Key Points
-
Bacterial species consist of genetically related strains that have evolved from a common ancestor and belong to the same clonal lineage.
-
Factors that are important for clonal evolution are horizontal gene transfer events or recombination and mutations in the bacterial genome. Host and environmental factors also influence clonality.
-
Molecular typing methods used to study clonality include restriction polymorphism patterns with PFGE (pulsed field gel electrophoresis), sequence-based methods, such as MLST (multilocus sequence typing), and more discriminatory methods, such as whole-genome microarrays and sequencing.
-
Clone-associated virulence of the pneumococcal pilus has been shown to influence the international spread of antibiotic resistant pneumococcal clones.
-
Even in highly recombinogenic species, such as Streptococcus pneumoniae, isolates that are non-transformable or have a reduced transformability are common, and frequently belong to clonal lineages that are associated with a high invasive-disease potential.
-
Comparative genomics and whole-genome sequencing of clinical isolates with known transmission and disease potentials will allow us to discover bacterial factors that influence spread and virulence and could be targeted by novel therapeutics and vaccines.
-
Vaccination against a limited number of pneumococcal polysaccharides in conjugated vaccines can lead to vaccine escape and selection, leading to replacement with non-vaccine pneumococcal clones that cause invasive disease.
Abstract
Globally spreading bacterial strains belong to clonal types that have the capacity to colonize, spread and cause disease in the community. Recent comparative genomic analyses of well-defined clinical isolates have led to the identification of bacterial properties that are required for the successful spread of bacterial clones. In this Review, we discuss the evolution of bacterial clones, the importance of recombination versus mutations for evolution of clones, common methods used to study clonal relationships among bacteria, factors that may contribute to the clonal spread of bacteria and the potential relevance of bacterial clones to clinical disease. We focus on the common pathogen Streptococcus pneumoniae, although other bacteria are also briefly discussed, such as Helicobacter pylori, Staphylococcus aureus and Mycobacterium tuberculosis.
Similar content being viewed by others
Main
Bacterial species consist of genetically related strains that have evolved from a common ancestor. These strains belong to the same clonal lineage, and evolve through mutation and horizontal gene transfer events that involve both homologous and non-homologous recombination1. In naturally competent Helicobacter pylori , which has a high rate of import of unusually short pieces of DNA through homologous recombination2, sequence variation is so extensive that within a chronically infected host who has been infected by more than one strain, so many different subclones emerge that virtually every progeny organism is genetically distinct2. Other species, such as Streptococcus pneumoniae and Neisseria spp., also have such a high rate of recombination that most of the genetic differences observed between different clonal types are thought to be due to recombination rather than mutation3,4. By contrast, Mycobacterium spp. have evolved primarily through mutation, and exhibit only a low level of horizontal gene transfer5,6 (Box 1). However, the contribution of horizontal gene transfer to the evolution of the Mycobacterium tuberculosis complex is not completely understood7.
Modelling the diversification of a uniform bacterial population8 revealed that when recombination is infrequent in a bacterial species, for example, in M. tuberculosis, distinct clonal populations appear within approximately 250,000 generations. However, for organisms in which recombination is much more frequent than mutation, transient, more diffuse clonal clusters emerge, but do not establish themselves as distinct clusters and instead become incorporated into the main cluster through recombination within the population. Only when sequence diversity markedly reduces homologous recombination can distinct clusters emerge and avoid being reabsorbed by recombination8. Consequently, clonal clusters evolve and disappear in this model in the absence of external selection. As homologous recombination involves the replacement of fragments in the bacterial chromosome, which leads to strain diversification, species in which such recombination events occur extremely rarely, such as M. tuberculosis, have a highly clonal population structure, whereas species in which recombination events occur frequently, such as H. pylori, are almost non-clonal9.
Mutation frequencies and mutation spectra also differ among bacterial species, and isolates of H. pylori have a higher spontaneous mutation frequency than most other bacterial species10. Furthermore, within a given species, different isolates can have significantly different spontaneous mutation frequencies11,12. In some cases, hypermutability among clinical isolates (bacteria with a mutator phenotype) have been associated with dysfunctional mismatch-repair systems13. Even though a mutator phenotype can increase adaptability to novel environmental conditions, it can also cause mutations that affect bacterial fitness to accumulate. It has been suggested that long-term maintenance of high bacterial mutation rates is driven by rapidly changing selection pressures, such as those imposed by bacteriophages14.
The presence of insertion (IS) elements and other repetitive sequences contributes to duplications, inversions and translocations within single genomes. One interesting feature that is unique to the pneumococcal genome, in contrast to 51 other analysed bacterial genomes, is the abundance of repeat regions, such as IS elements, which constitute approximately 5% of the pneumococcal genome, and non-IS elements, such as the repetitive DNA-sequence motifs Box and RUP elements15,16. Box elements are of unknown function, are typically ∼100–200 bp in length and are located in intergenic regions, whereas RUP repeats are usually ∼107 bp in length and are related to IS elements15,16. The high density of these repeats suggests they might have a role in intragenomic and intergenomic recombination events.
Clonal evolution and the ecological niche
In addition to mutation and recombination, clonal evolution is also affected by the population size, the size of the gene pool and the genetic content of the ecological niche. S. pneumoniae is primarily a human-specific organism that normally colonizes the nasopharynx of healthy preschool children17,18. The nasopharynx also contains non-virulent streptococci, such as Streptococcus oralis and Streptococcus mitis , as well as other competing organisms, such as Haemophilus influenzae and Moraxella catarralis , and could therefore provide genetic information that could facilitate the development of antibiotic resistance17,18. The clonal evolution of S. pneumoniae depends on selective forces in the upper airways, such as competition and collaboration with other pneumococcal strains or strains of other bacterial species to retrieve nutrients for growth, direct negative (for example, bacteriocins and hydrogen peroxide) or positive interactions for growth, as well as antibiotics taken by the host, and clear responses from the innate and adaptive immune system of the host. A particularly interesting example of pneumococcal competition in the nasopharynx is the phenomenon of pneumococcal fratricide, in which transformation-competent cells are able to kill non-competent cells, thereby allowing released DNA to be retrieved from lysed cells. This process has been shown to considerably increase horizontal gene transfer between different strains of pneumococci, as well as between related commensal streptococci19,20.
Selective pressures, in combination with available DNA from the same or different bacterial species, competence for transformation and mutation-promoting factors all contribute to the rise and fall of pneumococcal clonal lineages (Fig. 1). Furthermore, bacterial properties (for example, those that promote adhesion), environmental ecological factors and host responses all influence clonal diversification. The main factors that affect pneumococcal population structure (for example, clonal evolution) are depicted in Fig. 1.
Although recombination is the most important mechanism for the evolution of Streptococcus pneumoniae, spontaneous mutations also have a role in the clonal evolution of clinical isolates. The availability of DNA from the same or different bacterial species and bacterial factors that influence competence for transformation, as well as mutation-promoting factors, such as the production of reactive oxygen or nitrogen species from the bacteria and/or from infected immune cells, all contribute to the clonal population structure. Other bacterial factors, as well as environmental and host factors, also influence whether clonal diversification or clonal conservation (which leads to a genetically homogenous population) occurs.
Similar to pneumococci, H. pylori has a restricted habitat, but unlike pneumococci thrives in a genetically poorer environment, which acts as a constraint on genetic evolution. Although recombination through transformation is common in H. pylori, genetic diversity must be created before amplification can occur through horizontal gene transfer. This can be achieved by infection with multiple H. pylori strains during childhood. The decreased abundance of H. pylori in the human population, especially in developed countries21,22,23, is likely to decrease the incidence of multiple infections in the same individual, a factor that might seriously limit the genetic evolution and adaptability of this pathogen.
Transmission of both pneumococci and H. pylori requires close contact between individuals. The transmission of pneumococcal strains that belong to different clonal types is facilitated by keeping carriers and non-carriers in confined areas, such as at child-care centres, where the likelihood of transmission is dependent on the number of children who attend the centre, the number of bacteria in the nasopharynx of carriers, the ability of a clonal type to establish itself in a new host and host susceptibility. For H. pylori, transmission primarily occurs from mother to child during the first 1–2 years after birth24.
