Key Points
-
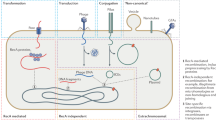
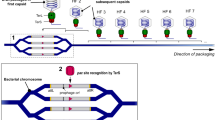
Prokaryotes transfer DNA between cells by three processes: transformation, transduction and conjugation. Transduction and conjugation depend on specialized mobile genetic elements (MGEs), which include most large plasmids and certain bacteriophages (phages).
-
Prokaryotes also possess a third class of MGEs called transposons. These elements can move and rearrange chromosomal DNA in the cell. Transposons move from cell to cell through plasmids, phages, or their derivatives called integrative conjugative elements (ICEs).
-
MGEs can mediate intra- or intercellular DNA trafficking because they have unique (core or backbone) genes that allow them to replicate independently of the cellular chromosome, to engage in homologous or non-homologous recombination, and to extrude (plasmid) or package (phage) DNA for efficient movement between cells.
-
In addition to their core genes, MGEs typically carry several different accessory genes that provide their host cell with a selective advantage, such as antibiotic resistance, virulence factors, or unusual metabolic pathways. Indeed, most medically and economically important bacterial phenotypes are encoded by MGEs.
-
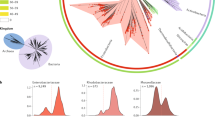
Although MGEs are the main agents of horizontal gene transfer (HGT), relatively few have been sequenced and analysis of their genomic and phylogenetic properties lags behind that of organismal chromosomes. Specifically, the major databases do not curate plasmid and phage nucleic acid or protein sequences. Sequencing MGE genomes presents unique challenges because phages require suitable hosts for propagation and plasmids must be physically separated from each other and from the host chromosomal DNA. The relatively small size of MGEs (∼5–500 kb) and their varied GC content thwart current automated annotation algorithms.
-
These challenges can be viewed as an opportunity to devise technical and bioinformatics tools for high throughput analysis of MGEs. This is important because understanding prokaryotic evolution requires knowledge of the agents that mediate this process. Such knowledge is essential for controlling problems such as the emergence of highly virulent antibiotic multi-resistant strains.
Abstract
Horizontal genomics is a new field in prokaryotic biology that is focused on the analysis of DNA sequences in prokaryotic chromosomes that seem to have originated from other prokaryotes or eukaryotes. However, it is equally important to understand the agents that effect DNA movement: plasmids, bacteriophages and transposons. Although these agents occur in all prokaryotes, comprehensive genomics of the prokaryotic mobile gene pool or 'mobilome' lags behind other genomics initiatives owing to challenges that are distinct from cellular chromosomal analysis. Recent work shows promise of improved mobile genetic element (MGE) genomics and consequent opportunities to take advantage — and avoid the dangers — of these 'natural genetic engineers'. This review describes MGEs, their properties that are important in horizontal gene transfer, and current opportunities to advance MGE genomics.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Burrus, V. & Waldor, M. K. Shaping bacterial genomes with integrative and conjugative elements. Res. Microbiol. 155, 376–386 (2004). A useful review of the role of ICEs in bacterial evolution.
Scott, J. R. & Churchward, G. G. Conjugative transposition. Annu. Rev. Microbiol. 49, 367–397 (1995).
Toussaint, A. & Merlin, C. Mobile elements as a combination of functional modules. Plasmid 47, 26–35 (2002).
Lawrence, J. G. & Hendrickson, H. Lateral gene transfer: when will adolescence end? Mol. Microbiol. 50, 739–749 (2003). A succinct framing of important questions in horizontal genomics research.
Chaconas, G. & Chen, C. W. in The Bacterial Chromosome (ed. Higgins, P. N.) 525–539 (ASM Press, Washington DC, 2004).
Stewart, P. E., Byram, R., Grimm, D., Tilly, K. & Rosa, P. A. The plasmids of Borrelia burgdorferi: essential genetic elements of a pathogen. Plasmid 53, 1–13 (2005).
Hinnebusch, J. & Tilly, K. Linear plasmids and chromosomes in bacteria. Mol. Microbiol. 10, 917–922 (1993).
Lilley, A., Young, P. & Bailey, M. J. in The Horizontal Gene Pool: Bacterial Plasmids and Gene Spread (ed. Thomas, C. M.) 287–300 (Harwood Academic, Amsterdam, Netherlands, 2000).
