Key Points
-
The ends of linear chromosomes are problematic for the eukaryotic DNA-replication machinery. Eukaryotes have evolved specialized structures at chromosome ends — known as telomeres — that are replicated by a unique mechanism using the reverse transcriptase, telomerase.
-
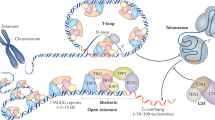
Telomeres are comprised of G-rich DNA repeats, the sequence of which varies among different organisms. Telomeres facilitate the complete replication of linear chromosomes and protect chromosome ends from degradation and end-to-end fusions.
-
In yeast and humans, the G-rich strand of telomeres extends in the 3′ direction to form a single-stranded G-tail. G-tails are telomerase substrates and bind single-strand-specific DNA-binding proteins that protect chromosome ends from degradation and from fusions with DNA breaks or other chromosome ends.
-
Telomerase action is highly regulated in vivo as inappropriate telomere addition is deleterious and can stabilize chromosomal aberrations.
-
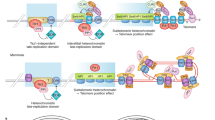
Telomerase can be regulated in various ways: by modulating the telomeric chromatin structure through single-stranded or double-stranded telomeric-DNA-binding proteins and their associated factors; by using telomerase accessory factors; by dissociating active telomerase from chromosome ends; and by sequestering active telomerase in specific subcellular compartments, such as the nucleolus.
-
Free single-stranded DNA can signal to DNA-damage-response pathways; so, telomeric G-tails must be shielded to avoid detection as damaged DNA. Since telomeres do not fuse with each other or with DNA breaks, telomeres are also protected from non-homologous end joining. Surprisingly, proteins such as Ku and the Mre11 complex that are involved in non-homologous end joining and DNA checkpoints are important for telomere maintenance.
-
Many of the components involved in regulating access of telomerase to the telomere are conserved from yeast to humans.
Abstract
In organisms with linear chromosomes, telomeres are essential to maintain genome integrity. However, inappropriate telomere addition, for example to double-stranded DNA breaks, might stabilize deleterious genetic changes. Therefore, telomere addition by telomerase is highly regulated, for example by mechanisms that determine the accessibility of telomeres to elongation by telomerase. These mechanisms, which have been studied mainly in budding yeast and human cell culture, can be subdivided into two classes: mechanisms that modulate the telomeric chromatin structure and those that sequester active telomerase from chromosome ends.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Wellinger, R. J., Wolf, A. J. & Zakian, V. A. Saccharomyces telomeres acquire single-strand TG1–3 tails late in S phase. Cell 72, 51–60 (1993).
Bourns, B. D., Alexander, M. K., Smith, A. M. & Zakian, V. A. Sir proteins, Rif proteins and Cdc13p bind Saccharomyces telomeres in vivo. Mol. Cell. Biol. 18, 5600–5608 (1998).
Taggart, A. K. P., Teng, S. -C. & Zakian, V. A. Est1p as a cell-cycle-regulated activator of telomere-bound telomerase. Science 297, 1023–1026 (2002). Shows that Est2 is telomere associated throughout most of the cell cycle, whereas Est1 is associated only when telomerase is active.
McElligott, R. & Wellinger, R. J. The terminal DNA structure of mammalian chromosomes. EMBO J. 16, 3705–3714 (1997).
Wright, W. E., Tesmer, V. M., Huffman, K. E., Levene, S. D. & Shay, J. W. Normal human chromosomes have long G-rich telomeric overhangs at one end. Genes Dev. 11, 2801–2809 (1997).
Makarov, V. L., Hirose, Y. & Langmore, J. P. Long G tails at both ends of human chromosomes suggest a C strand degradation mechanism for telomere shortening. Cell 88, 657–666 (1997).
Lundblad, V. & Szostak, J. W. A mutant with a defect in telomere elongation leads to senescence in yeast. Cell 57, 633–643 (1989).
Garvik, B., Carson, M. & Hartwell, L. Single-stranded DNA arising at telomeres in cdc13 mutants may constitute a specific signal for the RAD9 checkpoint. Mol. Cell. Biol. 15, 6128–6138 (1995).
Grandin, N., Damon, C. & Charbonneau, M. Ten1 functions in telomere end protection and length regulation in association with Stn1 and Cdc13. EMBO J. 20, 1173–1183 (2001).
Grandin, N., Reed, S. I. & Charbonneau, M. Stn1, a new Saccharomyces cerevisiae protein, is implicated in telomere size regulation in association with Cdc13. Genes Dev. 11, 512–527 (1997).
van Steensel, B., Smogorzewska, A. & de Lange, T. TRF2 protects human telomeres from end-to-end fusions. Cell 92, 401–413 (1998). Shows that telomeric fusions occur when G-tails are lost in human cells, despite the presence of duplex telomeric DNA.
Wellinger, R. J., Ethier, K., Labrecque, P. & Zakian, V. A. Evidence for a new step in telomere maintenance. Cell 85, 423–433 (1996).