Other bacterial pathogens have more complex ecological niches compared with pneumococci and H. pylori that involve both animate and inanimate environments. Staphylococcus aureus , for example, can dwell on a number of human and animal surfaces, such as the skin and nares, and is able to survive in the environment for an appreciable length of time, allowing both nosocomial spread and transmission within the community25 (Box 2).
H. pylori, S. aureus and pneumococci are examples of organisms that colonize humans and, in most cases, can be carried in healthy individuals. Disease is frequently associated with underlying genetic and/or predisposing factors in the host; for example, cancer, HIV-1, cardiovascular diseases26 and influenza virus infection27. Recent data suggest that the sensitization of influenza virus infection to subsequent pneumococcal pneumonia is mediated by viral induction of interferon-γ, which decreases the clearing responses of alveolar macrophages28, but it has also been suggested that influenza virus neuraminidase directly promotes pneumococcal adherence and infection29. However, it is not yet known if influenza virus infection of humans increases host susceptibility for all clonal types of pneumococci. Infection with pneumococci and other opportunistic pathogens can also occur in previously healthy individuals30, who can possess particular genetic haplotypes at loci that affect their susceptibility to infection and/or disease. For example, polymorphisms in the cytosolic muropeptide recognition proteins nucleotide-binding oligomerization domain 1 (NOD1) and NOD2 affect the outcome of H. pylori infection31, and specific human leukocyte antigen class II haplotypes affected severity of group A streptococcal disease32. Likewise, homozygosity for maltose-binding-protein codon variants, which are associated with low production of this innate immune mediator, is associated with a higher frequency of invasive pneumococcal disease33. However, a recent prospective cohort study of individuals from Denmark reveals that the genetic constitution of the human host only has a small role in the development of invasive pneumococcal disease34. Instead, other factors, such as premature birth and the level of crowdedness at child-care centres and in households, play major parts in predisposition towards invasive infection35,36.
Correlation of bacterial properties with disease
Our increased ability to compare clinical isolates from well defined carrier or patient groups genetically has allowed us to determine specific intraclonal (rapidly evolving through selection or high mutation frequency), clonal (evolving without external selection) or horizontally acquired bacterial traits that can be linked to infectivity or transmissibility, disease likelihood and disease type. Thus, isolates of H. pylori that express specific binding variants of the BabA adhesin can be correlated with the infection of different ethnic groups37. In another well-characterized example of functional adaptation, specific sequence variants of the pilus-associated FimH adhesin of uropathogenic Escherichia coli correlated with adhesion to the uroepithelium and subsequent infection of the lower urinary tract38. Such high-binding FimH variants are selected at a higher rate than housekeeping genes, which allows intraclonal FimH diversification39. Clonally related isolates of Neisseria meningitidis serotype C were recently shown to carry an insertion sequence at a site that controls capsular expression and were therefore resistant to human serum antibody killing40.
In S. pneumoniae, the capsular polysaccharide is used to categorize pneumococci into at least 91 capsular serotypes41,42,43. Several epidemiological studies have shown that serotype distribution differs depending on the geographical area and the time period studied, as well as on the isolation site of the bacteria (for example, blood, cerebrospinal fluid, ear or nasopharynx) and the disease being considered18,44,45,46,47,48,49. In addition, a number of molecular epidemiological studies have been performed on clinical strains of S. pneumoniae to compare the serotype distribution of nasopharyngeal isolates from healthy carriers and isolates from sterile sites, including blood and/or cerebrospinal fluid44,50,51,52. These studies, which included a meta-analysis, all showed that pneumococci which belong to different capsular serotypes have different odds ratios of causing invasive disease44,50,51. Thus, in these studies, pneumococci of serotypes 1, 4, 5 and 7F were more likely to cause invasive disease and were rarely found in the nasopharynx of healthy carriers. By contrast, isolates of serotypes 3, 6A, 19F and 23F had low invasive-disease potential and were predominantly found in healthy carriers44,50. Similarly, in the Finnish study by Hanage et al.52, serotypes 14, 18C, 19A and 6B had odds ratios of >1 for invasive disease, and were therefore associated with invasive disease, whereas serotypes 6A and 11A had odds ratios of <1. However, isolates with a low invasive-disease potential, such as type 19F isolates, may still cause invasive disease and are so predominant in the carrier population that they may be just as common in invasive disease as isolates that belong to serotypes associated with a higher invasive-disease potential53.
By including patient data in an invasive disease study it became apparent that S. pneumoniae serotypes with a higher invasive-disease potential (such as serotypes 1, 4 and 7F) were more likely to cause disease in previously healthy individuals, and therefore act as primary pathogens. This is in contrast to serotypes with low invasive-disease potential, which preferentially caused invasive disease in patients with underlying diseases and therefore act as opportunistic pathogens30. Interestingly, in this study, serotypes associated with the highest invasive-disease potential were associated with the lowest mortality30. This was partially because the healthier group of patients became infected by these serotypes. Mortality was also high in previously healthy individuals, however, at least for serotype 3, which is associated with a low odds ratio for invasive disease30. By contrast, an international study by Alanee et al.54, who studied pneumococcal bacteraemia in ten countries, did not find an association between disease severity or mortality and serotype, and suggested that host factors might be more important than serotype for disease outcome. However, in this study, clonal type (the genetic relatedness of strains within one serotype; discussed below) was not known, which might affect the interpretation of the results. In summary, these and other epidemiological studies reveal a more complex interpretation of bacterial virulence than that obtained through conventional virulence studies in animal models, and bring new parameters into play, such as the potential for invasive disease in healthy and compromised individuals and disease severity.
Clonal analyses of pneumococcal isolates have allowed studies of genetic relatedness between clinical isolates (a comparison of molecular typing schemes is provided in Fig. 2). Such studies have revealed that isolates from serotypes with high invasive-disease potential in Sweden, such as serotypes 1 and 7F, are more clonally related than isolates of serotypes with a lower invasive-disease potential30,51,53,55. Thus, based on multilocus sequence typing (MLST), Swedish isolates of invasive serotypes 1 and 7F seemed to belong mainly to clonal complex (CC) 306 and CC191, respectively (Figs 3, 4). Furthermore, using whole pneumococcus genome-based microarrays (TIGR4 (of serotype 4) and R6 (unencapsulated derivative of D39, a serotype 2 strain)), fewer genetic differences were found among isolates that belonged to sequence type (ST) 306 and ST191 than in other STs tested55. Interestingly, sub-Saharan isolates of serotype 1 belong to an unrelated epidemic CC (ST217)56 that, unlike the current Swedish CC ST306 (discussed above), is associated with severe disease (meningitis) and high mortality30,56.
Serotyping, which is based on the expression of capsular polysaccharides, is the most common method used to study relatedness between pneumococcal isolates. However, serotyping does not provide information about the genetic relatedness between pneumococcal strains, and therefore other molecular epidemiological methods are being used to study bacterial chromosome content. MLST (multilocus sequence typing), which relies on sequencing and comparing seven housekeeping genes, has no more resolving power than other DNA-based techniques described here, but can make national and international comparisons of strains. In pulsed-field gel electrophoresis (PFGE), the pneumococcal chromosomal DNA is cleaved by a restriction enzyme and then run on a pulsed-field gel, thereby separating large bands. This method has good discriminatory power, especially when used to investigate outbreaks. Using microarray technology, genes from sequenced pneumococcal genomes are spotted on an array, and the presence or absence of these genes can be studied by overlaying DNA from pneumococcal isolates of interest. The discriminatory power of microarrays depends on the number of genes to be investigated. Finally, the method with the most discriminatory power is whole-genome sequencing, which allows all genomic differences between clinical isolates to be identified.
Approximately 95% of CC306 and 98% of ST306 isolates are from serotype 1, and few serotype switches have therefore occurred within CC306. By contrast, only ∼50% of CC156 isolates and ∼70% of ST156 isolates are of serotype 9V and several serotype switches have occurred; for example, to serotypes 14 and 19F. In total, 30% of serotype 1 isolates belong to CC306 and ∼50% of CC306 isolates belong to ST306. By contrast, ∼80% of serotype 9V isolates belong to CC156 and ∼20% of CC156 isolates belong to ST156. Thus, there are major differences in the genetic relatedness between clinical pneumococcal isolates from different serotypes and clonal types, as determined using MLST (multilocus sequence typing), and this influences the interpretation of typing results. Based on data from the MLST database (see Further information).
