Key Points
-
MicroRNAs (miRNAs) are small non-coding RNAs that function as guide molecules in RNA silencing.
-
Biogenesis of miRNA is under tight temporal and spatial control.
-
Dysregulation of miRNA is associated with many human diseases, particularly cancer and neurodevelopmental disorders.
-
Regulation takes place at multiple levels including transcription, Drosha processing, Dicer processing, RNA editing, RNA methylation, uridylation, adenylation, Argonaute modification and RNA decay.
-
This Review summarizes our current understanding of how miRNAs are made and regulated, with a focus on animal systems.
Abstract
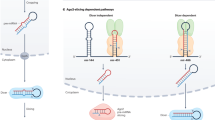
MicroRNAs (miRNAs) are small non-coding RNAs that function as guide molecules in RNA silencing. Targeting most protein-coding transcripts, miRNAs are involved in nearly all developmental and pathological processes in animals. The biogenesis of miRNAs is under tight temporal and spatial control, and their dysregulation is associated with many human diseases, particularly cancer. In animals, miRNAs are ∼22 nucleotides in length, and they are produced by two RNase III proteins — Drosha and Dicer. miRNA biogenesis is regulated at multiple levels, including at the level of miRNA transcription; its processing by Drosha and Dicer in the nucleus and cytoplasm, respectively; its modification by RNA editing, RNA methylation, uridylation and adenylation; Argonaute loading; and RNA decay. Non-canonical pathways for miRNA biogenesis, including those that are independent of Drosha or Dicer, are also emerging.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Ghildiyal, M. & Zamore, P. D. Small silencing RNAs: an expanding universe. Nature Rev. Genet. 10, 94–108 (2009).
Ishizu, H., Siomi, H. & Siomi, M. C. Biology of PIWI-interacting RNAs: new insights into biogenesis and function inside and outside of germlines. Genes Dev. 26, 2361–2373 (2012).
Huntzinger, E. & Izaurralde, E. Gene silencing by microRNAs: contributions of translational repression and mRNA decay. Nature Rev. Genet. 12, 99–110 (2011).
Bartel, D. P. MicroRNAs: target recognition and regulatory functions. Cell 136, 215–233 (2009).
Friedman, R. C., Farh, K. K., Burge, C. B. & Bartel, D. P. Most mammalian mRNAs are conserved targets of microRNAs. Genome Res. 19, 92–105 (2009).
Lujambio, A. & Lowe, S. W. The microcosmos of cancer. Nature 482, 347–355 (2012).
Im, H. I. & Kenny, P. J. MicroRNAs in neuronal function and dysfunction. Trends Neurosci. 35, 325–334 (2012).
Kim, V. N., Han, J. & Siomi, M. C. Biogenesis of small RNAs in animals. Nature Rev. Mol. Cell Biol. 10, 126–139 (2009).
Krol, J., Loedige, I. & Filipowicz, W. The widespread regulation of microRNA biogenesis, function and decay. Nature Rev. Genet. 11, 597–610 (2010).
Siomi, H. & Siomi, M. C. Posttranscriptional regulation of microRNA biogenesis in animals. Mol. Cell 38, 323–332 (2010).
Axtell, M. J., Westholm, J. O. & Lai, E. C. Vive la difference: biogenesis and evolution of microRNAs in plants and animals. Genome Biol. 12, 221 (2011).
Cuperus, J. T., Fahlgren, N. & Carrington, J. C. Evolution and functional diversification of miRNA genes. Plant Cell 23, 431–442 (2011).
Voinnet, O. Origin, biogenesis, and activity of plant microRNAs. Cell 136, 669–687 (2009).
Chang, S. S., Zhang, Z. & Liu, Y. RNA interference pathways in fungi: mechanisms and functions. Annu. Rev. Microbiol. 66, 305–323 (2012).
Griffiths-Jones, S., Saini, H. K., van Dongen, S. & Enright, A. J. miRBase: tools for microRNA genomics. Nucleic Acids Res. 36, D154–D158 (2008).
Chiang, H. R. et al. Mammalian microRNAs: experimental evaluation of novel and previously annotated genes. Genes Dev. 24, 992–1009 (2010).
Kozomara, A. & Griffiths-Jones, S. miRBase: annotating high confidence microRNAs using deep sequencing data. Nucleic Acids Res 42, D68–D73 (2014).
Hertel, J. et al. The expansion of the metazoan microRNA repertoire. BMC Genomics 7, 25 (2006).
Berezikov, E. Evolution of microRNA diversity and regulation in animals. Nature Rev. Genet. 12, 846–860 (2011).
Wheeler, B. M. et al. The deep evolution of metazoan microRNAs. Evol. Dev. 11, 50–68 (2009).
Ventura, A. et al. Targeted deletion reveals essential and overlapping functions of the miR-17 through 92 family of miRNA clusters. Cell 132, 875–886 (2008).
Kim, Y. K. et al. TALEN-based knockout library for human microRNAs. Nature Struct. Mol. Biol. 20, 1458–1464 (2013).
Lee, Y., Jeon, K., Lee, J. T., Kim, S. & Kim, V. N. MicroRNA maturation: stepwise processing and subcellular localization. EMBO J. 21, 4663–4670 (2002).
Roush, S. & Slack, F. J. The let-7 family of microRNAs. Trends Cell Biol. 18, 505–516 (2008).
Ozsolak, F. et al. Chromatin structure analyses identify miRNA promoters. Genes Dev. 22, 3172–3183 (2008).
Monteys, A. M. et al. Structure and activity of putative intronic miRNA promoters. RNA 16, 495–505 (2010).
Cai, X., Hagedorn, C. H. & Cullen, B. R. Human microRNAs are processed from capped, polyadenylated transcripts that can also function as mRNAs. RNA 10, 1957–1966 (2004).
Lee, Y. et al. MicroRNA genes are transcribed by RNA polymerase II. EMBO J. 23, 4051–4060 (2004).
Pfeffer, S. et al. Identification of microRNAs of the herpesvirus family. Nature Methods 2, 269–276 (2005).
Babiarz, J. E., Ruby, J. G., Wang, Y., Bartel, D. P. & Blelloch, R. Mouse ES cells express endogenous shRNAs, siRNAs, and other Microprocessor-independent, Dicer-dependent small RNAs. Genes Dev. 22, 2773–2785 (2008).
Davis-Dusenbery, B. N. & Hata, A. Mechanisms of control of microRNA biogenesis. J. Biochem. 148, 381–392 (2010).
Lee, Y. et al. The nuclear RNase III Drosha initiates microRNA processing. Nature 425, 415–419 (2003).
Denli, A. M., Tops, B. B., Plasterk, R. H., Ketting, R. F. & Hannon, G. J. Processing of primary microRNAs by the Microprocessor complex. Nature 432, 231–235 (2004).
Gregory, R. I. et al. The Microprocessor complex mediates the genesis of microRNAs. Nature 432, 235–240 (2004).