Dionne, I. & Wellinger, R. J. Cell-cycle-regulated generation of single-stranded G-rich DNA in the absence of telomerase. Proc. Natl Acad. Sci. USA 93, 13902–13907 (1996).
Hemann, M. & Greider, C. G-strand overhangs on telomeres in telomerase-deficient mouse cells. Nucl. Acids Res. 27, 3964–3969 (1999).
Bailey, S., Cornforth, M., Kurimasa, A., Chen, D. & Goodwin, E. Strand-specific postreplicative processing of mammalian telomeres. Science 293, 2462–2465 (2001).
Parenteau, J. & Wellinger, R. J. Accumulation of single-stranded DNA and destabilization of telomeric repeats in yeast mutant strains carrying a deletion of RAD27. Mol. Cell. Biol. 19, 4143–4152 (1999). References 15 and 16 show that telomeres that are replicated by lagging-strand versus leading-strand polymerases are processed differently.
Cimino-Reale, G., Pascale, E., Alvino, E., Starace, G. & D'Ambrosio, E. Long telomeric C-rich 5′-tails in human replicating cells. J. Biol. Chem. 278, 2136–2140 (2003).
Lingner, J. et al. Reverse transcriptase motifs in the catalytic subunit of telomerase. Science 276, 561–567 (1997).
Nakamura, T. M. et al. Telomerase catalytic subunit homologs from fission yeast and human. Science 277, 955–959 (1997).
Meyerson, M. et al. hEST2, the putative human telomerase catalytic subunit gene, is up-regulated in tumor cells and during immortalization. Cell 90, 785–795 (1997).
Singer, M. S. & Gottschling, D. E. TLC1, the template RNA component of the Saccharomyces cerevisiae telomerase. Science 266, 404–409 (1994).
Feng, J. et al. The RNA component of human telomerase. Science 269, 1236–1241 (1995).
Chakhparonian, M. & Wellinger, R. J. Telomere maintenance and DNA replication: how closely are these two connected? Trends Genet. 19, 439–446 (2003).
Conrad, M. N., Wright, J. H., Wolf, A. J. & Zakian, V. A. RAP1 protein interacts with yeast telomeres in vivo: overproduction alters telomere structure and decreases chromosome stability. Cell 63, 739–750 (1990).
Chong, L. et al. A human telomeric protein. Science 270, 1663–1667 (1995).
Broccoli, D., Smogorzewska, A., Chong, L. & de Lange, T. Human telomeres contain two distinct Myb-related proteins, TRF1 and TRF2. Nature Genet. 17, 231–235 (1997).
Tsukamoto, Y., Taggart, A. K. P. & Zakian, V. A. The role of the Mre11-Rad50-Xrs2 complex in telomerase-mediated lengthening of Saccharomyces cerevisiae telomeres. Curr. Biol. 11, 1328–1335 (2001).
Baumann, P. & Cech, T. R. Pot1, the putative telomere end-binding protein in fission yeast and humans. Science 292, 1171–1175 (2001).
Baumann, P., Podell, E. & Cech, T. R. Human Pot1 (protection of telomeres) protein: cytolocalization, gene structure, and alternative splicing. Mol. Cell. Biol. 22, 8079–8087 (2002).
Griffith, J. D. et al. Mammalian telomeres end in a large duplex loop. Cell 97, 503–514 (1999). Shows that mammalian telomeres form t-loops.
Strahl-Bolsinger, S., Hecht, A., Luo, K. & Grunstein, M. SIR2 and SIR4 interactions differ in core and extended telomeric heterochromatin in yeast. Genes Dev. 11, 83–93 (1997).
de Bruin, D., Zaman, Z., Liberatore, R. A. & Ptashne, M. Telomere looping permits gene activation by a downstream UAS in yeast. Nature 409, 109–113 (2001).
Zhou, J. -Q., Monson, E. M., Teng, S. -C., Schulz, V. P. & Zakian, V. A. The Pif1p helicase, a catalytic inhibitor of telomerase lengthening of yeast telomeres. Science 289, 771–774 (2000). Shows that the yeast Pif1 DNA helicase is a catalytic inhibitor of telomerase and describes a human homologue of Pif1.
Myung, K., Chen, C. & Kolodner, R. D. Multiple pathways cooperate in the suppression of genome instability in Saccharomyces cerevisiae. Nature 411, 1073–1076 (2001). Shows that the Pif1 DNA helicase suppresses gross chromosomal rearrangements by suppressing telomere addition.
Kyo, S. & Inoue, M. Complex regulatory mechanisms of telomerase activity in normal and cancer cells: how can we apply them for cancer therapy? Oncogene 21, 688–697 (2002).
Cong, Y. S., Wright, W. E. & Shay, J. W. Human telomerase and its regulation. Microbiol. Mol. Biol. Rev. 66, 407–425, (2002).
Cohn, M. & Blackburn, E. H. Telomerase in yeast. Science 269, 396–400 (1995).
Lingner, J., Cech, T. R., Hughes, T. R. & Lundblad, V. Three Ever Shorter Telomere (EST) genes are dispensable for in vitro yeast telomerase activity. Proc. Natl Acad. Sci. USA 94, 11190–11195 (1997).
