Key Points
-
Type III secretion systems (T3SSs) are protein transport nanomachines that are used by numerous important Gram-negative bacterial pathogens and symbionts to establish trans-kingdom interactions with different hosts. They are essential virulence factors for many notorious bacterial pathogens, including the agents of plague and typhoid fever.
-
T3SSs evolved from the flagellum, which is a key organelle for bacterial motility, and the core components that are involved in the assembly of these complex nanomachines are highly conserved. Although the flagellar systems are largely inherited vertically, the non-flagellar T3SSs can be transmitted through horizontal gene transfer.
-
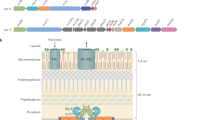
T3SSs are syringe-shaped, multi-megadalton complexes that are composed of more than 20 proteins and a series of ring structures embedded in the bacterial inner and outer membranes, as well as translocation pores in the host cell membrane. Encircled by these rings is a hollowed conduit that enables the delivery of partially unfolded virulence effector proteins into the host cell.
-
Integrative imaging technologies that combine cryo-electron microscopy (cryo-EM), X-ray crystallography, NMR and computer modelling have enabled the high-resolution visualization of key substructures of the T3SSs and the nanomachine in action. Assembly of the T3SS apparatus and substrate secretion occur in a defined temporal order and hierarchy, and genetic analyses and molecular biology have enabled the identification and functional characterization of the key regulators that control these processes.
-
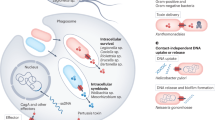
Effector proteins that are secreted through T3SSs carry out various functions within the host cell, including the manipulation of host immune responses and actin cytoskeletal dynamics, subverting gene expression and post-translational modifications, hijacking signal transduction pathways, and interrupting vesicle transport and endocytic trafficking, all of which can promote bacterial colonization, survival and replication.
-
T3SSs are attractive targets for vaccines and therapeutics owing to their essential roles in bacterial virulence and pathogenicity. By targeting bacterial virulence mechanisms instead of growth, inhibitors of T3SSs may exert less selective pressure on pathogens to develop drug resistance. Structural and functional characterization of T3SSs should facilitate mechanism-based drug design.
Abstract
Type III secretion systems (T3SSs) are protein transport nanomachines that are found in Gram-negative bacterial pathogens and symbionts. Resembling molecular syringes, T3SSs form channels that cross the bacterial envelope and the host cell membrane, which enable bacteria to inject numerous effector proteins into the host cell cytoplasm and establish trans-kingdom interactions with diverse hosts. Recent advances in cryo-electron microscopy and integrative imaging have provided unprecedented views of the architecture and structure of T3SSs. Furthermore, genetic and molecular analyses have elucidated the functions of many effectors and key regulators of T3SS assembly and secretion hierarchy, which is the sequential order by which the protein substrates are secreted. As essential virulence factors, T3SSs are attractive targets for vaccines and therapeutics. This Review summarizes our current knowledge of the structure and function of this important protein secretion machinery. A greater understanding of T3SSs should aid mechanism-based drug design and facilitate their manipulation for biotechnological applications.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Change history
12 May 2017
In the above article, a mistake was introduced in table 1; FliJ should have read 'FlgJ' in the row of 'Lytic transglycosylase'. This has now been corrected in the PDF and online. The authors apologize to readers for any confusion caused.
References
Costa, T. R. D. et al. Secretion systems in Gram-negative bacteria: structural and mechanistic insights. Nat. Rev. Microbiol. 13, 343–359 (2015).
Hueck, C. J. Type III protein secretion systems in bacterial pathogens of animals and plants. Microbiol. Mol. Biol. Rev. 62, 379–433 (1998).
Büttner, D. Protein export according to schedule: architecture, assembly, and regulation of type III secretion systems from plant- and animal-pathogenic bacteria. Microbiol. Mol. Biol. Rev. 76, 262–310 (2012).
Notti, R. Q. & Stebbins, C. E. The structure and function of type III secretion systems. Microbiol. Spectr. http://dx.doi.org/10.1128/microbiolspec.VMBF-0004-2015 (2015).
Portaliou, A. G., Tsolis, K. C., Loos, M. S., Zorzini, V. & Economou, A. Type III secretion: building and operating a remarkable nanomachine. Trends Biochem. Sci. 41, 175–189 (2016).
Abby, S. S. & Rocha, E. P. C. The non-flagellar type III secretion system evolved from the bacterial flagellum and diversified into host-cell adapted systems. PLoS Genet. 8, e1001983 (2012). This paper provides convincing bioinformatic evidence that T3SSs evolved from bacterial flagella.