Han, J. et al. The Drosha–DGCR8 complex in primary microRNA processing. Genes Dev. 18, 3016–3027 (2004).
Landthaler, M., Yalcin, A. & Tuschl, T. The human DiGeorge syndrome critical region gene 8 and its D. melanogaster homolog are required for miRNA biogenesis. Curr. Biol. 14, 2162–2167 (2004).
Chong, M. M. et al. Canonical and alternate functions of the microRNA biogenesis machinery. Genes Dev. 24, 1951–1960 (2010).
Wang, Y., Medvid, R., Melton, C., Jaenisch, R. & Blelloch, R. DGCR8 is essential for microRNA biogenesis and silencing of embryonic stem cell self-renewal. Nature Genet. 39, 380–385 (2007).
Shiohama, A., Sasaki, T., Noda, S., Minoshima, S. & Shimizu, N. Molecular cloning and expression analysis of a novel gene DGCR8 located in the DiGeorge syndrome chromosomal region. Biochem. Biophys. Res. Commun. 304, 184–190 (2003).
Goldberg, R., Motzkin, B., Marion, R., Scambler, P. J. & Shprintzen, R. J. Velo-cardio-facial syndrome: a review of 120 patients. Am. J. Med. Genet. 45, 313–319 (1993).
Filippov, V., Solovyev, V., Filippova, M. & Gill, S. S. A novel type of RNase III family proteins in eukaryotes. Gene 245, 213–221 (2000).
Fortin, K. R., Nicholson, R. H. & Nicholson, A. W. Mouse ribonuclease III. cDNA structure, expression analysis, and chromosomal location. BMC Genomics 3, 26 (2002).
Wu, H., Xu, H., Miraglia, L. J. & Crooke, S. T. Human RNase III is a 160-kDa protein involved in preribosomal RNA processing. J. Biol. Chem. 275, 36957–36965 (2000).
Tang, X., Zhang, Y., Tucker, L. & Ramratnam, B. Phosphorylation of the RNase III enzyme Drosha at Serine300 or Serine302 is required for its nuclear localization. Nucleic Acids Res. 38, 6610–6619 (2010).
Blaszczyk, J. et al. Crystallographic and modeling studies of RNase III suggest a mechanism for double-stranded RNA cleavage. Structure 9, 1225–1236 (2001).
Zhang, H., Kolb, F. A., Jaskiewicz, L., Westhof, E. & Filipowicz, W. Single processing center models for human Dicer and bacterial RNase III. Cell 118, 57–68 (2004).
Shiohama, A., Sasaki, T., Noda, S., Minoshima, S. & Shimizu, N. Nucleolar localization of DGCR8 and identification of eleven DGCR8-associated proteins. Exp. Cell Res. 313, 4196–4207 (2007).
Yeom, K. H., Lee, Y., Han, J., Suh, M. R. & Kim, V. N. Characterization of DGCR8/Pasha, the essential cofactor for Drosha in primary miRNA processing. Nucleic Acids Res. 34, 4622–4629 (2006).
Han, J. et al. Molecular basis for the recognition of primary microRNAs by the Drosha–DGCR8 complex. Cell 125, 887–901 (2006).
Sohn, S. Y. et al. Crystal structure of human DGCR8 core. Nature Struct. Mol. Biol. 14, 847–853 (2007).
Faller, M., Matsunaga, M., Yin, S., Loo, J. A. & Guo, F. Heme is involved in microRNA processing. Nature Struct. Mol. Biol. 14, 23–29 (2007).
Faller, M. et al. DGCR8 recognizes primary transcripts of microRNAs through highly cooperative binding and formation of higher-order structures. RNA 16, 1570–1583 (2010).
Barr, I. et al. Ferric, not ferrous, heme activates RNA-binding protein DGCR8 for primary microRNA processing. Proc. Natl Acad. Sci. USA 109, 1919–1924 (2012).
Weitz, S. H., Gong, M., Barr, I., Weiss, S. & Guo, F. Processing of microRNA primary transcripts requires heme in mammalian cells. Proc. Natl Acad. Sci. USA 111, 1861–1866 (2014).
Zeng, Y., Yi, R. & Cullen, B. R. Recognition and cleavage of primary microRNA precursors by the nuclear processing enzyme Drosha. EMBO J. 24, 138–148 (2005).
Auyeung, V. C., Ulitsky, I., McGeary, S. E. & Bartel, D. P. Beyond secondary structure: primary-sequence determinants license pri-miRNA hairpins for processing. Cell 152, 844–858 (2013).
Ma, H., Wu, Y., Choi, J. G. & Wu, H. Lower and upper stem-single-stranded RNA junctions together determine the Drosha cleavage site. Proc. Natl Acad. Sci. USA 110, 20687–20692 (2013).
Mori, M. et al. Hippo signaling regulates Microprocessor and links cell-density-dependent miRNA biogenesis to cancer. Cell 156, 893–906 (2014).
Kim, Y. K. & Kim, V. N. Processing of intronic microRNAs. EMBO J. 26, 775–783 (2007).
Morlando, M. et al. Primary microRNA transcripts are processed co-transcriptionally. Nature Struct. Mol. Biol. 15, 902–909 (2008).
Pawlicki, J. M. & Steitz, J. A. Primary microRNA transcript retention at sites of transcription leads to enhanced microRNA production. J. Cell Biol. 182, 61–76 (2008).
Han, J. et al. Posttranscriptional crossregulation between Drosha and DGCR8. Cell 136, 75–84 (2009).
Sundaram, G. M. et al. 'See-saw' expression of microRNA-198 and FSTL1 from a single transcript in wound healing. Nature 495, 103–106 (2013).
Knuckles, P. et al. Drosha regulates neurogenesis by controlling neurogenin 2 expression independent of microRNAs. Nature Neurosci. 15, 962–969 (2012).
Kadener, S. et al. Genome-wide identification of targets of the drosha-pasha/DGCR8 complex. RNA 15, 537–545 (2009).
Herbert, K. M., Pimienta, G., DeGregorio, S. J., Alexandrov, A. & Steitz, J. A. Phosphorylation of DGCR8 increases its intracellular stability and induces a progrowth miRNA profile. Cell Rep. 5, 1070–1081 (2013).
Tang, X., Li, M., Tucker, L. & Ramratnam, B. Glycogen synthase kinase 3 beta (GSK3β) phosphorylates the RNAase III enzyme Drosha at S300 and S302. PLoS ONE 6, e20391 (2011).
Wada, T., Kikuchi, J. & Furukawa, Y. Histone deacetylase 1 enhances microRNA processing via deacetylation of DGCR8. EMBO Rep. 13, 142–149 (2012).
Tang, X. et al. Acetylation of drosha on the N-terminus inhibits its degradation by ubiquitination. PLoS ONE 8, e72503 (2013).
