Key Points
-
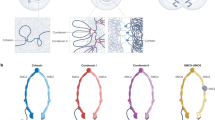
The six highly conserved structural maintenance of chromosomes (SMC) proteins form three types of heterodimer (SMC1–SMC3, SMC2–SMC4, SMC5–SMC6), which are core components of large multiprotein complexes. The best known complexes are cohesin, which is responsible for sister-chromatid cohesion, and condensin, which is required for full chromosome condensation in mitosis. Two variants of cohesin exist that differ in a non-SMC subunit.
-
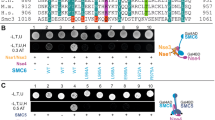
SMC proteins share five conserved domains. The amino-terminal domain contains a Walker A box, an NTP-binding motif, and, at the carboxyl terminus is a DA box with a Walker B-like sequence, and the LSGG signature motif that is typical for the ABC ATPase family of proteins. The central hinge domain, which is flanked by extended coiled–coil regions, is characterized by a set of four highly conserved glycine residues that are typical of flexible regions in a protein.
-
Dimerization of SMC monomers seems to be a function of the hinge domains. Protein interaction and microscopy data suggest that SMC dimers form a ring-like structure, in which the individual SMC protein folds back onto itself, allowing interaction between its amino and carboxyl termini. The ring might embrace DNA molecules. The non-SMC subunits associate with the SMC amino- and carboxy-terminal domains, giving the entire molecule a tadpole-like appearance.
-
Condesin gradually replaces cohesin in mitosis. Resolution of sister-chromatid cohesion requires cleavage of securin (Pds1) to activate separase (Esp1), which cleaves the phosphorylated Scc1 non-SMC subunit of cohesin, triggering its dissociation from the chromosomes.
-
Condensin introduces positive supercoils into relaxed circular DNA, and this is dependent on ATP and topoisomerase I. Knotted DNA is produced if toposiomerase II is present. The knotting activity is stimulated by cyclinB/cdc2 kinase phosphorylation of a non-SMC subunit of condensin.
-
Besides acting in sister-chromatid cohesion and in chromosome condensation, SMC proteins also function in DNA recombination and repair, and — in Caenorhabditis elegans — in gene dosage compensation.
-
SMC proteins and their complexes have also been found in meiotic cells and seem to act in meiotic sister-chromatid cohesion, condensation, and probably in DNA recombination. A meiosis-specific isoform of SMC1 has been described that largely replaces the canonical SMC1. Similarly, there are meiosis-specific variants of non-SMC subunits — for example the Rec8 protein.
Abstract
Members of the structural maintenance of chromosomes (SMC) family share a characteristic design and configuration of protein domains that provides the molecular basis for the various functions of this family in chromosome dynamics. SMC proteins have a role in chromosome condensation, sister-chromatid cohesion, DNA repair and recombination, and gene dosage compensation, and they function in somatic and meiotic cells. As more is learned about how their unique design affects their function, a picture of a dynamic and varied protein family is emerging.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Peterson, C. L. The SMC family: novel motor proteins for chromosome condensation? Cell 79, 389–392 (1994).
Gasser, S. M. Coiling up chromosomes. Curr. Biol. 5, 257–360 (1995).
Hirano, T., Mitchison, T. J. & Swedlow, J. R. The SMC family: from chromosome condensation to dosage compensation. Curr. Biol. 7, 329–336 (1995).
Strunnikov, A. V. & Jessberger, R. Structural maintenance of chromosomes (SMC) proteins: conserved molecular properties for multiple biological functions. Eur. J. Biochem. 263, 6–13 (1998).
Cobbe, N. & Heck, M. M. S. Review: SMCs in the world of chromosome biology — from prokaryotes to higher eukaryotes. J. Struct. Biol. 129, 123–143 (2000).
Nasmyth, K. Disseminating the genome: joining, resolving, and separating sister chromatids during mitosis and meiosis. Annu. Rev. Genet. 35, 673–745 (2001).
Hirano, T. The ABCs of SMC proteins: two-armed ATPases for chromosome condensation, cohesion, and repair. Genes Dev. 16, 399–414 (2002).
Soppa, J. Prokaryotic structural maintenance of chromosomes (SMC) proteins: distribution, phylogeny, and comparison with MukBs and additional prokaryotic and eukaryotic coiled-coil proteins. Gene 278, 253–264 (2001).