Dahlberg, C. & Chao, L. Amelioration of the cost of conjugative plasmid carriage in Eschericha coli K12. Genetics 165, 1641–1649 (2003). A discussion of the cost of maintaining plasmids: why do bacteria tolerate them?
Bradley, D. E., Taylor, D. E. & Cohen, D. R. Specification of surface mating systems among conjugative drug resistance plasmids in Escherichia coli K-12. J. Bacteriol. 143, 1466–1470 (1980).
Novick, R. P. Plasmid incompatibility. Microbiol. Rev. 51, 381–395 (1987).
Couturier, M., Bex, F., Bergquist, P. L. & Maas, W. K. Identification and classification of bacterial plasmids. Microbiol. Rev. 52, 375–395 (1988).
Helinski, D.R. in The Horizontal Gene Pool: Bacterial Plasmids and Gene Spread (ed. Thomas, C. M.) 1–21 (Harwood Academic, Amsterdam, Netherlands, 2000).
Redfield, R. J. et al. A novel CRP-dependent regulon controls expression of competence genes in Haemophilus influenzae. J. Mol. Biol. 347, 735–747 (2005).
Gomis-Ruth, F. X., Sola, M., de la Cruz, F. & Coll, M. Coupling factors in macromolecular type-IV secretion machineries. Curr. Pharm. Des. 10, 1551–1565 (2004).
Schroder, G. et al. TraG-like proteins of DNA transfer systems and of the Helicobacter pylori type IV secretion system: inner membrane gate for exported substrates? J. Bacteriol. 184, 2767–2779 (2002).
Frost, L. S. in Conjugation (ed. Clewell, D. B.) 189–221 (Plenum, New York, 1993).
Possoz, C., Ribard, C., Gagnat, J., Pernodet, J. L. & Guerineau, M. The integrative element pSAM2 from Streptomyces: kinetics and mode of conjugal transfer. Mol. Microbiol. 42, 159–66 (2001).
Pettis, G. S. & Cohen, S. N. Unraveling the essential role in conjugation of the Tra protein of Streptomyces lividans plasmid pIJ101. Antonie Van Leeuwenhoek 79, 247–250 (2001).
Garret, R. A. et al. in Plasmid Biology (eds Funnell, B. E. & Phillips, G. J.) 377–392 (ASM Press, Washington DC, 2004).
Wang, J., Parsons, L. M. & Derbyshire, K. M. Unconventional conjugal DNA transfer in mycobacteria. Nature Genet. 34, 80–84 (2003).
Lybarger, S. R. & Sandkvist, M. Microbiology. A hitchhiker's guide to type IV secretion. Science 304, 1122–1123 (2004).
Chen, I. & Dubnau, D. DNA uptake during bacterial transformation. Nature Rev. Microbiol. 2, 241–249 (2004).
Gomis-Ruth, F. X. et al. The bacterial conjugation protein TrwB resembles ring helicases and F1-ATPase. Nature 409, 637–641 (2001).
Zechner, E. L. et al. in The Horizontal Gene Pool: Bacterial Plasmids and Gene Spread (ed. Thomas, C. M.) 87–174 (Harwood Academic, Amsterdam, Netherlands, 2000).
Cascales, E. & Christie, P. J. The versatile bacterial type IV secretion systems. Nature Rev. Microbiol. 1, 137–149 (2003).
Lawley, T. D., Wilkins, B. M. & Frost, L. S. in Plasmid Biology (eds Funnell, B. E. & Phillips, G. J.) 203–226 (ASM Press, Washington DC, 2004).
Lawley, T. D., Klimke, W. A., Gubbins, M. J. & Frost, L. S. F factor conjugation is a true type IV secretion system. FEMS Microbiol. Lett. 224, 1–15 (2003). Defines the relationship of F-like T4SSs to P-like T4SSs.
Boltner, D. & Osborn, A. M. Structural comparison of the integrative and conjugative elements R391, pMERPH, R997, and SXT. Plasmid 51, 12–23 (2004).
Peabody, C. R. et al. Type II protein secretion and its relationship to bacterial type IV pili and archaeal flagella. Microbiology 149, 3051–3072 (2003).