Beernink, H. T., Miller, K., Deshpande, A., Bucher, P. & Cooper, J. P. Telomere maintenance in fission yeast requires an Est1 ortholog. Curr. Biol. 13, 575–580 (2003).
Friedman, K. L. & Cech, T. R. Essential functions of amino-terminal domains in the yeast telomerase catalytic subunit revealed by selection for viable mutants. Genes Dev. 13, 2863–2874 (1999).
Armbruster, B. N., Banik, S. S., Guo, C., Smith, A. C. & Counter, C. M. N-terminal domains of the human telomerase catalytic subunit required for enzyme activity in vivo. Mol. Cell. Biol. 21, 7775–7786 (2001).
Counter, C. M. et al. Dissociation among in vitro telomerase activity, telomere maintenance, and cellular immortalization. Proc. Natl Acad. Sci. USA 95, 14723–14728 (1998).
Xia, J., Peng, Y., Mian, I. S. & Lue, N. F. Identification of functionally important domains in the N-terminal region of telomerase reverse transcriptase. Mol. Cell. Biol. 20, 5196–5207 (2000).
Kim, M., Xu, L. & Blackburn, E. H. Catalytically active human telomerase mutants with allele-specific biological properties. Exp. Cell Res. 288, 277–287 (2003).
Evans, S. K. & Lundblad, V. Est1 and Cdc13 as comediators of telomerase access. Science 286, 117–120 (1999).
Hughes, T. R., Evans, S. K., Weilbaecher, R. G. & Lundblad, V. The Est3 protein is a subunit of yeast telomerase. Curr. Biol. 10, 809–812 (2000).
Armbruster, B. N., Etheridge, K. T., Broccoli, D. & Counter, C. M. Putative telomere-recruiting domain in the catalytic subunit of human telomerase. Mol. Cell. Biol. 23, 3237–3246 (2003).
Wright, J. H. & Zakian, V. A. Protein-DNA interactions in soluble telosomes from Saccharomyces cerevisiae. Nucl. Acids Res. 23, 1454–1460 (1995).
Gilson, E., Roberge, M., Giraldo, R., Rhodes, D. & Gasser, S. M. Distortion of the DNA double helix by RAP1 at silencers and multiple telomeric binding sites. J. Mol. Biol. 231, 293–310 (1993).
Konig, P., Giraldo, R., Chapman, L. & Rhodes, D. The crystal structure of the DNA-binding domain of yeast RAP1 in complex with telomeric DNA. Cell 85, 125–136 (1996).
Sussel, L. & Shore, D. Separation of transcriptional activation and silencing functions of the RAP1-encoded repressor/activator protein 1: isolation of viable mutants affecting both silencing and telomere length. Proc. Natl Acad. Sci. USA 88, 7749–7753 (1991).
Kyrion, G., Boakye, K. A. & Lustig, A. J. C-terminal truncation of RAP1 results in the deregulation of telomere size, stability, and function in Saccharomyces cerevisiae. Mol. Cell. Biol. 12, 5159–5173 (1992).
Marcand, S., Gilson, E. & Shore, D. A protein-counting mechanism for telomere length regulation in yeast. Science 275, 986–990 (1997).
Ray, A. & Runge, K. The yeast telomere length counting machinery is sensitive to sequences at the telomere-nontelomere junction. Mol. Cell. Biol. 19, 31–45 (1999).
Ray, A. & Runge, K. W. The C terminus of the major yeast telomere binding protein Rap1p enhances telomere formation. Mol. Cell. Biol. 1284–1295 (1998).
Bianchi, A., Smith, S., Chong, L., Elias, P. & de Lange, T. TRF1 is a dimer and bends telomeric DNA. EMBO J. 16, 1785–1794 (1997).
Bianchi, A. et al. TRF1 binds a bipartite telomeric site with extreme spatial flexibility. EMBO J. 18, 5735–5744 (1999).
Ancelin, K. et al. Targeting assay to study the cis functions of human telomeric proteins: evidence for inhibition of telomerase by TRF1 and for activation of telomere degradation by TRF2. Mol. Cell. Biol. 22, 3474–3487 (2002).
van Steensel, B. & de Lange, T. Control of telomere length by the human telomeric protein TRF1. Nature 385, 740–743 (1997).
Smogorzewska, A. et al. Control of human telomere length by TRF1 and TRF2. Mol. Cell. Biol. 20, 1659–1668 (2000).
Sharma, G. G. et al. hTERT associates with human telomeres and enhances genomic stability and DNA repair. Oncogene 22, 131–146 (2003).
de Bruin, D., Kantrow, S. M., Liberatore, R. A. & Zakian, V. A. Telomere folding is required for the stable maintenance of telomere position effects in yeast. Mol. Cell. Biol. 20, 7991–8000 (2000).
Hardy, C. F., Sussel, L. & Shore, D. A RAP1-interacting protein involved in transcriptional silencing and telomere length regulation. Genes Dev. 6, 801–814 (1992).