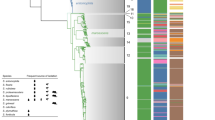
A population snap-shot based on the entire Streptococcus pneumoniae MLST (multilocus sequence typing) database, from May 2008. The 30 largest eBURST (see Further information for a link to eBURST on the MLST database) groups are highlighted. The predicted founders of the clonal complexes (CCs) are shown in blue and the size of the circles reflects the frequency of that sequence type (ST) in the data set. Connected STs differ at only one of the seven loci. An eBURST group (or CC) is defined as a group of STs in a population that share 6–7 alleles with at least one other ST in the group. Isolates from a CC are assumed to have a recent common ancestor. With eBURST it is possible to study strain microevolution based on the entire MLST database127,128.
Whether serotype-dependent differences in the likelihood of invasive pneumococcal disease depend only on different capsules that confer varying virulence or whether the different serotypes are composed of clonal types with different invasive-disease potentials, which would suggest that other bacterial factors are also important for disease outcome, has been debated. In a Finnish study of children aged ≤2 years old it was shown that strains of serotype 14 (ST156) had a higher invasive-disease potential than strains of serotype 9V (ST162), which are derived from the same CC. This suggests that the serotype 14 capsule provides increased invasiveness compared with the serotype 9V capsule52. However, isolates from the ST156 CC can differ by up to 40 genes57, and therefore a direct correlation between virulence and capsular type cannot be drawn.
Despite uncertainty in the human setting, clinical isolates of S. pneumoniae have been shown to possess dramatic differences in colonization and/or invasive disease after intranasal challenge of mice that correlate both with different capsular types and different clonal types of the same capsular type53. In this study, intraclonal differences were also found for two serotype 1 isolates of the same CC (ST228) that did not exhibit the same virulence after intraperitoneal challenge.
Thus, it seems reasonable to conclude that bacterial factors, such as the amount and type of pneumococcal capsule, affect bacterial virulence in humans in combination with other properties that may differ among different clonal lineages or even within single clones.
The diverse pneumococcal genome
Sequence analyses, as well as overall genome comparisons, have shown that S. pneumoniae belongs to a phylogenetic lineage from a large cluster of otherwise commensal streptococci. Included in this cluster are species such as S. mitis and Streptococcus pseudopneumoniae; S. pneumoniae is no more divergent from other members of the cluster than individual lineages of S. mitis are from each other58. Yet the S. pneumoniae lineage is by far the most virulent lineage of the cluster. The gene pool available for pneumococci might therefore be considerably larger than previously anticipated. The pneumococcal genome is highly diverse55,59,60. After complete sequencing of 17 pneumococcal strains, coding sequences from all strains were grouped into 3,170 orthologous gene clusters, of which only 1,454 (46%) were conserved among all 17 strains; the rest were accessory genes that could be present or absent in different isolates61. Using microarrays to study 40 pneumococcal clinical isolates that belonged to 12 serotypes and 33 STs, we found that most of the accessory genes were localized to 39 different regions of diversity or accessory regions55 that are localized in small gene clusters around the genome15,55,62,63. Unlike the well characterized pathogenicity islands of Gram-negative bacteria64, pneumococcal accessory regions are not integrated at the sites of tRNA genes, and only some differ significantly in G+C content from the rest of the genome. Some accessory regions might have evolved through phage integration, as in Streptococcus pyogenes , and some are flanked by insertion elements, which is indicative of site-specific recombination events. However, most cannot be distinguished from genes that belong to the core genome, suggesting that they represent a pool of genes that is common to both pneumococci and related streptococci55.
As expected from the finding of Feil and colleagues65 that diversity in the S. pneumoniae genus is primarily created by recombination, the pattern of accessory regions in individual pneumococcal strains from the same clonal cluster is similar or even identical55. The presence or absence of pneumococcal accessory regions can, for the most part, be explained by homologous recombination within the genus, but interspecies recombination probably also occurs. Strains that have been assigned to different clonal clusters based on MLST differ in their pattern of accessory regions, irrespective of whether they belong to the same capsular serotype or not55. Thus, it can be inferred that the distribution and expression of accessory regions and their expression pattern will affect virulence and disease outcome for a given isolate. However, there are substantial redundancies among virulence attributes, meaning that the genetic background of the bacteria will decide the impact of a single virulence factor.
Strains belonging to clonal types are frequently found in the nasopharynx, and are therefore associated with carriage and opportunistic infections and would be expected to harbour accessory regions that allow colonization and growth on mucosal surfaces. One example of such a region is the rlrA islet, which encodes pneumococcal pili66,67. rlrA is present in a restricted number of clonal lineages and, depending on differences in sequence, can be divided into three different clades, with an overall homology of 88–92%68. Pneumococcal pili have been shown to enhance adhesion to human respiratory epithelial cells, promote colonization of the murine upper airways and enhance the inflammatory response during the systemic phase of pneumococcal disease67. Despite the competitive advantage of pili expression, only approximately 20–30% of randomly selected isolates carry the rlrA pilus-encoding islet68,69,70. It is possible that immunogenic pili71 generate immunity in the population, thereby selecting against piliated pneumococcal strains in the community.
A second pilus type was recently discovered in S. pneumoniae72. As for rlrA-encoded pili (pilus islet 1 (PI-1)), PI-2 is present in a restricted number of clonal types that are associated with serotypes 1, 2, 7F, 19A and 19F, which are considered to be emerging serotypes. Furthermore, second-type pili contribute to epithelial adhesion, but their role in pneumonia and invasive disease has not been defined. However, the presence of PI-1 and/or PI-2 in isolates from the highly invasive CC ST306 (serotype 1), ST205 (serotype 4) and ST191 (serotype 7F) suggests that PI-1 and PI-2 pili play a part not only in adhesion but also in lung infection and invasive disease in humans.
Specific CCs of pneumococci can also carry other adhesive islets. For example, the pneumococcal serine-rich repeat protein (PsrP)73 is encoded in a large accessory region that is present in many isolates with high predicted invasiveness, such as ST306 of serotype 1, but is absent in most other isolates73. Pneumococcal accessory regions not only encode adhesins and other potential virulence attributes but also encode genes that are involved in transport and metabolism. Interestingly, a number of genes identified as being necessary for in vivo growth in mice belong to accessory regions that can be absent from clonal types which are capable of causing invasive disease in humans, suggesting significant redundancy in accessory gene functions (B.H-N., C.B., J.D. and S.N., unpublished observations). By comparing pneumococcal clonal types of serotypes 1, 4 and 7F, all of which have high invasive-disease potential, with all other clonal types, we found that only one accessory region is present in the invasive clonal types but missing in isolates of most other clonal types (B.H-N., C.B., J.D. and S.N., unpublished observations). This region, which was identified as important for invasive disease in a signature-tagged mutagenesis screen74 (a high-throughput method that identifies mutants with reduced or increased adaptation to certain environments), encodes a family 1 phospho-β-glucosidase, which suggests that complex-carbohydrate utilization contributes to high invasive-disease potential (B.H-N., C.B., J.D. and S.N., unpublished observations).
Even though accessory regions are only present in specific clonal lineages of S. pneumoniae, their expression and regulation is probably interconnected with regulatory networks that involve housekeeping genes. One recent example from the Tuomanen group75 showed that the rlrA regulatory locus of PI-1 is integrated into transcriptional regulatory networks that control the expression of virulence factors that are important in pneumococcal adhesion and invasion.
In addition to the presence and absence of accessory regions, sequenced isolates can contain genes that although present in all isolates possess considerable sequence variation. Such variation could be important for virulence, as shown by the ability of PspC (also called SpsA, CbpA, PbcA and Hic), a surface protein of S. pneumoniae, to bind secretory immunoglobulin A, C3 and complement factor H, and act as an adhesin76,77. However, whether or not allelic variation of pspC is associated with different clonal types has not yet been shown.
