Key Points
-
During the diversification of Fungi and the rise of conifer-dominated and angiosperm-dominated forests, ectomycorrhizal symbioses have enabled trees to colonize boreal and temperate regions.
-
Ectomycorrhizal fungi have evolved on several independent occasions from diverse saprotrophic lineages.
-
A large-scale loss of plant cell wall degradative enzymes — and, consequently, degradative abilities — has occurred in all ectomycorrhizal fungal lineages. However, the degradative enzymes that have been retained vary between each independent lineage and may correspond to a variation in capabilities for the decay of organic matter.
-
Genome analyses have shown how, through contraction and loss of major gene families, ectomycorrhizal fungi have become highly reliant on the availability of photoassimilates from their plant host, while preserving plant cell integrity by avoiding the release of degradative enzymes.
-
Ectomycorrhizal fungi use diffusible signalling molecules to manipulate the morphology and metabolism of host roots so that they provide a more suitable environment for fungal invasion.
-
The ectomycorrhizal fungus Laccaria bicolor has been used as a model for the study of protein effectors that manipulate host plant hormone receptors and related signalling pathways to dampen plant defences and facilitate fungal colonization.
Abstract
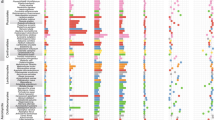
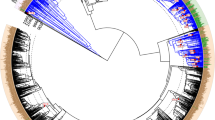
During the diversification of Fungi and the rise of conifer-dominated and angiosperm- dominated forests, mutualistic symbioses developed between certain trees and ectomycorrhizal fungi that enabled these trees to colonize boreal and temperate regions. The evolutionary success of these symbioses is evident from phylogenomic analyses that suggest that ectomycorrhizal fungi have arisen in approximately 60 independent saprotrophic lineages, which has led to the wide range of ectomycorrhizal associations that exist today. In this Review, we discuss recent genomic studies that have revealed the adaptations that seem to be fundamental to the convergent evolution of ectomycorrhizal fungi, including the loss of some metabolic functions and the acquisition of effectors that facilitate mutualistic interactions with host plants. Finally, we consider how these insights can be integrated into a model of the development of ectomycorrhizal symbioses.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Read, D. J., Leake, J. R. & Perez-Moreno, J. Mycorrhizal fungi as drivers of ecosystem processes in heathland and boreal forest biomes. Can. J. Bot. 82, 1243 (2004).
Smith, S. E. & Read, D. J. (eds) Mycorrhizal Symbiosis (Academic Press, 2008).
Pirozynski, K. A. & Malloch, D. W. The origin of land plants: a matter of mycotrophism. Biosystems 6, 153–164 (1975).
Simon, L., Bousquet, J., Lévesque, R. C. & Lalonde, M. Origin and diversification of endomycorrhizal fungi and coincidence with vascular land plants. Nature 363, 67–69 (1993).
Selosse, M. A. & Le Tacon, F. The land flora: a phototroph–fungus partnership? Trends Ecol. Evol. 13, 15–20 (1998).
Brundrett, M. C. Coevolution of roots and mycorrhizas of land plants. New Phytol. 154, 275–304 (2002).
Bidartondo, M. I. et al. The dawn of symbiosis between plants and fungi. Biol. Lett. 7, 574–577 (2011).
Strullu-Derrien, C. et al. Fungal associations in Horneophyton ligneri from the Rhynie Chert (c. 407 million year old) closely resemble those in extant lower land plants: novel insights into ancestral plant–fungus symbioses. New Phytol. 203, 964–979 (2014). This study is notable because it shows that early plants were colonized by species in the Mucoromycotina and Glomeromycota, which overturns the long-held paradigm that the early endophytes were exclusively in the Glomeromycota.
Van Der Heijden, M. G. A., Martin, F., Selosse, M. A. & Sanders, I. R. Mycorrhizal ecology and evolution: the past, the present, and the future. New Phytol. 205, 1406–1423 (2015).
Soudzilovskaia, N. A. et al. Global patterns of plant root colonization intensity by mycorrhizal fungi explained by climate and soil chemistry. Global Ecol. Biogeogr. 24, 371–382 (2015).