Michiels, T., Wattiau, P., Brasseur, R., Ruysschaert, J.-M. & Cornelis, G. Secretion of Yop proteins by Yersiniae. Infect. Immun. 58, 2840–2849 (1990).
Rosqvist, R., Forsberg, Å. & Wolf-Watz, H. Intracellular targeting of the Yersinia YopE cytotoxin in mammalian cells induces actin microfilament disruption. Infect. Immun. 59, 4562–4569 (1991).
van der Heijden, J. & Finlay, B. B. Type III effector-mediated processes in Salmonella infection. Future Microbiol. 7, 685–670 (2012).
Raymond, B. et al. Subversion of trafficking, apoptosis, and innate immunity by type III secretion system effectors. Trends Microbiol. 21, 430–441 (2013).
Büttner, D. Behind the lines — actions of bacterial type III effector proteins in plant cells. FEMS Microbiol. Rev. 40, 894–937 (2016).
Staehelin, C. & Krishnan, H. B. Nodulation outer proteins: double-edged swords of symbiotic rhizobia. Biochem. J. 470, 263–274 (2015).
Santos, A. S. & Finlay, B. B. Bringing down the host: enteropathogenic and enterohaemorrhagic Escherichia coli effector-mediated subversion of host innate immune pathways. Cell. Microbiol. 17, 318–332 (2015).
Bliska, J. B., Wang, X., Viboud, G. I. & Brodsky, I. E. Modulation of innate immune responses by Yersinia type III secretion system translocators and effectors. Cell. Microbiol. 15, 1622–1631 (2013).
Beeckman, D. S. & Vanrompay, D. C. Bacterial secretion systems with an emphasis on the chlamydial type III secretion system. Curr. Issues Mol. Biol. 12, 17–41 (2010).
Miyata, S., Casey, M., Frank, D. W., Ausubel, F. M. & Drenkard, E. Use of the Galleria mellonella caterpillar as a model host to study the role of the type III secretion system in Pseudomonas aeruginosa pathogenesis. Infect. Immun. 71, 2404–2413 (2003).
Block, A. & Alfano, J. R. Plant targets for Pseudomonas syringae type III effectors: virulence targets or guarded decoys? Curr. Opin. Microbiol. 14, 39–46 (2011).
Gaytán, M. O., Martinez-Santos, V. I., Soto, E. & González-Pedrajo, B. Type three secretion system in attaching and effacing pathogens. Front. Cell. Infect. Microbiol. 6, 129 (2016).
Gazi, A. D. et al. Phylogenetic analysis of a gene cluster encoding an additional, rhizobial-like type III secretion system that is narrowly distributed among Pseudomonas syringae strains. BMC Microbiol. 12, 188 (2012).
Diepold, A. & Wagner, S. Assembly of the bacterial type III secretion machinery. FEMS Microbiol. Rev. 38, 802–822 (2014).
Kubori, T. et al. Supramolecular structure of the Salmonella typhimurium type III protein secretion system. Science 280, 602–605 (1998). This paper provides the first electron microscope images of isolated Salmonella SPI-1 T3SS needle complexes.
Macnab, R. M. How bacteria assemble flagella. Annu. Rev. Microbiol. 57, 77–100 (2003).
Schraidt, O. & Marlovits, T. C. Three-dimensional model of Salmonella's needle complex at subnanometer resolution. Science 331, 1192–1195 (2011). This study describes cryo-EM structures of the Salmonella SPI-1 T3SS needle complex and determines the stoichiometry of its basal body components.
Cornelis, G. R. The type III secretion injectisome. Nat. Rev. Microbiol. 4, 811–825 (2006).
Burkinshaw, B. J. & Strynadka, N. C. J. Assembly and structure of the T3SS. Biochim. Biophys. Acta 1843, 1649–1663 (2014).
Schraidt, O. et al. Topology and organization of the Salmonella typhimurium type III secretion needle complex components. PLoS Pathog. 6, e1000824 (2010).
Hu, B. et al. Visualization of the type III secretion sorting platform of Shigella flexneri. Proc. Natl Acad. Sci. USA 112, 1047–1052 (2015). This paper characterizes the structure of the cytoplasmic sorting platform of the S. flexneri T3SS using cryo-EM.
Makino, F. et al. The architecture of the cytoplasmic region of type III secretion systems. Sci. Rep. 6, 33341 (2016). This study describes cryo-EM structures of the cytoplasmic sorting platform in both the Shigella spp. T3SS and the Salmonella SPI-1 T3SS and flagellum.
Mueller, C. A. et al. The V-antigen of Yersinia forms a distinct structure at the tip of injectisome needles. Science 310, 674–676 (2005).