Cheng, T. L. et al. MeCP2 suppresses nuclear microRNA processing and dendritic growth by regulating the DGCR8/Drosha complex. Dev. Cell 28, 547–560 (2014).
Fukuda, T. et al. DEAD-box RNA helicase subunits of the Drosha complex are required for processing of rRNA and a subset of microRNAs. Nature Cell Biol. 9, 604–611 (2007).
Davis, B. N., Hilyard, A. C., Lagna, G. & Hata, A. SMAD proteins control DROSHA-mediated microRNA maturation. Nature 454, 56–61 (2008).
Davis, B. N., Hilyard, A. C., Nguyen, P. H., Lagna, G. & Hata, A. Smad proteins bind a conserved RNA sequence to promote microRNA maturation by Drosha. Mol. Cell 39, 373–384 (2010).
Suzuki, H. I. et al. Modulation of microRNA processing by p53. Nature 460, 529–533 (2009).
Di Carlo, V. et al. TDP-43 regulates the microprocessor complex activity during in vitro neuronal differentiation. Mol. Neurobiol. 48, 952–963 (2013).
Kawahara, Y. & Mieda-Sato, A. TDP-43 promotes microRNA biogenesis as a component of the Drosha and Dicer complexes. Proc. Natl Acad. Sci. USA 109, 3347–3352 (2012).
Guil, S. & Caceres, J. F. The multifunctional RNA-binding protein hnRNP A1 is required for processing of miR-18a. Nature Struct. Mol. Biol. 14, 591–596 (2007).
Michlewski, G., Guil, S., Semple, C. A. & Caceres, J. F. Posttranscriptional regulation of miRNAs harboring conserved terminal loops. Mol. Cell 32, 383–393 (2008).
Trabucchi, M. et al. The RNA-binding protein KSRP promotes the biogenesis of a subset of microRNAs. Nature 459, 1010–1014 (2009).
Bohnsack, M. T. Exportin 5 is a RanGTP-dependent dsRNA-binding protein that mediates nuclear export of pre-miRNAs. RNA 10, 185–191 (2004).
Lund, E., Guttinger, S., Calado, A., Dahlberg, J. E. & Kutay, U. Nuclear export of microRNA precursors. Science 303, 95–98 (2004).
Yi, R., Qin, Y., Macara, I. G. & Cullen, B. R. Exportin-5 mediates the nuclear export of pre-microRNAs and short hairpin RNAs. Genes Dev. 17, 3011–3016 (2003).
Okada, C. et al. A high-resolution structure of the pre-microRNA nuclear export machinery. Science 326, 1275–1279 (2009).
Gwizdek, C. et al. Exportin-5 mediates nuclear export of minihelix-containing RNAs. J. Biol. Chem. 278, 5505–5508 (2003).
Zeng, Y. & Cullen, B. R. Structural requirements for pre-microRNA binding and nuclear export by Exportin 5. Nucleic Acids Res. 32, 4776–4785 (2004).
Iwasaki, Y. W. et al. Global microRNA elevation by inducible Exportin 5 regulates cell cycle entry. RNA 19, 490–497 (2013).
Wan, G. et al. DNA-damage-induced nuclear export of precursor microRNAs is regulated by the ATM–AKT pathway. Cell Rep. 3, 2100–2112 (2013).
Melo, S. A. et al. A genetic defect in exportin-5 traps precursor microRNAs in the nucleus of cancer cells. Cancer Cell 18, 303–315 (2010).
Xie, M. et al. Mammalian 5′-capped microRNA precursors that generate a single microRNA. Cell 155, 1568–1580 (2013).
Bernstein, E., Caudy, A. A., Hammond, S. M. & Hannon, G. J. Role for a bidentate ribonuclease in the initiation step of RNA interference. Nature 409, 363–366 (2001).
Grishok, A. et al. Genes and mechanisms related to RNA interference regulate expression of the small temporal RNAs that control C. elegans developmental timing. Cell 106, 23–34 (2001).
Hutvagner, G. et al. A cellular function for the RNA-interference enzyme Dicer in the maturation of the let-7 small temporal RNA. Science 293, 834–838 (2001).
Ketting, R. F. et al. Dicer functions in RNA interference and in synthesis of small RNA involved in developmental timing in C. elegans. Genes Dev. 15, 2654–2659 (2001).
Knight, S. W. & Bass, B. L. A role for the RNase III enzyme DCR-1 in RNA interference and germ line development in Caenorhabditis elegans. Science 293, 2269–2271 (2001).
Lee, Y. S. et al. Distinct roles for Drosophila Dicer-1 and Dicer-2 in the siRNA/miRNA silencing pathways. Cell 117, 69–81 (2004).
Xie, Z. et al. Genetic and functional diversification of small RNA pathways in plants. PLoS Biol. 2, E104 (2004).
Bernstein, E. et al. Dicer is essential for mouse development. Nature Genet. 35, 215–217 (2003).
Kanellopoulou, C. et al. Dicer-deficient mouse embryonic stem cells are defective in differentiation and centromeric silencing. Genes Dev. 19, 489–501 (2005).
Murchison, E. P., Partridge, J. F., Tam, O. H., Cheloufi, S. & Hannon, G. J. Characterization of Dicer-deficient murine embryonic stem cells. Proc. Natl Acad. Sci. USA 102, 12135–12140 (2005).
Yang, J. S. & Lai, E. C. Alternative miRNA biogenesis pathways and the interpretation of core miRNA pathway mutants. Mol. Cell 43, 892–903 (2011).
Kumar, M. S. et al. Dicer1 functions as a haploinsufficient tumor suppressor. Genes Dev. 23, 2700–2704 (2009).
Tsutsumi, A., Kawamata, T., Izumi, N., Seitz, H. & Tomari, Y. Recognition of the pre-miRNA structure by Drosophila Dicer-1. Nature Struct. Mol. Biol. 18, 1153–1158 (2011).
Gu, S. et al. The loop position of shRNAs and pre-miRNAs is critical for the accuracy of dicer processing in vivo. Cell 151, 900–911 (2012).
Macrae, I. J. et al. Structural basis for double-stranded RNA processing by Dicer. Science 311, 195–198 (2006).
Park, J. E. et al. Dicer recognizes the 5′ end of RNA for efficient and accurate processing. Nature 475, 201–205 (2011).
Tian, Y. et al. A phosphate-binding pocket within the platform-PAZ-connector helix cassette of human Dicer. Mol. Cell 53, 606–616 (2014).
Lau, P. W., Potter, C. S., Carragher, B. & MacRae, I. J. Structure of the human Dicer–TRBP complex by electron microscopy. Structure 17, 1326–1332 (2009).
Taylor, D. W. et al. Substrate-specific structural rearrangements of human Dicer. Nature Struct. Mol. Biol. 20, 662–670 (2013).
Wang, H. W. et al. Structural insights into RNA processing by the human RISC-loading complex. Nature Struct. Mol. Biol. 16, 1148–1153 (2009).
