Key Points
-
Mutation and horizontal gene transfer (HGT) continually give rise to new bacterial genotypes. Infrequently, such new bacterial genotypes establish and spread in the larger population through either positive selection or random genetic drift. Therefore, bacterial genomes are in a constant state of flux, and any segment of DNA in a large bacterial population might have the opportunity to be horizontally transferred.
-
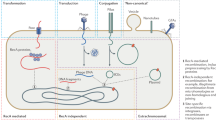
Three main mechanisms of HGT have been described: natural transformation, the uptake of free DNA in competent bacteria, exhibited by about 1% of validly described bacterial species; transduction, the transfer of bacterial DNA between a bacteriophage-infected bacterium and a bacteriophage-susceptible bacterium; and conjugation, the transfer of mobile genetic elements by pili structures assembled between two adjacently located bacteria.
-
A number of factors limit the transfer, uptake and stabilization of foreign DNA molecules acquired by bacteria. These include limited release and stability of adaptive DNA in the environment; limits on competence development; limits on host range of the transfer and maintenance mechanism of mobile genetic elements; recipient restriction enzyme activity; and limited ability of foreign DNA to integrate into a replicating genetic element owing to a lack of DNA sequence similarity.
-
Homologous recombination depends on the incoming DNA containing regions between 25 and 200 bp in length, depending on the system, of high similarity to the recipient genome. Dependence on DNA sequence similarity for recombination between species is relaxed in some mutator strains. DNA acquisition through double-stranded breaks and end-joining — illegitimate recombination — applies more to integration of circular DNA than linear fragments.
-
Most of the understanding of the processes facilitating HGT and their frequencies of occurrence have come from well designed laboratory studies of a few model bacterial species. These studies have proven suited to resolve the basic biological mechanisms involved, but fail to encompass the environmental variables involved. We have still to develop a quantitative and qualitative understanding of ongoing gene-transfer processes occurring under natural conditions. Biologically significant gene-transfer processes might occur at temporospatial scales that current methodology do not allow us to monitor.
Abstract
Bacteria evolve rapidly not only by mutation and rapid multiplication, but also by transfer of DNA, which can result in strains with beneficial mutations from more than one parent. Transformation involves the release of naked DNA followed by uptake and recombination. Homologous recombination and DNA-repair processes normally limit this to DNA from similar bacteria. However, if a gene moves onto a broad-host-range plasmid it might be able to spread without the need for recombination. There are barriers to both these processes but they reduce, rather than prevent, gene acquisition.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Griffith, F. The significance of pneumococcal types. J. Hyg. 27, 113–159 (1928).
Thomas, C. M. (ed.) The Horizontal Gene Pool: Bacterial Plasmids and Gene Spread (Harwood Academic Publishers, Amsterdam, 2000).
Nakamura, Y., Itoh, T., Matsuda, H. & Gojobori, T. Biased function of horizontally transferred genes in prokaryotic genomes. Nature Genet. 36, 760–766 (2004).
Jain, R., Rivera, M. C. & Lake, J. A. Horizontal gene transfer among genomes: the complexity hypothesis. Proc. Natl Acad. Sci. USA 96, 3801–3806 (1999).
Nielsen, K. M. & Townsend, J. P. Monitoring and modeling horizontal gene transfer. Nature Biotechnol. 22, 1110–1114 (2004).
Cohan, F. M., Roberts, M. S. & King, E. C. The potential for genetic exchange by transformation within a natural population of Bacillus subtilis. Evolution 45, 1383–1421 (1991).
Jonas, D. A. et al. Safety considerations of DNA in food. Ann. Nutr. Metab. 45, 235–254 (2001).
Paget, E. & Simonet, P. On the track of natural transformation in soil. FEMS Microbiol. Ecol. 15, 109–117 (1994).
Lorenz, M. G. & Wackernagel, W. Bacterial gene-transfer by natural genetic-transformation in the environment. Microbiol. Rev. 58, 563–602 (1994).
Moscoso, M. & Claverys, J. P. Release of DNA into the medium by competent Streptococcus pneumoniae: kinetics, mechanism and stability of the liberated DNA. Mol. Microbiol. 54, 783–794 (2004).
Lorenz, M. G., Gerjets, D. & Wackernagel, W. Release of transforming plasmid and chromosomal DNA from 2 cultured soil bacteria. Arch. Microbiol. 156, 319–326 (1991).
Ueda, S. & Hara, T. Studies on nucleic acid production and application. I. production of extracellular DNA by Pseudomonas sp. KYU-1. J. Appl. Biochem. 3, 1–10 (1981).
Whitchurch, C. B., Tolker-Nielsen, T., Ragas, P. C. & Mattick, J. S. Extracellular DNA required for bacterial biofilm formation. Science 295, 1487–1487 (2002).
Palmen, R. & Hellingwerf, K. J. Acinetobacter calcoaceticus liberates chromosomal DNA during induction of competence by cell lysis. Curr. Microbiol. 30, 7–10 (1995).