Epler, C. R., Dickenson, N. E., Bullitt, E. & Picking, W. L. Ultrastructural analysis of IpaD at the tip of the nascent MxiH type III secretion apparatus of Shigella flexneri. J. Mol. Biol. 420, 29–39 (2012).
Journet, L., Agrain, C., Broz, P. & Cornelis, G. R. The needle length of bacterial injectisomes is determined by a molecular ruler. Science 302, 1757–1760 (2003).
Erhardt, M., Singer, H. M., Wee, D. H., Keener, J. P. & Hughes, K. T. An infrequent molecular ruler controls flagellar hook length in Salmonella enterica. EMBO J. 30, 2948–2961 (2011).
Monjaras Feria, J. et al. Role of EscP (Orf16) in injectisome biogenesis and regulation of type III protein secretion in enteropathogenic Escherichia coli. J. Bacteriol. 194, 6029–6045 (2012).
Wee, D. H. & Hughes, K. T. Molecular ruler determines needle length for the Salmonella Spi-1 injectisome. Proc. Natl Acad. Sci. USA 112, 4098–4103 (2015).
Parsot, C., Hamiaux, C. & Page, A. L. The various and varying roles of specific chaperones in type III secretion systems. Curr. Opin. Microbiol. 6, 7–14 (2003).
Thomas, N. A., Ma, I., Prasad, M. E. & Rafuse, C. Expanded roles for multicargo and class 1B effector chaperones in type III secretion. J. Bacteriol. 194, 3767–3773 (2012).
Kimbrough, T. G. & Miller, S. I. Contribution of Salmonella typhimurium type III secretion components to needle complex formation. Proc. Natl Acad. Sci. USA 97, 11008–11013 (2000).
Gauthier, A., Puente, J. L. & Finlay, B. B. Secretin of the enteropathogenic Escherichia coli type III secretion system requires components of the type III apparatus for assembly and localization. Infect. Immun. 71, 3310–3319 (2003).
Wagner, S. et al. Organization and coordinated assembly of the type III secretion export apparatus. Proc. Natl Acad. Sci. USA 107, 17745–17750 (2010).
Diepold, A. et al. Deciphering the assembly of the Yersinia type III secretion injectisome. EMBO J. 29, 1928–1940 (2010).
Yip, C. K. et al. Structural characterization of the molecular platform for type III secretion system assembly. Nature 435, 702–707 (2005).
Marlovits, T. C. et al. Structural insights into the assembly of the type III secretion needle complex. Science 306, 1040–1042 (2004).
Zilkenat, S. et al. Determination of the stoichiometry of the complete bacterial type III secretion needle complex using a combined quantitative proteomic approach. Mol. Cell. Proteomics 15, 1598–1609 (2016).
Dietsche, T. et al. Structural and functional characterization of the bacterial type III secretion export apparatus. PLoS Pathog. 12, e1006071 (2016).
Worrall, L. J. et al. Near-atomic resolution cryo-EM analysis of the Salmonella T3S injectisome basal body. Nature 540, 597–601 (2016). This paper provides near-atomic resolution cryo-EM structures of the Salmonella SPI-1 T3SS basal body, revealing an unprecedented double β-barrel that is formed by the outer membrane secretin.
Hodgkinson, J. L. et al. Three-dimensional reconstruction of the Shigella T3SS transmembrane regions reveals 12-fold symmetry and novel features throughout. Nat. Struct. Mol. Biol. 16, 477–485 (2009).
Kowal, J. et al. Structure of the dodecameric Yersinia enterocolitica secretin YscC and its trypsin-resistant core. Structure 21, 2152–2161 (2013).
Spreter, T. et al. A conserved structural motif mediates formation of the periplasmic rings in the type III secretion system. Nat. Struct. Mol. Biol. 16, 468–476 (2009).
Bergeron, J. R. et al. A refined model of the prototypical Salmonella SPI-1 T3SS basal body reveals the molecular basis for its assembly. PLoS Pathog. 9, e1003307 (2013).
Abrusci, P. et al. Architecture of the major component of the type III secretion system export apparatus. Nat. Struct. Mol. Biol. 20, 99–104 (2013).
Zarivach, R. et al. Structural analysis of the essential self-cleaving type III secretion proteins EscU and SpaS. Nature 453, 124–127 (2008).
Deane, J. E. et al. Crystal structure of Spa40, the specificity switch for the Shigella flexneri type III secretion system. Mol. Microbiol. 69, 267–276 (2008).
Wiesand, U. et al. Structure of the type III secretion recognition protein YscU from Yersinia enterocolitica. J. Mol. Biol. 385, 854–866 (2009).
Kawamoto, A. et al. Common and distinct structural features of Salmonella injectisome and flagellar basal body. Sci. Rep. 3, 3369 (2013).