Wotton, D. & Shore, D. A novel Rap1p-interacting factor, Rif2p, cooperates with Rif1p to regulate telomere length in Saccharomyces cerevisiae. Genes Dev. 11, 748–760 (1997).
Alexander, M. K. & Zakian, V. A. Rap1p telomere association is not required for mitotic stability of a C(3)TA(2) telomere in yeast. EMBO J. 22, 1688–1696 (2003).
Smith, C. D., Smith, D. L., DeRisi, J. L. & Blackburn, E. H. Telomeric protein distributions and remodeling through the cell cycle in Saccharomyces cerevisiae. Mol. Biol. Cell. 14, 556–570 (2003).
Wiley, E. & Zakian, V. A. Extra telomeres, but not internal tracts of telomeric DNA, reduce transcriptional repression at Saccharomyces telomeres. Genetics 139, 67–79 (1995).
Teng, S. -C., Chang, J., McCowan, B. & Zakian, V. A. Telomerase-independent lengthening of yeast telomeres occurs by an abrupt Rad50p-dependent, Rif-inhibited recombinational process. Mol. Cell 6, 947–952 (2000).
Zhou, X. Z. & Lu, K. P. The Pin2/TRF1-interacting protein PinX1 is a potent telomerase inhibitor. Cell 107, 347–359 (2001).
Kim, S. H., Kaminker, P. & Campisi, J. TIN2, a new regulator of telomere length in human cells. Nature Genet. 23, 405–412 (1999).
Smith, S., Giriat, I., Schmitt, A. & de Lange, T. Tankyrase, a poly(ADP-ribose) polymerase at human telomeres. Science 282, 1484–1487 (1998). Shows that tankyrase might affect TRF1 and telomeres in humans through its PARP activity.
Hsu, H. L. et al. Ku acts in a unique way at the mammalian telomere to prevent end joining. Genes Dev. 14, 2807–2812 (2000).
Guglielmi, B. & Werner, M. The yeast homolog of human PinX1 is involved in rRNA and small nucleolar RNA maturation, not in telomere elongation inhibition. J. Biol. Chem. 277, 35712–35719 (2002).
Smith, S. & de Lange, T. Tankyrase promotes telomere elongation in human cells. Curr. Biol. 10, 1299–1302 (2000).
Chang, W., Dynek, J. N. & Smith, S. TRF1 is degraded by ubiquitin-mediated proteolysis after release from telomeres. Genes Dev. 17, 1328–1333 (2003). Shows that tankyrase 1 induces degradation of TRF1 by the proteasome.
Kaminker, P. G. et al. TANK2, a new TRF1-associated poly(ADP-ribose) polymerase, causes rapid induction of cell death upon overexpression. J. Biol. Chem. 276, 35891–35899 (2001).
Cook, B. D., Dynek, J. N., Chang, W., Shostak, G. & Smith, S. Role for the related poly(ADP-ribose) polymerases tankyrase 1 and 2 at human telomeres. Mol. Cell. Biol. 22, 332–342 (2002).
Li, B., Oestreich, S. & de Lange, T. Identification of human Rap1: implications for telomere evolution. Cell 101, 471–483 (2000).
Opresko, P. L. et al. Telomere-binding protein TRF2 binds to and stimulates the Werner and Bloom syndrome helicases. J. Biol. Chem. 277, 41110–41119 (2002).
Zhu, X. D., Kuster, B., Mann, M., Petrini, J. H. & de Lange, T. Cell-cycle-regulated association of RAD50/MRE11/NBS1 with TRF2 and human telomeres. Nature Genet. 25, 347–352 (2000).
Paull, T. T. & Gellert, M. Nbs1 potentiates ATP-driven DNA unwinding and endonuclease cleavage by the Mre11/Rad50 complex. Genes Dev. 13, 1276–1288 (1999).
Fry, M. & Loeb, L. A. Human Werner syndrome DNA helicase unwinds tetrahelical structures of the fragile X syndrome repeat sequence d(CGG)n. J. Biol. Chem. 274, 12797–12802 (1999).
Kruk, P. A., Rampino, N. J. & Bohr, V. A. DNA damage and repair in telomeres: relation to aging. Proc. Natl Acad. Sci. USA 92, 258–262 (1995).
Schulz, V. P. et al. Accelerated loss of telomeric repeats may not explain accelerated replicative decline of Werner syndrome cells. Hum. Genet. 97, 750–754 (1996).
Huang, P. et al. SGS1 is required for telomere elongation in the absence of telomerase. Curr. Biol. 11, 125–129 (2001).
Johnson, F. et al. The Saccharomyces cerevisiae WRN homolog Sgs1p participates in telomere maintenance in cells lacking telomerase. EMBO J. 20, 905–913 (2001).
Cohen, H. & Sinclair, D. Recombination-mediated lengthening of terminal telomeric repeats requires the Sgs1 DNA helicase. Proc. Natl Acad. Sci. USA 98, 3174–3179 (2001).