Friedlander, A. M. DNA release as a direct measure of microbial killing. J. Immunol. 115, 1404–1408 (1975).
Connolly, J. H., Herriott, R. M. & Gupta, S. Deoxyribonuclease in human blood and platelets. Br. J. Exp. Pathol. 43, 392–408 (1962).
Rozenberg-Arska, M., Salters, E. C., Vanstrijp, J. A., Hoekstra, W. P. M. & Verhoef, J. Degradation of Escherichia coli chromosomal and plasmid DNA in serum. J. Gen. Microbiol. 130, 217–222 (1984).
Doerfler, W. Foreign DNA in Mammalian Systems. (Wiley-VCH Verlag GmbH, Weinheim, 2000).
Worthey, A. L., Kane, J. F. & Orvos, D. R. Fate of pBR322 DNA in a wastewater matrix. J. Ind. Microbiol. Biotechnol. 22, 164–166 (1999).
Widmer, F., Seidler, R. J. & Watrud, L. S. Sensitive detection of transgenic plant marker gene persistence in soil microcosms. Mol. Ecol. 5, 603–613 (1996).
Widmer, F., Seidler, R. J., Donegan, K. K. & Reed, G. L. Quantification of transgenic plant marker gene persistence in the field. Mol. Ecol. 6, 1–7 (1997).
Romanowski, G., Lorenz, M. G., Sayler, G. S. & Wackernagel, W. Persistence of free plasmid DNA in soil monitored by various methods, including a transformation assay. Appl. Environ. Microbiol. 58 3012–3019 (1992).
Romanowski, G., Lorenz, M. G. & Wackernagel, W. Plasmid DNA in a groundwater aquifer microcosm — adsorption, DNAase resistance and natural genetic transformation of Bacillus subtilis. Mol. Ecol. 2, 171–181 (1993).
Landweber, L. in Genetics and the Extinction of Species: DNA and the Conservation of Biodiversity. (eds Landweber, L. & Dobson, A. P.) 163–186 (Princeton University Press, Princeton, 1999).
Hofreiter, M., Serre, D., Poinar, H. N., Kuch, M. & Paabo, S. Ancient DNA. Nature Rev. Genet. 2, 353–359 (2001).
Ogram, A., Sayler, G. S. & Barkay, T. The extraction and purification of microbial DNA from sediments. J. Microbiol. Methods 7, 57–66 (1987).
DeFlaun, M. F. & Paul, J. H. Detection of exogenous gene sequences in dissolved DNA from aquatic environments. Microb. Ecol. 18, 21–28 (1989).
Karl, D. M. & Bailiff, M. D. The measurement and distribution of dissolved nucleic acids in aquatic environments. Limnol. Oceanogr. 34, 543–558 (1989).
Schubbert, R., Renz, D., Schmitz, B. & Doerfler, W. Foreign (M13) DNA ingested by mice reaches peripheral leukocytes, spleen, and liver via the intestinal wall mucosa and can be covalently linked to mouse DNA. Proc. Natl Acad. Sci. USA 94, 961–966 (1997).
Einspanier, R. et al. The fate of forage plant DNA in farm animals: a collaborative case-study investigating cattle and chicken fed recombinant plant material. Eur. Food Res. Technol. 212, 129–134 (2001).
Chiter, A., Forbes, J. M. & Blair, G. E. DNA stability in plant tissues: implications for the possible transfer of genes from genetically modified food. FEBS Lett. 481, 164–168 (2000).
Paget, E., Lebrun, M., Freyssinet, G. & Simonet, P. The fate of recombinant plant DNA in soil. Eur. J. Soil Biol. 34, 81–88 (1998).
Gebhard, F. & Smalla, K. Monitoring field releases of genetically modified sugar beets for persistence of transgenic plant DNA and horizontal gene transfer. FEMS Microbiol. Ecol. 28, 261–272 (1999).
Ceccherini, M. et al. Degradation and transformability of DNA from transgenic leaves. Appl. Environ. Microbiol. 69, 673–678 (2003).
Graham, J. B. & Istock, C. A. Gene exchange and natural selection cause Bacillus subtilis to evolve in soil culture. Science 204, 637–639 (1979).
Nielsen, K. M., Bones, A. M. & van Elsas, J. D. Induced natural transformation of Acinetobacter calcoaceticus in soil microcosms. Appl. Environ. Microbiol. 63, 3972–3977 (1997).
Frischer, M. E., Stewart, G. J. & Paul, J. H. Plasmid transfer to indigenous marine bacterial-populations by natural transformation. FEMS Microbiol. Ecol. 15, 127–135 (1994).
Baur, B., Hanselman, K., Schlimme, W. & Jenni, B. Genetic transformation in freshwater: Escherichia coli is able to develop natural competence. Appl. Environ. Microbiol. 62, 3673–3678 (1996).
Mercer, D. K., Scott, K. P., Melville, C. M., Glover, L. A. & Flint, H. J. Transformation of an oral bacterium via chromosomal integration of free DNA in the presence of human saliva. FEMS Microbiol. Lett. 200, 163–167 (2001).