Strunnikov, A. V., Larionov, V. L. & Koshland D. SMC1: an essential yeast gene encoding a putative head-rod-tail protein is required for nuclear division and defines a new ubiquitous protein family. J. Cell Biol. 123, 1635–1648 (1993).The first definition of the SMC family and description of SMC1.
Lehmann, A. R. et al. The rad18 gene of Schizosaccharomyces pombe defines a new subgroup of the SMC superfamily involved in DNA repair. Mol. Cell. Biol. 15, 7067–7080 (1995).Identification of the SMC5–SMC6 subgroup and their role in DNA repair.
Jessberger, R., Frei, C. & Gasser, S. M. Chromosome dynamics: The SMC protein family. Curr. Opin. Genet. Dev. 8, 254–259 (1998).
Jones, S. & Sgouros, J. The cohesin complex: sequence homologies, interaction networks and shared motifs. Genome Biol. 2, 1–12 (2001). | Article |
Hagstrom, K. A., Holmes, V. F., Cozzarelli, N. R. & Meyer, B. J. C. elegans condensin promotes mitotic chromosome architecture, centromere organization, and sister chromatid segregation during mitosis and meiosis. Genes Dev. 16, 729–742 (2002).C. elegans has two SMC4 genes and specialized condensin-type complexes.
Revenkova, E., Eijpe, M., Heyting, C., Gross, B. & Jessberger, R. Novel meiosis-specific isoform of mammalian SMC1. Mol. Cell. Biol. 21, 6984–6998 (2001).The first cell-type- and meiosis-specific isoform of an SMC protein.
Holland, I. B. & Blight, M. A. ABC-ATPases, adaptable energy generators fuelling transmembrane movement of a variety of molecules in organisms from bacteria to humans. J. Mol. Biol. 293, 381–399 (1999).
Hirano, M., Anderson, D. E., Erickson, H. P. & Hirano, T. Bimodal activation of SMC ATPase by intra- and inter-molecular interactions. EMBO J. 20, 3238–3250 (2001).
Melby, T. E., Ciampaglio, C. N., Briscoe, G. & Erickson, H. P. The symmetrical structure of structural maintenance of chromosomes (SMC) and MukB proteins: long, antiparallel coiled coils, folded at a flexible hinge. J. Cell. Biol. 142, 1595–1604 (1998).The demonstration of hinge flexibility and antiparallelity of the SMC dimer.
Akhmedov, A. T. et al. Structural maintenance of chromosomes protein C-terminal domains bind preferentially to DNA with secondary structure. J. Biol. Chem. 273, 24088–24094 (1998).
Akhmedov, A. T., Gross, B. & Jessberger, R. Mammalian SMC3 C-terminal and coiled-coil protein domains specifically bind palindromic DNA, do not block DNA ends, and prevent DNA bending. J. Biol. Chem. 274, 38216–38224 (1999).
Blat, Y. & Kleckner, N. Cohesins bind to preferential sites along yeast chromosome III, with differential regulation along arm versus the centric region. Cell 98, 249–259 (1999).
Megee, P. C., Mistrot, C., Guacci, V. & Koshland, D. The centromeric sister chromatid cohesion site directs Mcd1p binding to adjacent sequences. Mol. Cell 4, 445–450 (1999).
Laloraya, S., Guacci, V. & Koshland, D. Chromosomal addresses of the cohesin component Mcd1p. J. Cell Biol. 151, 1047–1056 (2000).
Sutani, T. & Yanagida, M. DNA renaturation activity of the SMC complex implicated in chromosome condensation. Nature 388, 798–801 (1997).
Jessberger, R., Riwar, B., Baechtold, H. & Akhmedov, A. T. SMC proteins constitute two subunits of the mammalian recombination protein complex RC-1. EMBO J. 15, 4061–4068 (1996).The identification of an SMC1–SMC3 complex with recombination activity, indicating a role in the recombinational repair of DSBs.
van den Ent, F., Lockhart, A., Kendrick-Jones, J. & Lowe, J. Crystal structure of the N-terminal domain of MukB: a protein involved in chromosome partitioning. Structure Fold. Des. 15, 1181–1187 (1999).
