Key Points
-
There is growing concern about the reproducibility of scientific research, and neuroimaging research suffers from many features that are thought to lead to high levels of false results.
-
Statistical power of neuroimaging studies has increased over time but remains relatively low, especially for group comparison studies. An analysis of effect sizes in the Human Connectome Project demonstrates that most functional MRI studies are not sufficiently powered to find reasonable effect sizes.
-
Neuroimaging analysis has a high degree of flexibility in analysis methods, which can lead to inflated false-positive rates unless controlled for. Pre-registration of analysis plans and clear delineation of hypothesis-driven and exploratory research are potential solutions to this problem.
-
The use of appropriate corrections for multiple tests has increased, but some common methods can have highly inflated false-positive rates. The use of non-parametric methods is encouraged to provide accurate correction for multiple tests.
-
Software errors have the potential to lead to incorrect or irreproducible results. The adoption of improved software engineering methods and software testing strategies can help to reduce such problems.
-
Reproducibility will be improved through greater transparency in methods reporting and through increased sharing of data and code.
Abstract
Functional neuroimaging techniques have transformed our ability to probe the neurobiological basis of behaviour and are increasingly being applied by the wider neuroscience community. However, concerns have recently been raised that the conclusions that are drawn from some human neuroimaging studies are either spurious or not generalizable. Problems such as low statistical power, flexibility in data analysis, software errors and a lack of direct replication apply to many fields, but perhaps particularly to functional MRI. Here, we discuss these problems, outline current and suggested best practices, and describe how we think the field should evolve to produce the most meaningful and reliable answers to neuroscientific questions.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Poldrack, R. A. & Farah, M. J. Progress and challenges in probing the human brain. Nature 526, 371–379 (2015).
Logothetis, N. K. What we can do and what we cannot do with fMRI. Nature 453, 869–878 (2008).
Biswal, B. B. et al. Toward discovery science of human brain function. Proc. Natl Acad. Sci. USA 107, 4734–4739 (2010).
Kriegeskorte, N. et al. Matching categorical object representations in inferior temporal cortex of man and monkey. Neuron 60, 1126–1141 (2008).
Norman, K. A., Polyn, S. M., Detre, G. J. & Haxby, J. V. Beyond mind-reading: multi-voxel pattern analysis of fMRI data. Trends Cogn. Sci. 10, 424–430 (2006).
Poldrack, R. A. Inferring mental states from neuroimaging data: from reverse inference to large-scale decoding. Neuron 72, 692–697 (2011).
Ioannidis, J. P. A. Why most published research findings are false. PLoS Med. 2, e124 (2005). This landmark paper outlines the ways in which common practices can lead to inflated levels of false positives.
Simmons, J. P., Nelson, L. D. & Simonsohn, U. False-positive psychology: undisclosed flexibility in data collection and analysis allows presenting anything as significant. Psychol. Sci. 22, 1359–1366 (2011). This paper highlights the impact of common 'questionable research practices' on study outcomes and proposes a set of guidelines to prevent false-positive findings.
Gelman, A. & Loken, E. The statistical crisis in science. American Scientist 102, 40 (2014).
Ioannidis, J. P. A., Fanelli, D., Dunne, D. D. & Goodman, S. N. Meta-research: evaluation and improvement of research methods and practices. PLoS Biol. 13, e1002264 (2015).
Collins, F. S. & Tabak, L. A. Policy: NIH plans to enhance reproducibility. Nature 505, 612–613 (2014).
Button, K. S. et al. Power failure: why small sample size undermines the reliability of neuroscience. Nat. Rev. Neurosci. 14, 365–376 (2013). This paper sounded the first major alarm regarding low statistical power in neuroscience.
Yarkoni, T. Big correlations in little studies: inflated fMRI correlations reflect low statistical power — commentary on Vul et al. (2009). Perspect. Psychol. Sci. 4, 294–298 (2009).
David, S. P. et al. Potential reporting bias in fMRI studies of the brain. PLoS ONE 8, e70104 (2013).