Lin, J. J. & Zakian, V. A. The Saccharomyces CDC13 protein is a single-strand TG1–3 telomeric DNA binding protein in vitro that affects telomere behavior in vivo. Proc. Natl Acad. Sci. USA 93, 13760–13765 (1996).
Nugent, C. I., Hughes, T. R., Lue, N. F. & Lundblad, V. Cdc13p: a single-strand telomeric DNA-binding protein with a dual role in yeast telomere maintenance. Science 274, 249–252 (1996).
Pennock, E., Buckley, K. & Lundblad, V. Cdc13 delivers separate complexes to the telomere for end protection and replication. Cell 104, 387–396 (2001).
Qi, H. & Zakian, V. A. The Saccharomyces telomere-binding protein Cdc13p interacts with both the catalytic subunit of DNA polymerase α and the telomerase-associated Est1 protein. Genes Dev. 14, 1777–1778 (2000).
Lin, J. -J. & Zakian, V. A. An in vitro assay for Saccharomyces telomerase requires EST1. Cell 81, 1127–1135 (1995).
Livengood, A. J., Zaug, A. J. & Cech, T. R. Essential regions of Saccharomyces cerevisiae telomerase RNA: separate elements for Est1p and Est2p interaction. Mol. Cell. Biol. 22, 2366–2374 (2002).
Zhou, J., Hidaka, K. & Futcher, B. The Est1 subunit of yeast telomerase binds the Tlc1 telomerase RNA. Mol. Cell. Biol. 20, 1947–1955 (2000).
Virta-Pearlman, V., Morris, D. K. & Lundblad, V. Est1 has the properties of a single-stranded telomere end-binding protein. Genes Dev. 10, 3094–3104 (1996).
Marcand, S., Brevet, V., Mann, C. & Gilson, E. Cell cycle restriction of telomere elongation. Curr. Biol. 10, 487–490 (2000).
Taggart, A. K. P. & Zakian, V. A. Telomerase: what are the Est proteins doing? Curr. Opin. Cell Biol. 3, 275–280 (2003).
Stellwagen, A. E., Haimberger, Z. W., Veatch, J. R. & Gottschling, D. E. Ku interacts with telomerase RNA to promote telomere addition at native and broken chromosome ends. Genes Dev. 17, 2384–2395 (2003). Shows a physical interaction between yeast Ku and the telomerase RNA.
Chandra, A., Hughes, T. R., Nugent, C. I. & Lundblad, V. Cdc13 both positively and negatively regulates telomere replication. Genes Dev. 15, 404–414 (2001).
Reichenbach, P. et al. A human homolog of yeast est1 associates with telomerase and uncaps chromosome ends when overexpressed. Curr. Biol. 13, 568–574 (2003).
Snow, B. E. et al. Functional conservation of the telomerase protein est1p in humans. Curr. Biol. 13, 698–704 (2003).
Singh, S. M., Steinberg-Neifach, O., Mian, I. S. & Lue, N. F. Analysis of telomerase in Candida albicans: potential role in telomere end protection. Eukaryot. Cell 1, 967–977 (2002). References 100–102, as well as reference 39, describe Est1 homologues in humans and in other yeasts.
Gottschling, D. E. & Zakian, V. A. Telomere proteins: specific recognition and protection of the natural termini of Oxytricha macronuclear DNA. Cell 47, 195–205 (1986).
Mitton-Fry, R. M., Anderson, E. M., Hughes, T. R., Lundblad, V. & Wuttke, D. S. Conserved structure for single-stranded telomeric DNA recognition. Science 296, 145–147 (2002).
Horvath, M., Schweiker, V., Bevilacqua, J., Ruggles, J. & Schultz, S. Crystal structure of the Oxytricha nova telomere end binding protein complexed with single strand DNA. Cell 95, 963–974 (1998).
Loayza, D. & de Lange, T. POT1 as a terminal transducer of TRF1 telomere length control. Nature 424, 1013–1018 (2003).
Colgin, L. M. Human POT1 facilitates telomere elongation by telomerase. Curr. Biol. 13, 942–946 (2003). References 106 and 107 examine the effects of POT1 on human telomeres.
Sandell, L. L. & Zakian, V. A. Loss of a yeast telomere: arrest, recovery and chromosome loss. Cell 75, 729–739 (1993).
Karlseder, J., Broccoli, D., Dai, Y., Hardy, S. & de Lange, T. p53- and ATM-dependent apoptosis induced by telomeres lacking TRF2. Science 283, 1321–1325 (1999).
Liti, G. & Louis, E. NEJ1 prevents NHEJ-dependent telomere fusions in yeast without telomerase. Mol. Cell 11, 1373–1378 (2003). Shows that Nej1, in contrast to its role in promoting double-strand DNA break repair, protects telomeres from end fusions in telomerase-negative cells.
Jones, J. M., Gellert, M. & Yang, W. A Ku bridge over broken DNA. Structure 9, 881–884 (2001).
Gravel, S., Larrivee, M., Labrecque, P. & Wellinger, R. J. Yeast Ku as a regulator of chromosomal DNA end structure. Science 280, 741–744 (1998).