Kudryashev, M. et al. In situ structural analysis of the Yersinia enterocolitica injectisome. eLife 2, e00792 (2013).
Nans, A., Kudryashev, M., Saibil, H. R. & Hayward, R. D. Structure of a bacterial type III secretion system in contact with a host membrane in situ. Nat. Commun. 6, 10114 (2015).
Lara-Tejero, M., Kato, J., Wagner, S., Liu, X. & Galán, J. E. A sorting platform determines the order of protein secretion in bacterial type III systems. Science 331, 1188–1191 (2011).
Diepold, A., Kudryashev, M., Delalez, N. J., Berry, R. M. & Armitage, P. Composition, formation, and regulation of the cytosolic C-ring, a dynamic component of the type III secretion injectisome. PLoS Biol. 13, e1002039 (2015). This study demonstrates that the cytoplasmic C-ring and sorting platform of the Yersinia spp. T3SS are dynamic structures.
Notti, R. Q., Bhattacharya, S., Lilic, M. & Stebbins, C. E. A common assembly module in injectisome and flagellar type III secretion sorting platforms. Nat. Commun. 7, 8125 (2015).
McDowell, M. A. et al. Characterization of Shigella Spa33 and Thermotoga FliM/N reveals a new model for C-ring assembly in T3SS. Mol. Microbiol. 99, 749–766 (2016).
Zarivach, R., Vuckovic, M., Deng, W., Finlay, B. B. & Strynadka, N. C. J. Structural analysis of a prototypical ATPase from the type III secretion system. Nat. Struct. Mol. Biol. 14, 131–137 (2007).
Imada, K., Minamino, T., Tahara, A. & Namba, K. Structural similarity between the flagellar type III ATPase FliI and F1-ATPase subunits. Proc. Natl Acad. Sci. USA 104, 485–490 (2007).
Imada, K., Minamino, T., Uchida, Y., Kinoshita, M. & Namba, K. Insight into the flagella type III export revealed by the complex structure of the type III ATPase and its regulator. Proc. Natl Acad. Sci. USA 113, 3633–3638 (2016).
Akeda, Y. & Galan, J. E. Chaperone release and unfolding of substrates in type III secretion. Nature 437, 911–915 (2005). This paper shows the role of chaperones in unfolding substrates during type III secretion.
Marlovits, T. C. et al. Assembly of the inner rod determines needle length in the type III secretion injectisome. Nature 441, 637–640 (2006).
Loquet, A. et al. Atomic model of the type III secretion system needle. Nature 486, 276–279 (2012).
Demers, J.-P. et al. The common structural architecture of Shigella flexneri and Salmonella typhimurium type three secretion needles. PLoS Pathog. 9, e1003245 (2013). This study demonstrates that the needles of Salmonella Typhimurium and S. flexneri T3SSs adopt a common structural architecture.
Verasdonck, J. et al. Reassessment of MxiH subunit orientation and fold within native Shigella T3SS needles using surface labeling and solid-sate NMR. J. Struct. Biol. 192, 441–448 (2015).
Deane, J. E. et al. Molecular model of a type III secretion system needle: implications for host-cell sensing. Proc. Natl Acad. Sci. USA 103, 12529–12533 (2006).
Fujii, T. et al. Structure of a type III secretion needle at 7-Å resolution provides insights into its assembly and signaling mechanisms. Proc. Natl Acad. Sci. USA 109, 4461–4466 (2012).
Knutton, S. et al. A novel EspA-associated surface organelle of enteropathogenic Escherichia coli involved in protein translocation into epithelial cells. EMBO J. 17, 2166–2176 (1998).
Yip, C. K., Finlay, B. B. & Strynadka, N. C. J. Structural characterization of a type III secretion system filament protein in complex with its chaperone. Nat. Struct. Mol. Biol. 12, 75–81 (2005).
Jin, Q. & He, S.-Y. Role of the Hrp pilus in type III protein secretion in Pseudomonas syringae. Science 294, 2556–2558 (2001). This paper provides direct evidence that the Hrp pilus is probably the conduit for protein delivery in type III secretion.
Li, C.-M. et al. The Hrp pilus of Pseudomonas syringae elongates from its tip and acts as a conduit for translocation of the effector protein HrpZ. EMBO J. 21, 1909–1915 (2002). Together with reference 73, this paper shows the Hrp pilus as the conduit for protein delivery in type III secretion.
Romano, F. B. et al. Type 3 secretion translocators spontaneously assembles a hexadecameric transmembrane complex. J. Biol. Chem. 291, 6304–6315 (2016).