Yarkoni, T., Poldrack, R. A., Nichols, T. E., Van Essen, D. C. & Wager, T. D. Large-scale automated synthesis of human functional neuroimaging data. Nat. Methods 8, 665–670 (2011).
Friston, K. J., Frith, C. D., Liddle, P. F. & Frackowiak, R. S. Comparing functional (PET) images: the assessment of significant change. J. Cereb. Blood Flow Metab. 11, 690–699 (1991).
Jenkinson, M., Beckmann, C. F., Behrens, T. E. J., Woolrich, M. W. & Smith, S. M. FSL. Neuroimage 62, 782–790 (2012).
Worsley, K. J. et al. A unified statistical approach for determining significant signals in images of cerebral activation. Hum. Brain Mapp. 4, 58–73 (1996).
Cheng, D. & Schwartzman, A. Distribution of the height of local maxima of Gaussian random fields. Extremes 18, 213–240 (2015).
Van Essen, D. C. et al. The WU-Minn Human Connectome Project: an overview. Neuroimage 80, 62–79 (2013).
Tong, Y. et al. Seeking optimal region-of-interest (ROI) single-value summary measures for fMRI studies in imaging genetics. PLoS ONE 11, e0151391 (2016).
Devlin, J. T. & Poldrack, R. A. In praise of tedious anatomy. Neuroimage 37, 1033–1041 (2007).
Desikan, R. S. et al. An automated labeling system for subdividing the human cerebral cortex on MRI scans into gyral based regions of interest. Neuroimage 31, 968–980 (2006).
Durnez, J. et al. Power and sample size calculations for fMRI studies based on the prevalence of active peaks. Preprint at bioRxiv http://dx.doi.org/10.1101/049429 (2016).
Mumford, J. A. & Nichols, T. E. Power calculation for group fMRI studies accounting for arbitrary design and temporal autocorrelation. Neuroimage 39, 261–268 (2008).
Mennes, M., Biswal, B. B., Castellanos, F. X. & Milham, M. P. Making data sharing work: the FCP/INDI experience. Neuroimage 82, 683–691 (2013).
Thompson, P. M. et al. The ENIGMA Consortium: large-scale collaborative analyses of neuroimaging and genetic data. Brain Imaging Behav. 8, 153–182 (2014).
Rohlfing, T. & Poline, J.-B. Why shared data should not be acknowledged on the author byline. Neuroimage 59, 4189–4195 (2012).
Austin, M. A., Hair, M. S. & Fullerton, S. M. Research guidelines in the era of large-scale collaborations: an analysis of Genome-wide Association Study Consortia. Am. J. Epidemiol. 175, 962–969 (2012).
Savoy, R. L. Using small numbers of subjects in fMRI-based research. IEEE Eng. Med. Biol. Mag. 25, 52–59 (2006).
Poldrack, R. A. et al. Long-term neural and physiological phenotyping of a single human. Nat. Commun. 6, 8885 (2015).
Kerr, N. L. HARKing: hypothesizing after the results are known. Pers. Soc. Psychol. Rev. 2, 196–217 (1998).
Nosek, B. A. et al. Promoting an open research culture. Science 348, 1422–1425 (2015).
Chambers, C. D., Dienes, Z., McIntosh, R. D., Rotshtein, P. & Willmes, K. Registered reports: realigning incentives in scientific publishing. Cortex 66, A1–A2 (2015).
Sidén, P., Eklund, A., Bolin, D. & Villani, M. Fast Bayesian whole-brain fMRI analysis with spatial 3D priors. Neuroimage 146, 211–225 (2016).
Carp, J. On the plurality of (methodological) worlds: estimating the analytic flexibility of FMRI experiments. Front. Neurosci. 6, 149 (2012). This paper reports analyses of a single data set using 6,912 different analysis workflows, highlighting the large degree of variability in results across analyses in some brain regions.
Penny, W. D., Friston, K. J., Ashburner, J. T., Kiebel, S. J. & Nichols, T. E. Statistical Parametric Mapping: The Analysis of Functional Brain Images (Elsevier Science, 2011).