He, S. Y., Nomura, K. & Whittam, T. S. Type III protein secretion mechanism in mammalian and plant pathogens. Biochim. Biophys. Acta 1694, 181–206 (2004).
Planet, P. J., Kachlany, S. C., DeSalle, R. & Figurski, D. H. Phylogeny of genes for secretion NTPases: identification of the widespread tadA subfamily and development of a diagnostic key for gene classification. Proc. Natl Acad. Sci. USA 98, 2503–2508 (2001).
Savvides, S. N. et al. VirB11 ATPases are dynamic hexameric assemblies: new insights into bacterial type IV secretion. EMBO J. 22, 1969–1980 (2003).
Kim, S. R. & Komano, T. The plasmid R64 thin pilus identified as a type IV pilus. J. Bacteriol. 179, 3594–3603 (1997).
Model, P. & Russel, M. Prokaryotic secretion. Cell 61, 739–741 (1990).
Macnab, R. M. Type III flagellar protein export and flagellar assembly. Biochim. Biophys. Acta 1694, 207–217 (2004).
Averhoff, B. DNA transport and natural transformation in mesophilic and thermophilic bacteria. J. Bioenerg. Biomembr. 36, 25–33 (2004).
Cascales, E. & Christie, P. J. Definition of a bacterial type IV secretion pathway for a DNA substrate. Science 304, 1170–1173 (2004). Immunoprecipitation of Vir protein–DNA complexes defines the path of the DNA through the conjugative pore.
Hamilton, H. L., Dominguez, N. M., Schwartz, K. J., Hackett, K. T. & Dillard, J. P. Neisseria gonorrhoeae secretes chromosomal DNA via a novel type IV secretion system. Mol. Microbiol. 55, 1704–1721 (2005).
Cascales, E. & Christie, P. J. Agrobacterium VirB10, an ATP energy sensor required for type IV secretion. Proc. Natl Acad. Sci. USA 101, 17228–17233 (2004). VirB10 has TonB-like properties indicating that it is involved in signalling between the outer and inner membranes.
Christie, P. J. Type IV secretion:the Agrobacterium VirB/D4 and related conjugation systems. Biochim. Biophys. Acta 1694, 219–234 (2004).
Kalkum, M., Eisenbrandt, R. & Lanka, E. Protein circlets as sex pilus subunits. Curr. Protein Pept. Sci. 5, 417–424 (2004).
Lai, E. M., Eisenbrandt, R., Kalkum, M., Lanka, E. & Kado, C. I. Biogenesis of T pili in Agrobacterium tumefaciens requires precise VirB2 propilin cleavage and cyclization. J. Bacteriol. 184, 327–330 (2002).
Clewell, D. B. & Francia, M. V. in Plasmid Biology (eds. Funnell, B. E. & Phillips, G. J.) 227–256 (ASM Press, Washington DC, 2004).
Salyers, A. A., Shoemaker, N. B., Stevens, A. M. & Li, L. Y. Conjugative transposons: an unusual and diverse set of integrated gene transfer elements. Microbiol. Rev. 59, 579–90 (1995).
Charlebois, R. L., She, Q., Sprott, D. P., Sensen, C. W. & Garrett, R. A. Sulfolobus genome: from genomics to biology. Curr. Opin. Microbiol. 1, 584–588 (1998).
Wilkins, B. M. & Frost, L. S. in Molecular Medical Microbiology (ed. Sussman, M.) 355–400 (Academic, London, 2001).
Papke, R. T., Koenig, J. E., Rodriguez-Valera, F. & Doolittle, W. F. Frequent recombination in a saltern population of Halorubrum. Science 306, 1928–1929 (2004).
Ramirez-Arcos, S., Fernandez-Herrero, L. A., Marin, I. & Berenguer, J. Anaerobic growth, a property horizontally transferred by an Hfr-like mechanism among extreme thermophiles. J. Bacteriol. 180, 3137–3143 (1998).
Fiers, W. et al. Complete nucleotide sequence of bacteriophage MS2 RNA: primary and secondary structure of the replicase gene. Nature 260, 500–507 (1976). The first published genome sequence, which predates the advent of DNA sequencing techniques.