Sheahan, K.-L. & Isberg, R. R. Identification of mammalian proteins that collaborate with type III secretion system function: involvement of a chemokine receptor in supporting translocon activity. mBio 6, e02023-14 (2015).
Blondel, C. J. et al. CRISPR/Cas9 screens reveal requirements for host cell sulfation and fucosylation in bacterial type III secretion system-mediated cytotoxicity. Cell Host Microbe 20, 226–237 (2016).
Russo, B. C. et al. Intermediate filaments enable pathogen docking to trigger type 3 effector translocation. Nat. Microbiol. 1, 16025 (2016).
Deng, W. et al. Dissecting virulence: systematic and functional analyses of a pathogenicity island. Proc. Natl Acad. Sci. USA 101, 3597–3602 (2004).
Burkinshaw, B. J. et al. Structural analysis of a specialized type III secretion system peptidoglycan-cleaving enzyme. J. Biol. Chem. 290, 10406–10417 (2015).
Creasey, E. A., Delahay, R. M., Daniell, S. J. & Frankel, G. Yeast two-hybrid system survey of interactions between LEE-encoded proteins of enteropathogenic Escherichia coli. Microbiology 149, 2093–2106 (2003).
Sorg, I. et al. YscU recognizes translocators as export substrates of the Yersinia injectisome. EMBO J. 26, 3015–3024 (2007).
Wood, S. E., Jin, J. & Lloyd, S. A. YscP and YscU switch the substrate specificity of the Yersinia type III secretion system by regulating export of the inner rod protein YscI. J. Bacteriol. 190, 5252–4263 (2008).
Sal-Man, N., Deng, W. & Finlay, B. B. EscI: a crucial component of the type III secretion system forms the inner rod structure in enteropathogenic Escherichia coli. Biochem. J. 442, 119–125 (2012).
Makishima, S., Komoriya, K., Yamaguchi, S. & Aizawa, S. I. Length of the flagellar hook and the capacity of the type III export apparatus. Science 291, 2411–2413 (2001).
Lefebre, M. D. & Galán, J. E. The inner rod protein controls substrate switching and needle length in a Salmonella type III secretion system. Proc. Natl Acad. Sci. USA 111, 817–822 (2014).
Nariya, M. K., Israeli, J., Shi, J. J. & Deeds, E. J. Mathematical model for length control by the timing of substrate switching in the type III secretion system. PLoS Comput. Biol. 12, e1004851 (2016).
Wagner, S., Stenta, M., Metzger, L. C., Dal Perato, M. & Cornelis, G. R. Length control of the injectisome needle requires only one molecule of Yop secretion protein P (YscP). Proc. Natl Acad. Sci. USA 107, 13860–13865 (2010).
Bergeron, J. R. et al. The structure of a type 3-secretion system (T3SS) ruler protein suggests a molecular mechanism for needle length sensing. J. Biol. Chem. 391, 1676–1691 (2016).
Mizuno, S., Amida, H., Kobayashi, N., Aizawa, S. & Tate, S. The NMR structure of FliK, the trigger for the switch of substrate specificity in the flagellar type III secretion apparatus. J. Mol. Biol. 409, 558–573 (2011).
Ho, O. et al. Characterization of the ruler protein interaction interface on the substrate specificity switch protein in the Yersinia type III secretion system. J. Biol. Chem. 292, 3299–3311 (2017).
Login, F. H. & Wolf-Watz, H. YscU/FlhB of Yersinia pseudotuberculosis harbors a C-terminal T3S signal. J. Biol. Chem. 290, 26282–26291 (2015).
Monjaras Feria, J. V., Lefebre, M. D., Stierhof, Y.-D., Galán, J. E. & Wagner, S. Role of autocleavage in the function of a type III secretion specificity switch protein in Salmonella enterica serovar Typhimurium. mBio 6, e01459-15 (2015).
Frost, S. et al. Autoproteolysis and intramolecular dissociation of Yersinia YscU precedes secretion of its C-terminal polypeptide YscUCC . PLoS ONE 7, e49349 (2012).
Cherradi, Y. et al. Interplay between predicted inner-rod and gatekeeper in controlling substrate specificity of the type III secretion system. Mol. Microbiol. 87, 1183–1199 (2013).
Archuleta, T. L. & Spiller, B. W. A gatekeeper chaperone complex directs translocator secretion during type III secretion. PLoS Pathog. 10, e1004498 (2014).
Deng, W. et al. Regulation of type III secretion hierarchy of translocators and effectors in attaching and effacing bacterial pathogens. Infect. Immun. 73, 2135–2146 (2005).
Martinez-Argudo, I. & Blocker, A. J. The Shigella T3SS needle transmits a signal for MxiC release, which controls secretion of effectors. Mol. Microbiol. 78, 1365–1378 (2010).