Porter, S. E., Greenwell, P. W., Ritchie, K. B. & Petes, T. D. The DNA-binding protein Hdf1p (a putative Ku homologue) is required for maintaining normal telomere length in Saccharomyces cerevisiae. Nucl. Acids Res. 24, 582–585 (1996).
Polotnianka, R. M., Li, J. & Lustig, A. J. The yeast Ku heterodimer is essential for protection of the telomere against nucleolytic and recombinational activities. Curr. Biol. 8, 831–834 (1998).
Boulton, S. J. & Jackson, S. P. Identification of a Saccharomyces cerevisiae Ku80 homologue: roles in DNA double strand break rejoining and in telomeric maintenance. Nucl. Acids Res. 24, 4639–4648 (1996).
Laroche, T. et al. Mutation of yeast Ku genes disrupts the subnuclear organization of telomeres. Curr. Biol. 8, 653–656 (1998).
Nugent, C. I. et al. Telomere maintenance is dependent on activities required for end repair of double-strand breaks. Curr. Biol. 8, 657–660 (1998).
Peterson, S. E. et al. The function of a stem-loop in telomerase RNA is linked to the DNA repair protein Ku. Nature Genet. 27, 64–67 (2001).
Grandin, N., Damon, C. & Charbonneau, M. Cdc13 cooperates with the yeast Ku proteins and Stn1 to regulate telomerase recruitment. Mol. Cell. Biol. 22, 8397–8408 (2000).
Hsu, H. L., Gilley, D., Blackburn, E. H. & Chen, D. J. Ku is associated with the telomere in mammals. Proc. Natl Acad. Sci. USA 96, 12454–12458 (1999).
d'Adda di Fagagna, F. et al. Effects of DNA nonhomologous end-joining factors on telomere length and chromosomal stability in mammalian cells. Curr. Biol. 11, 1192–1196 (2001).
Chai, W., Ford, L. P., Lenertz, L., Wright, W. E. & Shay, J. W. Human Ku70/80 associates physically with telomerase through interaction with hTERT. J. Biol. Chem. 277, 47242–47247 (2002).
Bailey, S. M. et al. DNA double-strand break repair proteins are required to cap the ends of mammalian chromosomes. Proc. Natl Acad. Sci. USA 96, 14899–14904 (1999).
Samper, E., Goytisolo, F. A., Slijepcevic, P., van Buul, P. P. & Blasco, M. A. Mammalian Ku86 protein prevents telomeric fusions independently of the length of TTAGGG repeats and the G-strand overhang. EMBO Rep. 1, 244–252 (2000).
Espejel, S. et al. Mammalian Ku86 mediates chromosomal fusions and apoptosis caused by critically short telomeres. EMBO J. 21, 2207–2219 (2002).
D'Amours, D. & Jackson, S. P. The mre11 complex: at the crossroads of DNA repair and checkpoint signalling. Nature Rev. Mol. Cell Biol. 3, 317–327 (2002).
Costanzo, V. et al. Mre11 protein complex prevents double-strand break accumulation during chromosomal DNA replication. Mol. Cell 8, 137–147 (2001).
Kironmai, K. M. & Muniyappa, K. Alteration of telomeric sequences and senescence caused by mutations in RAD50 of Saccharomyces cerevisiae. Genes Cells 2, 443–455 (1997).
Boulton, S. J. & Jackson, S. P. Components of the Ku-dependent non-homologous end-joining pathway are involved in telomeric length maintenance and telomeric silencing. EMBO J. 17, 1819–1828 (1998).
Ritchie, K. B. & Petes, T. D. The Mre11p/Rad50p/Xrs2p complex and the Tel1p function in a single pathway for telomere maintenance in yeast. Genetics 155, 475–479 (2000).
Diede, S. J. & Gottschling, D. E. Exonuclease activity is required for sequence addition and Cdc13p loading at a de novo telomere. Curr. Biol. 11, 1336–1340 (2001).
Moreau, S., Ferguson, J. R. & Symington, L. S. The nuclease activity of Mre11 is required for meiosis but not for mating type switching, end joining, or telomere maintenance. Mol. Cell. Biol. 19, 556–566 (1999).
Carney, J. P. et al. The hMre11/hRad50 protein complex and Nijmegen breakage syndrome: linkage of double-strand break repair to the cellular DNA damage response. Cell 93, 477–486 (1998).
Varon, R. et al. Nibrin, a novel DNA double-strand break repair protein, is mutated in Nijmegen breakage syndrome. Cell 93, 467–476 (1998).
Kramer, K. M., Brock, J., Bloom, K. & Moore, K. Two different types of double-strand breaks in Saccharomyces cerevisiae are repaired by similar RAD52-independent, nonhomologous recombination events. Mol. Cell. Biol. 14, 1293–1301 (1994).
Mangahas, J. L., Alexander, M. K., Sandell, L. L. & Zakian, V. A. Repair of chromosome ends after telomere loss in Saccharomyces. Mol. Biol. Cell 12, 4078–4089 (2001).