Simard, S. W. et al. Net transfer of carbon between ectomycorrhizal tree species in the field. Nature 388, 579–582 (1997).
Selosse, M.-A., Richard, F., He, X. & Simard, S. W. Mycorrhizal networks: des liaisons dangereuses? Trends Ecol. Evol. 21, 621–628 (2006).
Klein, T., Siegwolf, R. T. W. & Körner, C. Belowground carbon trade among tall trees in a temperate forest. Science 352, 342–344 (2016).
Finlay, R. Ecological aspects of mycorrhizal symbiosis: with special emphasis on the functional diversity of interactions involving the extraradical mycelium. J. Exp. Bot. 59, 1115–1126 (2008).
Treseder, K. K., Torn, M. S. & Masiello, C. A. An ecosystem-scale radiocarbon tracer to test use of litter carbon by ectomycorrhizal fungi. Soil Biol. Biochem. 38, 1077 (2006).
Martin, F. et al. Developmental cross talking in the ectomycorrhizal symbiosis: signals and communication genes. New Phytol. 151, 145–154 (2001).
Spanu, P. D. The genomics of obligate (and nonobligate) biotrophs. Annu. Rev. Phytopathol. 50, 91–109 (2012).
Bonfante, P. & Genre, A. Mechanisms underlying beneficial plant–fungus interactions in mycorrhizal symbiosis. Nat. Commun. 1, 48 (2010).
Martin, F. et al. Sequencing the fungal tree of life. New Phytol. 190, 818–821 (2011).
Tedersoo, L., May, T. W. & Smith, M. E. Ectomycorrhizal lifestyle in fungi: global diversity, distribution, and evolution of phylogenetic lineages. Mycorrhiza 20, 217–263 (2010).
Peterson, R. L. & Massicotte, H. B. Exploring structural definitions of mycorrhizas, with emphasis on nutrient-exchange interfaces. Can. J. Bot. 82, 1074–1088 (2004).
Duplessis, S., Courty, P. E., Tagu, D. & Martin, F. Transcript patterns associated with ectomycorrhiza development in Eucalyptus globulus and Pisolithus microcarpus. New Phytol. 165, 599–611 (2005).
Larsen, P. E. et al. Multi-omics approach identifies molecular mechanisms of plant–fungus mycorrhizal interaction. Front. Plant Sci. 6, 1061 (2016).
Martin, F. & Selosse, M.-A. The Laccaria genome: a symbiont blueprint decoded. New Phytol. 180, 379–390 (2008).
Plett, J. M. & Martin, F. Blurred boundaries: lifestyle lessons from ectomycorrhizal fungal genomes. Trends Genet. 27, 14–22 (2011).
Grigoriev, I. V. et al. MycoCosm portal: gearing up for 1000 fungal genomes. Nucleic Acids Res. 42, D699–D704 (2014).
Matheny, P. B. & Hibbett, D. S. The relative ages of ectomycorrhizal mushrooms and their plant hosts estimated using Bayesian relaxed molecular clock analyses. BMC Biol. 7, 13 (2009). The Bayesian relaxed molecular clock analyses that are carried out in this study show that there have been at least eight independent origins of ectomycorrhizal associations involving angiosperms, and at least six-to-eight origins of associations with gymnosperms.
Matheny, P. B. et al. Out of the Palaeotropics? Historical biogeography and diversification of the cosmopolitan ectomycorrhizal mushroom family Inocybaceae. J. Biogeogr. 36, 577–592 (2009).
Skrede, I. et al. Evolutionary history of Serpulaceae (Basidiomycota): molecular phylogeny, historical biogeography and evidence for a single transition of nutritional mode. BMC Evol. Biol. 11, 230 (2011).
Rineau, F. et al. Carbon availability triggers the decomposition of plant litter and assimilation of nitrogen by an ectomycorrhizal fungus. ISME J. 7, 2010 (2013).
Bödeker, I. T. M. et al. Ectomycorrhizal Cortinarius species participate in enzymatic oxidation of humus in northern forest ecosystems. New Phytol. 203, 245 (2014).
Högberg, M. N. & Högberg, P. Extramatrical ectomycorrhizal mycelium contributes one-third of microbial biomass and produces, together with associated roots, half the dissolved organic carbon in a forest soil. New Phytol. 154, 791 (2002).
