Res. 43, 1231–1240 (2015).
Kitano, K., Kim, S.Y. & Hakoshima, T. Structural basis for DNA strand separation by the unconventional winged-helix domain of RecQ helicase WRN. Structure 18, 177–187 (2010).
Tammsalu, T. et al. Proteome-wide identification of SUMO2 modification sites. Sci. Signal. 7, rs2 (2014).
Hendriks, I.A. et al. Uncovering global SUMOylation signaling networks in a site-specific manner. Nat. Struct. Mol. Biol. 21, 927–936 (2014).
Becker, J. et al. Detecting endogenous SUMO targets in mammalian cells and tissues. Nat. Struct. Mol. Biol. 20, 525–531 (2013).
Fujimoto, M. et al. RPA assists HSF1 access to nucleosomal DNA by recruiting histone chaperone FACT. Mol. Cell 48, 182–194 (2012).
Anckar, J. & Sistonen, L. Regulation of HSF1 function in the heat stress response: implications in aging and disease. Annu. Rev. Biochem. 80, 1089–1115 (2011).
Westwood, J.T. & Wu, C. Activation of Drosophila heat shock factor: conformational change associated with a monomer-to-trimer transition. Mol. Cell. Biol. 13, 3481–3486 (1993).
Liu, X.D., Liu, P.C., Santoro, N. & Thiele, D.J. Conservation of a stress response: human heat shock transcription factors functionally substitute for yeast HSF. EMBO J. 16, 6466–6477 (1997).
Sistonen, L., Sarge, K.D. & Morimoto, R.I. Human heat shock factors 1 and 2 are differentially activated and can synergistically induce hsp70 gene transcription. Mol. Cell. Biol. 14, 2087–2099 (1994).
Fujimoto, M. & Nakai, A. The heat shock factor family and adaptation to proteotoxic stress. FEBS J. 277, 4112–4125 (2010).
Calderwood, S.K., Murshid, A. & Prince, T. The shock of aging: molecular chaperones and the heat shock response in longevity and aging: a mini-review. Gerontology 55, 550–558 (2009).
Chen, L., Glover, J.N., Hogan, P.G., Rao, A. & Harrison, S.C. Structure of the DNA-binding domains from NFAT, Fos and Jun bound specifically to DNA. Nature 392, 42–48 (1998).
Tan, K. et al. Mitochondrial SSBP1 protects cells from proteotoxic stresses by potentiating stress-induced HSF1 transcriptional activity. Nat. Commun. 6, 6580 (2015).
Zelin, E. & Freeman, B.C. Lysine deacetylases regulate the heat shock response including the age-associated impairment of HSF1. J. Mol. Biol. 427, 1644–1654 (2015).
Kalmar, B. & Greensmith, L. Activation of the heat shock response in a primary cellular model of motoneuron neurodegeneration-evidence for neuroprotective and neurotoxic effects. Cell. Mol. Biol. Lett. 14, 319–335 (2009).
Ashkenazy, H., Erez, E., Martz, E., Pupko, T. & Ben-Tal, N. ConSurf 2010: calculating evolutionary conservation in sequence and structure of proteins and nucleic acids. Nucleic Acids Res. 38, W529–W533 (2010).
Bernier-Villamor, V., Sampson, D.A., Matunis, M.J. & Lima, C.D. Structural basis for E2-mediated SUMO conjugation revealed by a complex between ubiquitin-conjugating enzyme Ubc9 and RanGAP1. Cell 108, 345–356 (2002).
Anckar, J. et al. Inhibition of DNA binding by differential sumoylation of heat shock factors. Mol. Cell. Biol. 26, 955–964 (2006).
Tateishi, Y. et al. Molecular basis for SUMOylation-dependent regulation of DNA binding activity of heat shock factor 2. J. Biol. Chem. 284, 2435–2447 (2009).
Niskanen, E.A. et al. Global SUMOylation on active chromatin is an acute heat stress response restricting transcription. Genome Biol. 16, 153 (2015).
Seifert, A., Schofield, P., Barton, G.J. & Hay, R.T. Proteotoxic stress reprograms the chromatin landscape of SUMO modification. Sci. Signal. 8, rs7 (2015).