Wang, D., Roe, A. J., McAteer, S., Shipston, M. J. & Gally, D. L. Hierarchal type III secretion of translocators and effectors from Escherichia coli O157:H7 requires the carboxy terminus of SepL that binds to Tir. Mol. Microbiol. 69, 1499–1512 (2008).
Yu, X. J., McGourty, K., Liu, M., Unsworth, K. E. & Holden, D. W. pH sensing by intracellular Salmonella induces effector translocation. Science 328, 1040–1043 (2010).
Armentrout, E. I. & Rietsch, A. The type III secretion translocation pore senses host cell contact. PLoS Pathog. 12, e1005530 (2016).
Mills, E., Baruch, K., Charpentier, X., Kobi, S. & Rosenshine, I. Real-time analysis of effector translocation by the type III secretion system of enteropathogenic Escherichia coli. Cell Host Microbe 3, 104–113 (2008).
Mills, E., Baruch, K., Aviv, G., Nitzan, M. & Rosenshine, I. Dynamics of the type III secretion system activity of enteropathogenic Escherichia coli. mBio 4, e00303-13 (2013). This study delineates the secretion hierarchy of EPEC effectors.
Dewoody, R., Merritt, P. M., Houppert, A. S. & Marketon, M. M. YopK regulates the Yersinia pestis type III secretion system from within host cells. Mol. Microbiol. 79, 1445–1461 (2011).
Berger, C. N. et al. EspZ of enteropathogenic and enterohemorrhagic Escherichia coli regulates type III secretion system protein translocation. mBio 3, e00317-12 (2012).
Radics, J., Königsmaier, L. & Marlovits, T. C. Structure of a pathogenic type 3 secretion system in action. Nat. Struct. Mol. Biol. 21, 82–87 (2014). This paper visualizes a trapped substrate in the Salmonella SPI-1 T3SS apparatus, which provides direct evidence for the needle lumen as the conduit for secretion.
Arnold, R. et al. Sequence-based prediction of type III secreted proteins. PLoS Pathog. 5, e1000376 (2009).
Samudrala, R., Heffron, F. & McDermott, J. E. Accurate prediction of secreted substrates and identification of a conserved putative secretion signal for type III secretion systems. PLoS Pathog. 5, 1000375 (2009).
McDermott, J. E. et al. Computational prediction of type III and IV secreted effectors in Gram-negative bacteria. Infect. Immun. 79, 23–32 (2011).
Anderson, D. M. & Schneewind, O. A. mRNA signal for the type III secretion of Yop proteins by Yersinia enterocolitica. Science 278, 1140–1143 (1997). This is the first report to show that some effectors use mRNA signals for type III secretion in Yersinia spp.
Anderson, D. M., Fouts, D. E., Collmer, A. & Schneewind, O. Reciprocal secretion of proteins by the bacterial type III machines of plant and animal pathogens suggests universal recognition of mRNA targeting signals. Proc. Natl Acad. Sci. USA 96, 12839–12843 (1999).
Lloyd, S. A., Norman, M., Rosqvist, R. & Wolf-Watz, H. Yersinia YopE is targeted for type III secretion by N-terminal, not mRNA, signals. Mol. Microbiol. 39, 520–531 (2001).
Lee, S. H. & Galan, J. E. Salmonella type III secretion-associated chaperones confer secretion-pathway specificity. Mol. Microbiol. 51, 483–495 (2004).
Deng, W., Yu, H. B., Li, Y. & Finlay, B. B. SepD/SepL-dependent secretion signals of the type III secretion system translocator proteins in enteropathogenic Escherichia coli. J. Bacteriol. 197, 1263–1275 (2015).
Tomalka, A. G., Stopford, C. M., Lee, P.-C. & Rietsch, A. A translocator-specific export signal establishes the translocator-effector secretion hierarchy that is important for type III secretion system function. Mol. Microbiol. 86, 1464–1481 (2012).
Niemann, G. S. et al. RNA type III secretion signals that require Hfq. J. Bacteriol. 195, 2119–2125 (2013). This study shows that some Salmonella spp. effectors also use RNA-based type III secretion signals.
Stebbins, C. E. & Galan, J. E. Maintenance of an unfolded polypeptide by a cognate chaperone in bacterial type III secretion. Nature 414, 77–81 (2001).
Izore, T., Job, V. & Dessen, A. Biogenesis, regulation, and targeting of the type III secretion system. Structure 19, 603–612 (2011).
Dohlich, K., Zumsteg, A. B., Goosmann, C. & Kolbe, M. A substrate-fusion protein is trapped inside the type III secretion system channel in Shigella flexneri. PLoS Pathog. 10, e1003881 (2014).