Clemmensen, K. E. et al. Roots and associated fungi drive long-term carbon sequestration in boreal forest. Science 339, 1615 (2013).
Rytioja, J. et al. Plant-polysaccharide-degrading enzymes from Basidiomycetes. Microbiol. Mol. Biol. Rev. 78, 614–649 (2014).
Floudas, D. et al. The Paleozoic origin of enzymatic lignin decomposition reconstructed from 31 fungal genomes. Science 336, 1715–1719 (2012). This study maps the detailed evolution of wood-degrading enzymes by using a large set of genomes from wood-decaying fungi and shows that a key peroxidase and other enzymes that are involved in lignin decay were present in the common ancestor of the Agaricomycetes.
Nagy, L. G. et al. Comparative genomics of early-diverging mushroom-forming fungi provides insights into the origins of lignocellulose decay capabilities. Mol. Biol. Evol. 33, 959–970 (2016).
Hori, C. et al. Genome wide analysis of polysaccharides degrading enzymes in 11 white- and brown-rot Polyporales provides insight into mechanisms of wood decay. Mycologia 105, 1412–1427 (2013).
Lundell, T. K., Mäkelä, M. R., de Vries, R. P. & Hildén, K. S. Genomics, lifestyles and future prospects of wood-decay and litter-decomposing basidiomycota. Adv. Bot. Res. 70, 329–370 (2014).
Ruiz-Dueñas, F. J. et al. Lignin-degrading peroxidases in Polyporales: an evolutionary survey based on 10 sequenced genomes. Mycologia 105, 1428–1444 (2013).
Martin, F. et al. The genome of Laccaria bicolor provides insights into mycorrhizal symbiosis. Nature 452, 88–92 (2008). This study describes the first genome from an ectomycorrhizal symbiont, and the predicted gene inventory of L. bicolor indicates previously unknown mechanisms of symbiosis that operate in biotrophic mycorrhizal fungi.
Martin, F. et al. Périgord black truffle genome uncovers evolutionary origins and mechanisms of symbiosis. Nature 464, 1033–1038 (2010).
Tisserant, E. et al. The genome of an arbuscular mycorrhizal fungus provides insights into the oldest plant symbiosis. Proc. Natl Acad. Sci. USA 110, 20117–20122 (2013).
Kohler, A. et al. Convergent losses of decay mechanisms and rapid turnover of symbiosis genes in mycorrhizal mutualists. Nat. Genet. 47, 410–415 (2015). This works takes advantage of large-scale comparative genomics to show that the ectomycorrhizal lifestyle evolved in fungi through the repeated evolution of a 'symbiosis toolkit', with decreased numbers of PCWDEs and lineage-specific suites of mycorrhiza-induced genes.
Peter, M. et al. Ectomycorrhizal ecology is imprinted in the genome of the dominant symbiotic fungus Cenococcum geophilum. Nat. Commun. 7, 12662 (2016).
Kracher, D. et al. Extracellular electron transfer systems fuel cellulose oxidative degradation. Science 352, 1098–1101 (2016).
Bödeker, I. T. M., Nygren, C. M. R., Taylor, A. F. S., Olson, A. & Lindahl, B. D. Class II peroxidase-encoding genes are present in a phylogenetically wide range of ectomycorrhizal fungi. ISME J. 3, 1387–1395 (2009).
Talbot, J. M., Allison, S. D. & Treseder, K. K. Decomposers in disguise: mycorrhizal fungi as regulators of soil C dynamics in ecosystems under global change. Funct. Ecol. 22, 955–963 (2008).
Shah, F. et al. Ectomycorrhizal fungi decompose soil organic matter using oxidative mechanisms adapted from saprotrophic ancestors. New Phytol. 209, 1705–1719 (2015). This works demonstrates that several ectomycorrhizal fungi are able to decompose plant litter through the use of the Fenton reaction.
Lindahl, B. D. & Tunlid, A. Ectomycorrhizal fungi — potential organic matter decomposers, yet not saprotrophs. New Phytol. 205, 1443–1447 (2015).
Wolfe, B. E., Tulloss, R. E. & Pringle, A. The irreversible loss of a decomposition pathway marks the single origin of an ectomycorrhizal symbiosis. PLoS ONE 7, e39597 (2012).