Schulz, V. P. & Zakian, V. A. The Saccharomyces PIF1 DNA helicase inhibits telomere elongation and de novo telomere formation. Cell 76, 145–155 (1994).
Mitchell, J. R., Cheng, J. & Collins, K. A box H/ACA small nucleolar RNA-like domain at the human telomerase RNA 3′ end. Mol. Cell. Biol. 19, 567–576 (1999).
Lukowiak, A. A., Narayanan, A., Li, Z. H., Terns, R. M. & Terns, M. P. The snoRNA domain of vertebrate telomerase RNA functions to localize the RNA within the nucleus. RNA 7, 1833–1844 (2001).
Etheridge, K. T. et al. The nucleolar localization domain of the catalytic subunit of human telomerase. J. Biol. Chem. 277, 24764–24770 (2002).
Yang, Y., Chen, Y., Zhang, C., Huang, H. & Weissman, S. M. Nucleolar localization of hTERT protein is associated with telomerase function. Exp. Cell Res. 277, 201–209 (2002).
Mitchell, J. R., Wood, E. & Collins, K. A telomerase component is defective in the human disease dyskeratosis congenita. Nature 402, 551–555 (1999). Shows that dyskerin is associated with human telomerase RNA and that dyskeratosis congenita cells have lower levels of telomerase RNA and activity.
Dragon, F., Pogacic, V. & Filipowicz, W. In vitro assembly of human H/ACA small nucleolar RNPs reveals unique features of U17 and telomerase RNAs. Mol. Cell. Biol. 20, 3037–3048 (2000).
Pogacic, V., Dragon, F. & Filipowicz, W. Human H/ACA small nucleolar RNPs and telomerase share evolutionarily conserved proteins NHP2 and NOP10. Mol. Cell. Biol. 20, 9028–9040 (2000).
Dez, C. et al. Stable expression in yeast of the mature form of human telomerase RNA depends on its association with the box H/ACA small nucleolar RNP proteins Cbf5p, Nhp2p and Nop10p. Nucl. Acids Res 29, 598–603 (2001).
Seto, A. G., Zaug, A. J., Sobel, S. G., Wolin, S. L. & Cech, T. R. Saccharomyces cerevisiae telomerase is an Sm small nuclear ribonucleoprotein particle. Nature 401, 177–180 (1999). Shows that the yeast telomerase RNA has structural features in common with small nuclear ribonucleoprotein particles.
Teixeira, M. T., Forstemann, K., Gasser, S. M. & Lingner, J. Intracellular trafficking of yeast telomerase components. EMBO Rep. 3, 652–659 (2002).
Wong, J. M., Kusdra, L. & Collins, K. Subnuclear shuttling of human telomerase induced by transformation and DNA damage. Nature Cell Biol. 4, 731–736 (2002). Shows that the intranuclear localization of active human telomerase depends on cell-cycle stage, transformation and DNA damage.
Broccoli, D., Young, J. W. & de Lange, T. Telomerase activity in normal and malignant hematopoietic cells. Proc. Natl Acad. Sci. USA 92, 9082–9086 (1995).
Hiyama, K. et al. Activation of telomerase in human lymphocytes and hematopoietic progenitor cells. J. Immunol. 155, 3711–3715 (1995).
Liu, K. et al. Constitutive and regulated expression of telomerase reverse transcriptase (hTERT) in human lymphocytes. Proc. Natl Acad. Sci. USA 96, 5147–5152 (1999).
Liu, K., Hodes, R. J. & Weng, N. Cutting edge: telomerase activation in human T lymphocytes does not require increase in telomerase reverse transcriptase (hTERT) protein but is associated with hTERT phosphorylation and nuclear translocation. J. Immunol. 166, 4826–4830 (2001).
Akiyama, M. et al. Nuclear factor-κB p65 mediates tumor necrosis factor α-induced nuclear translocation of telomerase reverse transcriptase protein. Cancer Res. 63, 18–21 (2003).
Haendeler, J., Hoffmann, J., Brandes, R. P., Zeiher, A. M. & Dimmeler, S. Hydrogen peroxide triggers nuclear export of telomerase reverse transcriptase via Src kinase family-dependent phosphorylation of tyrosine 707. Mol. Cell. Biol. 23, 4598–4610 (2003).
Spellman, P. T. et al. Comprehensive identification of cell cycle-regulated genes of the yeast Saccharomyces cerevisiae by microarray hybridization. Mol. Biol. Cell 9, 3273–3297 (1998).
Marcand, S., Wotton, D., Gilson, E. & Shore, D. Rap1p and telomere length regulation in yeast. Ciba Found. Symp. 211, 76–93 (1997).
Marcand, S., Brevet, V. & Gilson, E. Progressive cis-inhibition of telomerase upon telomere elongation. EMBO J. 18, 3509–3519 (1999).
Brevet, V. et al. The number of vertebrate repeats can be regulated at yeast telomeres by Rap1-independent mechanisms. EMBO J. 22, 1697–1706 (2003).