MacRae, I. J., Zhou, K. & Doudna, J. A. Structural determinants of RNA recognition and cleavage by Dicer. Nature Struct. Mol. Biol. 14, 934–940 (2007).
Vermeulen, A. et al. The contributions of dsRNA structure to Dicer specificity and efficiency. RNA 11, 674–682 (2005).
Zhang, H., Kolb, F. A., Brondani, V., Billy, E. & Filipowicz, W. Human Dicer preferentially cleaves dsRNAs at their termini without a requirement for ATP. EMBO J. 21, 5875–5885 (2002).
Forstemann, K. et al. Normal microRNA maturation and germ-line stem cell maintenance requires Loquacious, a double-stranded RNA-binding domain protein. PLoS Biol. 3, e236 (2005).
Jiang, F. et al. Dicer-1 and R3D1-L catalyze microRNA maturation in Drosophila. Genes Dev. 19, 1674–1679 (2005).
Saito, K., Ishizuka, A., Siomi, H. & Siomi, M. C. Processing of pre-microRNAs by the Dicer-1-Loquacious complex in Drosophila cells. PLoS Biol. 3, e235 (2005).
Fukunaga, R. et al. Dicer partner proteins tune the length of mature miRNAs in flies and mammals. Cell 151, 533–546 (2012).
Liu, X. et al. Dicer-1, but not Loquacious, is critical for assembly of miRNA-induced silencing complexes. RNA 13, 2324–2329 (2007).
Park, J. K., Liu, X., Strauss, T. J., McKearin, D. M. & Liu, Q. The miRNA pathway intrinsically controls self-renewal of Drosophila germline stem cells. Curr. Biol. 17, 533–538 (2007).
Chendrimada, T. P. et al. TRBP recruits the Dicer complex to Ago2 for microRNA processing and gene silencing. Nature 436, 740–744 (2005).
Haase, A. D. et al. TRBP, a regulator of cellular PKR and HIV-1 virus expression, interacts with Dicer and functions in RNA silencing. EMBO Rep. 6, 961–967 (2005).
Lee, Y. et al. The role of PACT in the RNA silencing pathway. EMBO J. 25, 522–532 (2006).
Lee, H. Y., Zhou, K., Smith, A. M., Noland, C. L. & Doudna, J. A. Differential roles of human Dicer-binding proteins TRBP and PACT in small RNA processing. Nucleic Acids Res. 41, 6568–6576 (2013).
Chakravarthy, S., Sternberg, S. H., Kellenberger, C. A. & Doudna, J. A. Substrate-specific kinetics of Dicer-catalyzed RNA processing. J. Mol. Biol. 404, 392–402 (2010).
Lee, H. Y. & Doudna, J. A. TRBP alters human precursor microRNA processing in vitro. RNA 18, 2012–2019 (2012).
Garcia, M. A., Meurs, E. F. & Esteban, M. The dsRNA protein kinase PKR: virus and cell control. Biochimie 89, 799–811 (2007).
Paroo, Z., Ye, X., Chen, S. & Liu, Q. Phosphorylation of the human microRNA-generating complex mediates MAPK/Erk signaling. Cell 139, 112–122 (2009).
Melo, S. A. et al. A TARBP2 mutation in human cancer impairs microRNA processing and DICER1 function. Nature Genet. 41, 365–370 (2009).
Tokumaru, S., Suzuki, M., Yamada, H., Nagino, M. & Takahashi, T. let-7 regulates Dicer expression and constitutes a negative feedback loop. Carcinogenesis 29, 2073–2077 (2008).
Forman, J. J., Legesse-Miller, A. & Coller, H. A. A search for conserved sequences in coding regions reveals that the let-7 microRNA targets Dicer within its coding sequence. Proc. Natl Acad. Sci. USA 105, 14879–14884 (2008).
Hammond, S. M., Boettcher, S., Caudy, A. A., Kobayashi, R. & Hannon, G. J. Argonaute2, a link between genetic and biochemical analyses of RNAi. Science 293, 1146–1150 (2001).
Mourelatos, Z. et al. miRNPs: a novel class of ribonucleoproteins containing numerous microRNAs. Genes Dev. 16, 720–728 (2002).
Tabara, H. et al. The rde-1 gene, RNA interference, and transposon silencing in C. elegans. Cell 99, 123–132 (1999).
Kawamata, T. & Tomari, Y. Making RISC. Trends Biochem. Sci. 35, 368–376 (2010).
Elkayam, E. et al. The structure of human Argonaute-2 in complex with miR-20a. Cell 150, 100–110 (2012).
Nakanishi, K., Weinberg, D. E., Bartel, D. P. & Patel, D. J. Structure of yeast Argonaute with guide RNA. Nature 486, 368–374 (2012).
Schirle, N. T. & MacRae, I. J. The crystal structure of human Argonaute2. Science 336, 1037–1040 (2012).
Song, J. J., Smith, S. K., Hannon, G. J. & Joshua-Tor, L. Crystal structure of Argonaute and its implications for RISC slicer activity. Science 305, 1434–1437 (2004).
Ma, J. B. et al. Structural basis for 5′-end-specific recognition of guide RNA by the A. fulgidus Piwi protein. Nature 434, 666–670 (2005).
Parker, J. S., Roe, S. M. & Barford, D. Structural insights into mRNA recognition from a PIWI domain-siRNA guide complex. Nature 434, 663–666 (2005).
Wang, Y., Sheng, G., Juranek, S., Tuschl, T. & Patel, D. J. Structure of the guide-strand-containing argonaute silencing complex. Nature 456, 209–213 (2008).
Frank, F., Sonenberg, N. & Nagar, B. Structural basis for 5′-nucleotide base-specific recognition of guide RNA by human AGO2. Nature 465, 818–822 (2010).
Wang, Y. et al. Structure of an argonaute silencing complex with a seed-containing guide DNA and target RNA duplex. Nature 456, 921–926 (2008).
Wang, Y. et al. Nucleation, propagation and cleavage of target RNAs in Ago silencing complexes. Nature 461, 754–761 (2009).
Liu, J. et al. Argonaute2 is the catalytic engine of mammalian RNAi. Science 305, 1437–1441 (2004).
Cheloufi, S., Dos Santos, C. O., Chong, M. M. & Hannon, G. J. A dicer-independent miRNA biogenesis pathway that requires Ago catalysis. Nature 465, 584–589 (2010).
Su, H., Trombly, M. I., Chen, J. & Wang, X. Essential and overlapping functions for mammalian Argonautes in microRNA silencing. Genes Dev. 23, 304–317 (2009).
Modzelewski, A. J., Holmes, R. J., Hilz, S., Grimson, A. & Cohen, P. E. AGO4 regulates entry into meiosis and influences silencing of sex chromosomes in the male mouse germline. Dev. Cell 23, 251–264 (2012).