Duggan, P. S., Chambers, P. A., Heritage, J. & Forbes, J. M. Survival of free DNA encoding antibiotic resistance from transgenic maize and the transformation activity of DNA in ovine saliva, ovine rumen fluid and silage effluent. FEMS Microbiol. Lett. 191, 71–77 (2000).
Brautigaum, M., Hertel, C. & Hammes, W. P. Evidence for natural transformation of Bacillus subtilis in foodstuffs. FEMS Microbiol. Lett. 155, 93–98 (1997).
Bauer, F., Hertel, C. & Hammes, W. P. Transformation of Escherichia coli in foodstuffs. Syst. Appl. Microbiol. 22 (1999).
Palmen, R. & Hellingwerf, K. J. Uptake and processing of DNA by Acinetobacter calcoaceticus — a review. Gene 192, 179–190 (1997).
Dubnau, D. DNA uptake in bacteria. Annu. Rev. Microbiol. 53, 217–244 (1999).
Dreiseikelmann, B. Translocation of DNA across bacterial-membranes. Microbiol. Rev. 58, 293–316 (1994).
Puyet, A., Greenberg, B. & Lacks, S. A. Genetic and structural characterization of EndA — a membrane-bound nuclease required for transformation of Streptococcus pneumoniae. J. Mol. Biol. 213, 727–738 (1990).
Chen, I. & Dubnau, D. DNA uptake during natural transformation. Nature Rev. Microbiol. 2, 241–249 (2004).
Nielsen, K. M. et al. Natural transformation and availability of transforming DNA to Acinetobacter calcoaceticus in soil microcosms. Appl. Environ. Microbiol. 63, 1945–1952 (1997).
Mejean, V. & Claverys, J. P. DNA processing during entry in transformation of Streptococcus pneumoniae. J. Biol. Chem. 268, 5594–5599 (1993).
Berndt, C., Meier, P. & Wackernagel, W. DNA restriction is a barrier to natural transformation in Pseudomonas stutzeri JM300. Microbiology 149, 895–901 (2003).
Maynard Smith, J., Feil, E. J. & Smith, N. H. Population structure and evolutionary dynamics of pathogenic bacteria. BioEssays 22, 1115–1122 (2000).
Feil, E. J. & Spratt, B. G. Recombination and the population structures of bacterial pathogens. Annu. Rev. Microbiol. 55, 561–590 (2001).
Townsend, J. P., Nielsen, K. M., Fisher, D. S. & Hartl, D. L. Horizontal acquisition of divergent chromosomal DNA in bacteria: effects of mutator phenotypes. Genetics 164, 13–21 (2003).
Matic, I. et al. Highly variable mutation rates in commensal and pathogenic Escherichia coli. Science 277, 1833–1834 (1997).
Vulic, M., Dionisio, F., Taddei, F. & Radman, M. Molecular keys to speciation: DNA polymorphism and the control of genetic exchange in enterobacteria. Proc. Natl Acad. Sci. USA 94, 9763–9767 (1997).
de Vries, J., Meier, P. & Wackernagel, W. The natural transformation of the soil bacteria Pseudomonas stutzeri and Acinetobacter sp by transgenic plant DNA strictly depends on homologous sequences in the recipient cells. FEMS Microbiol. Lett. 195, 211–215 (2001).
Majewski, J. & Cohan, F. M. The effect of mismatch repair and heteroduplex formation on sexual isolation in Bacillus. Genetics 148, 13–18 (1998).
Majewski, J., Zawadzki, P., Pickerill, P., Cohan, F. M. & Dowson, C. G. Barriers to genetic exchange between bacterial species: Streptococcus pneumoniae transformation. J. Bacteriol. 182, 1016–1023 (2000).
Heinemann, J. A. & Traavik, T. Problems in monitoring horizontal gene transfer in field trials of transgenic plants. Nature Biotechnol. 22, 1105–1109 (2004).
Ikeda, H., Shiraishi, K. & Ogata, Y. Illegitimate recombination mediated by double-strand break and end-joining in Escherichia coli. Adv. Biophys. 38, 3–20 (2004).
Ehrlich, S. D. et al. Mechanisms of illegitimate recombination. Gene 135, 161–166 (1993).
Nielsen, K. M. An assessment of factors affecting the likelihood of horizontal transfer of recombinant plant DNA to bacterial recipients in the soil and rhizosphere. Collection of Biosafety Reviews (Italy) 1, 96–149 (2003).
Kurland, C. G. What tangled web: barriers to rampant horizontal gene transfer. BioEssays 27, 741–747 (2005).
Dempsey, L. A. & Dubnau, D. A. Identification of plasmid and Bacillus subtilis chromosomal recombination sites used for pE194 integration. J. Bacteriol. 171, 2856–2865 (1989).
