Key Points
-
Emerging infectious diseases (including community-, hospital- and bioterrorism-acquired infections), emerging resistance to antimicrobial agents and increased social demand are increasing the volume and altering the nature of the activities required from clinical microbiology laboratories.
-
Centralization, an increase in automation and advances in bioinformatics allow clinical microbiology laboratories to keep up with these ever-increasing demands.
-
Technologies and techniques that are progressing at the moment include rapid molecular detection, identification and genotyping of bacteria; antimicrobial-resistance determination; rapid immunological detection of pathogens; easy-to-use electron microscopy; and data digitalization and the secure online exchange of information.
-
The future evolution of clinical microbiology might include the spread of 'at-doctor' tests and bedside tests at the same time as specialized diagnoses are centralized in reference laboratories that are connected on national and international scales. Centralization should allow the development of P3/P4 laboratories, molecular-biology platforms, including mass spectrometry, and serology platforms, including antigenic microarrays for serodiagnosis.
-
Sampling strategies might evolve towards pathology-based sampling kits in accordance with the development of multiplex platforms. In addition, data reporting could be based solely on digitalized figures and could include data interpretation and the addition of electronic links to up-to-date literature, which can be exchanged in a timely manner through the Internet.
-
Large clinical microbiology laboratories could engage in the regular reporting of epidemiological trends for pathogens, pathogen subtypes and antimicrobial resistance.
-
These anticipated changes will require the advanced training of technicians in bioinformatics, and the creation of posts for maintenance workers and engineers for the continuous implementation of new techniques. Clinical microbiologists will have an increased role in communicating with infectious-disease practitioners in the interpretation and delivery of results, with consultants in diagnosis and antimicrobial treatment, and in strategic laboratory management.
Abstract
In the past decade, clinical microbiology laboratories have undergone important changes with the introduction of molecular biology techniques and laboratory automation. In the future, there will be a need for more rapid diagnoses, increased standardization of testing and greater adaptability to cope with new threats from infectious microorganisms, such as agents of bioterrorism and emerging pathogens. The combination of the new tools that are now being developed in research laboratories, the general reorganization of clinical laboratories and improved communication between physicians and clinical microbiologists should lead to profound changes in the way that clinical microbiologists work.
Similar content being viewed by others
Main
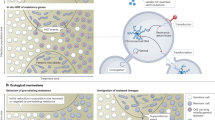
Clinical microbiology evolves in response to clinical needs1 (Fig. 1) and there have been many advances in this area during the past decade. For example, automated systems for the detection and identification of microorganisms and for the identification of resistant organisms have been developed2. These include automated blood-culture assays and systems for the phenotypic identification of bacteria. Antibiotic-sensitivity testing and computerized interpretation systems have also been developed3. These developments have not led to significant changes in the microbiology itself, but rather enable more samples to be processed by fewer personnel and provide increased intra- and inter-laboratory reproducibility. Similar developments have been seen with immunoassays: enzyme-linked immunosorbant assays (ELISAs) have been extensively automated and standardized in recent years and have also become cheaper. Many of the automated tests that are now available were initially designed for the mass testing of blood that is destined for blood banks, but later innovations led to western blots for the confirmation of HIV infection and Lyme disease and the development of tests for the rapid detection of anti-HIV antibodies in saliva or blood2. Worldwide — mainly because of the cost of containment measures for infectious diseases — there has been a general tendency over the past 10 years to centralize clinical microbiology laboratories such that they serve several hospitals, therefore maximizing the efficiency of testing at the lowest cost4. However, such centralization of clinical microbiology laboratories can result in a lack of consultation between physicians and clinical scientists.
There have also been rapid developments in the genetic detection and identification of microbial pathogens during this period, mainly through nucleic-acid amplification by polymerase chain reaction (PCR). PCR using universal primers followed by sequencing of the amplification products has enabled the rapid identification of cultured and uncultured bacteria and fungi. This procedure is now available in many large university hospital laboratories and is also carried out by new companies in the private sector. Genetic testing can also be used to identify DNA sequences that are associated with antimicrobial resistance. Viral loads can also be determined by PCR and this has been found to be a useful method for evaluating antiviral therapies. Finally, molecular methods allow typing of microorganisms, which can help to analyse the epidemiology of outbreaks and identify their sources, as well as distinguishing between relapse and re-infection in the case of persisting microorganisms. Indeed, infection control is now considered to be part of clinical microbiology.
In the future, new technologies, changes in the availability of skilled personnel and the prevailing economic environment will all influence clinical microbiology5. The use of molecular-biology techniques has dramatically changed clinical microbiology and the integration of the two disciplines is an important challenge for the future. In addition to detecting well-known pathogens, the laboratories of the future will also have to be able to recognize new pathogens and participate in food-safety monitoring and bioterrorism surveillance4. Recent experience has shown the necessity for laboratories to be able to rapidly develop diagnostic tests for new diseases that have a high social impact, as was the case with the outbreak of severe acute respiratory syndrome (SARS). Means must also be available for the rapid transfer of such technology to laboratories in which routine diagnostics are carried out. A current problem is that scientific responses to emerging threats are far more rapid than are administrative responses, and there are often prolonged delays in the approval of new diagnostic tests for use outside research laboratories.
In this article, we discuss the latest techniques that are available to clinical microbiologists (Fig. 2) and how they are applied, and speculate about what the future might hold for clinical microbiology laboratories.
Current technologies in clinical microbiology
Sequence-based identification of microorganisms. The availability of high-throughput thermal cyclers, new DNA polymerases and automated sequencers, together with reductions in the cost of reagents, has led to the sequence-based identification of microorganisms becoming a useful and reliable alternative to phenotypic methods of identification. Moreover, the patent on Taq polymerase will expire in March 2005 and this should further decrease the cost of PCR and sequencing. The development of 'molecular microbiology' — and therefore progress in clinical microbiology — has been linked to the development of online databases. Scientists worldwide can now send their own sequences to these databases and there has been an explosion in the amount of sequence data that are available. Of the databases that are available at present, GenBank6 is the largest and the most versatile. On 1 September 2003, GenBank contained 22,318,883 DNA sequences, including the complete genomes of 34 archaea, 128 bacteria, 19 eukaryotes and 1,174 viruses.