Forstemann, K., Horwich, M. D., Wee, L., Tomari, Y. & Zamore, P. D. Drosophila microRNAs are sorted into functionally distinct argonaute complexes after production by dicer-1. Cell 130, 287–297 (2007).
Okamura, K., Ishizuka, A., Siomi, H. & Siomi, M. C. Distinct roles for Argonaute proteins in small RNA-directed RNA cleavage pathways. Genes Dev. 18, 1655–1666 (2004).
Tomari, Y., Du, T. & Zamore, P. D. Sorting of Drosophila small silencing RNAs. Cell 130, 299–308 (2007).
Czech, B. et al. Hierarchical rules for Argonaute loading in Drosophila. Mol. Cell 36, 445–456 (2009).
Ghildiyal, M., Xu, J., Seitz, H., Weng, Z. & Zamore, P. D. Sorting of Drosophila small silencing RNAs partitions microRNA* strands into the RNA interference pathway. RNA 16, 43–56 (2010).
Kawamata, T., Seitz, H. & Tomari, Y. Structural determinants of miRNAs for RISC loading and slicer-independent unwinding. Nature Struct. Mol. Biol. 16, 953–960 (2009).
Okamura, K., Liu, N. & Lai, E. C. Distinct mechanisms for microRNA strand selection by Drosophila Argonautes. Mol. Cell 36, 431–444 (2009).
Jannot, G., Boisvert, M. E., Banville, I. H. & Simard, M. J. Two molecular features contribute to the Argonaute specificity for the microRNA and RNAi pathways in C. elegans. RNA 14, 829–835 (2008).
Steiner, F. A. et al. Structural features of small RNA precursors determine Argonaute loading in Caenorhabditis elegans. Nature Struct. Mol. Biol. 14, 927–933 (2007).
Azuma-Mukai, A. et al. Characterization of endogenous human Argonautes and their miRNA partners in RNA silencing. Proc. Natl Acad. Sci. USA 105, 7964–7969 (2008).
Meister, G. et al. Human Argonaute2 mediates RNA cleavage targeted by miRNAs and siRNAs. Mol. Cell 15, 185–197 (2004).
Dueck, A., Ziegler, C., Eichner, A., Berezikov, E. & Meister, G. microRNAs associated with the different human Argonaute proteins. Nucleic Acids Res. 40, 9850–9862 (2012).
Yoda, M. et al. ATP-dependent human RISC assembly pathways. Nature Struct. Mol. Biol. 17, 17–23 (2010).
Liu, Y. et al. C3PO, an endoribonuclease that promotes RNAi by facilitating RISC activation. Science 325, 750–753 (2009).
Pham, J. W., Pellino, J. L., Lee, Y. S., Carthew, R. W. & Sontheimer, E. J. A Dicer-2-dependent 80s complex cleaves targeted mRNAs during RNAi in Drosophila. Cell 117, 83–94 (2004).
Tomari, Y. et al. RISC assembly defects in the Drosophila RNAi mutant armitage. Cell 116, 831–841 (2004).
Gregory, R. I., Chendrimada, T. P., Cooch, N. & Shiekhattar, R. Human RISC couples microRNA biogenesis and posttranscriptional gene silencing. Cell 123, 631–640 (2005).
MacRae, I. J., Ma, E., Zhou, M., Robinson, C. V. & Doudna, J. A. In vitro reconstitution of the human RISC-loading complex. Proc. Natl Acad. Sci. USA 105, 512–517 (2008).
Maniataki, E. & Mourelatos, Z. A human, ATP-independent, RISC assembly machine fueled by pre-miRNA. Genes Dev. 19, 2979–2990 (2005).
Miyoshi, K., Tsukumo, H., Nagami, T., Siomi, H. & Siomi, M. C. Slicer function of Drosophila Argonautes and its involvement in RISC formation. Genes Dev. 19, 2837–2848 (2005).
Gredell, J. A., Dittmer, M. J., Wu, M., Chan, C. & Walton, S. P. Recognition of siRNA asymmetry by TAR RNA binding protein. Biochemistry 49, 3148–3155 (2010).
Noland, C. L., Ma, E. & Doudna, J. A. siRNA repositioning for guide strand selection by human Dicer complexes. Mol. Cell 43, 110–121 (2011).
Liu, X., Jin, D. Y., McManus, M. T. & Mourelatos, Z. Precursor microRNA-programmed silencing complex assembly pathways in mammals. Mol. Cell 46, 507–517 (2012).
Betancur, J. G. & Tomari, Y. Dicer is dispensable for asymmetric RISC loading in mammals. RNA 18, 24–30 (2012).
Ye, X. et al. Structure of C3PO and mechanism of human RISC activation. Nature Struct. Mol. Biol. 18, 650–657 (2011).
Diederichs, S. & Haber, D. A. Dual role for argonautes in microRNA processing and posttranscriptional regulation of microRNA expression. Cell 131, 1097–1108 (2007).
Leuschner, P. J., Ameres, S. L., Kueng, S. & Martinez, J. Cleavage of the siRNA passenger strand during RISC assembly in human cells. EMBO Rep. 7, 314–320 (2006).
Matranga, C., Tomari, Y., Shin, C., Bartel, D. P. & Zamore, P. D. Passenger-strand cleavage facilitates assembly of siRNA into Ago2-containing RNAi enzyme complexes. Cell 123, 607–620 (2005).
Rand, T. A., Petersen, S., Du, F. & Wang, X. Argonaute2 cleaves the anti-guide strand of siRNA during RISC activation. Cell 123, 621–629 (2005).
Nykanen, A., Haley, B. & Zamore, P. D. ATP requirements and small interfering RNA structure in the RNA interference pathway. Cell 107, 309–321 (2001).
Iwasaki, S. et al. Hsc70/Hsp90 chaperone machinery mediates ATP-dependent RISC loading of small RNA duplexes. Mol. Cell 39, 292–299 (2010).
Miyoshi, K., Miyoshi, T., Hartig, J. V., Siomi, H. & Siomi, M. C. Molecular mechanisms that funnel RNA precursors into endogenous small-interfering RNA and microRNA biogenesis pathways in Drosophila. RNA 16, 506–515 (2010).
Han, B. W., Hung, J. H., Weng, Z., Zamore, P. D. & Ameres, S. L. The 3′-to-5′ exoribonuclease Nibbler shapes the 3′ ends of microRNAs bound to Drosophila Argonaute1. Curr. Biol. 21, 1878–1887 (2011).
Liu, N. et al. The exoribonuclease Nibbler controls 3′ end processing of microRNAs in Drosophila. Curr. Biol. 21, 1888–1893 (2011).
Khvorova, A., Reynolds, A. & Jayasena, S. D. Functional siRNAs and miRNAs exhibit strand bias. Cell 115, 209–216 (2003).
Schwarz, D. S. et al. Asymmetry in the assembly of the RNAi enzyme complex. Cell 115, 199–208 (2003).
Hu, H. Y. et al. Sequence features associated with microRNA strand selection in humans and flies. BMC Genomics 10, 413 (2009).