Löwe, J., Cordell, S. C. & van den Ent, F. Crystal structure of the SMC head domain: an ABC ATPase with 900 residues antiparallel coiled-coil inserted. J. Mol. Biol. 306, 25–35 (2001).
Hopfner, K. P. et al. Structural biology of Rad50 ATPase: ATP-driven conformational control in DNA double-strand break repair and the ABC-ATPase superfamily. Cell 101, 789–800 (2000).
Haering, C. H., Löwe, J., Hochwagen, A. & Nasmyth, K. Molecular architecture of SMC proteins and the yeast cohesion complex. Mol. Cell 9, 773–788 (2002).Evidence for SMC dimerization through the hinge domain and for an 'embrace model' of cohesin's association with DNA.
Anderson, D. E., Losada, A., Erickson, H. P. & Hirano, T. Condensin and cohesin display different arm conformations with characteristic hinge angles. J. Cell Biol. 156, 419–424 (2002).
Beaseley, M., Xu, H., Warren, W. & McKay, M. Conserved disruptions in the predicted coiled-coil domains of eukaryotic SMC complexes: implications for structure and function Genome Res. 12, 1201–1209 (2002).
Yoshimura, S. H. et al. Condensin architecture and interaction with DNA. Regulatory non-SMC subunits bind to the head of SMC heterodimer. Curr Biol. 12, 508–513 (2002).
Guacci, V., Koshland, D. & Strunnikov, A. A direct link between sister chromatid cohesion and chromosome condensation revealed through the analysis of MCD1 in S. cerevisiae. Cell 91, 47–58 (1997).Mcd1–Scc1, a component of cohesin, is essential for sister-chromatid cohesion.
Michaelis, C., Ciosk, R. & Nasmyth, K. Cohesins: chromosomal proteins that prevent premature separation of sister chromatids. Cell 91, 35–46 (1997).Cohesin, Scc1 and Smc1 are essential for sister-chromatid cohesion.
Losada, A., Hirano, M. & Hirano, T. Identification of Xenopus SMC protein complexes required for sister chromatid cohesion. Genes Dev. 12, 1986–1997 (1998).The identification of vertebrate cohesin and demonstration of its essential function.
Hirano, T. & Mitchison, T. J. A heterodimeric coiled-coil protein required for mitotic chromosome condensation in vitro. Cell 79, 449–458 (1994).The first demonstration of SMC proteins required for chromosome condensation.
Hirano, T., Kobayashi, R. & Hirano, M. Condensins, chromosome condensation protein complexes containing XCAP-C, XCAP-E and a Xenopus homolog of the Drosophila barren protein. Cell 89, 511–521 (1997).The identification and characterization of the condensin holocomplex.
Fousteri, M. I. & Lehmann, A. R. A novel SMC protein complex in Schizosaccharomyces pombe contains the Rad18 DNA repair protein. EMBO J. 19, 1691–1702 (2000).
Jessberger, R., Podust, V., Hübscher, U. & Berg, P. A mammalian protein complex that repairs double-strand breaks and deletions by recombination. J. Biol. Chem. 268, 15070–15079 (1993).
Lieb, J. D., Albrecht, M. R., Chuang, P. T. & Meyer, B. J. MIX-1: an essential component of the C. elegans mitotic machinery executes X chromosome dosage compensation. Cell 92, 265–277 (1998).
Losada, A., Yokochi, T., Kobayashi, R. & Hirano, T. Identification and characterization of SA/Scc3p subunits in the Xenopus and human cohesin complexes. J. Cell Biol. 150, 405–416 (2000).
Sumara, I., Vorlaufer, E., Gieffers, C., Peters, B. H. & Peters, J. M. Characterization of vertebrate cohesin complexes and their regulation in prophase. J. Cell Biol. 151, 749–762 (2000).
Toth, A. et al. Yeast cohesin complex requires a conserved protein Eco1p(Ctf7), to establish cohesion between sister chromatids during DNA replication. Genes Dev. 13, 320–333 (1999).
Tomonaga, T. et al. Characterization of fission yeast cohesin: essential anaphase proteolysis of Rad21 phosphorylated in the S phase. Genes Dev. 14, 2757–2770 (2000).
Carson, D. R. & Christman, M. F. Evidence that replication fork components catalyze establishment of cohesion between sister chromatids. Proc. Natl Acad. Sci. USA 98, 8270–8275 (2001).