Little is known about the contribution to virulence of naturally occurring mutations in single genes. The pore-forming toxin pneumolysin of S. pneumoniae is thought to be particularly important for virulence78. Yet one of the most successful invasive clones of serotype 1 (ST306) expresses a non-haemolytic pneumolysin79,80. This suggests that a deficiency in active pneumolysin production in the context of the ST306 genetic background is neutral or perhaps even advantageous for human pneumonia, which is frequently associated with serotype 1 infections. Researchers have recently observed an increased incidence of empyemas, which are associated with childhood pneumonia and are preferentially caused by serotype 1 pneumococci both in England and France81,82, but it is unknown whether these more severe pneumonias are caused by serotype 1 isolates that produce a non-haemolytic form of pneumolysin.
Spread of penicillin non-susceptible clones
S. pneumoniae, like Neisseria spp., has evolved decreased susceptibility to penicillin by remodelling the penicillin-binding proteins through a series of horizontal gene transfer events and mutations83,84. The occurrence of penicillin-non-susceptible pneumococci (PNSP) in several different geographical areas, which led to resistance in >50% of the pneumococcal population in some regions, was mainly due to the spread of a limited number of clonal types85,86,87. Such clones appear both in countries with high and low antibiotic consumption. In the few cases in which the proposed susceptible variants of the same clones were studied, the susceptible ancestors also demonstrate high transmission ability in the community. Therefore, when antibiotic resistance evolves in pneumococcal strains that can be transmitted to and colonize the human nasopharynx there is a risk that such strains could emerge as international clones for global spread.
One such PNSP clonal cluster is ST156 (also referred to as Spain9V-3 by the Pneumococcal Molecular Epidemiology Network86 (PMEN; see Further information)). This clonal cluster is associated with several capsular serotypes, such as 9V, 14 and 19F, and has been isolated from humans on four continents57,88. In several countries, including Sweden, ST156 has been the dominating PNSP clone for several years57,88,89. Both PNSP and penicillin-susceptible pneumococci isolates (from ST162) that belong to the ST156 clonal cluster were found to carry PI-1, the rlrA pilus-encoding islet (Fig. 4; Table 1). Interestingly, the internationally spread Spain6B-2 clone, which caused a dramatic increase of penicillin non-susceptibility in Iceland when it spread among children87, also carries PI-1, but produces pili of a different clade than pili produced by pneumococci of the ST156 clonal cluster. When all PNSP isolates in Sweden were characterized, ∼70% belonged to piliated clones of the three different clades57, which is higher than the incidence of piliation among randomly collected strains57,68,69,70. To study the in vivo importance of pneumococcal pili for colonization and spread, mice were inoculated intranasally with a low dose of piliated and non-piliated pneumococci of two clinical isolates that differed only in their possession of PI-1, as determined by microarray analysis. In this competition experiment, the piliated clinical isolate 'outcompeted' the non-piliated isolate for colonization57. No correlation to penicillin non-susceptibility has been observed between the second pneumococcal pilus islet, PI-2, even though the globally spreading PNSP clone Taiwan (19F)-14 contains both pilus islets68,72. Available data therefore suggest that penicillin non-susceptibility is built up in clones that already carry the rlrA (PI-1) islet, resulting in PNSP strains that are particularly capable of global dissemination.
Competence and clonal evolution
S. pneumoniae is a naturally transformable species. This competence for uptake and incorporation of DNA is induced by the secretion of competence-stimulating peptide (CSP), of which two (CSP1 and CSP2) have been reported to date90,91. CSP interacts with the membrane-bound histidine kinase receptor (ComD), which leads to phosphorylation of the cognate response regulator ComE. In its phosphorylated state, ComE activates 20 so-called early genes, of which the alternative sigma factor ComX directs the transcription of around 60 genes, including those involved in DNA uptake92,93. Microarray analyses revealed that most, if not all, of the genes involved in transformation belong to the core genome (B.H-N., C.B., J.D. and S.N., unpublished observations). Nevertheless, as many as 50% of the randomly selected encapsulated clinical isolates were poorly transformable or non-transformable under laboratory conditions after induction with either CSP1 or CSP2 (Refs 90, 94, 95). These strains belonged to several serotypes, and isolates of the same serotype often differed in their competence induction by CSP1 or CSP2 (Ref. 90). We used clonally defined pneumococcal isolates to revisit this issue and found that a number of isolates that could not be transformed after induction by CSP1 and CSP2 belonged to highly successful clonal clusters, such as ST306 of serotype 1, that exhibit epidemic behaviour when causing invasive disease but rarely colonize the nasopharynx96. Furthermore, isolates of ST124 from serotype 14, one of the most prevalent penicillin-susceptible clones in Sweden for a number of years, and isolates of ST180 from serotype 3, a serotype that has one of the highest rate of mortality in humans, had low transformability30,45 (B.H-N., P.B. and S.N., unpublished observations). Clonal clusters associated with low transformability were also rarely found to be PNSP. These results suggest that a subset of non-transformable pneumococcal clonal clusters may not have access to the gene pool that is available for fully transformable pneumococcal isolates, a factor that might affect their evolution through recombination.
Clonal selection after vaccination
The seven-valent pneumococcal conjugate polysaccharide vaccine (PCV7) was licensed for use in 2000 for infants and young children in the United States97. It is clear that owing to herd immunity in the United States and in other countries where PCV7 has been introduced, the vaccine has been beneficial and has significantly reduced the incidence of pneumococcal invasive disease and pneumococcal carriage caused by vaccine serotypes (serotypes 4, 6B, 9V, 14, 19F, 18C and 23F) in both vaccinated children and in the unvaccinated community97,98,99,100,101,102. However, an increase in non-vaccine serotypes, such as serotypes 3, 15, 19A, 22F and 33F, among children has been noted, and the non-vaccine serotype 19A has become the predominant cause of invasive pneumococcal disease in children in the United States103. In a study from 2003–2004, Beall and colleagues104 showed that CC199 predominated among serotype 19A isolates in children less than 5 years old, which represented approximately 70% of invasive serotype 19A isolates. Brueggemann et al.105 found that this increase in serotype 19A was partly due to the prevalence of this genotype prior to introduction of the vaccine and partly owing to the emergence of a genotype that had previously only been associated with serotype 4. This novel vaccine-escape strain evolved following a single genetic event in which a 39 kb fragment was transferred from the serotype 19A strain into the type 4 strain, resulting in both a capsular switch and penicillin non-susceptibility; two penicillin-binding proteins were also involved in this recombination event105. In Alaska, an increase was observed in non-vaccine types, predominantly of serotype 19A, that cause invasive disease. Most of the expansion of this serotype was attributed to an increase in one genotype, ST199 (Ref. 101). Furthermore, serotype replacement after vaccine introduction has emerged in invasive pneumococcal disease with serotypes not included in the vaccine; for example, serotypes 3, 7F, 15B/C/F, 19A, 22F, 33F and 38 have been described in various surveillance systems103,106,107,108,109,110,111. In Portugal, the most common non-vaccine invasive serotypes in children after vaccine introduction were types 1, 19A, 7F, 3 and 33F112.
Particularly worrisome has been the resistant nature of some non-vaccine isolates111,113. The incidence of invasive pneumococcal disease owing to penicillin-resistant 19A isolates has increased considerably in the post-vaccine era in the United States. Of 151 penicillin-resistant 19A isolates, 111 (73.5%) belonged to the same CC as the multidrug-resistant Taiwan (19F)-14 described above114. Clonal analysis of emerging 19A acute otitis media isolates that are resistant to all Food and Drug Administration (FDA)-approved antibiotics showed that these isolates belonged to ST2722, a single-locus sequence variant of the globally spread ST156 discussed above111. Other PNSP clones of non-vaccine serotypes have also expanded in the post-vaccine era in the United States; for example, ST558 of serotype 35B115. These emerging PNSP non-vaccine isolates belong to clonal clusters that possess PI-1 (Refs 67, 68). We might therefore be witnessing clonal selection of pre-existing PNSP isolates of non-vaccine serotypes that were minor lineages in the carriage population, but have the capacity to expand, potentially promoted by adhesive properties, such as pili.