Lau, N. C., Lim, L. P., Weinstein, E. G. & Bartel, D. P. An abundant class of tiny RNAs with probable regulatory roles in Caenorhabditis elegans. Science 294, 858–862 (2001).
Wu, H., Ye, C., Ramirez, D. & Manjunath, N. Alternative processing of primary microRNA transcripts by Drosha generates 5′ end variation of mature microRNA. PLoS ONE 4, e7566 (2009).
Qi, H. H. et al. Prolyl 4-hydroxylation regulates Argonaute 2 stability. Nature 455, 421–424 (2008).
Wu, C. et al. Hypoxia potentiates microRNA-mediated gene silencing through posttranslational modification of Argonaute2. Mol. Cell. Biol. 31, 4760–4774 (2011).
Zeng, Y., Sankala, H., Zhang, X. & Graves, P. R. Phosphorylation of Argonaute 2 at serine-387 facilitates its localization to processing bodies. Biochem. J. 413, 429–436 (2008).
Horman, S. R. et al. Akt-mediated phosphorylation of argonaute 2 downregulates cleavage and upregulates translational repression of microRNA targets. Mol. Cell 50, 356–367 (2013).
Shen, J. et al. EGFR modulates microRNA maturation in response to hypoxia through phosphorylation of AGO2. Nature 497, 383–387 (2013).
Rudel, S. et al. Phosphorylation of human Argonaute proteins affects small RNA binding. Nucleic Acids Res. 39, 2330–2343 (2011).
Mazumder, A., Bose, M., Chakraborty, A., Chakrabarti, S. & Bhattacharyya, S. N. A transient reversal of miRNA-mediated repression controls macrophage activation. EMBO Rep. 14, 1008–1016 (2013).
Leung, A. K. et al. Poly(ADP-ribose) regulates stress responses and microRNA activity in the cytoplasm. Mol. Cell 42, 489–499 (2011).
Seo, G. J. et al. Reciprocal inhibition between intracellular antiviral signaling and the RNAi machinery in mammalian cells. Cell Host Microbe 14, 435–445 (2013).
Rybak, A. et al. The let-7 target gene mouse lin-41 is a stem cell specific E3 ubiquitin ligase for the miRNA pathway protein Ago2. Nature Cell Biol. 11, 1411–1420 (2009).
Johnston, M., Geoffroy, M. C., Sobala, A., Hay, R. & Hutvagner, G. HSP90 protein stabilizes unloaded argonaute complexes and microscopic P-bodies in human cells. Mol. Biol. Cell 21, 1462–1469 (2010).
Smibert, P., Yang, J. S., Azzam, G., Liu, J. L. & Lai, E. C. Homeostatic control of Argonaute stability by microRNA availability. Nature Struct. Mol. Biol. 20, 789–795 (2013).
Bronevetsky, Y. et al. T cell activation induces proteasomal degradation of Argonaute and rapid remodeling of the microRNA repertoire. J. Exp. Med. 210, 417–432 (2013).
Loedige, I., Gaidatzis, D., Sack, R., Meister, G. & Filipowicz, W. The mammalian TRIM-NHL protein TRIM71/LIN-41 is a repressor of mRNA function. Nucleic Acids Res. 41, 518–532 (2013).
Chang, H. M. et al. Trim71 cooperates with microRNAs to repress Cdkn1a expression and promote embryonic stem cell proliferation. Nature Commun. 3, 923 (2012).
Chen, J., Lai, F. & Niswander, L. The ubiquitin ligase mLin41 temporally promotes neural progenitor cell maintenance through FGF signaling. Genes Dev. 26, 803–815 (2012).
Martinez, N. J. & Gregory, R. I. Argonaute2 expression is post-transcriptionally coupled to microRNA abundance. RNA 19, 605–612 (2013).
Ryan, B. M., Robles, A. I. & Harris, C. C. Genetic variation in microRNA networks: the implications for cancer research. Nature Rev. Cancer 10, 389–402 (2010).
Calin, G. A. et al. A microRNA signature associated with prognosis and progression in chronic lymphocytic leukemia. N. Engl. J. Med. 353, 1793–1801 (2005).
Tian, T. et al. A functional genetic variant in microRNA-196a2 is associated with increased susceptibility of lung cancer in Chinese. Cancer Epidemiol. Biomarkers Prev. 18, 1183–1187 (2009).
Jazdzewski, K. et al. Polymorphic mature microRNAs from passenger strand of pre-miR-146a contribute to thyroid cancer. Proc. Natl Acad. Sci. USA 106, 1502–1505 (2009).
Jazdzewski, K. et al. Common SNP in pre-miR-146a decreases mature miR expression and predisposes to papillary thyroid carcinoma. Proc. Natl Acad. Sci. USA 105, 7269–7274 (2008).
Ameres, S. L. & Zamore, P. D. Diversifying microRNA sequence and function. Nature Rev. Mol. Cell Biol. 14, 475–488 (2013).
Pasquinelli, A. E. et al. Conservation of the sequence and temporal expression of let-7 heterochronic regulatory RNA. Nature 408, 86–89 (2000).
Suh, M. R. et al. Human embryonic stem cells express a unique set of microRNAs. Dev. Biol. 270, 488–498 (2004).
Thomson, J. M. et al. Extensive post-transcriptional regulation of microRNAs and its implications for cancer. Genes Dev. 20, 2202–2207 (2006).
Wulczyn, F. G. et al. Post-transcriptional regulation of the let-7 microRNA during neural cell specification. FASEB J. 21, 415–426 (2007).
Heo, I. et al. Lin28 mediates the terminal uridylation of let-7 precursor microRNA. Mol. Cell 32, 276–284 (2008).
Newman, M. A., Thomson, J. M. & Hammond, S. M. Lin-28 interaction with the Let-7 precursor loop mediates regulated microRNA processing. RNA 14, 1539–1549 (2008).
Rybak, A. et al. A feedback loop comprising lin-28 and let-7 controls pre-let-7 maturation during neural stem-cell commitment. Nature Cell Biol. 10, 987–993 (2008).
Viswanathan, S. R., Daley, G. Q. & Gregory, R. I. Selective blockade of microRNA processing by Lin28. Science 320, 97–100 (2008).
Loughlin, F. E. et al. Structural basis of pre-let-7 miRNA recognition by the zinc knuckles of pluripotency factor Lin28. Nature Struct. Mol. Biol. 19, 84–89 (2012).
Nam, Y., Chen, C., Gregory, R. I., Chou, J. J. & Sliz, P. Molecular basis for interaction of let-7 microRNAs with Lin28. Cell 147, 1080–1091 (2011).
Hagan, J. P., Piskounova, E. & Gregory, R. I. Lin28 recruits the TUTase Zcchc11 to inhibit let-7 maturation in mouse embryonic stem cells. Nature Struct. Mol. Biol. 16, 1021–1025 (2009).
