Abstract
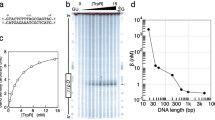
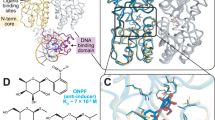
Crystal structures of the Lac repressor, with and without isopropylthiogalactoside (IPTG), and the repressor bound to operator have provided a model for how the binding of the inducer reduces the affinity of the repressor for the operator. However, because of the low resolution of the operator-bound structure (4.8 Å), the model for the allosteric transition was presented in terms of structural elements rather than in terms of side chain interactions. Here we have constructed a dimeric Lac repressor and determined its structure at 2.6 Å resolution in complex with a symmetric operator and the anti-inducer orthonitrophenylfucoside (ONPF). The structure enables the induced (IPTG-bound) and repressed (operator-bound) conformations of the repressor to be compared in atomic detail. An extensive network of interactions between the DNA-binding and core domains of the repressor suggests a possible mechanism for the allosteric transition.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Jacob, F. & Monod, J. J. Mol. Biol. 3, 318–356 (1961).
Monod, J., Wyman, J. & Changeux, J.-P. J. Mol. Biol. 12, 88– 118 (1965).
Gilbert, W. & Muller-Hill, B. Proc. Natl. Acad. Sci. USA 56, 1891–1898 (1966).
Gilbert, W. In Protein-Ligand Interactions (eds Sund, H. & Blauer, G.) 193 (Walterde Gruyter, Berlin; 1975).
Mueller-Hill, B., Rickenberg, H.V. & Wallenfels, K. J. Mol. Biol. 10, 303– 318 (1964).
Jacob, F. & Monod, J. Cold Spring Harbor Symp. Quant. Biol. 26, 193–211 ( 1961).
Riggs, A.D., Newby, R.F. & Bourgeois, S. J. Mol. Biol. 51, 303– 314 (1970).
Lewis, M. et al. Science 271, 1247–1254 (1996).
Friedman, A.M., Fischmann, T.O. & Steitz, T.A. Science 268, 1721– 1727 (1995).
Chuprina, V.P. et al. J. Mol. Biol. 234, 446– 462 (1993).
Slijper, M., Bonvin, A.M., Boelens, R. & Kaptein, R. J. Mol. Biol. 259, 761–773 (1996).
Spronk, C.A., Slijper, M., van Boom, J.H., Kaptein, R. & Boelens, R. Nature Struct. Biol. 3, 916–919 (1996).
Lehming, N., Sartorius, J., Oehler, S., von Wilcken-Bergmann, B. & Muller-Hill, B. Proc. Natl. Acad. Sci. USA 85, 7947–7951 (1988).
Alberti, S., Oehler, S., von Wilcken-Bergmann, B., Kramer, H. & Muller-Hill, B. New Biol. 3, 57–62 (1991).
Chen, J. & Matthews, K.S. J. Biol. Chem. 267 , 13843–13850 (1992).
Markiewicz, P., Kleina, L., Cruz, C., Ehret, S. & Miller, J.H. J. Mol. Biol. 240, 421– 433 (1994).
Spronk, C.A.E.M. et al. Structure 7, 1483– 1492 (1999).
Pace, H.C. et al. Trends Biochem. Sci. 22, 334– 339 (1997).
Barry, J.K. & Matthews, K.S. Biochemistry 38, 3579–3590 (1999).
Falcon, C.M. & Matthews K.S., J. Biol. Chem. 274 , 30849–30857 (1999).
Schumacher, M.A., Choi, K.Y., Zalkin, H. & Brennan, R.G. Science 266, 763–770 (1994).
Schumacher, M.A., Choi, K.Y., Lu, F., Zalkin, H. & Brennan, R.G. Cell 83, 147– 155 (1995).
Navaza, J. Acta Crystallogr. D50, 157–163 (1994).
Jones, T.A., Zhou, J.-Y., Cowan, S.W. & Kjeldgaard, M. Acta Crystallogr. A47, 110–119 ( 1991).
Brünger, A.T. X-PLOR Version 3.1: A system for X-ray Crystallography and NMR. (Yale University Press New Haven, Connecticut; 1992).
Brünger, A.T. et al. Acta Crystallogr. D54, 905– 921 (1998).
Adams, P.D., Pannu, N.S., Read, R.J. & Brunger, A.T. Proc. Natl. Acad. Sci. USA 94, 5018–5023 (1997).
Brünger, A.T. Nature 355, 472–475 ( 1992).
Lee, B. & Richards, F.M. J. Mol. Biol. 55, 379–400 (1971).
Kraulis, P. J. Appl. Crystallogr. 24, 946–950 (1991).
Merritt, E.A. & Bacon, D.J. Methods Enzymol. 277 , 505–524 (1997).
Acknowledgements
We thank G. Van Duyne and P. Lu for useful discussions. This work was supported by an NIH grant to M.L. and an NIH NRSA to C.E.B. The work is based upon research conducted at the NSLS beamline X25 and at the Cornell High Energy Synchrotron Source (CHESS) with the use of the Macromolecular Diffraction at CHESS (MacCHESS) facility.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Bell, C., Lewis, M. A closer view of the conformation of the Lac repressor bound to operator . Nat Struct Mol Biol 7, 209–214 (2000). https://doi.org/10.1038/73317
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/73317
This article is cited by
-
Highly tunable TetR-dependent target gene expression in the acetic acid bacterium Gluconobacter oxydans
Applied Microbiology and Biotechnology (2021)
-
Sequence specificity in DNA–drug intercalation: MD simulation and density functional theory approaches
Journal of Computer-Aided Molecular Design (2020)
-
Computational predictors fail to identify amino acid substitution effects at rheostat positions
Scientific Reports (2017)
-
Design and application of a lactulose biosensor
Scientific Reports (2017)
-
Insights into genome architecture deduced from the properties of short Lac repressor-mediated DNA loops
Biophysical Reviews (2016)