Sanger, F., Coulson, A. R., Hong, G. F., Hill, D. F. & Petersen, G. B. Nucleotide sequence of bacteriophage lambda DNA. J. Mol. Biol. 162, 729–773 (1982).
Canchaya, C., Fournous, G. & Brussow, H. The impact of prophages on bacterial chromosomes. Mol. Microbiol. 53, 9–18 (2004).
Canchaya, C., Fournous, G., Chibani-Chennoufi, S., Dillmann, M. L. & Brussow, H. Phage as agents of lateral gene transfer. Curr. Opin. Microbiol. 6, 417–424 (2003). A good perspective on how bacterial genomics reveals the main impact of phages on bacterial chromosome evolution.
Pedulla, M. L. et al. Origins of highly mosaic mycobacteriophage genomes. Cell 113, 171–182 (2003).
Merril, C. R., Scholl, D. & Adhya, S. L. The prospect for bacteriophage therapy in Western medicine. Nature Rev. Drug Discov. 2, 489–497 (2003).
Zhang, S. Fabrication of novel biomaterials through molecular self-assembly. Nature Biotechnol. 21, 1171–1178 (2003).
Lwoff, A. Lysogeny. Bacteriol. Rev. 17, 269–337 (1953).
Freeman, V. J. Studies on the virulence of bacteriophage-infected strains of Corynebacterium diphtheriae. J. Bacteriol. 61, 675–688 (1951).
Hendrix, R. W. Bacteriophage genomics. Curr. Opin. Microbiol. 6, 506–511 (2003).
Zinder, N. D. & Lederberg, J. Genetic exchange in Salmonella. J. Bacteriol. 64, 679–699 (1952).
Mizuuchi, K. & Baker, T. in Mobile DNA II (eds. Craig, N. L., Craigie, R., Gellert, M. & Lambowitz, A.) 12–23 (ASM press, Washington DC, 2002).
Hughes, V. M. & Datta, N. Conjugative plasmids in bacteria of the pre-antibiotic era. Nature 302, 725–726 (1983).
Mazel, D. & Davies, J. Antibiotic resistance in microbes. Cell. Mol. Life Sci. 56, 742–754. (1999).
Bennett, P. M. Genome plasticity: insertion sequence elements, transposons and integrons, and DNA rearrangement. Methods Mol. Biol. 266, 71–113 (2004).
Hall, R. M. Mobile gene cassettes and integrons: moving antibiotic resistance genes in Gram-negative bacteria. Ciba Found. Symp. 207, 192–202; discussion 202–205 (1997).
Liebert, C. A., Hall, R. M. & Summers, A. O. Transposon Tn21, flagship of the floating genome. Microbiol. Mol. Biol. Rev. 63, 507–522 (1999).
Novick, R. P. Mobile genetic elements and bacterial toxinoses: the superantigen-encoding pathogenicity islands of Staphylococcus aureus. Plasmid 49, 93–105 (2003).
Shipley, P. L., Gyles, C. L. & Falkow, S. Characterization of plasmids that encode for the K88 colonization antigen. Infect. Immun. 20, 559–566 (1978). Early recognition of role for plasmids in the bacterial colonization of animal hosts.
Schell, J. et al. Interactions and DNA transfer between Agrobacterium tumefaciens, the Ti-plasmid and the plant host. Proc. R. Soc. Lond., B, Biol. Sci. 204, 251–266 (1979). Early demonstration of pathogenesis that involves plasmid-directed transfer of DNA from a bacterium to a plant.
Brussow, H., Canchaya, C. & Hardt, W. D. Phages and the evolution of bacterial pathogens: from genomic rearrangements to lysogenic conversion. Microbiol. Mol. Biol. Rev. 68, 560–602, (2004).
Banks, D. J., Beres, S. B. & Musser, J. M. The fundamental contribution of phages to GAS evolution, genome diversification and strain emergence. Trends Microbiol. 10, 515–521 (2002).
Boyd, E. F. & Brussow, H. Common themes among bacteriophage-encoded virulence factors and diversity among the bacteriophages involved. Trends Microbiol. 10, 521–529 (2002).
Koehler, T. M. Bacillus anthracis genetics and virulence gene regulation. Curr. Top. Microbiol. Immunol. 271, 143–164 (2002).
