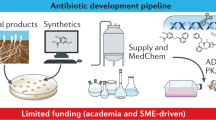
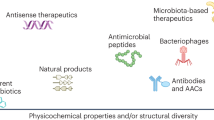
Abstract
There is a constant need for new antibacterial drugs owing to the inevitable development of resistance that follows the introduction of antibiotics to the clinic. When a new class of antibiotic is introduced, it is effective at first, but will eventually select for survival of the small fraction of bacterial populations that have an intrinsic or acquired resistance mechanism. Pathogens that are resistant to multiple drugs emerge around the globe, so how robust are antibiotic discovery processes?
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Thornsberry, C. et al. Regional trends in antimicrobial resistance among clinical isolates of Streptococcus pneumoniae, Haemophlius influenzae, and Moraxella catarrhalis in the United States: results from the TRUST Surveillance Program, 1999–2000. Clin. Infect. Dis. 34, S4–S16 (2002).
Palumbi, S. R. Humans as the world's greatest evolutionary force. Science 293, 1786–1790 (2001).
Walsh, C. T. Antibiotics: Actions, Origins, Resistance. (ASM Press, Washington DC, USA, 2003).
Coates, A., Hu, Y., Bax, R. & Page,C. The future challenges facing the development of new antimicrobial drugs. Nature Rev. Drug Discov. 1, 895–910 (2002).
Tally, F. P. & DeBruin, M. F. Development of daptomycin for gram-positive infections. J. Antimicrob. Chemother. 46, 523–526 (2000).
MacNeil, I. A. et al. Expression and isolation of small molecules from soil DNA libraries. J. Mol. Microbiol. Biotechnol. 3, 301–308 (2001).
Khosla, C. Harnessing the biosynthetic potential of modular polyketide synthases. Chem. Rev. 97, 2577–2590 (1997).
Cane, D. E., Walsh, C. T. & Khosla, C. Harnessing the biosynthetic code; combinations, permutations, and mutations. Science 282, 63–68 (1998).
Howe, R. A. & Spencer, R. C. Cotrimoxazole, rationale for re-examining its indications for use. Drug Saf. 14, 213–218 (1996).
Scholar, E. M. & Pratt, W. B. The Antimicrobial Drugs 2nd edn (Oxford Univ. Press, New York, 2000).
Maxwell, A. DNA Gyrase as a drug target. Trends Microbiol 5, 102–109 (1997).
Arya, P. D., Chou, T. H. & Baek, M. G. Diversity-based organic synthesis in the era of genomics and proteomics. Angew. Chem. Int. Ed. 40, 339–346 (2001).
Arya, P. D., Joseph, R. & Chou, D. T. Toward high-throughput synthesis of complex natural product-like compounds in the genomics and proteomics age. Chem. Biol. 9, 145–146 (2002).
Schreiber, S. L. Target-oriented and diversity-oriented organic synthesis in drug discovery. Science 287, 1964–1969 (2000).
Teague, S., Leason, D., Oprea, P. D. & Davis, T. The design of leadlike combinatorial libraries Angew. Chem. Int. Ed. 38, 3743–3748 (1999).
McDaniel, R. A. et al. Multiple genetic modifications of the erythromycin polyketide synthase to produce a library of “unnatural” natural products. Proc. Natl Acad. Sci. USA 96, 1846–1851 (1999).
Walsh, C. T. Combinatorial biosynthesis of antibiotics: challenges and opportunities. Chem. Biochem. 3, 124–134 (2002).
Carter, A. P. et al. Functional insights from the structure of the 30S ribosomal subunit and its interactions with antibiotics. Nature 407, 340–348 (2000).
Hansen, J. L. et al. The structures of four macrolide antibiotics bound to the large ribosomal subunit. Mol. Cell 10, 117–128 (2002).
Pioletti, M. et al. Crystal structures of complexes of the small ribosomal subunit with tetracycline, edeine, and IF3. EMBO J. 20, 1829–1839 (2001).
Schlunzen, F. R. et al. Structural basis for the interaction of antibiotics with the peptidyl transferase centre in eubacteria. Nature 413, 814–821 (2001).
Peterson, T. D. et al. The comprehensive microbial resource. Nucl. Acids Res. 29, 123–125 (2001).