Strict criteria must be used when interpreting sequence data. A major drawback of online databases is that they are dependent on the quality of the sequences that are submitted by scientists. Inaccurately reported sequences can lead to individual microorganisms being misidentified and even to whole groups of organisms requiring reclassification. Moreover, the availability of the current data could potentially become compromised in the future and common international repositories would be useful.
For bacteria, studies of 'universal' genes — such as the 16S ribosomal RNA gene — have enabled the correct classification of numerous microorganisms, including some that were previously unidentifiable7. Unfortunately, it is not possible to discriminate between all species by comparing their 16S rRNA gene sequences. At present, we know of 75 genes that are present in all bacterial genomes8. The sequences of some of these genes — for example, rpoB (which encodes the RNA polymerase β-subunit)9 and the SIGMA-FACTOR genes10 — differ between bacterial species and can therefore be used for bacterial identification. Large databases of the sequences of these genes are now available. For eukaryotes, the 18S rRNA gene has also been used as a 'universal' gene and there is an increasing number of 18S rRNA sequences in databases11. By contrast, there is no common gene for the detection of all viruses, although PCR targets have been developed for virus families, such as the herpesviridae12.
'Universal' genes cannot be used when a higher level of identification is required, such as strain typing in investigations of disease outbreaks, antibiotic resistance, nosocomial infections or suspected bioterrorism attacks. Different gene sequences enable different levels of identification. Depending on the gene that is used, the genus, species, subspecies or even strain of an organism can be established. When it is crucial to determine whether one or more clinical isolates are of the same strain, a comparison of several genome sequences might be required. Such studies have identified highly variable genome fragments and have led to the development of several techniques, including MULTILOCUS SEQUENCE TYPING13. Similar identification tools have also been developed for fungi14 and protozoa15. Other sequence-based typing techniques have used intergenic, non-coding spacer sequences16 that are more likely to be variable. The one that has been the most widely used is the 16S–23S intergenic spacer for bacteria16 and the INTERNAL TRANSCRIBED SPACER (ITS) for fungi17. Specific, fully automated bioinformatics tools, such as BIBI (Bioinformatics Bacterial Identification)20, identify microorganisms by combining sequence databases and phylogeny-display programmes.
Quantitative PCR. The development of molecular beacons and other fluorescent probes (such as the TaqMan® probe that is produced by Roche Molecular Systems, Inc.) has made real-time monitoring of PCR amplification and quantification of nucleic acids in clinical specimens possible21. Beacons are hairpin-shaped oligonucleotide probes that have a fluorophore that emits photons and is closely associated with a quencher, which absorbs the photons. In the presence of a complementary strand of DNA, the hairpin structure unfolds and fluorescence is emitted, and this can be measured to determine the amount of DNA that is present. Multiple targets can be monitored22 using fluorophores with different emission spectra. This technology, which is highly sensitive, is commercially available as the TaqMan® (Applied Biosystems; Perkin-Elmer Corp.) and LightCycler (Roche Diagnostics Corp.) systems. The advantages of real-time PCR are the rapidity of the assay (30–40 min), the ability to quantify and identify PCR products directly without the use of agarose gels, and the fact that contamination of the nucleic acids can be limited. Kits are available for the detection and quantification of DNA and RNA in clinical samples, and the technique has been specifically developed to enable the follow-up of patients with HIV and hepatitis C infections (Amplitech AME Bioscience; Bayer Diagnostics; Roche Diagnostics).
DNA microarrays. The development of microarray analysis has been a milestone in several areas of microbiology, and in clinical microbiology microarrays are used for microorganism detection and identification and gene-expression analysis. Microarrays consist of many probes that are discretely located on a solid substrate, such as glass. The probes can be PCR products or oligonucleotides and the 'targets' can be PCR products, genomic DNA, total RNA, amplified RNA (aRNA), complementary DNA, plasmid DNA, bacterial suspensions or clinical specimens23.
When PCR is used to detect DNA in clinical specimens, microarrays can then be used to identify the amplified products by hybridization to an array that is composed of pathogen-specific probes. Using broad-range primers, such as those that amplify the 16S rRNA gene, a single PCR can be used to detect multiple pathogens simultaneously. DNA microarrays have been used to detect bacteria and viruses24,25 and, because the number of possible hybridizations is now very high, sequencing of a PCR product on an array is possible. DNA microarrays can also be used for strain identification or genotyping if the arrays incorporate species-specific gene sequences and the reactivity profiles of isolates and reference strains are compared26.
Antigen microarrays for serology. Assays with the ability to detect specific antibodies against several pathogens simultaneously have a wide application in the diagnosis of infectious diseases27,28. The indirect immunofluorescence assay (IFA) is commonly used to detect antibodies to infectious agents29 as it is easy to use, sensitive and specific. Recently, protein arrays have been used to develop comparative fluorescence assays to measure the concentrations of specific proteins and antibodies in complex solutions30. The lack of stability of fluorescent dyes is no longer a problem with the use of new-generation Alexa Fluor® dyes (Molecular Probes), which are available in several emission colours and allow multiple stainings of the same spot. Specific fluorescence is detected by confocal scanning microscopy and is quantified using internal calibration curves. The dyes have enabled the development of prototypes of antigenic microarrays to detect antibodies in human sera that are specific for one or several pathogens. An antigen microarray for Lyme disease has been described31. Similarly, a microarray with specific antigens of Toxoplasma gondii, cytomegalovirus, herpesvirus 1 and 2 and rubella virus gave favourable results when used to detect reactive immunoglobulin G32. Such pioneering studies provide evidence that antigen-microarray analysis could become an important diagnostic tool. The technique will enable IFAs to be standardized and inter-operator and inter-laboratory variation will be minimized. It will also be possible to digitalize IFA photographs, which will enable them to be stored and exchanged more easily.