Okinaka, R. T. et al. Sequence and organization of pXO1, the large Bacillus anthracis plasmid harbouring the anthrax toxin genes. J. Bacteriol. 181, 6509–6515 (1999).
Crossman, L. C. Plasmid replicons of Rhizobium. Biochem. Soc. Trans. 33, 157–158 (2005).
Sullivan, J. T. et al. Comparative sequence analysis of the symbiosis island of Mesorhizobium loti strain R7A. J. Bacteriol. 184, 3086–3095 (2002).
Paul, J. H. & Sullivan, M. B. Marine phage genomics: what have we learned? Curr. Opin. Biotechnol. 16, 299–307 (2005).
Wade, N. Court says lab-made life can be patented. Science 208, 1445 (1980).
Kellogg, S. T., Chatterjee, D. K. & Chakrabarty, A. M. Plasmid-assisted molecular breeding: new technique for enhanced biodegradation of persistent toxic chemicals. Science 214, 1133–1135 (1981).
Lindstrom, J. E. et al. Microbial populations and hydrocarbon biodegradation potentials in fertilized shoreline sediments affected by the T/V Exxon Valdez oil spill. Appl. Environ. Microbiol. 57, 2514–2522 (1991).
von Canstein, H., Li, Y. & Wagner-Dobler, I. Long-term performance of bioreactors cleaning mercury-contaminated wastewater and their response to temperature and mercury stress and mechanical perturbation. Biotechnol. Bioeng. 74, 212–219 (2001).
van der Meer, J. R. & Sentchilo, V. Genomic islands and the evolution of catabolic pathways in bacteria. Curr. Opin. Biotechnol. 14, 248–254 (2003).
Schluter, A. et al. The 64,508 bp IncP-1b antibiotic multiresistance plasmid pB10 isolated from a waste-water treatment plant provides evidence for recombination between members of different branches of the IncP-1b group. Microbiology 149, 3139–3153 (2003).
Gogarten, J. P., Doolittle, W. F. & Lawrence, J. G. Prokaryotic evolution in light of gene transfer. Mol. Biol. Evol. 19, 2226–2238 (2002).
Frank, A. C., Amiri, H. & Andersson, S. G. Genome deterioration: loss of repeated sequences and accumulation of junk DNA. Genetica 115, 1–12 (2002).
Mira, A., Ochman, H. & Moran, N. A. Deletional bias and the evolution of bacterial genomes. Trends Genet. 17, 589–596 (2001).
Botstein, D. A theory of modular evolution for bacteriophages. Ann. N. Y. Acad. Sci. 354, 484–490 (1980). A seminal paper on the mosaic nature of lambdoid phages, which is now clearly applicable to several other phage families.
Casjens, S., Hatfull, G. & Hendrix, R. Evolution of the dsDNA tailed-bacteriophage genomes. Semin. Virol. 3, 383–397 (1992).
Canchaya, C., Proux, C., Fournous, G., Bruttin, A. & Brussow, H. Prophage genomics. Microbiol. Mol. Biol. Rev. 67, 238–276 (2003).
Burge, C. B. & Karlin, S. Finding the genes in genomic DNA. Curr. Opin. Struct. Biol. 8, 346–354 (1998).
Claverie, J. M. Computational methods for exon detection. Mol. Biotechnol. 10, 27–48 (1998).
Guigo, R., Agarwal, P., Abril, J. F., Burset, M. & Fickett, J. W. An assessment of gene prediction accuracy in large DNA sequences. Genome Res. 10, 1631–1642 (2000).
Guigo, R., Knudsen, S., Drake, N. & Smith, T. Prediction of gene structure. J. Mol. Biol. 226, 141–157 (1992).
Borodovsky, M. & McIninch, J. Recognition of genes in DNA sequence with ambiguities. Biosystems 30, 161–171 (1993).
Snyder, E. E. & Stormo, G. D. Identification of protein coding regions in genomic DNA. J. Mol. Biol. 248, 1–18 (1995).
Burge, C. & Karlin, S. Prediction of complete gene structures in human genomic DNA. J. Mol. Biol. 268, 78–94 (1997).
Lukashin, A. V. & Borodovsky, M. GeneMark.hmm: new solutions for gene finding. Nucleic Acids Res. 26, 1107–1115 (1998).