Lovett, S. T., Hurley, R. L., Sutera, V. A., Aubuchon, R. H. & Lebedeva, M. A. Crossing over between regions of limited homology in Escherichia coli: RecA-dependent and RecA-independent pathways. Genetics 160, 851–859 (2002).
Shen, P. & Huang, H. V. Homologous recombination in Escherichia coli — dependence on substrate length and homology. Genetics 112, 441–457 (1986).
Majewski, J. & Cohan, F. M. DNA sequence similarity requirements for interspecific recombination in Bacillus. Genetics 153, 1525–1533 (1999).
Meier, P. & Wackernagel, W. Mechanisms of homology-facilitated illegitimate recombination for foreign DNA acquisition in transformable Pseudomonas stutzeri. Mol. Microbiol. 48, 1107–1118 (2003).
de Vries, J., Herzfeld, T. & Wackernagel, W. Transfer of plastid DNA from tobacco to the soil bacterium Acinetobacter sp. by natural transformation. Mol. Microbiol. 53, 323–334 (2004).
de Vries, J. & Wackernagel, W. Integration of foreign DNA during natural transformation of Acinetobacter sp. by homology-facilitated illegitimate recombination. Proc. Natl Acad. Sci. USA 99, 2094–2099 (2002).
Prudhomme, M., Libante, V. & Claverys, J. P. Homologous recombination at the border: insertion-deletions and the trapping of foreign DNA in Streptococcus pneumoniae. Proc. Natl Acad. Sci. USA 99, 2100–2105 (2002).
Nielsen, K. M., van Elsas, J. D. & Smalla, K. Transformation of Acinetobacter sp. strain BD413(pFG4 Delta nptII) with transgenic plant DNA in soil microcosms and effects of kanamycin on selection of transformants. Appl. Environ. Microbiol. 66, 1237–1242 (2000).
Kay, E., Vogel, T. M., Bertolla, F., Nalin, R. & Simonet, P. In situ transfer of antibiotic resistance genes from transgenic (transplastomic) tobacco plants to bacteria. Appl. Environ. Microbiol. 68, 3345–3351 (2002).
Funchain, P., Yeung, A., Stewart, J., Clendenin, W. M. & Miller, J. H. Amplification of mutator cells in a population as a result of horizontal transfer. J. Bacteriol. 183, 3737–3741 (2001).
Feng, G., Tsui, H. C. T. & Winkler, M. E. Depletion of the cellular amounts of the MutS and MutH methyl-directed mismatch repair proteins in stationary-phase Escherichia coli K-12 cells. J. Bacteriol. 178, 2388–2396 (1996).
LeClerc, J. E., Li, B. G., Payne, W. L. & Cebula, T. A. High mutation frequencies among Escherichia coli and Salmonella pathogens. Science 274, 1208–1211 (1996).
Richardson, A. R., Yu, Z., Popovic, T. & Stojiljkovic, I. Mutator clones of Neisseria meningitidis in epidemic serogroup A disease. Proc. Natl Acad. Sci. USA 99, 6103–6107 (2002).
Young, D. M. & Ornston, L. N. Functions of the mismatch repair gene mutS from Acinetobacter sp strain ADP1. J. Bacteriol. 183, 6822–6831 (2001).
Burrus, V., Pavlovic, G., Decaris, B. & Guedon, G. Conjugative transposons: the tip of the iceberg. Mol. Microbiol. 46, 601–610 (2002).
Flores, M. et al. Prediction, identification, and artificial selection of DNA rearrangements in Rhizobium: Toward a natural genomic design. Proc. Natl Acad. Sci. USA 97, 9138–9143 (2000).
Mavingui, P. et al. Dynamics of genome architecture in Rhizobium sp strain NGR234. J. Bacteriol. 184, 171–176 (2002).
Mahillon, J. & Chandler, M. Insertion sequences. Microbiol. Mol. Biol. Rev. 62, 725–774. (1998).
Reimmann, C. & Haas, D. Mode of replicon fusion mediated by the duplicated insertion-sequence IS21 in Escherichia coli. Genetics 115, 619–625 (1987).
Reimmann, C. et al. Genetic structure, function and regulation of the transposable element IS21. Mol. Gen. Genetics 215, 416–424 (1989).
Bao, T. H., Betermier, M., Polard, P. & Chandler, M. Assembly of a strong promoter following IS911 circularization and the role of circles in transposition. EMBO J. 16, 3357–3371 (1997).
Duval-Valentin, G., Marty-Cointin, B. & Chandler, M. Requirement of IS911 replication before integration defines a new bacterial transposition pathway. EMBO J. 23, 3897–3906 (2004).
Duval-Valentin, G., Normand, C., Khemici, V., Marty, B. & Chandler, M. Transient promoter formation: a new feedback mechanism for regulation of IS911 transposition. EMBO J. 20, 5802–5811 (2001).
McGrath, B. M. & Pembroke, J. T. Detailed analysis of the insertion site of the mobile elements R997, pMERPH, R392, R705 and R391 in E. coli K12. FEMS Microbiol. Lett. 237, 19–26 (2004).