Felten, J. et al. The ectomycorrhizal fungus Laccaria bicolor stimulates lateral root formation in poplar and Arabidopsis through auxin transport and signaling. Plant Physiol. 151, 1991–2005 (2009).
Martin, F., Kohler, A. & Duplessis, S. Living in harmony in the wood underground ectomycorrhizal genomics. Curr. Opin. Plant Biol. 10, 204–210 (2007).
Luo, Z. B. et al. Upgrading root physiology for stress tolerance by ectomycorrhizas: insights from metabolite and transcriptional profiling into reprogramming for stress anticipation. Plant Physiol. 151, 1902–1917 (2009).
Tschaplinski, T. J. et al. Populus trichocarpa and Populus deltoides exhibit different metabolomic responses to colonization by the symbiotic fungus Laccaria bicolor. Mol. Plant Microbe Interact. 27, 546–556 (2014).
Veneault-Fourrey, C. & Martin, F. Mutualistic interactions on a knife-edge between saprotrophy and pathogenesis. Curr. Opin. Plant Biol. 14, 444–450 (2011).
Vayssieres, A. et al. Development of the poplar–Laccaria bicolor ectomycorrhiza modifies root auxin metabolism, signaling, and response. Plant Physiol. 169, 890–902 (2015).
Krause, K. et al. Biosynthesis and secretion of indole-3-acetic acid and its morphological effects on Tricholoma vaccinum–spruce ectomycorrhiza. Appl. Environ. Microbiol. 81, 7003–7011 (2015).
Plett, J. M. et al. Ethylene and jasmonic acid act as negative modulators during mutualistic symbiosis between Laccaria bicolor and Populus roots. New Phytol. 202, 270–286 (2014).
Ditengou, F. A. et al. Volatile signalling by sesquiterpenes from ectomycorrhizal fungi reprogrammes root architecture. Nat. Commun. 6, 6279 (2015). This works suggests that volatile sesquiterpenes have a key role in the development of the ectomycorrhizal symbiosis by stimulating the formation of short roots.
Sukumar, P. et al. Involvement of auxin pathways in modulating root architecture during beneficial plant–microorganism interactions. Plant Cell Environ. 36, 909–919 (2013).
Doré, J. et al. Comparative genomics, proteomics and transcriptomics give new insight into the exoproteome of the basidiomycete Hebeloma cylindrosporum and its involvement in ectomycorrhizal symbiosis. New Phytol. 208, 1169–1187 (2015).
Plett, J. M. et al. The mutualist Laccaria bicolor expresses a core gene regulon during the colonization of diverse host plants and a variable regulon to counteract host-specific defenses. Mol. Plant Microbe Interact. 28, 261–273 (2014).
Garcia, K., Doidy, J., Zimmermann, S. D., Wipf, D. & Courty, P. E. Take a trip through the plant and fungal transportome of mycorrhiza. Trends Plant Sci. http://dx.doi.org/10.1016/j.tplants.2016.07.010 (2016).
Doré, J., Marmeisse, R., Combier, J. P. & Gay, G. A fungal conserved gene from the Basidiomycete Hebeloma cylindrosporum is essential for efficient ectomycorrhiza formation. Mol. Plant Microbe Interact. 27, 1059–1069 (2014).
Liao, H. L. et al. Metatranscriptomic analysis of ectomycorrhizal roots reveals genes associated with Piloderma–Pinus symbiosis: improved methodologies for assessing gene expression in situ. Environ. Microbiol. 16, 3730–3742 (2014).
Plett, J. M. et al. A secreted effector protein of Laccaria bicolor is required for symbiosis development. Curr. Biol. 21, 1197 (2011).
Plett, J. M. et al. The effector MiSSP7 of the mutualistic fungus Laccaria bicolor stabilizes the Populus JAZ6 protein and represses JA-responsive genes. Proc. Natl Acad. Sci. USA 111, 8299 (2014). This study shows that the symbiotic effector MiSSP7 from the ectomycorrhizal fungus L. bicolor interacts with the jasmonate co-receptor in planta to enable the development of symbiosis.