Delcher, A. L., Harmon, D., Kasif, S., White, O. & Salzberg, S. L. Improved microbial gene identification with GLIMMER. Nucleic Acids Res. 27, 4636–4641 (1999).
Fischer, D. & Eisenberg, D. Finding families for genomic ORFans. Bioinformatics 15, 759–762 (1999).
Amiri, H., Davids, W. & Andersson, S. G. Birth and death of orphan genes in Rickettsia. Mol. Biol. Evol. 20, 1575–1587 (2003).
Domazet-Loso, T. & Tautz, D. An evolutionary analysis of orphan genes in Drosophila. Genome Res. 13, 2213–2219 (2003).
Daubin, V. & Ochman, H. Bacterial genomes as new gene homes: the genealogy of ORFans in E. coli. Genome Res. 14, 1036–1042 (2004).
Morgenstern, B. et al. Exon discovery by genomic sequence alignment. Bioinformatics 18, 777–787 (2002).
Meyer, I. M. & Durbin, R. Comparative ab initio prediction of gene structures using pair HMMs. Bioinformatics 18, 1309–1318 (2002).
Crollius, H. R. et al. Characterization and repeat analysis of the compact genome of the freshwater pufferfish Tetraodon nigroviridis. Genome Res. 10, 939–949 (2000).
Badger, J. H. & Olsen, G. J. CRITICA: coding region identification tool invoking comparative analysis. Mol. Biol. Evol. 16, 512–524 (1999).
Wiehe, T., Gebauer-Jung, S., Mitchell-Olds, T. & Guigo, R. SGP-1: prediction and validation of homologous genes based on sequence alignments. Genome Res. 11, 1574–1583 (2001).
Bateman, A. et al. The Pfam protein families database. Nucleic Acids Res. 30, 276–280 (2002).
Gough, J. & Chothia, C. SUPERFAMILY: HMMs representing all proteins of known structure. SCOP sequence searches, alignments and genome assignments. Nucleic Acids Res. 30, 268–272 (2002).
Andreeva, A. et al. SCOP database in 2004: refinements integrate structure and sequence family data. Nucleic Acids Res. 32, D226–D229 (2004).
Berriman, M. & Rutherford, K. Viewing and annotating sequence data with Artemis. Brief. Bioinformatics 4, 124–132 (2003).
Delcher, A. L., Phillippy, A., Carlton, J. & Salzberg, S. L. Fast algorithms for large-scale genome alignment and comparison. Nucleic Acids Res. 30, 2478–2483 (2002).
Harris, M. A. et al. The Gene Ontology (GO) database and informatics resource. Nucleic Acids Res. 32, D258–D261 (2004).
Leplae, R., Hebrant, A., Wodak, S. J. & Toussaint, A. ACLAME: a CLAssification of Mobile genetic Elements. Nucleic Acids Res 32 (Database issue), D45–D49 (2004).
Eddy, S. R. A model of the statistical power of comparative genome sequence analysis. PLoS Biol. 3, e10 (2005).
Eckhardt, T. A rapid method for the identification of plasmid desoxyribonucleic acid in bacteria. Plasmid 1, 584–588 (1978).
Williams, L., Miller, D., Summers, A. O. & Detter, C. Fast, cheap, and easy preparation of library-quality DNA from 100+ kb, low copy eubacterial plasmids. Plasmid 53, 45–46 (2005).
Guo, X. H., Huff, E. J. & Schwartz, D. C. Sizing of large DNA molecules by hook formation in a loose matrix. J. Biomol. Struct. Dyn. 11, 1–10 (1993).
Funnell, B. E. & Phillips, G. J. (eds) Plasmid Biology (ASM Press, Washington DC, 2004).
Craig, N. L., Craigie, R., Gellert, M. & Lambowitz, A. M. (eds) Mobile DNA II (ASM Press, Washington DC, 2002).
Molbak, L. et al. The plasmid genome database. Microbiology 149, 3043–3045 (2003)
Chandler, M. & Mahillon, J. in Mobile DNA II (eds. Craig, N. L., Craigie, R., Gellert, M. & Lambowitz, A. M.) 305–366 (ASM Press, Washington DC, 2003).
Mantri, Y. & Williams, K. P. Islander: a database of integrative islands in prokaryotic genomes, the associated integrases and their DNA site specificities. Nucleic Acids Res. 32 (Database issue), D55–D58 (2004).