Mateos, L. M., Schafer, A., Kalinowski, J., Martin, J. F. & Puhler, A. Integration of narrow-host-range vectors from Escherichia coli into the genomes of amino acid-producing Corynebacteria after intergeneric conjugation. J. Bacteriol. 178, 5768–5775 (1996).
Ikeda, H., Shimizu, H., Ukita, T. & Kumagai, M. A novel assay for illegitimate recombination in Escherichia coli — stimulation of λ-bio transducing phage formation by ultraviolet-light and its independence from RecA function. Adv. Biophys. 31, 197–208 (1995).
Shiraishi, K., Hanada, K., Iwakura, Y. & Ikeda, H. Roles of recJ, RecO, and RecR in RecET-mediated illegitimate recombination in Escherichia coli. J. Bacteriol. 184, 4715–4721 (2002).
Chiu, C.-M. & Thomas, C. M. Evidence for the past integration of IncP-1 plasmids into bacterial chromosomes. FEMS Lett. 241, 163–169 (2004).
Cascales, E. & Christie, P. J. The versatile bacterial type IV secretion systems. Nature Rev. Microbiol. 1, 137–149 (2003).
Anthony, K. G., Sherburne, C., Sherburne, R. & Frost, L. S. The role of the pilus in recipient cell recognition during bacterial conjugation mediated by F-like plasmids. Mol. Microbiol. 13, 939–953 (1994).
Ishiwa, A. & Komano, T. PilV adhesins of plasmid R64 thin pili specifically bind to the lipopolysaccharides of recipient cells. J. Mol. Biol. 343, 615–625 (2004).
Samuels, A. L., Lanka, E. & Davies, J. E. Conjugative junctions in RP4-mediated mating of Escherichia coli. J. Bacteriol. 182, 2709–2715 (2000).
Kelly, B. A. & Kado, C. I. Agrobacterium-mediated T-DNA transfer and integration into the chromosome of Streptomyces lividans. Mol. Plant Pathol. 3, 125–134 (2002).
Giebelhaus, L. A. et al. The Tra2 core of the IncP alpha plasmid RP4 is required for intergeneric mating between Escherichia coli and Streptomyces lividans. J. Bacteriol. 178, 6378–6381 (1996).
Bingle, L. E. H., Zatyka, M., Manzoor, S. E. & Thomas, C. M. Co-operative interactions control conjugative transfer of broad host-range plasmid RK2: full effect of minor changes in TrbA operator depends on KorB. Mol. Microbiol. 49, 1095–1108 (2003).
Grohmann, E., Muth, G. & Espinosa, M. Conjugative plasmid transfer in Gram-positive bacteria. Microbiol. Mol. Biol. Rev. 67, 277–301 (2003).
Hirt, H., Schlievert, P. M. & Dunny, G. M. In vivo induction of virulence and antibiotic resistance transfer in Enterococcus faecalis mediated by the sex pheromone-sensing system of pCF10. Infect. Immun. 70, 716–723 (2002).
Chandler, J. R. & Dunny, G. M. Enterococcal peptide sex pheromones: synthesis and control of biological activity. Peptides 25, 1377–1388 (2004).
Waters, C. M. & Dunny, G. M. Analysis of functional domains of the Enterococcus faecalis pheromone-induced surface protein aggregation substance. J. Bacteriol. 183, 5659–5667 (2001).
Flannagan, S. E. & Clewell, D. B. Identification and characterization of genes encoding sex pheromone cAM373 activity in Enterocccus faecalis and Staphylococcus aureus. Mol. Microbiol. 44, 803–817 (2002).
Clewell, D. B., Francia, M. V., Flannagan, S. E. & An, F. Y. Enterococcal plasmid transfer: sex pheromones, transfer origins, relaxases, and the Staphylococcus aureus issue. Plasmid 48, 193–201 (2002).
Kurenbach, B. R. et al. Intergeneric transfer of the Enterococcus faecalis plasmid pIP501 to Escherichia coli and Streptomyces lividans and sequence analysis of its tra region. Plasmid 50, 86–93 (2003).
Maas, R. M., Gotz, J., Wohlleben, W. & Muth, G. The conjugative plasmid pSG5 from Streptomyces ghanaensis DSM 2932 differs in its transfer functions from other Streptomyces rolling-circle-type plasmids. Microbiology 144, 2809–2817 (1998).
Pettis, G. S. & Cohen, S. N. Mutational analysis of the tra locus of the broad-host-range Streptomyces plasmid pIJ101. J. Bacteriol. 182, 4500–4504 (2000).
Bentley, S. D. et al. SCP1, a 356,023 bp linear plasmid adapted to the ecology and developmental biology of its host, Streptomyces coelicolor A3(2). Mol. Microbiol. 51, 1615–1628 (2004).
Haug, I. et al. Streptomyces coelicolor A3(2) plasmid SCP2*: deductions from the complete sequence. Microbiology 149, 505–513 (2003).