Tanaka, T., Cosma, M. P., Wirth, K. & Nasmyth, K. Identification of cohesin association sites at centromeres and along chromosome arms. Cell 98, 847–858 (1999).
Donze, D., Adams, C. R., Rine, J. & Kamakaka, R. T. The boundaries of the silenced HMR domain in Saccharomyces cerevisiae. Genes Dev. 13, 698–708 (1999).
Uhlmann, F. & Nasmyth, K. Cohesion between sister chromatids must be established during DNA replication. Curr. Biol. 8, 1095–1101 (1998).
Ciosk, R. et al. Cohesin's binding to chromosomes depends on a separate complex consisting of Scc2 and Scc4 proteins. Mol. Cell 5, 243–254 (2000).
Uhlmann, F. Secured cutting: controlling separase at the metaphase to anaphase transition. EMBO Rep. 2, 487–492 (2001).
Waizenegger, I. C., Hauf, S., Meinke, A. & Peters, J. M. Two distinct pathways remove mammalian cohesin from chromosome arms in prophase and from centromeres in anaphase. Cell 103, 399–410 (2000).
Uhlmann, F., Wernic, D., Poupart, M. A., Koonin, E. V. & Nasmyth, K. Cleavage of cohesin by the CD clan protease separin triggers anaphase in yeast. Cell 103, 375–386 (2000).
Alexandru, G., Uhlmann, F., Mechtler, K., Poupart, M. A. & Nasmyth, K. Phosphorylation of the cohesin subunit Scc1 by Polo/Cdc5 kinase regulates sister chromatid separation in yeast. Cell 105, 459–472 (2001).
Sumara, I. et al. The dissociation of cohesin from chromosomes in prophase is regulated by Polo-like kinase. Mol. Cell 9, 515–525 (2002).
Losada, A. & Hirano, T. Shaping the metaphase chromosome: coordination of cohesion and condensation. Bioessays 23, 924–935 (2001).
Wang, Z., Castano, I. B., De Las Penas, A., Adams, C. & Christman, M. F. Pol kappa: a DNA polymerase required for sister chromatid cohesion. Science 289, 774–779 (2000).
Schmiesing, J. A., Gregson, H. C., Zhou, S. & Yokomori, K. A human condensin complex containing hCAP-C-hCAP-E and CNAP1, a homolog of Xenopus XCAP-D2, colocalizes with phosphorylated histone H3 during the early stage of mitotic chromosome condensation. Mol. Cell. Biol. 20, 6996–6700 (2000).
Saitoh, N., Goldberg, I. G., Wood, E. R. & Earnshaw, W. C. ScII: an abundant chromosome scaffold protein is a member of a family of putative ATPases with an unusual predicted tertiary structure. J. Cell Biol. 127, 303–318 (1994).The first description of a vertebrate SMC protein.
Freeman, L., Aragon-Alcaide, L. & Strunnikov, A. The condensin complex governs chromosome condensation and mitotic transmission of rDNA. J. Cell Biol. 149, 811–824 (2000).
Kimura, K., Cuvier, O. & Hirano, T. Chromosome condensation by a human condensin complex in Xenopus egg extracts. J. Biol. Chem. 276, 5417–5420 (2001).
Saka, Y. et al. Fission yeast cut3 and cut14, members of a ubiquitous protein family, are required for chromosome condensation and segregation in mitosis. EMBO J. 13, 4938–4952 (1994).The first documentation of SMC2–SMC4-type proteins acting in condensation.
Strunnikov, A. V., Hogan, E. & Koshland, D. SMC2, a Saccharomyces cerevisiae gene essential for chromosome segregation and condensation defines a subgroup within the SMC family. Genes Dev. 9, 587–599 (1995).
Bhat, M. A., Philp, A. V., Glover, D. M. & Bellen, H. J. Chromatid segregation at anaphase requires the barren product, a novel chromosome-associated protein that interacts with topoisomerase II. Cell 87, 1103–1114 (1996).
Britton, R. A., Lin, D. C. & Grossman, A. D. Characterization of a prokaryotic SMC protein involved in chromosome partitioning. Genes Dev. 12, 1254–1259 (1998).
Sutani, T. et al. Fission yeast condensin complex: essential roles of non-SMC subunits for condensation and Cdc2 phosphorylation of Cut3/SMC4. Genes Dev. 13, 2271–2283 (1999).