Ji, Y. B. et al. Identification of critical staphylococcal genes using conditional phenotypes generated by antisense RNA. Science 293, 2266–2269 (2001).
McDevitt, D. & Rosenberg, M. Exploiting genomics to discover new antibiotics. Trends Microbiol. 9, 611–617 (2001).
Ye, X. Y. et al. Better substrates for bacterial transglycosylases. J. Am. Chem. Soc. 123, 3155–3156 (2001).
Yusupov, M. M. et al. Crystal structure of the ribosome at 5.5-Å resolution. Science 292, 883–896 (2001).
Navarre, W. W. & Schneewind, O. Surface proteins of gram-positive bacteria and mechanisms of their targeting to the cell wall envelope. Microbiol. Mol. Biol. Rev. 63, 174–229 (1999).
Pallen, M. J., Lam, A. C., Antonio, M. & Dunbar, K. An embarrassment of sortases-a richness of substrates? Trends Microbiol. 9, 97–102 (2001).
Rohdich, F., Kis, K., Bacher, A. & Eisenreich, W. The non-mevalonate pathway of isoprenoids: genes, enzymes, and intermediates. Curr. Opin. Chem. Biol. 5, 535–540 (2001).
Murray, B. E. Vancomcyin-resistant enterococcal infections New Eng. J. Med. 342, 710–721 (2000).
Bonhoeffer, S., Lipsitch, M. & Levin, B. R. Evaluating treatment protocols to prevent antibiotic resistance. Proc. Natl Acad. Sci. USA 94, 12106–12111 (1997).
Shlaes, D. et al. Guidelines for the prevention of antimicrobial resistance in hospitals. Clin. Infect. Dis. 25, 584–599 (1997).
Author information
Authors and Affiliations
Related links
Related links
FURTHER INFORMATION
Christopher Walsh's laboratory
United States Center for Disease Control
Glossary
- ANTIMETABOLITE
-
A structural analogue of metabolites that inhibit enzymes that normally recognize the metabolite as substrate.
- COMBINATORIAL CHEMISTRY
-
The application of synthetic methods, often in high-throughput format, to combine molecular fragments to create many structurally related congeners.
- HETEROCYCLIC DRUGS
-
A drug molecule that contains carbon, nitrogen, oxygen and sulphur atoms in ring structures — typically five- or six-membered rings.
- MACROLIDE
-
Macrocyclic (ring greater than six carbons) structures in which the cyclization occurs by lactone formation between the two ends of the molecule. The macrolactone erythronolide provides the scaffold in the antibiotic erythromycin.
- METAGENOME
-
The collective genomes of soil bacteria.
- NATURAL-PRODUCT SCAFFOLD
-
The structural and architectural platform of natural products that can be decorated, for example, by glycosylation or with synthetic modifications.
- NON-RIBOSOMAL PEPTIDE
-
A peptide that is made from amino acids, often amino acids that are not found in proteins, that are assembled by multimodular enzymes not involving RNA templates or the ribosome for peptide-bond formation.
- POLYKETIDE
-
A natural product that is assembled from malonyl CoA units through intermediates with many ketone (polyketonic) groups that allow for directed reactivity to product structures.
- SORTASE
-
An enzyme in the cell membrane of Gram-positive bacteria that sorts proteins to be displayed in the outer membrane by covalently connecting those proteins to the peptidoglycan layer.
Rights and permissions
About this article
Cite this article
Walsh, C. Where will new antibiotics come from?. Nat Rev Microbiol 1, 65–70 (2003). https://doi.org/10.1038/nrmicro727
Issue Date:
DOI: https://doi.org/10.1038/nrmicro727
This article is cited by
-
Discovery of a structural class of antibiotics with explainable deep learning
Nature (2024)
-
Deep learning-guided discovery of an antibiotic targeting Acinetobacter baumannii
Nature Chemical Biology (2023)
-
Antimicrobial efficacy of Mentha piperata-derived biogenic zinc oxide nanoparticles against UTI-resistant pathogens
Scientific Reports (2023)
-
A cell-free strategy for host-specific profiling of intracellular antibiotic sensitivity and resistance
npj Antimicrobials and Resistance (2023)
-
Visualizing the membrane disruption action of antimicrobial peptides by cryo-electron tomography
Nature Communications (2023)