Mass spectrometry. Mass spectrometry is an analytical technique that is used for the detection of minute quantities of a molecule in a complex medium33. Refinements in instrumentation have facilitated the use of mass spectrometry in clinical microbiology laboratories. It can be used to sequence PCR amplification products instead of SANGER'S METHOD, which is used at present. The molecular identification of bacteria can be based on the sequencing of the universal 16S rRNA gene34, which is a two-day task (including culture time) that can only be performed using pure cultures of bacteria. Mass-spectrometry analysis of base-specific fragmentation patterns of PCR-amplified DNA has recently been studied as a technique for the rapid identification of bacterial isolates and for the detection of specific 16S rRNA gene fragments that are amplified from complex environmental samples35. In this study, 300 base pairs were determined, and the technique shows promise of becoming the standard method for identification of pathogens, strain typing and determination of mutations that are associated with resistance in bacteria and viruses.
Mass spectrometry has also been important in the development of proteomics, in which it has been used to identify and, increasingly, quantify large numbers of proteins in complex samples. A particular form of protein-chip array technology combined with time-of-flight mass spectrometry — called surface-enhanced laser-desorption/ionization time-of-flight mass spectrometry (SELDI-TOF-MS) — allows the determination of complex protein patterns in serum, body fluids and tissue samples36. This technique offers the advantages of speed, simplicity and sensitivity in conditions that are hard to diagnose, such as Alzheimer's disease37, cancers38,39,40 and autoimmune diseases37,41. Its use for the diagnosis of infectious diseases has been anticipated42, but no experimental data have been published so far.
Microscopy. Microscopy techniques have evolved with respect to three factors: the nature of the signal, the user-friendliness of the equipment, and the digitalization and computerized processing of the image. Electron microscopes — which were once complicated instruments that were only used in research — have been simplified and are now increasingly being used in clinical microbiology laboratories. They are useful for the rapid screening of routine clinical samples, such as stool samples, and also for detecting viruses in tissue cultures. With negative staining, a viral pathogen can be identified within 10 minutes of the arrival of a clinical or tissue-culture specimen in an electron-microscopy laboratory43. Electron microscopy was used to rapidly identify Bacillus anthracis during the recent anthrax letters attack in the United States43 and also to identify the genus (Coronavirus) of the SARS virus. The implementation of new technologies, such as automated pattern recognition44, remote operation of microscopes and telemicroscopy — which uses digital image acquisition and review of images through the internet45 — will further increase the usefulness of electron microscopy.
Flow cytometry. Flow cytometry (FCM) was originally developed as an automated method for measuring the optical or fluorescence characteristics of cells or particles in suspension46. It was used as early as during the Second World War by the United States army to detect bacteria and spores. Flow cytometers are now more user-friendly and less expensive, and are becoming increasingly important in clinical microbiology47.
Almost all microorganisms can be detected directly in clinical specimens by FCM. Bacteria can be detected as particles in liquid samples in the case of bacteraemia and bacteriuria, and can also be identified using polyclonal or monoclonal antibodies that are coated with fluorescent molecules. FCM has been applied successfully to the detection of a wide range of bacteria and to patient specimens, including blood, urine, EXUDATES and EFFUSIONS, bile, BRONCHIAL-LAVAGE FLUID and even faeces47. Flow cytometers that are designed specifically for urine analysis are now available commercially, and these simultaneously determine the concentrations of bacteria, red and white blood cells and squamous cells (for example, the UF100 flow cytometer that is manufactured by Sysmex Corp.). In eukaryotes, FCM studies have mostly been used for Candida spp. and Giardia lamblia.
Because of their small size, viruses cannot be detected directly using FCM, but can be detected indirectly in infected cells in clinical samples or after inoculation and culture of viruses in cell culture. FCM detects and quantifies viral antigens on the surface of, or within, infected cells. FCM can also detect and quantify viral nucleic acids by in situ hybridization of specific viral nucleic acids in cell suspensions and can simultaneously identify the infected cells by cell phenotyping. This application of FCM has been used for several viruses, including HIV and cytomegalovirus47.
For serological studies, antigen-coated microspheres can be incubated with sera, and a fluorescence-conjugated antibody against human IgG added in a second step. FCM can then be used to detect the fluorescent microspheres47. FCM can also simultaneously detect antibodies to more than one antigen by coating different antigens on different-sized particles. FCM can even analyse the antibody response to HIV by simultaneously detecting specific antibodies to the various antigens of the virus.
Given the versatility of FCM, it is likely that the number of applications of this technique in clinical microbiology will increase in the future. The growth of microorganisms could be evaluated and FLUOROGENIC-SUBSTRATE-HYDROLYSIS FCM3 could also be used. Similarly, the development of portable flow cytometers could be useful in detecting acts of biowarfare.
New techniques and adaptations
Detection of microorganisms in clinical specimens. One of the main challenges of clinical microbiology is the identification of microorganisms in clinical specimens. This can be achieved using the following methods: i) growth of microorganisms in culture, using media or cell lines, with varying incubation times, temperatures and atmospheres. Experience has shown that the hypothesis that some microorganisms cannot be grown in the laboratory might not necessarily be true48,49,50; ii) detection of microorganisms within infected tissues by light or electron microscopy; and iii) detection and identification of previously unknown DNA or RNA fragments or antigens. The most appropriate test depends on the incidence of infection, how contagious the agent is and the health consequences for the patient and the community.
Cell-culture systems have made a crucial contribution to the diagnosis of infectious diseases that are caused by viruses and intracellular bacteria. Such systems have not only made the direct isolation and identification of microorganisms possible, but have also enabled the production of antigens for serological assays, evaluation of the efficacy of antimicrobial agents and the production of vaccines50. Because no single cell-culture system has so far been found to be suitable for the isolation of all intracellular microorganisms, the search for other susceptible cell lines will continue. Another source of progress in the development of culture media is the analysis of genome sequences. The availability of these sequences allows any important metabolic pathways that are missing to be identified, and could therefore permit the design of culture media that are specific for a particular microorganism49.