Stecker, C., Johann, A., Herzberg, C., Averhoff, B. & Gottschalk, G. Complete nucleotide sequence and genetic organization of the 210-kilobase linear plasmid of Rhodococcus erythropolis BD2. J. Bacteriol. 185, 5269–5274 (2003).
Cabezon, E., Sastre, J. I. & de la Cruz, F. Genetic evidence of a coupling role for the TraG protein family in bacterial conjugation. Mol. Gen. Genet. 254, 400–406 (1997).
Llosa, M., Zunzunegui, S. & de la Cruz, F. Conjugative coupling proteins interact with cognate and heterologous VirB10-like proteins while exhibiting specificity for cognate relaxosomes. Proc. Natl Acad. Sci. USA 100, 10465–10470 (2003).
Frost, L. S., Ippen-Ihler, K. & Skurray, R. A. Analysis of the sequence and gene products of the transfer region of the F sex factor. Microbiol. Rev. 58, 162–210 (1994).
Haase, J., Kalkum, M. & Lanka, E. TrbK, a small cytoplasmic membrane lipoprotein, functions in entry exclusion of the IncP alpha plasmid RP4. J. Bacteriol. 178, 6720–6729 (1996).
Hochhut, B., Beaber, J. W., Woodgate, R. & Waldor, M. K. Formation of chromosomal tandem arrays of the SXT element and R391, two conjugative chromosomally integrating elements that share an attachment site. J. Bacteriol. 183, 1124–1132 (2001).
Pohlman, R. F., Genetti, H. D. & Winans, S. C. Entry exclusion of the IncN plasmid-pKM101 is mediated by a single hydrophilic protein containing a lipid attachment motif. Plasmid 31, 158–165 (1994).
Possoz, C., Gagnat, J., Sezonov, G., Guerineau, M. & Pernodet, J. L. Conjugal immunity of Streptomyces strains carrying the integrative element pSAM2 is due to the pif gene (pSAM2 immunity factor). Mol. Microbiol. 47, 1385–1393 (2003).
Boyd, E. F., Hill, C. W., Rich, S. M. & Hartl, D. L. Mosaic structure of plasmids from natural populations of Escherichia coli. Genetics 143, 1091–1100 (1996).
Schluter, A. et al. The 64 508 bp IncP-1β antibiotic multiresistance plasmid pB10 isolated from a waste-water treatment plant provides evidence for recombination between members of different branches of the IncP-1β group. Microbiology 149, 3139–3153 (2003).
Peters, J. E., Bartoszyk, I. M., Dheer, S. & Benson, S. A. Redundant homosexual F transfer facilitates selection-induced reversion of plasmid mutations. J. Bacteriol. 178, 3037–3043 (1996).
Ghigo, J. M. Natural conjugative plasmids induce bacterial biofilm development. Nature 412, 442–445 (2001).
Reisner, A., Haagensen, J. A. J., Schembri, M. A., Zechner, E. L. & Molin, S. Development and maturation of Escherichia coli K-12 biofilms. Mol. Microbiol. 48, 933–946 (2003).
Sukulpovi, S. & O'Connor, C. D. TraT lipoprotein, a plasmid-specified mediator of interactions between Gram-negative bacteria and their environment. Microbiol. Rev. 54, 331–341 (1990).
Jeltsch, A. Maintenance of species identity and controlling speciation of bacteria: a new function for restriction/modification systems? Gene 317, 13–16 (2003).
Lacks, S. A. & Springhorn, S. S. Transfer of recombinant plasmids containing the gene for DpnII DNA methylase into strains of Streptococcus pneumoniae that produce DpnI or DpnII restirction endonucleases. J. Bacteriol. 158, 905–909 (1984).
Moser, D. P., Zarka, D. & Kallas, T. Characterization of a restriction barrier and electrotransformation of the Cyanobacterium nostoc Pcc-7121. Arch. Microbiol. 160, 229–237 (1993).
Purdy, D. et al. Conjugative transfer of clostridial shuttle vectors from Escherichia coli to Clostridium difficile through circumvention of the restriction barrier. Mol. Microbiol. 46, 439–452 (2002).
Pinedo, C. A. & Smets, B. F. Conjugal TOL transfer from Pseudomonas putida to Pseudomonas aeruginosa: effects of restriction porificiency, toxicant exposure, cell density ratios, and conjugation detection method on observed transfer efficiencies. Appl. Environ. Microbiol. 71, 51–57 (2005).
Wilkins, B. M., Chilley, P. M., Thomas, A. T. & Pocklington, M. J. Distribution of restriction enzyme recognition sequences on broad host range plasmid RP4: molecular and evolutionary implications. J. Mol. Biol. 258, 447–456 (1996).
Bassett, C. L. & Janisiewicz, W. J. Electroporation and stable maintenance of plasmid DNAs in a biocontrol strain of Pseudomonas syringae. Biotechnol. Lett. 25, 199–203 (2003).