Shu, W., Liu, J., Ji, H. & Lu, M. Core structure of the outer membrane lipoprotein from Escherichia coli at 1.9 A resolution. J. Mol. Biol. 299, 1101–1112 (2000).
Sorrells, T.R., Booth, L.N., Tuch, B.B. & Johnson, A.D. Intersecting transcription networks constrain gene regulatory evolution. Nature 523, 361–365 (2015).
Innan, H. & Kondrashov, F. The evolution of gene duplications: classifying and distinguishing between models. Nat. Rev. Genet. 11, 97–108 (2010).
Force, A. et al. Preservation of duplicate genes by complementary, degenerative mutations. Genetics 151, 1531–1545 (1999).
Baker, C.R., Hanson-Smith, V. & Johnson, A.D. Following gene duplication, paralog interference constrains transcriptional circuit evolution. Science 342, 104–108 (2013).
Björk, J.K. et al. Heat-shock factor 2 is a suppressor of prostate cancer invasion. Oncogene doi:10.1038/onc.2015.241 29 June (2015).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Matthews, B.W. Solvent content of protein crystals. J. Mol. Biol. 33, 491–497 (1968).
Terwilliger, T.C. et al. Iterative model building, structure refinement and density modification with the PHENIX AutoBuild wizard. Acta Crystallogr. D Biol. Crystallogr. 64, 61–69 (2008).
Emsley, P., Lohkamp, B., Scott, W.G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Acknowledgements
We thank M. Schumacher (Duke University) and N. Tonthat (Duke University) for valuable reagents and insights into protein expression and purification, B. Wacker for experimental assistance, J. Joutsen for valuable SUMOylation insights, A. Masoudi for assistance in calculating distance difference matrices, R. Davidowitz for developing an artistic representation of HSF models, N. Nicely and the Duke University Crystallography Core Facility, the Duke University Proteomics Core Facility and I. Benjamin (University of Wisconsin at Madison) for the Hsf1−/− Hsf2−/− MEFs. This work was funded by the Duke University Core Facility Voucher Program, United States National Institutes of Health grants T32 GM007105 and R01 NS065890 (D.J.T.), a Senior Visiting Professorship from the Sigrid Jusélius Foundation, Helsinki, Finland (D.J.T.) and The Academy of Finland and The Finnish Cancer Organization (L.S.).
Author information
Authors and Affiliations
Contributions
A.M.J. conceived and performed experiments, analyzed data and wrote the manuscript. C.W.P. assisted in crystallographic data analysis and presentation. L.S. and D.J.T. conceived experiments, analyzed data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Overview of the HSF2 DBD two-site HSE structure.
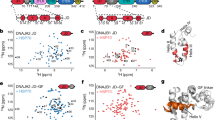
a) Crystals of HSF2 DBD bound to DNA grown overnight at room temperature. Arrow indicates the crystal used for data collection. b) Stereo image of the 2Fo-Fc electron density map of the sequence-specific interaction of HSF2 Arg63 and the guanine of the nGAAn motif contoured at 2.0 σ. c) Alignment of human HSF2 DBD (blue) and K. lactis DBD (gold). d) Illustration of the interaction of HSF2 Lys72, analogous to HSF1 Lys80, interacting with the phosphate backbone through the epsilon amino group in the side chain as well as the peptide backbone
Supplementary Figure 2 The HSF2 wing domain does not contact DNA.
Stereo image of an Fo-Fc simulated annealing omit map contoured at 2.0 σ (green) overlaid with the final 2Fo-Fc electron density map contoured at 0.8 σ (blue). These electron density maps confirm the topology of the wing domain and support the conclusion that the wing domain does not engage in DNA contacts.