Lavoie, B. D., Hogan, E. & Koshland, D. In vivo dissection of the chromosome condensation machinery: reversibility of condensation distinguishes contributions of condensin and cohesin. J. Cell Biol. 156, 805–815 (2002).
Biggins, S., Bhalla, N., Chang, A., Smith, D. L. & Murray, A. W. Genes involved in sister chromatid separation and segregation in the budding yeast Saccharomyces cerevisiae. Genetics 159, 453–470 (2001).
Steffensen, S. et al. A role for Drosophila SMC4 in the resolution of sister chromatids in mitosis. Curr. Biol. 11, 295–307 (2001).
Kimura, K. & Hirano, T. ATP-dependent positive supercoiling of DNA by 13S condensin: a biochemical implication for chromosome condensation. Cell 90, 625–634 (1997).
Kimura, K. & Hirano, T. Dual roles of the 11S regulatory subcomplex in condensin functions. Proc. Natl Acad. Sci. USA 97, 11972–11977 (2000).
Kimura, K., Rybenkov, V. V., Crisona, N. J., Hirano, T. & Cozzarelli, N. R. 13S condesin actively reconfigures DNA by introducing global positive writhe: implications for chromosome condensation. Cell 98, 239–248 (1999).
Bazett-Jones, D. P., Kimura, K. & Hirano, T. Efficient supercoiling of DNA by a single condensin complex as revealed by electron spectroscopic imaging. Mol. Cell 9, 1183–1190 (2002).
Kimura, K., Hirano, M., Kobayashi, R. & Hirano, T. Phosphorylation and activation of 13S condensin by Cdc2 in vitro. Science 282, 487–490 (1998).
Sutani, T. et al. Fission yeast condensin complex: essential roles of non-SMC subunits for condensation and Cdc2 phosphorylation of Cut3/SMC4. Genes Dev. 13, 2271–2283 (1999).
Giet, R. & Glover, D. M. Drosophila aurora B kinase is required for histone H3 phosphorylation and condensin recruitment during chromosome condensation and to organize the central spindle during cytokinesis. J. Cell Biol. 152, 669–682 (2001).
Hirano, T. Chromosome cohesion, condensation, and separation. Annu. Rev. Biochem. 69, 115–144 (2000).
Castano, I. B., Brzoska, P. M., Sadoff, B. U., Chen, H. & Christman, M. F. Mitotic chromosome condensation in the rDNA requires TRF4 and DNA topoisomerase I in Saccharomyces cerevisiae. Genes Dev. 10, 2564–2576 (1996).
Hartman, T., Stead, K., Koshland, D. & Guacci, V. Pds5p is an essential chromosomal protein required for both sister chromatid cohesion and condensation in Saccharomyces cerevisiae. J. Cell Biol. 151, 613–626 (2000).
Sonoda, E. et al. Scc1/Rad21/Mcd1 is required for sister chromatid cohesion and kinetochore function in vertebrate cells. Dev. Cell. 1, 759–770 (2001).
Chuang, P. T., Albertson, D. G. & Meyer, B. J. DPY-27: a chromosome condensation protein homolog that regulates C. elegans dosage compensation through association with the X chromosome. Cell 79, 459–474 (1994).An SMC4-like protein in C. elegans acts in gene-dosage compensation, a new function for SMC proteins.
Chuang, P. T., Lieb, J. D. & Meyer, B. J. Sex-specific assembly of a dosage compensation complex on the nematode X chromosome. Science 274, 1736–1738 (1996).
Lieb, J. D., Capowski, E. E., Meneely, P. & Meyer, B. J. DPY-26, a link between dosage compensation and meiotic chromosome segregation in the nematode. Science 274, 1732–1736 (1996).
Davis, T. L. & Meyer, B. J. SDC-3 coordinates the assembly of a dosage compensation complex on the nematode X chromosome. Development 124, 1019–1031 (1997).
Jessberger, R., Chui, G., Linn, S. & Kemper, B. Analysis of the mammalian recombination protein complex RC-1. Mutat. Res. 350, 217–222 (1996).
Hübscher, U., Nasheuer, H.-P. & Syväoja, J. Eukaryotic DNA polymerases, a growing family. Trends Biochem. Sci. 25, 143–147 (2000).