Cox, R. W. AFNI: what a long strange trip it's been. Neuroimage 62, 743–747 (2012).
Heininga, V. E., Oldehinkel, A. J., Veenstra, R. & Nederhof, E. I just ran a thousand analyses: benefits of multiple testing in understanding equivocal evidence on gene-environment interactions. PLoS ONE 10, e0125383 (2015).
Chambers, C. D., Feredoes, E., Muthukumaraswamy, S. D. & Etchells, J. P. Instead of 'playing the game' it is time to change the rules: Registered Reports at AIMS Neuroscience and beyond. AIMS Neurosci. 1, 4–17 (2014).
Muthukumaraswamy, S. D., Routley, B., Droog, W., Singh, K. D. & Hamandi, K. The effects of AMPA blockade on the spectral profile of human early visual cortex recordings studied with non-invasive MEG. Cortex 81, 266–275 (2016).
Hobson, H. M. & Bishop, D. V. M. Mu suppression — a good measure of the human mirror neuron system? Cortex 82, 290–310 (2016).
Churchill, N. W. et al. Optimizing preprocessing and analysis pipelines for single-subject fMRI: 2. Interactions with ICA, PCA, task contrast and inter-subject heterogeneity. PLoS ONE 7, e31147 (2012).
Bennett, C. M., Miller, M. B. & Wolford, G. L. Neural correlates of interspecies perspective taking in the post-mortem Atlantic Salmon: an argument for multiple comparisons correction. Neuroimage 47, S125 (2009).
Eklund, A., Nichols, T. E. & Knutsson, H. Cluster failure: why fMRI inferences for spatial extent have inflated false-positive rates. Proc. Natl Acad. Sci. USA 113, 7900–7905 (2016). This paper shows that some commonly used methods for cluster-based multiple-comparison correction can exhibit inflated false-positive rates.
Nichols, T. & Hayasaka, S. Controlling the familywise error rate in functional neuroimaging: a comparative review. Stat. Methods Med. Res. 12, 419–446 (2003).
Wager, T. D., Lindquist, M. & Kaplan, L. Meta-analysis of functional neuroimaging data: current and future directions. Soc. Cogn. Affect. Neurosci. 2, 150–158 (2007).
Lieberman, M. D. & Cunningham, W. A. Type I and Type II error concerns in fMRI research: re-balancing the scale. Soc. Cogn. Affect. Neurosci. 4, 423–428 (2009).
Bennett, C. M., Wolford, G. L. & Miller, M. B. The principled control of false positives in neuroimaging. Soc. Cogn. Affect. Neurosci. 4, 417–422 (2009).
Hayasaka, S. & Nichols, T. E. Validating cluster size inference: random field and permutation methods. Neuroimage 20, 2343–2356 (2003).
Gorgolewski, K. J. et al. NeuroVault.org: a web-based repository for collecting and sharing unthresholded statistical maps of the human brain. Front. Neuroinform. 9, 8 (2015).
Hunt, L. T., Dolan, R. J. & Behrens, T. E. J. Hierarchical competitions subserving multi-attribute choice. Nat. Neurosci. 17, 1613–1622 (2014).
Shehzad, Z. et al. A multivariate distance-based analytic framework for connectome-wide association studies. Neuroimage 93 (Pt.1), 74–94 (2014).
Rubinov, M. & Sporns, O. Complex network measures of brain connectivity: uses and interpretations. Neuroimage 52, 1059–1069 (2010).
Craddock, R. C., Milham, M. P. & LaConte, S. M. Predicting intrinsic brain activity. Neuroimage 82, 127–136 (2013).
Butler, R. W. & Finelli, G. B. The infeasibility of quantifying the reliability of life-critical real-time software. IEEE Trans. Software Eng. 19, 3–12 (1993).
Cox, R. W., Reynolds, R. C. & Taylor, P. A. AFNI and clustering: false positive rates redux. Preprint at bioRxiv http://dx.doi.org/10.1101/065862 (2016).
Waskom, M. L., Kumaran, D., Gordon, A. M., Rissman, J. & Wagner, A. D. Frontoparietal representations of task context support the flexible control of goal-directed cognition. J. Neurosci. 34, 10743–10755 (2014).