The most spectacular advance in the diagnosis of infectious diseases has been the recent introduction of molecular detection methods, especially PCR and RT-PCR (PCR after reverse transcription of RNA). Molecular techniques also enable the detection of multiple infectious agents — including bacteria, viruses and fungi — by MULTIPLEX PCR74, microarrays (which can detect all potential agents in an infection51) and quantitative assays, such as real-time PCR52.
The availability of complete genome sequences in computerized databases has facilitated the rational selection of PCR and sequencing primers. Primer specificity can be tested using in silico tools such as BLAST (basic local alignment search tool)53. Repeated sequences can be selected to increase the sensitivity of detection54. Another advantage is the facility to select new, specific sets of primers when contamination is encountered in a PCR assay (for example, for SUICIDE PCR55).
A major pitfall in the molecular detection of microorganisms is the number of false-positive results due to contamination in 'home-made' PCRs. This problem can be overcome by fully automating the process, from processing the sample to sequence or molecular identification. The other current limitation is the presence in clinical samples of inhibitors that are not removed completely by boiling or adding bovine serum albumin56.
In the case of viruses, the discovery of the hepatitis C virus in 1989 triggered the use of molecular methods to search for unknown viral agents in clinical specimens57 and raised the suggestion that many human diseases of unknown aetiology might be caused by unrecognized viruses. The hepatitis C and E viruses were identified by immunoscreening or differential hybridization of libraries that were constructed without any amplification57,58. The human TT virus was characterized using REPRESENTATIONAL DIFFERENCE ANALYSIS59. Other methods that might be useful for detecting unknown viruses include the DNase treatment of serum followed by SEQUENCE-INDEPENDENT SINGLE-PRIMER AMPLIFICATION (SISPA)60.
Detection of antibodies. Serum is easy to obtain, store and send on filter paper for analysis61. The indirect diagnosis of infectious diseases by the detection of specific antibodies will continue to be a key approach for clinical microbiologists. Both individual, ready-to-use assays that would be available in the doctor's surgery and platform tests for high-throughput screening and diagnosis could be developed. 'At-doctor' serology tests would be useful for the rapid determination of the status of a patient and for speedy decision-making regarding post-exposure prophylaxis and treatment. In addition, the rapid detection of specific IgMs during epidemics could help to improve isolation measures for patients who are suspected to be infected. Antigenic microarrays will enable testing for several pathogens in parallel using little serum and few reagents. All the pathogens that are known to be causative agents of a particular clinical syndrome could therefore be tested for simultaneously using an antigenic microarray. Moreover, the versatility of this new assay will allow emerging pathogens to be quickly incorporated into the test. Future developments in antigenic microarray technology will include the development of automation for the indirect diagnosis of infectious diseases. The microarray apparatus could be fully automated up to the result output; generation of output would be rapid, as the overall process will probably take less than 2 hours, and automated microarrays could test several parameters simultaneously (for example, testing for antigens and different classes of antibody), thus allowing pathology-driven testing instead of the pathogen-driven testing that is carried out at present. As several antigens can be tested for simultaneously, this development will lead to a severalfold decrease in labour time and, consequently, a reduction in cost. These automated microarrays will be suitable both for mass screening of sera in epidemiology studies and in blood banks, and for diagnostics that are carried out on single serum samples in clinical microbiology laboratories.
Antimicrobial-susceptibility testing. Two strategies are available for determining antimicrobial susceptibility: phenotypic and genomic methods. Phenotypic methods for bacteria include dilution and disc-diffusion methods, and real-time PCR has been used for fastidious bacteria that can only be grown in tissue culture62,63,64.
FCM has proved to be very useful in studying the physiological effects of antimicrobial agents on bacterial cells. It can also be used for antimicrobial-susceptibility testing and can indicate BACTERICIDAL and BACTERIOSTATIC effects. Results are obtained rapidly — often in less than 4 hours — and mixed populations that might respond to antimicrobial agents in different ways can be studied. Several commercial companies have produced FCM kits that enable antibiotic-susceptibility testing to be carried out rapidly in clinical microbiology laboratories. New developments in the field of FCM antibiotic-susceptibility testing include assays for slow-growing bacteria, such as mycobacteria65.
Drug resistance in parasites, although it is an increasingly recognized phenomenon, is not generally tested for in clinical laboratories at present because of a lack of standardized methods. FCM has been used successfully to detect resistance to antimalarial drugs in Plasmodium spp.66 In the case of viruses, FCM has been used to screen and understand the activity of antiviral compounds and to test the susceptibility of clinical isolates of HIV.
Detection of the genes or mutations that are responsible for resistance to antimicrobial agents is an alternative to phenotypic detection and can be carried out directly on clinical specimens. Molecular tests for resistance have been developed commercially for bacteria such as Staphylococcus aureus (LightCycler® Staphylococcus Kit M Grade; Roche), in which resistance to oxacillin is primarily mediated by the mecA gene67. They have also been developed for viruses — for example, for HIV-1 (Trugene HIV-1 genotyping kit; Bayer)68 to detect mutations of the genes that encode the HIV protease and reverse transcriptase, and for cytomegalovirus, in which resistance to ganciclovir is mainly caused by mutations of the gene that encodes phosphotransferase69. The panel of genes that are known to be associated with resistance is expected to increase considerably in the future.
Communication and organization
Microbial surveillance and reporting. Automated methods for microbial identification will allow epidemiological trends to be followed more easily in hospitals and in the community. Similarly, automation in the molecular diagnosis of resistance to key antibiotics could allow bacterial resistance in hospitals to be surveyed online. This information can be updated weekly to provide data that are pertinent to the infection-control department (where one exists), and also to clinical physicians. For example, weekly tabulation of the documented cases of respiratory syncitial virus, influenza virus and rotavirus could help to establish prevention measures. Likewise, regular tabulation of microbial antibiotic resistance trends will assist hospitals in making empiric recommendations for antibiotic treatment.
Clinical microbiology laboratories. The general organization of the clinical microbiology laboratory of the future is the subject of much controversy4,70,71 (Table 1; Fig. 3).