Heo, I. et al. TUT4 in concert with Lin28 suppresses microRNA biogenesis through pre-microRNA uridylation. Cell 138, 696–708 (2009).
Chang, H. M., Triboulet, R., Thornton, J. E. & Gregory, R. I. A role for the Perlman syndrome exonuclease Dis3l2 in the Lin28-let-7 pathway. Nature 497, 244–248 (2013).
Ustianenko, D. et al. Mammalian DIS3L2 exoribonuclease targets the uridylated precursors of let-7 miRNAs. RNA 19, 1632–1638 (2013).
Heo, I. et al. Mono-uridylation of pre-microRNA as a key step in the biogenesis of group II let-7 microRNAs. Cell 151, 521–532 (2012).
Katoh, T. et al. Selective stabilization of mammalian microRNAs by 3′ adenylation mediated by the cytoplasmic poly(A) polymerase GLD-2. Genes Dev. 23, 433–438 (2009).
Backes, S. et al. Degradation of host microRNAs by poxvirus poly(A) polymerase reveals terminal RNA methylation as a protective antiviral mechanism. Cell Host Microbe 12, 200–210 (2012).
Yang, W. et al. Modulation of microRNA processing and expression through RNA editing by ADAR deaminases. Nature Struct. Mol. Biol. 13, 13–21 (2006).
Kawahara, Y., Zinshteyn, B., Chendrimada, T. P., Shiekhattar, R. & Nishikura, K. RNA editing of the microRNA-151 precursor blocks cleavage by the Dicer–TRBP complex. EMBO Rep. 8, 763–769 (2007).
Xhemalce, B., Robson, S. C. & Kouzarides, T. Human RNA methyltransferase BCDIN3D regulates microRNA processing. Cell 151, 278–288 (2012).
Ruegger, S. & Grosshans, H. MicroRNA turnover: when, how, and why. Trends Biochem. Sci. 37, 436–446 (2012).
Suzuki, H. I. et al. MCPIP1 ribonuclease antagonizes dicer and terminates microRNA biogenesis through precursor microRNA degradation. Mol. Cell 44, 424–436 (2011).
Upton, J. P. et al. IRE1α cleaves select microRNAs during ER stress to derepress translation of proapoptotic caspase-2. Science 338, 818–822 (2012).
Ramachandran, V. & Chen, X. Degradation of microRNAs by a family of exoribonucleases in Arabidopsis. Science 321, 1490–1492 (2008).
Chatterjee, S., Fasler, M., Bussing, I. & Grosshans, H. Target-mediated protection of endogenous microRNAs in C. elegans. Dev. Cell 20, 388–396 (2011).
Chatterjee, S. & Grosshans, H. Active turnover modulates mature microRNA activity in Caenorhabditis elegans. Nature 461, 546–549 (2009).
Krol, J. et al. Characterizing light-regulated retinal microRNAs reveals rapid turnover as a common property of neuronal microRNAs. Cell 141, 618–631 (2010).
Das, S. K. et al. Human polynucleotide phosphorylase selectively and preferentially degrades microRNA-221 in human melanoma cells. Proc. Natl Acad. Sci. USA 107, 11948–11953 (2010).
Thomas, M. F. et al. Eri1 regulates microRNA homeostasis and mouse lymphocyte development and antiviral function. Blood 120, 130–142 (2012).
Libri, V. et al. Murine cytomegalovirus encodes a miR-27 inhibitor disguised as a target. Proc. Natl Acad. Sci. USA 109, 279–284 (2012).
Ameres, S. L. et al. Target RNA-directed trimming and tailing of small silencing RNAs. Science 328, 1534–1539 (2010).
Baccarini, A. et al. Kinetic analysis reveals the fate of a microRNA following target regulation in mammalian cells. Curr. Biol. 21, 369–376 (2011).
Xie, J. et al. Long-term, efficient inhibition of microRNA function in mice using rAAV vectors. Nature Methods 9, 403–409 (2012).
Cazalla, D., Yario, T. & Steitz, J. A. Down-regulation of a host microRNA by a Herpesvirus saimiri noncoding RNA. Science 328, 1563–1566 (2010).
Lee, S. et al. Selective degradation of host MicroRNAs by an intergenic HCMV noncoding RNA accelerates virus production. Cell Host Microbe 13, 678–690 (2013).
Xie, M. & Steitz, J. A. Versatile microRNA biogenesis in animals and their viruses. RNA Biol. http://dx.doi.org/10.4161/rna.28985 (2014).
Berezikov, E., Chung, W. J., Willis, J., Cuppen, E. & Lai, E. C. Mammalian mirtron genes. Mol. Cell 28, 328–336 (2007).
Okamura, K., Hagen, J. W., Duan, H., Tyler, D. M. & Lai, E. C. The mirtron pathway generates microRNA-class regulatory RNAs in Drosophila. Cell 130, 89–100 (2007).
Ruby, J. G., Jan, C. H. & Bartel, D. P. Intronic microRNA precursors that bypass Drosha processing. Nature 448, 83–86 (2007).
Flynt, A. S., Greimann, J. C., Chung, W. J., Lima, C. D. & Lai, E. C. MicroRNA biogenesis via splicing and exosome-mediated trimming in Drosophila. Mol. Cell 38, 900–907 (2010).
Ender, C. et al. A human snoRNA with microRNA-like functions. Mol. Cell 32, 519–528 (2008).
Cazalla, D., Xie, M. & Steitz, J. A. A primate herpesvirus uses the integrator complex to generate viral microRNAs. Mol. Cell 43, 982–992 (2011).
Bogerd, H. P. et al. A mammalian herpesvirus uses noncanonical expression and processing mechanisms to generate viral microRNAs. Mol. Cell 37, 135–142 (2010).
Cifuentes, D. et al. A novel miRNA processing pathway independent of Dicer requires Argonaute2 catalytic activity. Science 328, 1694–1698 (2010).
Yang, J. S. et al. Conserved vertebrate mir-451 provides a platform for Dicer-independent, Ago2-mediated microRNA biogenesis. Proc. Natl Acad. Sci. USA 107, 15163–15168 (2010).
Yoda, M. et al. Poly(A)-specific ribonuclease mediates 3′-end trimming of Argonaute2-cleaved precursor microRNAs. Cell Rep. 5, 715–726 (2013).
Shenoy, A. & Blelloch, R. Genomic analysis suggests that mRNA destabilization by the microprocessor is specialized for the auto-regulation of Dgcr8. PLoS ONE 4, e6971 (2009).
Triboulet, R., Chang, H. M., Lapierre, R. J. & Gregory, R. I. Post-transcriptional control of DGCR8 expression by the Microprocessor. RNA 15, 1005–1011 (2009).
Karginov, F. V. et al. Diverse endonucleolytic cleavage sites in the mammalian transcriptome depend upon microRNAs, Drosha, and additional nucleases. Mol. Cell 38, 781–788 (2010).