Kazan, K. & Manners, J. M. JAZ repressors and the orchestration of phytohormone crosstalk. Trends Plant Sci. 17, 22–31 (2012).
Goossens, J., Fernandez-Calvo, P., Schweizer, F. & Goossens, A. Jasmonates: signal transduction components and their roles in environmental stress responses. Plant Mol. Biol. 91, 673–689 (2016).
Zuccaro, A. et al. Endophytic life strategies decoded by genome and transcriptome analyses of the mutualistic root symbiont Piriformospora indica. PLoS Pathog. 7, e1002290 (2011).
Lahrmann, U. et al. Mutualistic root endophytism is not associated with the reduction of saprotrophic traits and requires a noncompromised plant innate immunity. New Phytol. 207, 841–857. This works demonstrate the importance of indole-carboxylic acid derivatives as potential key players in the maintenance of a mutualistic interaction with root endophytes.
Fesel, P. H. & Zuccaro, A. Dissecting endophytic lifestyle along the parasitism/mutualism continuum in Arabidopsis. Curr. Opin. Microbiol. 32, 103–112 (2016).
Kloppholz, S., Kuhn, H. & Requena, N. A secreted fungal effector of Glomus intraradices promotes symbiotic biotrophy. Curr. Biol. 21, 1204–1209 (2011). This study characterizes the symbiotic secreted effector SP7 from the arbuscular mycorrhizal fungus G. intraradices (now classified as R. irregularis ) and shows that it promotes the colonization of the host plant.
Garcia, K., Delaux, P. M., Cope, K. R. & Ané, J. M. Molecular signals required for the establishment and maintenance of ectomycorrhizal symbiosis. New Phytol. 208, 79–87 (2015).
Weßling, R. et al. Convergent targeting of a common host protein-network by pathogen effectors from three kingdoms of life. Cell Host Microbe 16, 364–375 (2014).
Lo Presti, L. et al. Fungal effectors and plant susceptibility. Annu. Rev. Plant Biol. 66, 513–545 (2015).
Plett, J. M. & Martin, F. Reconsidering mutualistic plant–fungal interactions through the lens of effector biology. Curr. Opin. Plant Biol. 26, 45–50 (2015).
Veneault-Fourrey, C. et al. Genomic and transcriptomic analysis of Laccaria bicolor CAZome reveals insights into polysaccharides remodelling during symbiosis establishment. Fungal Genet. Biol. 72, 168–181 (2014).
Mankel, A., Krause, K. & Kothe, E. Identification of a hydrophobin gene that is developmentally regulated in the ectomycorrhizal fungus Tricholoma terreum. Appl. Environ. Microbiol. 68, 1408–1413 (2002).
Tagu, D. et al. Immuno-localization of hydrophobin HYDPt-1 from the ectomycorrhizal basidiomycete Pisolithus tinctorius during colonization of Eucalyptus globulus roots. New Phytol. 149, 127–135 (2000).
Plett, J. M. et al. Phylogenetic, genomic organization and expression analysis of hydrophobin genes in the ectomycorrhizal basidiomycete Laccaria bicolor. Fungal Genet. Biol. 49, 199–209 (2012).
Pellegrin, C., Morin, E., Martin, F. & Veneault-Fourrey, C. Comparative analysis of secretomes from ectomycorrhizal fungi with an emphasis on small secreted proteins. Front. Microbiol. 6, 1278 (2015).
Lionetti, V. & Métraux, J.-P. Plant cell wall in pathogenesis, parasitism and symbiosis. Front. Plant Sci. 5, 612 (2014).
Cope, K. et al. Poplar as a model for dissecting early mycorrhizal signaling in woody perennials. 2016 International Society for Molecular Plant–Microbe Interactions Congress http://www.ismpmi.org/congress/2016/abstracts/pages/abstractdetail.aspx?LID=358 (2016).
Delaux, P. M., Séjalon-Delmas, N., Bécard, G. & Ané, J. M. Evolution of the plant–microbe symbiotic 'toolkit'. Trends Plant Sci. 18, 298–304 (2013).
Parniske, M. Arbuscular mycorrhiza: the mother of plant-root endosymbioses. Nat. Rev. Microbiol. 6, 763–775 (2008).