Poldrack, R. A. et al. Guidelines for reporting an fMRI study. Neuroimage 40, 409–414 (2008).
Carp, J. The secret lives of experiments: methods reporting in the fMRI literature. Neuroimage 63, 289–300 (2012).
Guo, Q. et al. The reporting of observational clinical functional magnetic resonance imaging studies: a systematic review. PLoS ONE 9, e94412 (2014).
Nichols, T. E. et al. Best practices in data analysis and sharing in neuroimaging using MRI. Preprint at bioRxiv http://dx.doi.org/10.1101/054262 (2016).
Poldrack, R. A. Can cognitive processes be inferred from neuroimaging data? Trends Cogn. Sci. 10, 59–63 (2006).
Gelman, A. & Stern, H. The difference between 'significant' and 'not significant' is not itself statistically significant. Am. Stat. 60, 328–331 (2006).
Nieuwenhuis, S., Forstmann, B. U. & Wagenmakers, E.-J. Erroneous analyses of interactions in neuroscience: a problem of significance. Nat. Neurosci. 14, 1105–1107 (2011).
Boekel, W. et al. A purely confirmatory replication study of structural brain–behavior correlations. Cortex 66, 115–133 (2015).
Begley, C. G. & Ellis, L. M. Drug development: raise standards for preclinical cancer research. Nature 483, 531–533 (2012).
Open Science Collaboration. Estimating the reproducibility of psychological science. Science 349, aac4716 (2015). This paper reports a large-scale collaboration that quantified the replicability of research in psychology, showing that less than half of the published findings were replicable.
Zuo, X.-N. et al. An open science resource for establishing reliability and reproducibility in functional connectomics. Sci. Data 1, 140049 (2014).
Poldrack, R. A. et al. Toward open sharing of task-based fMRI data: the OpenfMRI project. Front. Neuroinform. 7, 1–12 (2013).
Gil, Y. et al. Toward the geoscience paper of the future: best practices for documenting and sharing research from data to software to provenance. Earth Space Sci. 3, 388–415 (2016).
Boulesteix, A.-L. Ten simple rules for reducing overoptimistic reporting in methodological computational research. PLoS Comput. Biol. 11, e1004191 (2015).
Gorgolewski, K. J. et al. The brain imaging data structure, a format for organizing and describing outputs of neuroimaging experiments. Sci. Data 3, 160044 (2016).
Flint, J. & Munafò, M. R. Candidate and non-candidate genes in behavior genetics. Curr. Opin. Neurobiol. 23, 57–61 (2013).
Ioannidis, J. P., Tarone, R. & McLaughlin, J. K. The false-positive to false-negative ratio in epidemiologic studies. Epidemiology 22, 450 (2011).
Burgess, S. et al. Using published data in Mendelian randomization: a blueprint for efficient identification of causal risk factors. Eur. J. Epidemiol. 30, 543–552 (2015).
Stein, J. L. et al. Identification of common variants associated with human hippocampal and intracranial volumes. Nat. Genet. 44, 552–561 (2012).
Barch, D. M. et al. Function in the human connectome: task-fMRI and individual differences in behavior. Neuroimage 80, 169–189 (2013).