Lin, Y. T. & Sullivan, C. S. Expanding the role of Drosha to the regulation of viral gene expression. Proc. Natl Acad. Sci. USA 108, 11229–11234 (2011).
Heras, S. R. et al. The Microprocessor controls the activity of mammalian retrotransposons. Nature Struct. Mol. Biol. 20, 1173–1181 (2013).
Macias, S. et al. DGCR8 HITS-CLIP reveals novel functions for the Microprocessor. Nature Struct. Mol. Biol. 19, 760–766 (2012).
Luhur, A., Chawla, G., Wu, Y. C., Li, J. & Sokol, N. S. Drosha-independent DGCR8/Pasha pathway regulates neuronal morphogenesis. Proc. Natl Acad. Sci. USA 111, 1421–1426 (2014).
Kaneko, H. et al. DICER1 deficit induces Alu RNA toxicity in age-related macular degeneration. Nature 471, 325–330 (2011).
Hu, Q. et al. DICER- and AGO3-dependent generation of retinoic acid-induced DR2 Alu RNAs regulates human stem cell proliferation. Nature Struct. Mol. Biol. 19, 1168–1175 (2012).
Meister, G. Argonaute proteins: functional insights and emerging roles. Nature Rev. Genet. 14, 447–459 (2013).
Karamyshev, A. L. et al. Inefficient SRP interaction with a nascent chain triggers a mRNA quality control pathway. Cell 156, 146–157 (2014).
Ipsaro, J. J., Haase, A. D., Knott, S. R., Joshua-Tor, L. & Hannon, G. J. The structural biochemistry of Zucchini implicates it as a nuclease in piRNA biogenesis. Nature 491, 279–283 (2012).
Nishimasu, H. et al. Structure and function of Zucchini endoribonuclease in piRNA biogenesis. Nature 491, 284–287 (2012).
Siomi, M. C., Sato, K., Pezic, D. & Aravin, A. A. PIWI-interacting small RNAs: the vanguard of genome defence. Nature Rev. Mol. Cell Biol. 12, 246–258 (2011).
Kataoka, N., Fujita, M. & Ohno, M. Functional association of the Microprocessor complex with the spliceosome. Mol. Cell Biol. 29, 3243–3254 (2009).
Acknowledgements
The authors are grateful to the members of their laboratory for helpful discussions and comments. Work in the authors laboratory was supported by the Research Center Programme (EM1402) of the Institute for Basic Science IBS) from the Ministry of Science, Information and Communication Technology (ICT) and Future Planning of Korea (M.H. and V.N.K.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
FURTHER INFORMATION
Glossary
- Argonaute family proteins
-
(AGO family proteins). Proteins that associate with small RNAs and function as effectors in RNA silencing. AGO proteins carry two characteristic domains — PIWI (an endoribonuclease domain) and PAZ (PIWI–AGO–ZWILLE; the 3′ end-binding module).
- PIWI-interacting RNA
-
(piRNA). Small silencing RNAs (24–30 nucleotides long) that bind PIWI clade Argonaute proteins in animals and silence germline transposons. They are thought to derive from single-stranded RNA precursors and do not require RNase III enzymes for their maturation.
- Pachytene
-
The stage of meiotic prophase that immediately follows the zygotene. It is characterized by paired chromosomes that are condensed and visibly divided into chromatids, and by the occurrence of crossing-over.
- Let-7 family
-
The let-7 gene was initially discovered as an essential developmental gene in Caenorhabditis elegans and, later, as one of the first two microRNAs (miRNAs). The let-7 miRNA family is highly conserved throughout bilaterian animals, and it suppresses cell proliferation and promotes cell differentiation. It is also a tumour suppressor.
- ChIP–seq
-
(Chromatin immunoprecipitation followed by sequencing). A method used to analyse protein interactions with DNA. It combines ChIP with parallel DNA sequencing to identify the binding sites of DNA-associated proteins.
- DiGeorge syndrome
-
An autosomal recessive genetic disorder caused by a deletion in chromosome 22 that is commonly associated with heart defects, poor immune system function, a cleft palate and behavioural disorders.
- RNA editing
-
A molecular process through which some cells make discrete changes to specific nucleotide sequences within an RNA molecule after it has been generated by the RNA polymerase. Editing events may include the insertion, deletion and substitution of nucleotides within the edited RNA molecule.
- RNA-induced silencing complex
-
(RISC). A ribonucleoprotein complex that consists of a small RNA guide strand bound to an Argonaute protein. RISC mediates all RNA-silencing pathways, and it can also include auxiliary proteins that extend or modify its function.
- RISC-loading complex
-
(RLC). A protein complex containing Dicer, a double-stranded-RNA-binding protein, an Argonaute (AGO) protein and potentially other proteins that are required for loading of small RNAs onto the AGO protein.
- Processing bodies
-
(P-bodies). Distinct foci within the cytoplasm of the eukaryotic cell that consist of many enzymes involved in mRNA turnover. They are thought to be sites for translational suppression and/or mRNA decay, and to be involved in RNA silencing. Some mRNAs can exit P-bodies and reinitiate translation.
- Terminal uridylyl transferases
-
Nucleotidyl transferases that covalently add one or more UMP moieties from UTP to the 3′-OH group of an RNA substrate in a template-independent manner.
- Short hairpin RNAs
-
Sequences of RNA that have a tight hairpin turn that can be processed by Dicer.
- Small nucleolar RNAs
-
(snoRNAs). A class of small RNA molecules that primarily guide chemical modifications of other RNAs. This mainly includes rRNAs, tRNAs and small nuclear RNAs.
- Retrotransposon
-
A transposable element that replicates through an RNA intermediate, which is converted by a reverse transcriptase to cDNA. The cDNA can be inserted into genomic DNA, which increases the number of copies of the retrotransposon in the genome.
- Alu
-
A short (∼300 bp long) stretch of repetitive elements that are classified as short interspersed elements among the class of repetitive DNA elements. Alu elements of different kinds occur in large numbers in primate genomes, accounting for over 10% of the human genome.
Rights and permissions
About this article
Cite this article
Ha, M., Kim, V. Regulation of microRNA biogenesis. Nat Rev Mol Cell Biol 15, 509–524 (2014). https://doi.org/10.1038/nrm3838
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrm3838
This article is cited by
-
The potential of microRNAs in cancer diagnostic and therapeutic strategies: a narrative review
The Journal of Basic and Applied Zoology (2024)
-
DiCleave: a deep learning model for predicting human Dicer cleavage sites
BMC Bioinformatics (2024)
-
Unraveling the microRNAs, key players in folliculogenesis and ovarian diseases
Middle East Fertility Society Journal (2024)
-
Breaking the mold with RNA—a “RNAissance” of life science
npj Genomic Medicine (2024)
-
Maintaining Drosha expression with Cdk5 inhibitors as a potential therapeutic strategy for early intervention after TBI
Experimental & Molecular Medicine (2024)