Kottke, I. & Oberwinkler, F. The cellular structure of the Hartig net: coenocytic & transfer cell-like organization. Nordic J. Bot. 7, 85–95 (1987).
Agerer, R. (ed) Descriptions of Ectomycorrhizae (Einhorn-Verlag, 1996–2012).
Krings, M., Taylor, T. N., Taylor, E. L., Dotzler, N. & Walker, C. Arbuscular mycorrhizal-like fungi in Carboniferous arborescent lycopsids. New Phytol. 191, 311–314 (2011).
LePage, B. A., Currah, R. S., Stockey, R. A. & Rothwell, G. W. Fossil ectomycorrhizae from the Middle Eocene. Am. J. Bot. 84, 410–412 (1997).
Wang, H. et al. Rosid radiation and the rapid rise of angiosperm-dominated forest. Proc. Natl Acad. Sci. USA 106, 3853–3858 (2009).
Rice, A. V. & Currah, R. S. Oidiodendron maius: saprobe in Sphagnum peat, mutualist in ericaceous roots. Soil Biol. 9, 227–246 (2006).
Sasaki-Sekimoto, Y. et al. Basic helix–loop–helix transcription factors jasmonate-associated MYC2-like1 (JAM1), JAM2, and JAM3 are negative regulators of jasmonate responses in Arabidopsis. Plant Physiol. 163, 291–304 (2013).
Xin, X. F. & He, S. Y. Pseudomonas syringae pv. tomato DC3000: a model pathogen for probing disease susceptibility and hormone signaling in plants. Annu. Rev. Phytopathol. 51, 473–498 (2013).
Gimenez-Ibanez, S. et al. The bacterial effector HopX1 targets JAZ transcriptional repressors to activate jasmonate signaling and promote infection in Arabidopsis. PLoS Biol. 12, e1001792 (2014).
Jiang, S. S. et al. Bacterial effector activates jasmonate signaling by directly targeting JAZ transcriptional repressors. PLoS Pathog. 9, e1003715 (2013).
Plett, J. M. & Martin, F. Poplar root exudates contain compounds that induce the expression of MiSSP7 in Laccaria bicolor. Plant Signal. Behav. 7, 12–15 (2012).
Hacquard, S. et al. Laser microdissection and microarray analysis of Tuber melanosporum ectomycorrhizas reveal functional heterogeneity between mantle and Hartig net compartments. Environ. Microbiol. 15, 1853–1869 (2013).
Garcia, K. & Zimmermann, S. D. The role of mycorrhizal associations in plant potassium nutrition. Front. Plant Sci. 5, 337 (2014).
Becquer, A. et al. From soil to plant, the journey of P through trophic relationships and ectomycorrhizal association. Front. Plant Sci. 5, 548 (2014).
López, M. F. et al. The sugar porter gene family of Laccaria bicolor: function in ectomycorrhizal symbiosis and soil-growing hyphae. New Phytol. 180, 365–378 (2008).
Xu, H. et al. Overexpression of Laccaria bicolor aquaporin JQ585595 alters root water transport properties in ectomycorrhizal white spruce (Picea glauca) seedlings. New Phytol. 205, 757–770 (2015).
Acknowledgements
Many of the findings discussed in this manuscript were obtained within the framework of the Mycorrhizal Genomics Initiative (MGI) consortium and the Oak Ridge National Laboratory Plant–Microbe Interfaces project. The authors thank I. Grigoriev, J. Tuskan, M. Doktycz, J. Plett, E. Morin, Y. Daguerre, C. Pellegrin, L. Nagy, D. Floudas, M. Peter and J. Labbé for exciting discussions and interactions in relation to this manuscript. The MGI is supported by the French National Institute for Agricultural Research (INRA), the US Department of Energy (DOE) Joint Genome Institute (JGI; Office of Science of the US Department of Energy under contract number DE-AC02-05CH11231), the Region Lorraine Research Council and the European Commission (European Regional Development Fund (ERDF)). Research in the laboratory of F.M. is funded by the Laboratory of Excellence Advanced Research on the Biology of Tree and Forest Ecosystems (ARBRE; grant ANR-11-LABX-0002-01), the US DOE through the Oak Ridge National Laboratory Scientific Focus Area for Genomics Foundational Sciences (Plant Microbe Interfaces Project).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Glossary
- Rhizoid-based rooting systems
-
Simple hair-like protuberances that extend from the epidermal cells of certain plants. Rhizoids are similar in structure and function to the root hairs of vascular land plants.