Acknowledgements
R.A.P., J.D., J.-B.P. and K.J.G. are supported by the Laura and John Arnold Foundation. J.D. has received funding from the European Union's Horizon 2020 research and innovation programme under the Marie Sklodowska-Curie grant agreement No 706561. M.R.M. is supported by the Medical Research Council (MRC) (MC UU 12013/6) and is a member of the UK Centre for Tobacco and Alcohol Studies, a UK Clinical Research Council Public Health Research Centre of Excellence. Funding from the British Heart Foundation, Cancer Research UK, the Economic and Social Research Council, the MRC and the National Institute for Health Research, under the auspices of the UK Clinical Research Collaboration, is gratefully acknowledged. C.I.B. is supported by the Intramural Research Program of the US National Institutes of Health (NIH)–National Institute of Mental Health (NIMH) (ZIA-MH002909). T.Y. is supported by the NIMH (R01MH096906). P.M.M. acknowledges personal support from the Edmond J. Safra Foundation and Lily Safra and research support from the MRC, the Imperial College Healthcare Trust Biomedical Research Centre and the Imperial Engineering and Physical Sciences Research Council Mathematics in Healthcare Centre. T.E.N. is supported by the Wellcome Trust (100309/Z/12/Z), NIH–National Institute of Neurological Disorders and Stroke (R01NS075066) and NIH–National Institute of Biomedical Imaging and Bioengineering (NIBIB) (R01EB015611). J.-B.P. is supported by the NIBIB (P41EB019936) and by NIH–National Institute on Drug Abuse (U24DA038653). Data were provided (in part) by the Human Connectome Project, WU-Minn Consortium (principal investigators: D. Van Essen and K. Ugurbil; 1U54MH091657), which is funded by the 16 Institutes and Centers of the NIH that support the NIH Blueprint for Neuroscience Research, and by the McDonnell Center for Systems Neuroscience at Washington University. The authors thank J. Wexler for performing annotation of Neurosynth data, S. David for providing sample-size data, and R. Cox and P. Taylor for helpful comments on a draft of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
PowerPoint slides
Supplementary information
Supplementary information S1 (figure)
A depiction of the data from Figure 1 showing all data points. (PDF 177 kb)
Glossary
- Linear mixed-effects analysis
-
An analysis in which some measured independent variables are treated as randomly sampled from the population, in contrast to a traditional fixed-effects analysis, in which all predictors are treated as fixed and known.
- Familywise error
-
(FWE). The probability of at least one false positive among multiple statistical tests.
- Random field theory
-
The theory describing the behaviour of geometric points on a random topological space.
- Euler characteristic
-
A topological measure that is used to describe the set of thresholded voxels in the context of random field theory.
- False discovery rate
-
(FDR). The expected proportion of false positives among all significant findings when performing multiple statistical tests.
- Functional localizer
-
An independent scan that is used to identify regions on the basis of their functional response; for example, for the responses of face-responsive regions to faces.
- Bayesian methods
-
An approach to statistical analysis focusing on updating beliefs via probability distributions and symmetrically comparing candidate models.
- Mass univariate testing
-
An approach to the analysis of multivariate data in which the same model is fit to each element of the observed data (for example, each voxel).
- Permutation tests
-
Also known as randomization tests. Approaches for testing statistical significance by comparing to a null distribution that is obtained by rearranging the labels of the observed data.
- 'Not invented here' philosophy
-
The philosophy that any solution to a problem that was developed by someone else is necessarily inferior and must be re-engineered from scratch.
- Interpolation
-
The operation by which a function is applied to the sampled data to obtain estimates of the data at positions where data have not been sampled.
- Software container
-
A self-contained software tool that encompasses all of the necessary software and dependencies to run a particular program.
Rights and permissions
About this article
Cite this article
Poldrack, R., Baker, C., Durnez, J. et al. Scanning the horizon: towards transparent and reproducible neuroimaging research. Nat Rev Neurosci 18, 115–126 (2017). https://doi.org/10.1038/nrn.2016.167
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrn.2016.167
This article is cited by
-
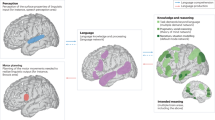
The language network as a natural kind within the broader landscape of the human brain
Nature Reviews Neuroscience (2024)
-
The PREMISE database of 20 Macaca fascicularis PET/MRI brain images available for research
Lab Animal (2024)
-
A research agenda for understanding how social inequality is linked to brain structure and function
Nature Human Behaviour (2024)
-
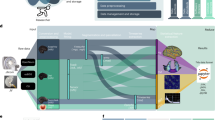
ezBIDS: Guided standardization of neuroimaging data interoperable with major data archives and platforms
Scientific Data (2024)
-
A neurofunctional signature of subjective disgust generalizes to oral distaste and socio-moral contexts
Nature Human Behaviour (2024)