We have not found any published report on post-vaccine expansion in the United States of non-vaccine serotypes that are associated with a high invasive-diseasepotential, such as serotypes 1 and 7F. Furthermore, no capsular switches for these non-vaccine serotypes of high invasive-disease potential have been reported. However, these serotypes are not as common in the United States as they are in some European countries, and therefore vaccination could have different effects on the emergence of more invasive non-vaccine type strains in Europe compared with the United States. In a recent study by Lipsitch et al.116, only one example of capsular switching in which a novel ST was associated with a non-vaccine serotype was noted (a serotype 35F strain of ST124, which is usually associated with serotype 14). As ST124 of serotype 14 represents one of the most successful invasive clones, it will be important to monitor whether its disease-causing capacity is retained after capsular switching to serotype 35F, a serotype that is rarely associated with invasive disease in children116. In this study, other examples of emerging non-vaccine serotype strains could have been selected from an already existing pool of clones with non-vaccine serotype capsules116.
Future perspectives
We are far from understanding the precise nature of clonal success and clonal dynamics for any bacterial species. Recent advances in sequence technology are likely to lead to extensive whole-genome sequencing of clinical isolates and comparative genomics combined with in vivo studies of potential virulence markers. Mathematical modelling based on observed data will also give us some clues to the mechanism behind successful clonal spread within the community, taking into account bacterial, host and environmental factors. These kinds of analyses, in which information from molecular epidemiological studies of clinical isolates can be combined with clinical information about the patients infected, will lead to a new understanding of why some bacterial clones spread successfully whereas others disappear. This will eventually lead to better strategies for the prevention of clonal transmission in our society.
A high number of S. pneumoniae genomes are being sequenced, which will provide information about the so-called pan-genome of this species and on whether this genome is open (whether additional new genes will be discovered within each genome). The pattern of presence or absence of variable genes, polymorphisms in individual genes and gene-expression data needs to be combined with clinical, epidemiological and human susceptibility data to fully understand the interplay between clone and host properties that promote invasive disease by an organism that normally behaves as an innocent colonizer of the nasopharynx. Because the pneumococcus is not the only inhabitant of its ecological niche, we need more information about the bacterial population structure and dynamics in the nasopharynx, a task that can be approached by metagenomic analyses.
References
Fraser, C., Hanage, W. P. & Spratt, B. G. Recombination and the nature of bacterial speciation. Science 315, 476–480 (2007).Discusses conceptual models of bacterial speciation and suggests that the rate of recombination and its relationship to genetic divergence have a strong influence on the outcome of speciation.
Falush, D. et al. Recombination and mutation during long-term gastric colonization by Helicobacter pylori: estimates of clock rates, recombination size, and minimal age. Proc. Natl Acad. Sci. USA 98, 15056–15061 (2001).
Feil, E. J., Enright, M. C. & Spratt, B. G. Estimating the relative contributions of mutation and recombination to clonal diversification: a comparison between Neisseria meningitidis and Streptococcus pneumoniae. Res. Microbiol. 151, 465–469 (2000).
Spratt, B. G., Hanage, W. P. & Feil, E. J. The relative contributions of recombination and point mutation to the diversification of bacterial clones. Curr. Opin. Microbiol. 4, 602–606 (2001).
Cole, S. T. et al. Deciphering the biology of Mycobacterium tuberculosis from the complete genome sequence. Nature 393, 537–544 (1998).
Dos Vultos, T. et al. Evolution and diversity of clonal bacteria: the paradigm of Mycobacterium tuberculosis. PLoS ONE 3, e1538 (2008).
Becq, J. et al. Contribution of horizontally acquired genomic islands to the evolution of the tubercle bacilli. Mol. Biol. Evol. 24, 1861–1871 (2007).
Hanage, W. P., Spratt, B. G., Turner, K. M. & Fraser, C. Modelling bacterial speciation. Philos. Trans. R. Soc. Lond. B 361, 2039–2044 (2006).
Fraser, C., Hanage, W. P. & Spratt, B. G. Neutral microepidemic evolution of bacterial pathogens. Proc. Natl Acad. Sci. USA 102, 1968–1973 (2005).Shows that the genetic structure of three important human–pathogen populations can be explained by using a simple evolutionary model that is based on neutral mutational drift, modulated by recombination, and incorporates the impact of epidemic transmission in local populations.
Bjorkholm, B. et al. Mutation frequency and biological cost of antibiotic resistance in Helicobacter pylori. Proc. Natl Acad. Sci. USA 98, 14607–14612 (2001).
Hall, L. M. & Henderson-Begg, S. K. Hypermutable bacteria isolated from humans — a critical analysis. Microbiology 152, 2505–2514 (2006).
Gould, C. V., Sniegowski, P. D., Shchepetov, M., Metlay, J. P. & Weiser, J. N. Identifying mutator phenotypes among fluoroquinolone-resistant strains of Streptococcus pneumoniae using fluctuation analysis. Antimicrob. Agents Chemother. 51, 3225–3229 (2007).
Hogardt, M. et al. Stage-specific adaptation of hypermutable Pseudomonas aeruginosa isolates during chronic pulmonary infection in patients with cystic fibrosis. J. Infect. Dis. 195, 70–80 (2007).
Pal, C., Macia, M. D., Oliver, A., Schachar, I. & Buckling, A. Coevolution with viruses drives the evolution of bacterial mutation rates. Nature 450, 1079–1081 (2007).
Tettelin, H. et al. Complete genome sequence of a virulent isolate of Streptococcus pneumoniae. Science 293, 498–506 (2001).Describes the whole genome sequence for the invasive strain TIGR4.
Mrazek, J., Gaynon, L. H. & Karlin, S. Frequent oligonucleotide motifs in genomes of three streptococci. Nucleic Acids Res. 30, 4216–4221 (2002).
Henriqus Normark, B. et al. Clonal analysis of Streptococcus pneumoniae nonsusceptible to penicillin at day-care centers with index cases, in a region with low incidence of resistance: emergence of an invasive type 35B clone among carriers. Microb. Drug Resist. 9, 337–344 (2003).
Sa-Leao, R. et al. High rates of transmission of and colonization by Streptococcus pneumoniae and Haemophilus influenzae within a day care center revealed in a longitudinal study. J. Clin. Microbiol. 46, 225–234 (2008).
Claverys, J. P., Martin, B. & Havarstein, L. S. Competence-induced fratricide in streptococci. Mol. Microbiol. 64, 1423–1433 (2007).
Johnsborg, O., Eldholm, V., Bjornstad, M. L. & Havarstein, L. S. A predatory mechanism dramatically increases the efficiency of lateral gene transfer in Streptococcus pneumoniae and related commensal species. Mol. Microbiol. 69, 245–253 (2008).
Haruma, K. Trend toward a reduced prevalence of Helicobacter pylori infection, chronic gastritis, and gastric cancer in Japan. Gastroenterol. Clin. North Am. 29, 623–631 (2000).
Manuel, D., Cutler, A., Goldstein, J., Fennerty, M. B. & Brown, K. Decreasing prevalence combined with increasing eradication of Helicobacter pylori infection in the United States has not resulted in fewer hospital admissions for peptic ulcer disease-related complications. Aliment. Pharmacol. Ther. 25, 1423–1427 (2007).
Suerbaum, S. & Josenhans, C. Helicobacter pylori evolution and phenotypic diversification in a changing host. Nature Rev. Microbiol. 5, 441–452 (2007).
Magalhaes Queiroz, D. M. & Luzza, F. Epidemiology of Helicobacter pylori infection. Helicobacter 11 (Suppl. 1), 1–5 (2006).
Nygaard, T. K., Deleo, F. R. & Voyich, J. M. Community-associated methicillin-resistant Staphylococcus aureus skin infections: advances toward identifying the key virulence factors. Curr. Opin. Infect. Dis. 21, 147–152 (2008).
Madhi, S. A., Schoub, B., Simmank, K., Blackburn, N. & Klugman, K. P. Increased burden of respiratory viral associated severe lower respiratory tract infections in children infected with human immunodeficiency virus type-1. J. Pediatr. 137, 78–84 (2000).
Klugman, K. P. & Madhi, S. A. Pneumococcal vaccines and flu preparedness. Science 316, 49–50 (2007).