Gauthier, A. & Finlay, B. B. Translocated intimin receptor and its chaperone interact with ATPase of the type III secretion apparatus of enteropathogenic Escherichia coli. J. Bacteriol. 185, 6747–6755 (2003).
Chen, L. et al. Substrate-activated conformational switch on chaperones encodes a targeting signal in type III secretion. Cell Rep. 3, 709–715 (2013).
Sorg, J. A., Blaylock, B. & Schneewind, O. Secretion signal recognition by YscN, the Yersinia type III secretion ATPase. Proc. Natl Acad. Sci. USA 103, 16490–16496 (2006).
Wilharm, G. et al. Yersinia enterocolitica type III secretion depends on the proton motive force but not on the flagellar motor components MotA and MotB. Infect. Immun. 72, 4004–4009 (2004).
Erhardt, M., Mertens, M. E., Fabiani, F. D. & Hughes, K. T. ATPase-independent type-III protein secretion in Salmonella enterica. PLoS Genet. 10, e1004800 (2014).
Rathinavelan, T. et al. A repulsive electrostatic mechanism for protein export through the type III secretion apparatus. Biophys. J. 98, 452–461 (2010).
Evans, L. D., Poulter, S., Terentjev, E. M., Hughes, C. & Fraser, G. M. A chain mechanism for flagellum growth. Nature 504, 287–290 (2013).
Galán, J. E., Lara-Tejero, M., Marlovits, T. C. & Wagner, S. Bacterial type III secretion systems: specialized nanomachines for protein delivery into target cells. Annu. Rev. Microbiol. 68, 415–438 (2014).
Crepin, V. F., Shaw, R., Abe, C. M., Knutton, S. & Frankel, G. Polarity of enteropathogenic Escherichia coli EspA filament assembly and protein secretion. J. Bacteriol. 187, 2881–2889 (2005). This paper provides direct evidence that the EspA filament is the conduit for protein delivery in the T3SS of EPEC.
Akopyan, K. et al. Translocation of surface-localized effectors in type III secretion. Proc. Natl Acad. Sci. USA 108, 1639–1644 (2011). This paper demonstrates that surface-associated Yersinia effectors can be translocated into host cells and proposes an alternative two-step model to the direct injection model of T3SSs.
Edgren, T., Forsberg, Å., Rosqvist, R. & Wolf-Watz, H. Type III secretion in Yersinia: injectisome or not? PLoS Pathog. 8, e1002669 (2012).
Marteyn, B. et al. Modulation of Shigella virulence in response to available oxygen in vivo. Nature 465, 355–358 (2010).
Wang, H. et al. Increased plasmid copy number is essential for Yersinia T3SS function and virulence. Science 353, 492–495 (2016).
Bahrani, F. K., Sansonetti, P. J. & Parsot, C. Secretion of Ipa proteins by Shigella flexneri: inducer molecules and kinetics of activation. Infect. Immun. 65, 4005–4010 (1997).
Kim, J. et al. Factors triggering type III secretion in Pseudomonas aeruginosa. Microbiology 151, 3575–3587 (2005).
Murillo, I., Martinez-Argudo, I. & Blocker, A. J. Genetic dissection of the signaling cascade that controls activation of the Shigella type III secretion system from the needle tip. Sci. Rep. 6, 27649 (2016).
Kenjale, R. et al. The needle component of the type III secretion of Shigella regulates the activity of the secretion apparatus. J. Biol. Chem. 280, 42929–42937 (2005).
Fernandez-Leiro, R. & Scheres, S. H. W. Unraveling biological macromolecules with cryo-electron microscopy. Nature 537, 339–346 (2016).
Marshall, N. C. & Finlay, B. B. Targeting the type III secretion system to treat bacterial infections. Expert Opin. Ther. Targets 18, 137–152 (2014).
Anantharajah, A., Mingeot-Leclercq, M.-P. & Van Bambeke, F. Targeting the type three secretion system in Pseudomonas aeruginosa. Trends Pharmacol. Sci. 37, 734–749 (2016).
Pallen, M. J., Beatson, S. A. & Bailey, C. M. Bioinformatics, genomics, and evolution of non-flagellar type-III secretion systems: a Darwinian perspective. FEMS Microbiol. Rev. 29, 201–229 (2005).
Sun, Y. H., Rolán, H. G. & Tsolis, R. M. Injection of flagellin into the host cell cytosol by Salmonella enterica serotype Typhimurium. J. Biol. Chem. 282, 33897–33901 (2007).
Du, J. et al. The type III secretion system apparatus determines the intracellular niche of bacterial pathogens. Proc. Natl Acad. Sci. USA 113, 4794–4799 (2016).