Morishita, T., Tsutsui, Y., Iwasaki, H. & Shinagawa, H. The Schizosaccharomyces pombe rad60 gene is essential for repairing double-strand DNA breaks spontaneously occurring during replication and induced by DNA-damaging agents. Mol. Cell. Biol. 22, 3537–3548 (2002).
Fujioka, Y., Kimata, Y., Nomaguchi, K., Watanabe, K. & Kohno, K. Identification of a novel non-SMC component of the SMC5/SMC6 complex involved in DNA repair. J. Biol. Chem. 377, 21585–21591 (2002).
Taylor, E. M. et al. Characterization of a novel human SMC heterodimer homologous to the Schizosaccharomyces pombe Rad18/Spr18 complex. Mol. Biol. Cell 12, 1583–1594 (2001).
Mengiste, T., Revenkova, E., Bechtold, N. & Paszkowski, J. An SMC-like protein is required for efficient homologous recombination in Arabidopsis. EMBO J. 18, 4505–4512 (1999).
Hanin, M., Mengiste, T., Bogucki, A. & Paszkowski, J. Elevated levels of intrachromosomal homologous recombination in Arabidopsis overexpressing the MIM gene. Plant J. 24, 183–189 (2000).
Sjøgren, C. & Nasmyth, K. Sister chromatid cohesion is required for postreplicative double-strand break repair in Saccharomyces cerevisiae. Curr. Biol. 11, 991–995 (2001).
Birkenbihl, R. P. & Subramani, S. Cloning and characterization of rad21 an essential gene of Schizosaccharomyces pombe involved in DNA double-strand-break repair. Nucl. Acids Res. 20, 6605–6611 (1992).
Johnson, R. D. & Jasin, M. Sister chromatid gene conversion is a prominent double-strand break repair pathway in mammalian cells. EMBO J. 19, 3398–3407 (2000).
Kim, S. T., Xu, B. & Kastan, M. B. Involvement of the cohesin protein, Smc1, in Atm-dependent and independent responses to DNA damage. Genes Dev. 16, 560–570 (2002).
Yazdi, P. T. et al. SMC1 is a downstream effector in the ATM/NBS1 branch of the human S-phase checkpoint. Genes Dev. 16, 571–582 (2002).
Shiloh, Y. ATM and ATR: networking cellular responses to DNA damage. Curr. Opin. Genet. Dev. 11, 71–77 (2001).
Klein, F. et al. A central role for cohesins in sister chromatid cohesion, formation of axial elements, and recombination during yeast meiosis. Cell 98, 91–103 (1999).SMC3 is required for meiotic sister-chromatid cohesion and reciprocal recombination.
Eijpe, M., Heyting, C., Gross, B. & Jessberger, R. Association of mammalian SMC1 and SMC3 proteins with meiotic chromosomes and synaptonemal complexes. J. Cell Sci. 113, 673–682 (2000).
Pelttari, J. et al. A meiotic chromosomal core consisting of cohesin complex proteins recruits DNA recombination proteins and promotes synapsis in the absence of an axial element in mammalian meiotic cells. Mol Cell Biol. 21, 5667–5677 (2001).
Prieto, I. et al. Mammalian STAG3 is a cohesin specific to sister chromatid arms in meiosis I. Nature Cell Biol. 3, 761–766 (2001).
Watanabe, Y. & Nurse, P. Cohesin Rec8 is required for reductional chromosome segregation at meiosis. Nature 400, 461–464 (1999).
Watanbe, Y., Yokobayashi, S., Yamamoto, M. & Nurse, P. Pre-meiotic S phase is linked to reductional chromosome segregation and recombination. Nature 409, 359–363 (2001).A pre-meiotic role for a meiosis-specific component of cohesin, Rec8, that conveys meiotic behaviours to centromeres.
Pasierbek, P. et al. A Caenorhabditis elegans cohesion protein with functions in meiotic chromosome pairing and disjunction. Genes Dev. 15, 1349–1360 (2001).
Ghiselli, G. & Iozzo, R. V. Overexpression of bamacan/SMC3 causes transformation. J. Biol. Chem. 275, 20235–20238 (2000).
Hopfner, K. P. et al. Structural biochemistry and interaction architecture of the DNA double-strand break repair Mre11 nuclease and Rad50-ATPase. Cell 105, 473–485 (2001).