Sun, K. & Metzger, D. W. Inhibition of pulmonary antibacterial defense by interferon-γ during recovery from influenza infection. Nature Med. 14, 558–564 (2008).
Peltola, V. T., Murti, K. G. & McCullers, J. A. Influenza virus neuraminidase contributes to secondary bacterial pneumonia. J. Infect. Dis. 192, 249–257 (2005).
Sjostrom, K. et al. Clonal and capsular types decide whether pneumococci will act as a primary or opportunistic pathogen. Clin. Infect. Dis. 42, 451–459 (2006).
Rosenstiel, P. et al. Influence of polymorphisms in the NOD1/CARD4 and NOD2/CARD15 genes on the clinical outcome of Helicobacter pylori infection. Cell. Microbiol. 8, 1188–1198 (2006).
Kotb, M. et al. An immunogenetic and molecular basis for differences in outcomes of invasive group A streptococcal infections. Nature Med. 8, 1398–1404 (2002).
Roy, S. et al. MBL genotype and risk of invasive pneumococcal disease: a case-control study. Lancet 359, 1569–1573 (2002).
Hjuler, T. et al. Genetic susceptibility to severe infection in families with invasive pneumococcal disease. Am. J. Epidemiol. 167, 814–819 (2008).
Hjuler, T. et al. Perinatal and crowding-related risk factors for invasive pneumococcal disease in infants and young children: a population-based case-control study. Clin. Infect. Dis. 44, 1051–1056 (2007).
Karlsson, D., Jansson, A., Normark, B. H. & Nilsson, P. An individual-based network model to evaluate interventions for controlling pneumococcal transmission. BMC Infect. Dis. 8, 83 (2008).
Aspholm-Hurtig, M. et al. Functional adaptation of BabA, the H. pylori ABO blood group antigen binding adhesin. Science 305, 519–522 (2004).
Sokurenko, E. V. et al. Pathogenic adaptation of Escherichia coli by natural variation of the FimH adhesin. Proc. Natl Acad. Sci. USA 95, 8922–8926 (1998).
Weissman, S. J. et al. Clonal analysis reveals high rate of structural mutations in fimbrial adhesins of extraintestinal pathogenic Escherichia coli. Mol. Microbiol. 59, 975–988 (2006).
Uria, M. J. et al. A generic mechanism in Neisseria meningitidis for enhanced resistance against bactericidal antibodies. J. Exp. Med. 205, 1423–1434 (2008).
Heidelberger, M. Precipitating cross-reactions among pneumococcal types. Infect. Immun. 41, 1234–1244 (1983).
Henrichsen, J. Six newly recognized types of Streptococcus pneumoniae. J. Clin. Microbiol. 33, 2759–2762 (1995).
Park, I. H. et al. Discovery of a new capsular serotype (6C) within serogroup 6 of Streptococcus pneumoniae. J. Clin. Microbiol. 45, 1225–1233 (2007).
Brueggemann, A. B. et al. Temporal and geographic stability of the serogroup-specific invasive disease potential of Streptococcus pneumoniae in children. J. Infect. Dis. 190, 1203–1211 (2004).A meta-analysis study in which seven data sets of invasive and carriage pneumococcal isolates recovered from children were used to determine invasive disease potential for each pneumococcal serotype.
Henriques, B. et al. Molecular epidemiology of Streptococcus pneumoniae causing invasive disease in 5 countries. J. Infect. Dis. 182, 833–839 (2000).
Henriques Normark, B. et al. Dynamics of penicillin-susceptible clones in invasive pneumococcal disease. J. Infect. Dis. 184, 861–869 (2001).
Normark, B. H. et al. Changes in serotype distribution may hamper efficacy of pneumococcal conjugate vaccines in children. Scand. J. Infect. Dis. 33, 848–850 (2001).
Foster, D. et al. Invasive pneumococcal disease: epidemiology in children and adults prior to implementation of the conjugate vaccine in the Oxfordshire region, England. J. Med. Microbiol. 57, 480–487 (2008).
Jefferson, T., Ferroni, E., Curtale, F., Giorgi Rossi, P. & Borgia, P. Streptococcus pneumoniae in western Europe: serotype distribution and incidence in children less than 2 years old. Lancet Infect. Dis. 6, 405–410 (2006).
Brueggemann, A. B. et al. Clonal relationships between invasive and carriage Streptococcus pneumoniae and serotype- and clone-specific differences in invasive disease potential. J. Infect. Dis. 187, 1424–1432 (2003).
Sandgren, A. et al. Effect of clonal and serotype-specific properties on the invasive capacity of Streptococcus pneumoniae. J. Infect. Dis. 189, 785–796 (2004).
Hanage, W. P. et al. Invasiveness of serotypes and clones of Streptococcus pneumoniae among children in Finland. Infect. Immun. 73, 431–435 (2005).
Sandgren, A. et al. Virulence in mice of pneumococcal clonal types with known invasive disease potential in humans. J. Infect. Dis. 192, 791–800 (2005).
Alanee, S. R. et al. Association of serotypes of Streptococcus pneumoniae with disease severity and outcome in adults: an international study. Clin. Infect. Dis. 45, 46–51 (2007).
Dagerhamn, J. et al. Determination of accessory gene patterns predicts the same relatedness among strains of Streptococcus pneumoniae as sequencing of housekeeping genes does and represents a novel approach in molecular epidemiology. J. Clin. Microbiol. 46, 863–868 (2008).Sequence variations in housekeeping genes of S. pneumoniae assessed by MLST are correlated to whole-genome microarray analyses.
Yaro, S. et al. Epidemiological and molecular characteristics of a highly lethal pneumococcal meningitis epidemic in Burkina Faso. Clin. Infect. Dis. 43, 693–700 (2006).
Sjöström, K. et al. Clonal success of piliated penicillin nonsusceptible pneumococci. Proc. Natl Acad. Sci. USA 104, 12907–12912 (2007).Describes the pilus, a pneumococcal virulence factor that is important for the spread of successful PNSP clones, such as ST156.
Kilian, M. et al. Evolution of Streptococcus pneumoniae and its close commensal relatives. PLoS ONE 3, e2683 (2008).
Hanage, W. P., Fraser, C. & Spratt, B. G. The impact of homologous recombination on the generation of diversity in bacteria. J. Theor. Biol. 239, 210–219 (2006).
Obert, C. et al. Identification of a candidate Streptococcus pneumoniae core genome and regions of diversity correlated with invasive pneumococcal disease. Infect. Immun. 74, 4766–4777 (2006).
Hiller, N. L. et al. Comparative genomic analyses of seventeen Streptococcus pneumoniae strains: insights into the pneumococcal supragenome. J. Bacteriol. 189, 8186–8195 (2007).Discusses sequence data from eight nasopharyngeal strains and nine publicly available genomes of S. pneumoniae.
Bruckner, R., Nuhn, M., Reichmann, P., Weber, B. & Hakenbeck, R. Mosaic genes and mosaic chromosomes — genomic variation in Streptococcus pneumoniae. Int. J. Med. Microbiol. 294, 157–168 (2004).
Hakenbeck, R. et al. Mosaic genes and mosaic chromosomes: intra- and interspecies genomic variation of Streptococcus pneumoniae. Infect. Immun. 69, 2477–2486 (2001).
Gal-Mor, O. & Finlay, B. B. Pathogenicity islands: a molecular toolbox for bacterial virulence. Cell. Microbiol. 8, 1707–1719 (2006).
Feil, E. J., Smith, J. M., Enright, M. C. & Spratt, B. G. Estimating recombinational parameters in Streptococcus pneumoniae from multilocus sequence typing data. Genetics 154, 1439–1450 (2000).
Hava, D. L., Hemsley, C. J. & Camilli, A. Transcriptional regulation in the Streptococcus pneumoniae rlrA pathogenicity islet by RlrA. J. Bacteriol. 185, 413–421 (2003).
Barocchi, M. A. et al. A pneumococcal pilus influences virulence and host inflammatory responses. Proc. Natl Acad. Sci. USA 103, 2857–2862 (2006).The first report to describe a pilus on the surface of pneumococci.