Some of the work that is now carried out in laboratories could be ceded to patients and doctors with the commercial development of self-testing kits, such as those that detect HIV infection using saliva or blood spots and those that diagnose group A streptococcus throat infections. The number of available 'at-doctor' tests might increase and could potentially include the detection of group B streptococci in the vaginas of pregnant women and detection of S. aureus in the nasal passages of patients and healthcare workers. These tests could be made available at outpatient clinics.
The organization of clinical microbiology laboratories in the future might follow the current trend in the life sciences for the development of large, centralized laboratories with more organized and efficient transport of samples to these laboratories and computer-based communication of results. Hospital-based laboratories have largely disappeared in the United States in recent times, although this has resulted in increased physician dissatisfaction with clinical microbiology services (Table 1). Large, centralized laboratories, however, are more able to meet the increasing demand for specific tests that are performed infrequently, should enable increased processing of samples at night and at weekends and also provide improved access to highly specialized experts. The creation of large, centralized laboratories will require the training of technicians and engineers in the use of sophisticated techniques and instruments. In addition, the molecular identification of microorganisms requires training in bioinformatics.
The roles of different types of personnel in clinical microbiology laboratories in the future might be as follows: highly qualified, dedicated technicians or engineers would be required to maintain instrument platforms, owing to their increasing complexity; diagnostic tests would be performed by technicians; new diagnostic tests would be developed by engineers. The activity of the clinical microbiologist would therefore be re-oriented towards the evaluation and interpretation of tests, including the determination of their sensitivity, specificity and predictive values; communication of results to clinicians, in particular, infectious-disease specialists; advising on antibiotic therapy and sampling strategy in conjunction with infectious-disease specialists; following trends in disease epidemiology and reporting these trends to infection-control departments; coordinating the work of the technicians and engineers and the overall strategy of the laboratory, such as when to implement new tests or to stop using others; and contributing to the international scientific community by reporting cases of rare infectious diseases.
It is anticipated that the capacity of laboratories to handle highly pathogenic microorganisms with the appropriate levels of safety will increase in the future. Certainly, if larger laboratories are created, they would be expected to have the capacity to grow hazardous microorganisms at biosafety level 3, to detect agents that might be used in bioterrorism, to deal with emerging pathogens and to detect uncultured viruses by electron microscopy. As already mentioned, the necessity for laboratories with these advanced capabilities was shown recently in the cases of the SARS outbreak and the anthrax bioterrorism attacks. These laboratories could work as reference centres and could be connected in national and international networks, servicing not only local and regional areas, but also remote areas — including developing countries.
Sampling strategy and choice of laboratory tests. The sampling strategies that are used in clinical microbiology need to improve in the future for several reasons. Requests for laboratory tests to diagnose infectious diseases depend largely on the knowledge of the physicians who are caring for patients. As a result, there is considerable variability in the appropriateness of the samples submitted and, therefore, in the accuracy of the resulting laboratory diagnoses. For example, the standardization of sampling for infectious endocarditis (for which three blood cultures and serological testing are required for diagnosis) could dramatically reduce the number of culture-negative endocarditis patients72. In diseases such as pneumonia and meningoencephalitis, the number of known aetiological agents is increasing and depends on many epidemiological factors. The identification of aetiological agents should benefit from the use of versatile diagnostic tools.
To be optimally efficient, sampling strategies and laboratory tests should be primarily evidence-based. To optimize diagnostic efficiency, a possible strategy would be to sample by syndrome — for example, nosocomial pneumonia, endocarditis, infectious arthritis, meningoencephalitis, uveitis or fever on return from tropical areas. Such a strategy should be based on an extensive review of the international literature that identifies the potential aetiologies of a given syndrome. The most appropriate clinical samples and laboratory tests could be deduced from this information, and diagnostic kits could be developed for each syndrome. This strategy would enable a rational search for aetiological agents, limit the use of unnecessary tests and improve the cost-effectiveness of diagnosis. Another advantage of such a strategy is that it could be coupled to a versatile diagnostic method that enables the identification of a large panel of microorganisms. Such versatile methods are available for culture, molecular detection (with broad-range standardized PCR assays), FCM detection assays and broad-range microarrays.
Digital pictures. A key development in clinical microbiology has been in the technology that is available to produce digital pictures, which enables image libraries to be established and readily exchanged. This includes pictures taken during clinical examinations when cutaneous signs are present, light-, confocal- and electron-microscopy pictures, and images of microarray analyses. Although it is still rare, microbiology reports can be enhanced with digital images of the organisms that are involved. The inclusion of digital photographs of Gram-stain and acid-fast-stain preparations is technically possible, and precedents for providing such images have already been set by current pathology and radiology information systems. Picture digitalization enables information to be sent instantly by e-mail. The feasibility of using static-image telepathology to assess Gram-stained microbiological preparations has been evaluated positively, although the selection of informative microscopic fields before their transmission was a prerequisite for success73. A major application of these images is that they can be used to obtain expert advice. This is of particular importance with P3 emerging and re-emerging agents and with group A and B agents of bioterrorism. In such cases, microbial agents must be confined in secure laboratories, but their digital images can be freely exchanged. The e-mailing of digital pictures was recently helpful in the management of patients with suspected smallpox or anthrax. In a patient presenting with multiple haemorrhagic vesicles on the face, anonymous digital pictures of the clinical features were e-mailed to the United States Centers for Disease Control and Prevention to obtain an expert opinion74 and the usefulness of such pictures was emphasized during the SARS outbreak73. In addition, appropriate microbiology pictures can be e-mailed to clinicians for incorporation into the patient's medical records.
Conclusions
There have been substantial changes in the role of clinical microbiology laboratories over the past decade. The ongoing technological revolution has rapidly transformed research, diagnostic and therapeutic tools. However, there is still a requirement for the reliability and the standardization of sampling and biological testing to be increased as information is exchanged internationally. In future, the organization of clinical microbiology laboratories will probably follow that of other areas of the life sciences — that is, with large centralized laboratories and a local head. The position and the role of experts in clinical microbiology processes will become an important question. The interpretation of diagnostic tests can require different levels of expertise. The predictive value of a test depends on its intrinsic quality and on the prevalence of the disease in the population. Therefore, the final interpretation relies on both the microbiologist and the physician. In some instances, the indications of the clinical and epidemiological data to the laboratory will make it possible for a diagnosis to be made. In other cases, communication of the intrinsic values of the test (for example, sensitivity and specificity) might allow an infectious-disease specialist to make a diagnosis on-site (Fig. 1).