Belogurov, A. A., Delver, E. P. & Rodzevich, O. V. IncN plasmid pKM101 and IncI1 plasmid ColIb-P9 encode homologous antirestriction proteins in their leading regions. J. Bacteriol. 174, 5079–5085 (1992).
Belogurov, A. A., Delver, E. P. & Rodzevich, O. V. Plasmid pKM101 encodes 2 nonhomologous antirestriction proteins (ArdA and ArdB) whose expression is controlled by homologous regulatory sequences. J. Bacteriol. 175, 4843–4850 (1993).
Belogurov, A. A. et al. Antirestriction protein ard (TypeC) encoded by IncW plasmid pSa has a high similarity to the “protein transport” domain of the TraC1 primase of promiscous plasmid RP4. J. Mol. Biol. 296, 969–977 (2000).
Chilley, P. M. & Wilkins, B. M. Distribution of the ArdA family of antirestriction gene on conjugative plasmids. Microbiology 141, 2157–2164 (1995).
Larsen, M. H. & Figurski, D. H. Structure, expression, and regulation of the kilC operon of promiscuous IncP-α plasmids. J. Bacteriol. 176, 5022–5032 (1994).
Thorsted, P.A et al. Complete sequence of the IncP β plasmid R751: implications for evolution and organisation of the IncP. backbone. J. Mol. Biol. 282, 969–990 (1998).
Kobayashi, I. Behavior of restriction-modification systems as selfish mobile elements and their impact on genome evolution. Nucleic Acids Res. 29, 3742–3756 (2001).
Nakayama, Y. & Kobayashi, I. Restriction-modification gene complexes as selfish gene entities: roles of a regulatory system in their establishment, maintenance, and apoptotic mutual exclusion. Proc. Natl Acad. Sci. USA 95, 6442–6447 (1998).
Adamczyk, M. & Jagura-Burdzy, G. Spread and survival of promiscuous IncP-1 plasmids. Acta Biochim. Pol. 50, 425–453 (2003).
Rawlings, D. E. & Tietze, E. Comparative biology of IncQ and IncQ-like plasmids. Microbiol. Mol. Biol. Rev. 65, 481–496 (2001).
Scherzinger, E., Haring, V., Lurz, R. & Otto, S. Plasmid RSF1010 DNA-replication in vitro promoted by purified RSF1010 RepA, RepB and RepC proteins. Nucleic Acids Res. 19, 1203–1211 (1991).
Becker, E. C. & Meyer, R. J. Acquisition of resistance genes by the IncQ plasmid R1162 is limited by its high copy number and lack of a partitioning mechanism. J. Bacteriol. 179, 5947–5950 (1997).
Haines, A. S., Jones, K., Cheung, M. & Thomas, C. M. The IncP-6 plasmid Rms149 consists of a small mobilizable backbone with multiple large insertions. J. Bacteriol. 187, 4728–4738 (2005).
Kramer, M. G., Espinosa, M., Misra, T. K. & Khan, S. A. Lagging strand replication of rolling-circle plasmids: specific recognition of the ssoA-type origins in different Gram-positive bacteria. Proc. Natl Acad. Sci. USA 95, 10505–10510 (1998).
Anand, S. P., Mitra, P., Naqvi, A. & Khan, S. A. Bacillus anthracis and Bacillus cereus PcrA helicases can support DNA unwinding and in vitro rolling-circle replication of plasmid pT181 of Staphylococcus aureus. J. Bacteriol. 186, 2195–2199 (2004).
Doran, K. S., Helinski, D. R. & Konieczny, I. Host-dependent requirement for specific DnaA boxes for plasmid RK2 replication. Mol. Microbiol. 33, 490–498 (1999).
Caspi, R. et al. A broad host range replicon with different requirements for replication initiation in three bacterial species. EMBO J. 20, 3262–3271 (2001).
Zhong, Z. P., Helinski, D. & Toukdarian, A. A specific region in the N terminus of a replication initiation protein of plasmid RK2 is required for recruitment of Pseudomonas aeruginosa DnaB helicase to the plasmid origin. J. Biol. Chem. 278, 45305–45310 (2003).
Jiang, Y., Pacek, M., Helinski, D. R., Konieczny, I. & Toukdarian, A. A multifunctional plasmid-encoded replication initiation protein both recruits and positions an active helicase at the replication origin. Proc. Natl Acad. Sci. USA 100, 8692–8697 (2003).
Bignell, C. R., Haines, A. S., Khare, D. & Thomas, C. M. Effect of growth rate and incC mutation on symmetric plasmid distribution by the IncP-1 partitioning apparatus. Mol. Microbiol. 34, 205–216 (1999).
Siddique, A. & Figurski, D. H. The active partition gene incC of IncP plasmids is required for stable maintenance in a broad range of hosts. J. Bacteriol. 184, 1788–1793 (2002).
Zhong, Z. P., Helinski, D. & Toukdarian, A. Plasmid host-range: restrictions to F replication in Pseudomonas. Plasmid 54, 48–56 (2005).
Maestro, B. et al. Modulation of pPS10 host range by DnaA. Mol. Microbiol. 46, 223–234 (2002).