Moschioni, M. et al. Streptococcus pneumoniae contains 3 rlrA pilus variants that are clonally related. J. Infect. Dis. 197, 888–896 (2008).
Aguiar, S. I., Serrano, I., Pinto, F. R., Melo-Cristino, J. & Ramirez, M. The presence of the pilus locus is a clonal property among pneumococcal invasive isolates. BMC Microbiol. 8, 41 (2008).
Basset, A. et al. Association of the pneumococcal pilus with certain capsular serotypes but not with increased virulence. J. Clin. Microbiol. 45, 1684–1689 (2007).
Gianfaldoni, C. et al. Streptococcus pneumoniae pilus subunits protect mice against lethal challenge. Infect. Immun. 75, 1059–1062 (2007).
Bagnoli, F. et al. A second pilus type in Streptococcus pneumoniae is prevalent in emerging serotypes and mediates adhesion to host cells. J. Bacteriol. 190, 5480–5492 (2008).
Rose, L. et al. Antibodies against PsrP, a novel Streptococcus pneumoniae adhesin, block adhesion and protect mice against pneumococcal challenge. J. Infect. Dis. 198, 375–383 (2008).
Hava, D. L. & Camilli, A. Large-scale identification of serotype 4 Streptococcus pneumoniae virulence factors. Mol. Microbiol. 45, 1389–1406 (2002).
Rosch, J. W., Mann, B., Thornton, J., Sublett, J. & Tuomanen, E. Convergence of regulatory networks on the pilus locus of Streptococcus pneumoniae. Infect. Immun. 76, 3187–3196 (2008).
Dave, S., Carmicle, S., Hammerschmidt, S., Pangburn, M. K. & McDaniel, L. S. Dual roles of PspC, a surface protein of Streptococcus pneumoniae, in binding human secretory IgAand factor H. J. Immunol. 173, 471–477 (2004).
Iannelli, F., Oggioni, M. R. & Pozzi, G. Allelic variation in the highly polymorphic locus pspC of Streptococcus pneumoniae. Gene 284, 63–71 (2002).
Hirst, R. A., Kadioglu, A., O'Callaghan, C. & Andrew, P. W. The role of pneumolysin in pneumococcal pneumonia and meningitis. Clin. Exp. Immunol. 138, 195–201 (2004).
Jefferies, J. M. et al. Presence of nonhemolytic pneumolysin in serotypes of Streptococcus pneumoniae associated with disease outbreaks. J. Infect. Dis. 196, 936–944 (2007).
Kirkham, L. A. et al. Identification of invasive serotype 1 pneumococcal isolates that express nonhemolytic pneumolysin. J. Clin. Microbiol. 44, 151–159 (2006).
Bekri, H. et al. Streptococcus pneumoniae serotypes involved in children with pleural empyemas in France. Arch. Pediatr. 14, 239–243 (2007).
Fletcher, M., Leeming, J., Cartwright, K. & Finn, A. Childhood empyema: limited potential impact of 7-valent pneumococcal conjugate vaccine. Pediatr. Infect. Dis. J. 25, 559–560 (2006).
Laible, G. & Hakenbeck, R. Penicillin-binding proteins in β-lactam-resistant laboratory mutants of Streptococcus pneumoniae. Mol. Microbiol. 1, 355–363 (1987).
Laible, G., Spratt, B. G. & Hakenbeck, R. Interspecies recombinational events during the evolution of altered PBP 2x genes in penicillin-resistant clinical isolates of Streptococcus pneumoniae. Mol. Microbiol. 5, 1993–2002 (1991).
Decousser, J. W. et al. Invasive Streptococcus pneumoniae in France: antimicrobial resistance, serotype, and molecular epidemiology findings from a monthly national study in 2000 to 2002. Antimicrob. Agents Chemother. 48, 3636–3639 (2004).
McGee, L. et al. Nomenclature of major antimicrobial-resistant clones of Streptococcus pneumoniae defined by the pneumococcal molecular epidemiology network. J. Clin. Microbiol. 39, 2565–2571 (2001).
Soares, S., Kristinsson, K. G., Musser, J. M. & Tomasz, A. Evidence for the introduction of a multiresistant clone of serotype 6B Streptococcus pneumoniae from Spain to Iceland in the late 1980s. J. Infect. Dis. 168, 158–163 (1993).
Zemlickova, H., Melter, O. & Urbaskova, P. Epidemiological relationships among penicillin non-susceptible Streptococcus pneumoniae strains recovered in the Czech Republic. J. Med. Microbiol. 55, 437–442 (2006).
Hogberg, L. et al. Penicillin-resistant pneumococci in Sweden 1997–2003: increased multiresistance despite stable prevalence and decreased antibiotic use. Microb. Drug Resist. 12, 16–22 (2006).
Pozzi, G. et al. Competence for genetic transformation in encapsulated strains of Streptococcus pneumoniae: two allelic variants of the peptide pheromone. J. Bacteriol. 178, 6087–6090 (1996).
Havarstein, L. S., Coomaraswamy, G. & Morrison, D. A. An unmodified heptadecapeptide pheromone induces competence for genetic transformation in Streptococcus pneumoniae. Proc. Natl Acad. Sci. USA 92, 11140–11144 (1995).
Claverys, J. P., Prudhomme, M. & Martin, B. Induction of competence regulons as a general response to stress in gram-positive bacteria. Annu. Rev. Microbiol. 60, 451–475 (2006).
Johnsborg, O., Eldholm, V. & Havarstein, L. S. Natural genetic transformation: prevalence, mechanisms and function. Res. Microbiol. 158, 767–778 (2007).
Iannelli, F., Oggioni, M. R. & Pozzi, G. Sensor domain of histidine kinase ComD confers competence pherotype specificity in Streptoccoccus pneumoniae. FEMS Microbiol. Lett. 252, 321–326 (2005).
Whatmore, A. M., Barcus, V. A. & Dowson, C. G. Genetic diversity of the streptococcal competence (com) gene locus. J. Bacteriol. 181, 3144–3154 (1999).
Brueggemann, A. B. & Spratt, B. G. Geographic distribution and clonal diversity of Streptococcus pneumoniae serotype 1 isolates. J. Clin. Microbiol. 41, 4966–4970 (2003).
Whitney, C. G. et al. Decline in invasive pneumococcal disease after the introduction of protein-polysaccharide conjugate vaccine. N. Engl. J. Med. 348, 1737–1746 (2003).Shows that the seven-valent pneumococcal conjugate vaccine prevents disease in vaccinated young children, which could reduce the rate of pneumococcal disease in adults in the United States.
Hammitt, L. L. et al. Indirect effect of conjugate vaccine on adult carriage of Streptococcus pneumoniae: an explanation of trends in invasive pneumococcal disease. J. Infect. Dis. 193, 1487–1494 (2006).
Hsu, K. K. & Pelton, S. I. Heptavalent pneumococcal conjugate vaccine: current and future impact. Expert. Rev. Vaccines 2, 619–631 (2003).
Poehling, K. A. et al. Invasive pneumococcal disease among infants before and after introduction of pneumococcal conjugate vaccine. JAMA 295, 1668–1674 (2006).
Singleton, R. J. et al. Invasive pneumococcal disease caused by nonvaccine serotypes among alaska native children with high levels of 7-valent pneumococcal conjugate vaccine coverage. JAMA 297, 1784–1792 (2007).
Whitney, C. G. et al. Effectiveness of seven-valent pneumococcal conjugate vaccine against invasive pneumococcal disease: a matched case-control study. Lancet 368, 1495–1502 (2006).
Hicks, L. A. et al. Incidence of pneumococcal disease due to non-pneumococcal conjugate vaccine (PCV7) serotypes in the United States during the era of widespread PCV7 vaccination, 1998–2004. J. Infect. Dis. 196, 1346–1354 (2007).Provides data on serotype replacement in invasive pneumococcal disease after vaccine introduction in the United States.
Beall, B. et al. Pre- and postvaccination clonal compositions of invasive pneumococcal serotypes for isolates collected in the United States in 1999, 2001, and 2002. J. Clin. Microbiol. 44, 999–1017 (2006).