Finally, the need for consultation on rare diseases with experts around the world using the Internet will increase. In our experience, we receive queries about the diagnosis and treatment for Whipple's disease and Q fever several times a week. Such queries for assistance will probably increase in number as experts in particular fields become more readily identified by physicians, and even patients, through the Internet.
References
Isenberg, H. D. Clinical microbiology: past, present, and future. J. Clin. Microbiol. 41, 917–918 (2003).
O'Hara, C. M., Weinstein, M. P. & Miller, J. M. In Manual of Clinical Microbiology (eds Murray, P. R., Baron, E. J., Jorgensen, J. H., Pfaller, M. A. & Yolken, R. H.), 185–207 (ASM, Washington DC, 2003).
Ferraro, M. J. & Jorgensen, J. H. In Manual of Clinical Microbiology (eds Murray, P. R., Baron, E. J., Jorgensen, J. H., Pfaller, M. A. & Yolken, R. H.) 208–217 (ASM, Washington DC, 2003).
Peterson, L. R. et al. Role of clinical microbiology laboratories in the management and control of infectious diseases and the delivery of health care. Clin. Infect. Dis. 32, 605–611 (2001).
Baron, E. J. Speculations on the microbiology laboratory of the future. Clin. Infect. Dis. 35 (Suppl. 1), S84–S87 (2002).
Benson, D. A., Karsch-Mizrachi, I., Lipman, D. J., Ostell, J. & Wheeler, D. L. GenBank. Nucleic Acids Res. 31, 23–27 (2003).
Drancourt, M. et al. 16S ribosomal DNA sequence analysis of a large collection of environmental and clinical unidentifiable bacterial isolates. J. Clin. Microbiol. 38, 3623–3630 (2000).
Koonin, E. V. & Galperin, M. Y. Sequence — Evolution — Function: Computational Approaches in Comparative Genomics (Kluwer Academic, Boston, 2003).
Mollet, C., Drancourt, M. & Raoult, D. rpoB sequence analysis as a novel basis for bacterial identification. Mol. Microbiol. 26, 1005–1011 (1997).
Comincini, S., Barbarini, D., Telecco, S., Bono, L. & Marone, P. Rapid identification of Mycobacterium tuberculosis and Mycobacterium avium by polymerase chain reaction and restriction enzyme analysis within sigma factor regions. New Microbiol. 21, 391–395 (1998).
Le, T. H., Blair, D. & McManus, D. P. Mitochondrial genomes of parasitic flatworms. Trends Parasitol. 18, 206–213 (2002).
Calvario, A. et al. Herpes consensus PCR test: a useful diagnostic approach to the screening of viral diseases of the central nervous system. J. Clin. Virol. 25 (Suppl. 1), S71–S78 (2002).
Maiden, M. C. et al. Multilocus sequence typing: a portable approach to the identification of clones within populations of pathogenic microorganisms. Proc. Natl Acad. Sci. USA 95, 3140–3145 (1998).
Tavanti, A., Gow, N. A., Senesi, S., Maiden, M. C. & Odds, F. C. Optimization and validation of multilocus sequence typing for Candida albicans. J. Clin. Microbiol. 41, 3765–3776 (2003).
Desquesnes, M. & Davila, A. M. Applications of PCR-based tools for detection and identification of animal trypanosomes: a review and perspectives. Vet. Parasitol. 109, 213–231 (2002).
Gurtler, V. & Stanisich, V. A. New approaches to typing and identification of bacteria using the 16S–23S rDNA spacer region. Microbiology 142, 3–16 (1996).
Iwen, P. C., Hinrichs, S. H. & Rupp, M. E. Utilization of the internal transcribed spacer regions as molecular targets to detect and identify human fungal pathogens. Med. Mycol. 40, 87–109 (2002).
Devulder, G., Perrière, G., Baty, F. & Flandrois, J. P. BIBI, a bioinformatics bacterial identification tool. J. Clin. Microbiol. 41, 1785–1787 (2003).
Tyagi, S. & Kramer, F. R. Molecular beacons: probes that fluoresce upon hybridization. Nature Biotechnol. 14, 303–308 (1996).
Tyagi, S., Bratu, D. P. & Kramer, F. R. Multicolor molecular beacons for allele discrimination. Nature Biotechnol. 16, 49–53 (1998).
Call, D. R., Borucki, M. K. & Loge, F. J. Detection of bacterial pathogens in environmental samples using DNA microarrays. J. Microbiol. Methods 53, 235–243 (2003).
Hwang, T. S. et al. Detection and typing of HPV genotypes in various cervical lesions by HPV oligonucleotide microarray. Gynecol. Oncol. 90, 51–56 (2003).
Ye, R. W., Wang, T., Bedzyk, L. & Croker, K. M. Applications of DNA microarrays in microbial systems. J. Microbiol. Methods 47, 257–272 (2001).
Liu, W. T., Mirzabekov, A. D. & Stahl, D. A. Optimization of an oligonucleotide microchip for microbial identification studies: a non-equilibrium dissociation approach. Environ. Microbiol. 3, 619–629 (2001).
Kricka, L. J. Ultrasensitive immunoassay techniques. Clin. Biochem. 26, 325–331 (1993).
Silzel, J. W., Cercek, B., Dodson, C., Tsay, T. & Obremski, R. J. Mass-sensing, multianalyte microarray immunoassay with imaging detection. Clin. Chem. 44, 2036–2043 (1998).
Constantine, C. C. Importance and pitfalls of molecular analysis to parasite epidemiology. Trends Parasitol 19, 346–348 (2003).
Haab, B. B. Advances in protein microarray technology for protein expression and interaction profiling. Curr. Opin. Drug Discov. Devel. 4, 116–123 (2001).