- Basidiomycetes
-
(Formally known as Basidiomycota). A division or phylum within the kingdom Fungi that, together with the ascomycetes (formally known as Ascomycota), constitute the subkingdom Dikarya (often referred to as 'higher fungi'). Basidiomycetes reproduce sexually through the formation of specialized club-shaped end cells, known as basidia, that contain meiospores.
- Ascomycetes
-
(Formally known as Ascomycota). A division or phylum in the kingdom Fungi that, together with the basidiomycetes, form the subkingdom Dikarya. Members of the Ascomycota are commonly known as the sac fungi. The defining feature of ascomycetes is the ascus, a microscopic sexual structure in which meiospores, known as ascospores, are formed.
- Glomeromycetes
-
(Formally known as Glomeromycota). One of the seven currently recognized phyla in the kingdom Fungi. The 230 recognized species are all obligate symbionts of land plants that form arbuscular mycorrhizal associations.
- Mycorrhizal networks
-
Underground networks of hyphae that are produced by mycorrhizal fungi. Mycorrhizal networks connect individual plants together and transfer water, carbon and other nutrients.
- Saprotrophic fungi
-
Fungi that obtain their nutrition from non-living organic material.
- Rhizosphere
-
The soil that surrounds and is influenced by the roots of a plant.
- Protocorms
-
Tuber-shaped bodies with trichomes that are produced by the young seedlings of various orchids and other plants that have associated mycorrhizal fungi.
- Apoplastic space
-
In plants, the apoplastic space, or apoplast, is formed by the continuum of cell walls of adjacent cells as well as the extracellular space. It is the space outside of the plasma membrane.
- Rhizodermis
-
The epidermis that is formed by the outermost layer of primary cells in the plant root.
- Root cortex
-
The outermost layer of the plant root, which is bound on the outside by the epidermis (or rhizodermis) and on the inside by the endodermis. The root cortex is usually composed of large thin-walled parenchyma cells.
- Laccases
-
Enzymes that carry out a one-electron oxidation on phenols and similar molecules. Laccases are part of a larger group of enzymes that are termed the multicopper enzymes and that occur widely in fungi (but are also found in many plants and bacteria).
- White-rot fungi
-
Fungi that decay wood by breaking down lignin and cellulose.
- Auriculariales
-
An order of the kingdom Fungi in the class Agaricomycetes. Species in the Auriculariales often differentiate gelatinous fruit bodies and are thus commonly named 'jelly fungi'.
- Brown-rot fungi
-
Fungi that decay wood by breaking down hemicellulose and cellulose, leaving the lignin behind.
- Endophyte
-
A bacterium or fungus that lives inside a plant host without causing apparent disease.
- Agaricomycetidae
-
A subclass of fungi in the phylum Basidiomycota.
- Sesquiterpenes
-
A class of volatile hydrocarbons that consist of three isoprene units.
- Middle lamella
-
In plants, the middle lamella is formed by a pectin layer that cements the cell walls of adjoining cells together.
Rights and permissions
About this article
Cite this article
Martin, F., Kohler, A., Murat, C. et al. Unearthing the roots of ectomycorrhizal symbioses. Nat Rev Microbiol 14, 760–773 (2016). https://doi.org/10.1038/nrmicro.2016.149
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrmicro.2016.149
This article is cited by
-
Emerging insights into nitrogen assimilation in gymnosperms
Trees (2024)
-
Mass Production of Arbuscular Mycorrhizal Fungi on the Sorghum Plants Inoculated with Burkholderia arboris Using Soybean Mill Waste and Vermicompost-Amended Soil–Sand Substrate
Current Microbiology (2024)
-
Tropical tree ectomycorrhiza are distributed independently of soil nutrients
Nature Ecology & Evolution (2024)
-
Underutilized wild edible fungi and their undervalued ecosystem services in Africa
CABI Agriculture and Bioscience (2023)
-
Mycorrhizal C/N ratio determines plant-derived carbon and nitrogen allocation to symbiosis
Communications Biology (2023)