Dong, X., Stothard, P., Forsythe, I. J. & Wishart, D. S. PlasMapper: a web server for drawing and auto-annotating plasmid maps. Nucleic Acids Res. W365–W371 (2004).
Galperin, M.Y. The molecular biology database collection. Nucleic Acids Res. 33, (Database issue) Entry no. 750 (2005).
Acknowledgements
We thank M. Syvanen for the public domain software metaphor for MGEs and the reviewers for their thoughtful critiques. Work in the laboratories of R.L. and A.T. is supported by the Fonds National de la Recherche Scientifique, the Université Libre de Bruxelles, Belgium, and the European Space Agency. L.S.F. acknowledges support from the Canadian Institutes of Health Research and Natural Sciences and Engineering Research Council of Canada. A.O.S. acknowledges support from the US Department of Energy (DOE) Genomes-To-Life Program, the assistance of L. Williams, the staff of the DOE Joint Genome Institute, Walnut Creek, California and Oak Ridge National Laboratory, Tennessee.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
Glossary
- TRANSFORMATION
-
Gene transfer that is mediated by the uptake of free DNA.
- CONJUGATION
-
Gene transfer that is mediated by certain plasmids or ICEs with relevant transfer genes. Cell–cell contact is required for conjugation, unlike transduction or transformation.
- TRANSDUCTION
-
Gene transfer that is mediated by certain types of bacteriophage.
- INTEGRATIVE CONJUGATIVE ELEMENTS
-
(ICEs). Together with conjugative transposons (CTns) and genomic islands, these are chromosomally located gene clusters that encode phage-linked integrases and conjugation proteins as well as other genes associated with an observable phenotype such as virulence or symbiosis. ICEs and CTns are gene clusters that can be transferred between cells, whereas genomic islands have not been shown to transfer. Although these gene clusters have some phage-like genes, they do not lyse the cell or form extracellular particles.
- HOMOLOGOUS RECOMBINATION
-
DNA recombination that requires extensive sequence similarity in the involved DNA segments. It is usually effected by chromosomally encoded genes, but some phages also have orthologues of such chromosomal genes.
- MOBILIZATION
-
Transfer by a conjugative element of a plasmid or part of the bacterial cellular chromosome that cannot effect self transfer. Mediated by the trans-acting proteins of the conjugative plasmid that function on cognate mobilization (oriT) sites in the mobilized plasmid to direct it to the conjugation pore built by the conjugative element.
- INTEGRON
-
A genetic element that encodes an integrase enzyme, which can assemble tandem arrays of genes or gene fragments and provide them with a promoter for expression. Often associated with antibiotic multi-resistance.
- NON-HOMOLOGOUS RECOMBINATION
-
DNA recombination that requires little or no similarity between the DNA segments involved. This process is carried out by specialized enzymes that are encoded by transposons and phages.
- METAGENOMICS
-
Sequencing of a clone library derived from the total DNA purified from a complex microbial ecosystem. This is followed by computer assembly of the reads into multiple linkage groups assumed to represent the organisms present in the community, including those that cannot be cultured.
Rights and permissions
About this article
Cite this article
Frost, L., Leplae, R., Summers, A. et al. Mobile genetic elements: the agents of open source evolution. Nat Rev Microbiol 3, 722–732 (2005). https://doi.org/10.1038/nrmicro1235
Issue Date:
DOI: https://doi.org/10.1038/nrmicro1235
This article is cited by
-
Metals are overlooked in the evolution of antibiotic resistance
Soil Ecology Letters (2024)
-
Cemeteries and graveyards as potential reservoirs of antibiotic resistance genes and bacteria: a review
Environmental Chemistry Letters (2024)
-
Tn3-like structures co-harboring of blaCTX-M-65, blaTEM-1 and blaOXA-10 in the plasmids of two Escherichia coli ST1508 strains originating from dairy cattle in China
BMC Veterinary Research (2023)
-
Gill-associated bacteria are homogeneously selected in amphibious mangrove crabs to sustain host intertidal adaptation
Microbiome (2023)
-
Plasmid-encoded toxin defence mediates mutualistic microbial interactions
Nature Microbiology (2023)