Pavlickova, P. et al. Antibody detection in human serum using a versatile protein chip platform constructed by applying nanoscale self-assembled architectures on gold. J. Proteome Res 1, 227–231 (2002).
Mezzasoma, L. et al. Antigen microarrays for serodiagnosis of infectious diseases. Clin. Chem. 48, 121–130 (2002). A pioneering work on antigenic microarrays for the serological analysis of infectious diseases.
Gershon, D. Mass spectrometry goes mainstream. Nature 424, 581 (2003).
Woese, C. R. Bacterial evolution. Microbiol. Rev. 51, 221–271 (1987).
von Wintzingerode, F. et al. Base-specific fragmentation of amplified 16S rRNA genes analyzed by mass spectrometry: a tool for rapid bacterial identification. Proc. Natl Acad. Sci. USA 99, 7039–7044 (2002).
Issaq, H. J., Conrads, T. P., Prieto, D. A., Tirumalai, R. & Veenstra, T. D. SELDI–TOF MS for diagnostic proteomics. Anal. Chem. 75, 148A–155A (2003).
Carrette, O. et al. A panel of cerebrospinal fluid potential biomarkers for the diagnosis of Alzheimer's disease. Proteomics 3, 1486–1494 (2003). Highlights the use of mass spectrometry for diagnosis.
Lehrer, S. et al. Putative protein markers in the sera of men with prostatic neoplasms. BJU Int. 92, 223–225 (2003).
Ardekani, A. M., Liotta, L. A. & Petricoin, E. F. Clinical potential of proteomics in the diagnosis of ovarian cancer. Expert Rev. Mol. Diagn. 2, 312–320 (2002).
Issaq, H. J., Veenstra, T. D., Conrads, T. P. & Felschow, D. The SELDI–TOF MS approach to proteomics: protein profiling and biomarker identification. Biochem. Biophys. Res. Commun. 292, 587–592 (2002).
Grus, F. H., Joachim, S. C. & Pfeiffer, N. Analysis of complex autoantibody repertoires by surface-enhanced laser desorption/ionization–time of flight mass spectrometry. Proteomics 3, 957–961 (2003).
Rubin, R. B. & Merchant, M. A rapid protein profiling system that speeds study of cancer and other diseases. Am. Clin. Lab. 19, 28–29 (2000).
Hazelton, P. R. & Gelderblom, H. R. Electron microscopy for rapid diagnosis of infectious agents in emergent situations. Emerg. Infect. Dis. 9, 294–303 (2003). Illustrates the necessity of electron microscopy in clinical microbiology laboratories.
Utagawa, E. T. et al. Application of an automated specimen search system installed in a transmission electron microscope for the detection of caliciviruses in clinical specimens. J. Virol. Methods 100, 49–56 (2002).
Schroeder, J. A., Voelkl, E. & Hofstaedter, F. Ultrastructural telepathology — remote EM-diagnostic via Internet. Ultrastruct. Pathol. 25, 301–307 (2001).
Shapiro, H. M. Practical Flow Cytometry (Wiley–Liss, New York, 1995).
Alvarez-Barrientos, A., Arroyo, J., Canton, R., Nombela, C. & Sanchez–Perez, M. Applications of flow cytometry to clinical microbiology. Clin. Microbiol. Rev. 13, 167–195 (2000).
Raoult, D. et al. Cultivation of the bacillus of Whipple's disease. New Engl. J. Med. 342, 620–625 (2000).
Renesto, P. et al. Genome-based design of a cell-free culture medium for Tropheryma whipplei. Lancet 362, 447–449 (2003). The first illustration of genome-driven axenic-culture technique.
Houpikian, P. & Raoult, D. Traditional and molecular techniques for the study of emerging bacterial diseases: one laboratory's perspective. Emerg. Infect. Dis. 8, 122–131 (2002).
Murray, A. E. et al. DNA/DNA hybridization to microarrays reveals gene-specific differences between closely related microbial genomes. Proc. Natl Acad. Sci. USA 98, 9853–9858 (2001).
Dworkin, L. L., Gibler, T. M. & van Gelder, R. N. Real-time quantitative polymerase chain reaction diagnosis of infectious posterior uveitis. Arch. Ophthalmol. 120, 1534–1539 (2002).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Kengne, P., Veas, F., Vidal, N., Rey, J. L. & Cuny, G. Trichomonas vaginalis: repeated DNA target for highly sensitive and specific polymerase chain reaction diagnosis. Cell. Mol. Biol. 40, 819–831 (1994).
Raoult, D. et al. Molecular identification by 'suicide PCR' of Yersinia pestis as the agent of Medieval Black Death. Proc. Natl Acad. Sci. USA. 97, 12800–12803 (2000).
Abu-Al-Soud, W. & Radstrom, P. Effects of amplification facilitators on diagnostic PCR in the presence of blood, feces, and meat. J. Clin. Microbiol. 38, 4463–4470 (2000).
Choo, Q. L. et al. Isolation of a cDNA clone derived from a blood-borne non-A, non-B viral hepatitis genome. Science 244, 359–362 (1989).
Reyes, G. R. et al. Isolation of a cDNA from the virus responsible for enterically transmitted non-A, non–B hepatitis. Science 247, 1335–1339 (1990).
Lisitsyn, N. & Wigler, M. Cloning the differences between two complex genomes. Science 259, 946–951 (1993).
Allander, T., Emerson, S. U., Engle, R. E., Purcell, R. H. & Bukh, J. A virus discovery method incorporating DNase treatment and its application to the identification of two bovine parvovirus species. Proc. Natl Acad. Sci. USA 98, 11609–11614 (2001).
Fenollar, F. & Raoult, D. Diagnosis of rickettsial diseases using samples dried on blotting paper. Clin. Diagn. Lab. Immunol. 6, 483–488 (1999).
Rolain, J. M., Sthul, L., Maurin, M. & Raoult, D. Evaluation of antibiotic susceptibilities of three Rickettsial species including Rickettsia felis by a quantitative PCR DNA assay. Antimicrob. Agents Chemother. 46, 2747–2751 (2002).