Maestro, B., Sanz, J. M., Diaz-Orejas, R. & Fernandez-Tresguerres, E. Modulation of pPS10 host range by plasmid-encoded RepA initiator protein. J. Bacteriol. 185, 1367–1375 (2003).
Greated, A., Titok, M., Krasowiak, R., Fairclough, R. J. & Thomas, C. M. The replication and stable-inheritance functions of IncP-9 plasmid pM3. Microbiology 146, 2249–2258 (2000).
Sevastsyanovich, Y. R., Titok, M. A., Krasowiak, R., Bingle, L. E. H. & Thomas, C. M. Ability of IncP-9 plasmid pM3 to replicate in E. coli is dependent on both rep and par functions. Mol. Microbiol. 57, 819–833 (2005).
Wu, L. T. & Tseng, Y. H. Characterization of the IncW cryptic plasmid pXV2 from Xanthomonas campestris pv. vesicatoria. Plasmid 44, 163–172 (2000).
Nielsen, K. M. Barriers to horizontal gene transfer by natural transformation in soil bacteria. APMIS Suppl. 84, 77–84 (1998).
Nielsen, K. M., Bones, A. M., Smalla, K. & van Elsas, J. D. Horizontal gene transfer from plants to terrestrial bacteria — a rare event? FEMS Microbiol. Rev. 22, 79–103 (1998).
Acknowledgements
Current work related to this review in the laboratory of C.M.T. is supported by The Wellcome Trust, INTAS, BBSRC and the Darwin Trust of Edinburgh. K.M.N. acknowledge support from The Research Council of Norway.
Author information
Authors and Affiliations
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
Related links
DATABASES
Entrez
FURTHER INFORMATION
Glossary
- COMPETENCE
-
The ability of bacteria to take up extracellular DNA.
- TRANSFORMATION FREQUENCY
-
The number of bacteria carrying the horizontally acquired DNA divided by the total number of bacteria exposed, per given time unit.
- HOMOLOGOUS RECOMBINATION
-
Recombination that depends on extensive segments of high sequence similarity between two DNA molecules.
- INSERTION SEQUENCE
-
A transposable DNA segment that normally only encodes the enzymes that mediate its own transposition and has no phenotypic marker.
- TRANSPOSASE
-
The enzyme that promotes cutting the DNA at the ends of a transposable element and joining to the DNA molecule into which the element is to be inserted.
- PHAGE
-
An abbreviation of bacteriophage — a virus that specifically infects bacteria.
- SPECIALIZED TRANSDUCING PHAGE
-
A bacteriophage that integrates into a host-cell chromosome and then is excised again, bringing with it (as part of the phage genome) part of the host chromosome that can be transferred across to a new host.
- PILUS
-
The proteinaceous fibre made from multiple subunits of a protein called pilin that mediates contact between donor and recipient bacteria prior to conjugative transfer.
- PHEROMONE
-
A diffusible small molecule that can act as a chemical signal.
- HYPHAE
-
The tube-like cellular growth associated with mycelial organisms.
- MOBILIZATION
-
The process of a non-self-transmissible element being allowed to tranfer by the presence of a self-transmissible element.
- RELAXASOME
-
The protein–DNA complex at the transfer origin that results in nicking of the DNA when the proteins are denatured chemically or cleaved proteolytically.
- SURFACE EXCLUSION
-
The reduction of transfer frequency during conjugative transfer to recipients already carrying a related plasmid.
- PARALOGUES
-
Homologous genes in the same organism that have evolved from a gene duplication and a subsequent divergence of function.
- ORTHOLOGUES
-
Homologues that are related to each other through a speciation event.
- HELICASE
-
Enzyme that unwinds DNA duplexes.
- LAGGING-STRAND SYNTHESIS
-
Discontinuous synthesis on the strand running back from the replication fork, dependent on regular synthesis of primers by a primase and other primosome proteins.
Rights and permissions
About this article
Cite this article
Thomas, C., Nielsen, K. Mechanisms of, and Barriers to, Horizontal Gene Transfer between Bacteria. Nat Rev Microbiol 3, 711–721 (2005). https://doi.org/10.1038/nrmicro1234
Issue Date:
DOI: https://doi.org/10.1038/nrmicro1234
This article is cited by
-
Horizontal gene transfer after faecal microbiota transplantation in adolescents with obesity
Microbiome (2024)
-
Plasmids, a molecular cornerstone of antimicrobial resistance in the One Health era
Nature Reviews Microbiology (2024)
-
Ser/Leu-swapped cell-free translation system constructed with natural/in vitro transcribed-hybrid tRNA set
Nature Communications (2024)
-
Virulence and antimicrobial resistance genes occurring in Salmonella spp. isolated from aquatic food
Journal of Consumer Protection and Food Safety (2024)
-
Genomic insights into plasmid-mediated antimicrobial resistance in the bacterium Bhargavaea beijingensis strain PS04
Archives of Microbiology (2024)