Hopfner, K. P., Putnam, C. D. & Tainer, J. DNA double-strand break repair from head to tail. Curr. Opin. Struct. Biol. 12, 115–122 (2002).
Hopfner, K. P. et al. The Rad50 zinc-hook is a structure joining Mre11 complexes in DNA recombination and repair. Nature 418, 562–566 (2002).
de Jager, M. et al. Human Rad50/Mre11 is a flexible complex that can tether DNA ends. Mol. Cell 8, 1129–1135 (2001).
Acknowledgements
I thank members of my laboratory for discussions, in particular E. Revenkova and A. Solovicova for comments on the manuscript. Research in the author's laboratory has been supported by grants from the HFSP and the NIH.
Author information
Authors and Affiliations
Related links
Related links
DATABASES
LocusLink
Saccharomyces genome database
Wormbase
FURTHER INFORMATION
Glossary
- WALKER A AND B MOTIFS
-
ATP-interaction motifs defined by Walker and colleagues in 1982. The A box contains a P-loop that is involved in phosphate binding; a B-like sequence is contained in the highly conserved DA box at the SMC carboxyl termini and could be involved in binding Mg2+. A and B sites can interact to form an ATP-binding and hydrolysis fold.
- CHROMATIN IMMUNOPRECIPITATION
-
(ChIP). A technique that isolates sequences from soluble DNA chromatin extracts (complexes of DNA and protein) using antibodies that recognize specific chromosomal proteins.
- CATENATION/DECATENATION
-
Topoisomerases catenate (join) or decatenate (separate) two circular DNA molecules by cutting one DNA strand, passing a second strand through the break and resealing the break in DNA.
- DNA POLYMERASE σ
-
One of a series of newly described DNA polymerases. Polσ (previously named Polσ) is required to establish cohesion in S phase. It is the product of the TRF4 gene, which has redundant homologues.
- RNA INTERFERENCE
-
The process by which an introduced double-stranded RNA specifically silences the expression of genes through degradation of their cognate mRNAs.
- DNA LIGASE III
-
One of three DNA ligases found in eukaryotic cells. Acts in repair of single-stranded DNA breaks and forms a complex with the repair gene XRCC1. There is a meiotic isoform, DNA ligase IIIβ.
- DNA POLYMERASE ɛ
-
A DNA polymerase that associates with the replication fork, acts in nucleotide-excision repair, possibly in recombinational repair, and might contribute an S-phase checkpoint function. In yeast the enzyme is not essential for many of these functions.
- NUCLEOTIDE-EXCISION REPAIR
-
The main pathway for removal of UV-damaged bases.
- SYNTHETIC INTERACTIONS
-
These occur when a double mutant has a phenotype different from either single mutant parent. For suppressors (synthetic viable), the double mutant is viable when at least one of the single mutants is not. For synthetic lethal mutants, the double mutant is inviable under conditions in which both parents are viable.
- ATM KINASE
-
The protein that is mutated in ataxia-telangiectasia, with major functions in signal transduction in the response to DNA damage.
- AXIAL ELEMENTS
-
Two axial elements form the side elements of the ladder-like synaptonemal complex. They are connected by numerous transverse filaments along their length. Each axial element supports the two sister chromatids of one homologue.
- SYNAPTONEMAL COMPLEX
-
A characteristic, zipper- or ladder-like protein structure, formed during meiotic prophase between paired homologous chromosomes.
Rights and permissions
About this article
Cite this article
Jessberger, R. The many functions of smc proteins in chromosome dynamics. Nat Rev Mol Cell Biol 3, 767–778 (2002). https://doi.org/10.1038/nrm930
Issue Date:
DOI: https://doi.org/10.1038/nrm930
This article is cited by
-
Protein phosphatase 2A interacts with Verthandi/Rad21 to regulate mitosis and organ development in Drosophila
Scientific Reports (2019)
-
Not just gene expression: 3D implications of chromatin modifications during sexual plant reproduction
Plant Cell Reports (2018)
-
NCAPH plays important roles in human colon cancer
Cell Death & Disease (2017)
-
Can corruption of chromosome cohesion create a conduit to cancer?
Nature Reviews Cancer (2011)
-
Chromatin configuration and epigenetic landscape at the sex chromosome bivalent during equine spermatogenesis
Chromosoma (2011)