Rolain, J. M., Maurin, M., Bryskier, A. & Raoult, D. In vitro activities of telithromycin (HMR 3647) against Rickettsia rickettsii, Rickettsia conorii, Rickettsia africae, Rickettsia typhi, Rickettsia prowazekii, Coxiella burnetii, Bartonella henselae, Bartonella quintana, Bartonella bacilliformis, and Ehrlichia chaffeensis. Antimicrob. Agents Chemother. 44, 1391–1393 (2000).
Masselot, F., Boulos, A., Maurin, M., Rolain, J. M. & Raoult, D. Molecular evaluation of antibiotic susceptibility: Tropheryma whipplei paradigm. Antimicrob. Agents Chemother. 47, 1658–1664 (2003).
Ozanne, V., Ortalo-Magne, A., Vercellone, A., Fournie, J. J. & Daffe, M. Cytometric detection of mycobacterial surface antigens: exposure of mannosyl epitopes and of the arabinan segment of arabinomannans. J. Bacteriol. 178, 7254–7259 (1996).
Pattanapanyasat, K., Webster, H. K., Udomsangpetch, R., Wanachiwanawin, W. & Yongvanitchit, K. Flow cytometric two-color staining technique for simultaneous determination of human erythrocyte membrane antigen and intracellular malarial DNA. Cytometry 13, 182–187 (1992).
Berger-Bachi, B. Expression of resistance to methicillin. Trends Microbiol. 2, 389–393 (1994).
Hirsch, M. S. et al. Antiretroviral drug resistance testing in adults infected with human immunodeficiency virus type 1: 2003 recommendations of an International AIDS Society USA Panel. Clin. Infect. Dis. 37, 113–128 (2003).
Emery, V. C. & Griffiths, P. D. Prediction of cytomegalovirus load and resistance patterns after antiviral chemotherapy. Proc. Natl Acad. Sci. USA 97, 8039–8044 (2000).
Baron, E. J. Microbiology on the merge: the business of clinical practice. ASM News 67, 191–195 (2001).
Kolmos, H. J. Role of the clinical microbiology laboratory in infection control — a Danish perspective. J. Hosp. Infect. 48 (Suppl. A), S50–S54 (2001).
Werner, M., Andersson, R., Olaison, L. & Hogevik, H. A Clinical study of culture-negative endocarditis. Medicine (Baltimore) 82, 263–273 (2003).
McLaughlin, W. J. et al. Telemicrobiology: feasibility study. Telemed. J. 4, 11–17 (1998).
Hanrahan, J. A., Jakubowycz, M. & Davis, B. R. A smallpox false alarm. New Engl. J. Med. 348, 467–468 (2003).
Damon, I., Rotz, L., Seward, J. & Hughes, J. The CDC reply: a smallpox false alarm. New Engl. J. Med. 348, 468 (2003).
Elnifro, E. M. et al. Multiplex PCR optimization and application in diagnostic virology. Clin. Microbiol. Rev. 13, 559–570 (2000).
Acknowledgements
We thank C. Corona for secretarial assistance.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
D. Raoult and M. Drancourt have shares in a biotechnology start-up company and own patents on the rpoB-based identification of bacteria.
Related links
Related links
DATABASES
Entrez
Infectious Disease Information
FURTHER INFORMATION
Glossary
- SIGMA FACTOR
-
The subunit of the RNA polymerase holoenzyme that is required for promoter-sequence recognition and the ability to initiate transcription.
- MULTILOCUS SEQUENCE TYPING
-
A DNA-sequence-based approach that is used to characterize bacterial isolates of bacteria through the Internet.
- INTERNAL TRANSCRIBED SPACER
-
The 18S–28S intergenic spacer.
- SANGER'S METHOD
-
The first efficient method for DNA sequencing, which uses dideoxynucleotides
- EXUDATES
-
Fluids, cells or other cellular substances that are discharged from inflamed tissues.
- EFFUSIONS
-
Fluids that are discharged from inflamed tissues.
- BRONCHIAL-LAVAGE FLUID
-
Fluid obtained by washing the alveolar compartment of the lung.
- FLUOROGENIC-SUBSTRATE-HYDROLYSIS FCM
-
Technique based on the measurement of fluorescence emitted during the hydrolysis of a fluorogenic substrate by a microorganism.
- MULTIPLEX PCR
-
Detection of multiple microorganisms in a single assay.
- SUICIDE PCR
-
PCR protocol in which the positive control is omitted and the DNA target is changed for each experiment.
- REPRESENTATIONAL DIFFERENCE ANALYSIS
-
A strategy that analyses the differences between two complex genomes.
- SEQUENCE-INDEPENDENT SINGLE-PRIMER AMPLIFICATION
-
Technique that uses a single linker/primer, which is ligated to blunt-ended molecules.
- BACTERICIDAL
-
An antibiotic that kills 99.99% of a bacterial population.
- BACTERIOSTATIC
-
An antibiotic that inhibits the growth of a bacterial population.
Rights and permissions
About this article
Cite this article
Raoult, D., Fournier, P. & Drancourt, M. What does the future hold for clinical microbiology?. Nat Rev Microbiol 2, 151–159 (2004). https://doi.org/10.1038/nrmicro820
Issue Date:
DOI: https://doi.org/10.1038/nrmicro820
This article is cited by
-
Rapid antibiotic susceptibility testing of bacteria from patients’ blood via assaying bacterial metabolic response with surface-enhanced Raman spectroscopy
Scientific Reports (2020)
-
Quadruplex PCR assay for identification of Corynebacterium pseudotuberculosis differentiating biovar Ovis and Equi
BMC Veterinary Research (2017)
-
leBIBIQBPP: a set of databases and a webtool for automatic phylogenetic analysis of prokaryotic sequences
BMC Bioinformatics (2015)
-
High-fat diet alters gut microbiota physiology in mice
The ISME Journal (2014)
-
Feasibility of matrix-assisted laser desorption/ionisation time-of-flight mass spectrometry (MALDI-TOF MS) networking in university hospitals in Brussels
European Journal of Clinical Microbiology & Infectious Diseases (2014)