Supplementary Figure 3 HSF2 DBD C-terminal contacts with DNA.
a) Lys110 reaches toward the phosphate backbone to engage in water-mediated hydrogen bond network to contact the phosphate of A7, which is located in between the two nGAAn motifs. b) Arg109 utilizes three water molecules to create a hydrogen bond network with the phosphate of G1 and the bases of G2 and T3 of the 2-site HSE sequence. Arg109 also makes a direct contact to the base of G2. c) Sequence alignment of HSFs. The sequence encoding for the carboxyl terminal helix shown in purple in Fig 3 is underlined. Specific residues discussed in Fig 3 are indicated by an asterisk. d) Crystals of HSF2 bound to a 3-site HSE DNA sequence grew in 3-4 days at room temperature. Arrow indicates the crystal that was used for data collection. e) Stereo image of 2Fo-Fc electron density maps for the sequence-specific Arg63 interaction with guanine of the nGAAn motif contoured at 2.0 σ. All four monomers in the asymmetric unit utilize this interaction.
Supplementary Figure 4 In vitro analysis of HSF DBD wing-domain chimeras.
a) Immunoblot analysis of in vitro SUMOylated 6xHis-tagged HSF1 DBD and HSF2 DBD using anti-His tag antibody. HSF1 DBD is not modified by SUMO1 or SUMO2, whereas HSF2 DBD is efficiently modified with the SUMO E3 ligases RANBP2ΔFG or PIAS1, and also in the absence of E3 ligase (UBC9 Only). b) Wing domain sequences of HSF1, HSF2 and HSF4. Underlined sequence delineates the wing domain and the sequences that were used to create the HSF1 and HSF2 DBD chimeras. Shown in red are putative SUMOylation sites. c) In vitro DNA binding analysis of the HSF wing chimera DBDs. DBD protein was titrated into 1 nM FITC-labeled HSE DNA and changes in relative polarization of the sample were measured and converted to binding Kd values. All DBD variants exhibit similar affinity for HSE DNA in vitro.
Supplementary Figure 5 HSF1 and HSF2 directly interact and are cross-linked in a complex.
a) Illustration of the dual cassette used for HSF1 and HSF2 co-expression in E. coli. A bi-directional, lac operated, IPTG- inducible promoter was used to simultaneously express StrepII-tagged HSF1 and 6xHis-tagged HSF2 in the pET15b vector. b) Coomassie staining of size exclusion fractions containing HSF1/2 hetero-complexes from Fig. 7b. c) Coomassie staining of HSF1/2 hetero-complexes following DSS crosslinking. Increasing concentrations of DSS result in the appearance of higher molecular weight species. Numbered boxes indicate bands that were excised for mass spectrometry analysis for the data described in Supplementary Table 1. d) Immunoblot analysis of HSF1 in the purification of HSF1/2 heterocomplexes. Co-expression of WT HSF1 and WT HSF2 results in elution of HSF1 following tandem streptactin and NiNTA affinity chromatography (NE, DUAL). Expression of HSF1ΔLZ1-3 and WT HSF2 does not result in elution of HSF1 following tandem affinity chromatography (NE, DUALΔ1). However, HSF1 is observed in the Streptactin Elution suggesting that HSF1 is lost during NiNTA purification since it is not interacting with 6xHis-HSF2.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5 (PDF 1083 kb)
Supplementary Data Set 1
Mass spectrometry data for Supplementary Fig. 5c (PDF 2367 kb)
Supplementary Data Set 2
Uncropped blot images (XLS 11 kb)
Source data
Rights and permissions
About this article
Cite this article
Jaeger, A., Pemble, C., Sistonen, L. et al. Structures of HSF2 reveal mechanisms for differential regulation of human heat-shock factors. Nat Struct Mol Biol 23, 147–154 (2016). https://doi.org/10.1038/nsmb.3150
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3150
This article is cited by
-
Mitochondria in Cryptococcus: an update of mitochondrial transcriptional regulation in Cryptococcus
Current Genetics (2023)
-
HSF2BP protects against acute liver injury by regulating HSF2/HSP70/MAPK signaling in mice
Cell Death & Disease (2022)
-
Functional diversification of heat shock factors
Biologia Futura (2022)
-
Evolution and co-evolution: insights into the divergence of plant heat shock factor genes
Physiology and Molecular Biology of Plants (2022)
-
CBP-HSF2 structural and functional interplay in Rubinstein-Taybi neurodevelopmental disorder
Nature Communications (2022)