Brueggemann, A. B., Pai, R., Crook, D. W. & Beall, B. Vaccine escape recombinants emerge after pneumococcal vaccination in the United States. PLoS Pathog. 3, e168 (2007).
Hanage, W. P. Serotype replacement in invasive pneumococcal disease: where do we go from here? J. Infect. Dis. 196, 1282–1284 (2007).
Jacobs, M. R. et al. Changes in serotypes and antimicrobial susceptibility of invasive Streptococcus pneumoniae strains in Cleveland: a quarter century of experience. J. Clin. Microbiol. 46, 982–990 (2008).
Kaplan, S. L. et al. Decrease of invasive pneumococcal infections in children among 8 children's hospitals in the United States after the introduction of the 7-valent pneumococcal conjugate vaccine. Pediatrics 113, 443–449 (2004).
Pai, R. et al. Postvaccine genetic structure of Streptococcus pneumoniae serotype 19A from children in the United States. J. Infect. Dis. 192, 1988–1995 (2005).
Pelton, S. I. et al. Emergence of 19A as virulent and multidrug resistant Pneumococcus in Massachusetts following universal immunization of infants with pneumococcal conjugate vaccine. Pediatr. Infect. Dis. J. 26, 468–472 (2007).
Pichichero, M. E. & Casey, J. R. Emergence of a multiresistant serotype 19A pneumococcal strain not included in the 7-valent conjugate vaccine as an otopathogen in children. JAMA 298, 1772–1778 (2007).Describes a non-vaccine pneumococcal serotype 19A strain that emerged in the United States as an otopathogen that is resistant to all FDA-approved antibiotics for treatment of acute otitis media in children.
Dias, R. & Canica, M. Invasive pneumococcal disease in Portugal prior to and after the introduction of pneumococcal heptavalent conjugate vaccine. FEMS Immunol. Med. Microbiol. 51, 35–42 (2007).
Farrell, D. J., Klugman, K. P. & Pichichero, M. Increased antimicrobial resistance among nonvaccine serotypes of Streptococcus pneumoniae in the pediatric population after the introduction of 7-valent pneumococcal vaccine in the United States. Pediatr. Infect. Dis. J. 26, 123–128 (2007).
Moore, M. R. et al. Population snapshot of emergent Streptococcus pneumoniae serotype 19A in the United States, 2005. J. Infect. Dis. 197, 1016–1027 (2008).
Hanage, W. P. et al. Diversity and antibiotic resistance among nonvaccine serotypes of Streptococcus pneumoniae carriage isolates in the post-heptavalent conjugate vaccine era. J. Infect. Dis. 195, 347–352 (2007).
Lipsitch, M. et al. Strain characteristics of Streptococcus pneumoniae carriage and invasive disease isolates during a cluster-randomized clinical trial of the 7-valent pneumococcal conjugate vaccine. J. Infect. Dis. 196, 1221–1227 (2007).
Diep, B. A. et al. Complete genome sequence of USA300, an epidemic clone of community-acquired meticillin-resistant Staphylococcus aureus. Lancet 367, 731–739 (2006).
Kennedy, A. D. et al. Epidemic community-associated methicillin-resistant Staphylococcus aureus: recent clonal expansion and diversification. Proc. Natl Acad. Sci. USA 105, 1327–1332 (2008).Describes a recent clonal expansion and diversification of a subset of S. aureus isolates classified as USA300 and suggests that small genetic changes in the bacterial genome can have profound effects on virulence.
Fowler, V. G. Jr et al. Potential associations between hematogenous complications and bacterial genotype in Staphylococcus aureus infection. J. Infect. Dis. 196, 738–747 (2007).
Ernst, J. D., Trevejo-Nunez, G. & Banaiee, N. Genomics and the evolution, pathogenesis, and diagnosis of tuberculosis. J. Clin. Invest. 117, 1738–1745 (2007).Reviews some of the progress in tuberculosis research that has resulted from knowledge of its genome sequence.
Gandotra, S., Schnappinger, D., Monteleone, M., Hillen, W. & Ehrt, S. In vivo gene silencing identifies the Mycobacterium tuberculosis proteasome as essential for the bacteria to persist in mice. Nature Med. 13, 1515–1520 (2007).
Dormans, J. et al. Correlation of virulence, lung pathology, bacterial load and delayed type hypersensitivity responses after infection with different Mycobacterium tuberculosis genotypes in a BALB/c mouse model. Clin. Exp. Immunol. 137, 460–468 (2004).
Lopez, B. et al. A marked difference in pathogenesis and immune response induced by different Mycobacterium tuberculosis genotypes. Clin. Exp. Immunol. 133, 30–37 (2003).
Newton, S. M. et al. A deletion defining a common Asian lineage of Mycobacterium tuberculosis associates with immune subversion. Proc. Natl Acad. Sci. USA 103, 15594–15598 (2006).
Reed, M. B. et al. A glycolipid of hypervirulent tuberculosis strains that inhibits the innate immune response. Nature 431, 84–87 (2004).
Thwaites, G. et al. Relationship between Mycobacterium tuberculosis genotype and the clinical phenotype of pulmonary and meningeal tuberculosis. J. Clin. Microbiol. 46, 1363–1368 (2008).
Feil, E. J., Li, B. C., Aanensen, D. M., Hanage, W. P. & Spratt, B. G. eBURST: inferring patterns of evolutionary descent among clusters of related bacterial genotypes from multilocus sequence typing data. J. Bacteriol. 186, 1518–1530 (2004).
Feil, E. J. Small change: keeping pace with microevolution. Nature Rev. Microbiol. 2, 483–495 (2004).
Acknowledgements
We thank present and past members of our laboratory for their contributions to the work discussed in this Review. Work done in our laboratory was supported by the Swedish Research Council, the Torsten and Ragnar Söderbergs foundation, the Swedish Royal Academy of Sciences, the EU Sixth Framework Programme and the Swedish Foundation for Strategic Research.
Author information
Authors and Affiliations
Corresponding author
Related links
Related links
DATABASES
Entrez Genome
S. aureus subsp. aureus USA300
Entrez Genome Project
Entrez Protein
FURTHER INFORMATION
Glossary
- Clonal cluster
-
Strains belong to a clonal cluster if they share at least five out of the seven housekeeping genes according to multilocus sequence typing.
- Serotype
-
Bacteria can be divided into serotypes depending on differences in their capsular polysaccharides.
- Odds ratio
-
(OR). An estimate of the likelihood that a serotype is associated with invasive disease or carriage. A serotype with an OR of >1 is more likely to cause invasive disease than colonize the nasopharynx.
- Clonal complex
-
(CC). Strains from a particular CC have the same ancestor according to multilocus sequence typing and the algorithm eBURST (>250 groups and >700 singletons have been described so far).
- Clone
-
Can be defined using molecular epidemiological methods, such as pulsed field gel electrophoresis or multilocus sequence typing.
- Empyema
-
The presence of pus in a body cavity, especially the pleural cavity.
- Genotype
-
The entire genetic constitution of an organism or the genetic composition at a specific gene locus or set of loci.
- Single-locus sequence variant
-
A strain that differs in only one of the seven alleles detected with multilocus sequence typing.
Rights and permissions
About this article
Cite this article
Henriques-Normark, B., Blomberg, C., Dagerhamn, J. et al. The rise and fall of bacterial clones: Streptococcus pneumoniae. Nat Rev Microbiol 6, 827–837 (2008). https://doi.org/10.1038/nrmicro2011
Issue Date:
DOI: https://doi.org/10.1038/nrmicro2011
This article is cited by
-
Molecular analyses identifies new domains and structural differences among Streptococcus pneumoniae immune evasion proteins PspC and Hic
Scientific Reports (2021)
-
The histone demethylase KDM6B fine-tunes the host response to Streptococcus pneumoniae
Nature Microbiology (2020)
-
Frequency-dependent selection in vaccine-associated pneumococcal population dynamics
Nature Ecology & Evolution (2017)
-
Lineage structure of Streptococcus pneumoniae may be driven by immune selection on the groEL heat-shock protein
Scientific Reports (2017)
-
A longitudinal study of natural antibody development to pneumococcal surface protein A families 1 and 2 in Papua New Guinean Highland children: a cohort study
Pneumonia (2016)