Soto, E. et al. Functional characterization of EscK (Orf4), a sorting platform component of the enteropathogenic Escherichia coli injectisome. J. Bacteriol. 199, e00538-16 (2017).
Tampakaki, A. P. Commonalities and differences of T3SSs in rhizobia and plant pathogenic bacteria. Front. Plant Sci. 5, 114 (2014).
Acknowledgements
The authors apologize to those colleagues whose studies were not highlighted here owing to space constraints. This work was funded by operating grants from the Canadian Institutes of Health Research (to B.B.F. and N.C.J.S.) and the Howard Hughes International Senior Scholar program (to N.C.J.S.). N.C.J.S. is a Tier I Canada Research Chair in Antibiotic Discovery. B.B.F. is the UBC Peter Wall Distinguished Professor.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Glossary
- Cryo-electron microscopy
-
(Cryo-EM). A transmission electron microscopy (TEM) technique that is used in structural biology for imaging unstained and unfixed frozen-hydrated specimens at cryogenic (generally liquid nitrogen) temperatures, thus enabling the preservation of their native state.
- Basal body
-
A component of the type III secretion apparatus that is composed of highly oligomerized concentric rings that are embedded in the bacterial inner and outer membranes, excluding any extracellular appendages such as the needle filament.
- Needle
-
The hollow filamentous structure that is formed by helical polymerization of a single protein that lines the inside of the type III secretion system basal body and protrudes to the bacterial surface.
- Translocation pore
-
Hetero-oligomeric complexes of two hydrophobic membrane proteins that contain a central pore and form in the host cell membrane, and that enable the injection of effector proteins.
- Secretin
-
A family of proteins that form large and extremely stable multimeric complexes and channels in the outer membrane of Gram-negative bacteria. Secretins are essential for protein transport across membranes in type II and type III secretion systems, the type IV pilus system, and filamentous bacteriophage assembly and extrusion.
- Sorting platform
-
A substructure that includes the cytoplasmic ring and the ATPase complex. It is located on the bacterial cytoplasmic side of the type III secretion apparatus, where dynamic substrate uploading and secretion takes place.
- Secretion hierarchy
-
The temporal order by which the different categories (early, middle and late) of substrate proteins are secreted by type III secretion systems.
- Chaperones
-
Small proteins in type III secretion systems that maintain substrates in a partially unfolded secretion-competent state and/or prevent undesired premature protein–protein interactions or aggregation. They may also be involved in substrate targeting and secretion hierarchy.
- Injectisome
-
A syringe-like protein secretion apparatus that forms a channel across both the inner membrane and the outer membrane of Gram-negative bacteria, as well as the host cell membrane, and is used by bacteria to deliver bacterial proteins into eukaryotic host cells through type III secretion.
- General secretory pathway
-
(Sec pathway). An essential protein export machinery that transports proteins across the plasma membrane and is evolutionarily conserved in all domains of life. It often recognizes its substrates through an amino-terminal signal peptide, which is cleaved during secretion. The majority of secreted proteins in bacteria are exported through this pathway.
- β-Barrel
-
A closed structure that is formed by twisted and coiled β-sheets that are composed of β-strands. It is a common assembly by certain outer membrane proteins, such as porins and secretins of protein secretion systems, in Gram-negative bacteria.
- F/V-type ATPases
-
Phosphorylation factor-type (F-type) F0F1 ATPases and vacuolar-type (V-type) ATPases are multisubunit protein complexes that couple transmembrane movement and the pumping of ions (protons or Na+) with ATP hydrolysis or synthesis. They are involved in providing the energy for various cellular activities, including protein trafficking and the movement of metabolites.
Rights and permissions
About this article
Cite this article
Deng, W., Marshall, N., Rowland, J. et al. Assembly, structure, function and regulation of type III secretion systems. Nat Rev Microbiol 15, 323–337 (2017). https://doi.org/10.1038/nrmicro.2017.20
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrmicro.2017.20
This article is cited by
-
Cytosolic sorting platform complexes shuttle type III secretion system effectors to the injectisome in Yersinia enterocolitica
Nature Microbiology (2024)
-
Microbial life in 25-m-deep boreholes in ancient permafrost illuminated by metagenomics
Environmental Microbiome (2023)
-
Assembly mechanism of a Tad secretion system secretin-pilotin complex
Nature Communications (2023)
-
Salmonella T3SS effector SseK1 arginine-glycosylates the two-component response regulator OmpR to alter bile salt resistance
Scientific Reports (2023)
-
Genome wide analysis revealed conserved domains involved in the effector discrimination of bacterial type VI secretion system
Communications Biology (2023)