Wellinger, R. J., Wolf, A. J. & Zakian, V. A. Origin activation and formation of single-strand TG1–3 tails occur sequentially in late S phase on a yeast linear plasmid. Mol. Cell. Biol. 13, 4057–4065 (1993).
Acknowledgements
We thank T. Fisher and M. Sabourin for critical reading of the manuscript, S. Schnakenberg for help with the figures and B. Lenzmeier and J. Bessler for thoughtful discussions. Work in the Zakian lab is supported by grants from the National Institutes of Health (NIH). M.K.M. is supported by the Damon Runyon Cancer Research Foundation. L.R.V. was funded in part by the Helen Hay Whitney Foundation and by the NIH.
Author information
Authors and Affiliations
Corresponding author
Glossary
- TELOMERASE
-
Specialized ribonucleoprotein, the catalytic core of which is composed of an RNA subunit and a reverse transcriptase subunit that facilitates the replication of linear chromosome ends or telomeres. The RNA subunit contains the template for sequence addition (3′ CACACACCCACACCAC 5′ in S. cerevisiae and 3′ CAAUCCCAAUC 5′ in humans).
- OKAZAKI FRAGMENT
-
Short DNA fragment that is formed during DNA replication due to the discontinuous synthesis of the lagging strand. Okazaki fragments are initiated with an 8–12-base stretch of RNA.
- REVERSE TRANSCRIPTASE
-
An enzyme that copies single-stranded RNA into a single-stranded DNA.
- T-LOOP
-
Duplex telomeric loop that results from invasion of 3′ G-rich overhangs into duplex telomeric regions. T-loops have been found on eukaryotic telomeres and range in size from 0.3 kb to >30 kb.
- MYB-LIKE DOMAIN
-
Highly conserved DNA-binding domain that is composed of tandem repeats of a helix-turn-helix motif.
- KU PROTEIN
-
A highly conserved heterodimer consisting of ∼70- and ∼80-kDa subunits that binds at double-stranded DNA breaks and at telomeres and is important for DNA repair and telomere functions.
- ADENOSINE-DIPHOSPHATE-RIBOSE POLYMERASE
-
An enzyme that uses NAD+ as a substrate to produce peptidyl-glutamic acid poly-ADP-ribose-modified proteins. This modification regulates various processes such as differentiation, proliferation and the repair of single-stranded DNA breaks.
- PROTEASOME
-
A multi-protein complex that degrades proteins marked for destruction by ubiquitylation.
- UBIQUITYLATION
-
The addition of the small evolutionarily conserved polypeptide, ubiquitin, to proteins that are targeted for destruction.
- HELICASE
-
An enzyme that uses the energy of ATP hydrolysis to unwind duplex nucleic acids
- EXONUCLEASE
-
An enzyme that hydrolyzes ester linkages within nucleic acids. They can remove nucleotides from either the 3′ or 5′ end of the molecule.
- MRE11 COMPLEX
-
A highly conserved protein complex that is composed of MRE11, RAD50 and NBS1 (in humans) and Mre11, Rad50 and Xrs2 (also known as the MRX complex, in yeast) and that is involved in detection, signalling and repair of DNA damage. In humans, mutations in ATM, MRE11 and NBS1 are associated with increased predisposition to cancer and cause ataxia–telangiectasia (AT), AT-like disorder (AT-LD) and Nijmegen breakage syndrome (NBS), respectively.
- WRN HELICASE/EXONUCLEASE
-
WRN is a member of the RecQ helicase subfamily and has 3′→5′ helicase and 3′→5′ exonuclease activities. Mutations in human WRN result in Werner syndrome, an autosomal-recessive disease that is characterized by premature ageing, chromosome instability and telomere–telomere fusions.
- NON-HOMOLOGOUS END JOINING
-
(NHEJ). A double-stranded DNA break (DSB) repair pathway that involves the largely homology-independent ligation of two DNA ends.
- INTRA-S-PHASE CHECKPOINT
-
Pathway that responds to stalled replication forks and other DNA damage during S phase by activating the ATM-like kinases Mec1 and Rad53. Checkpoint activation results in delayed S-phase progression and inhibits spindle elongation.
- SMALL NUCLEOLAR RNA
-
(snoRNA). Stable RNA species in the eukaryotic nucleolus, most of which function to target the major nucleotide modifications in ribosomal RNA or are involved in rRNA processing. There are three classes of snoRNAs: box H/ACA snoRNAs, box C/D snoRNAs and 7-2/MRP snoRNAs.
Rights and permissions
About this article
Cite this article
Vega, L., Mateyak, M. & Zakian, V. Getting to the end: telomerase access in yeast and humans. Nat Rev Mol Cell Biol 4, 948–959 (2003). https://doi.org/10.1038/nrm1256
Issue Date:
DOI: https://doi.org/10.1038/nrm1256
This article is cited by
-
Topoisomerase II inhibition suppresses the proliferation of telomerase-negative cancers
Cellular and Molecular Life Sciences (2015)
-
PP2A and Aurora differentially modify Cdc13 to promote telomerase release from telomeres at G2/M phase
Nature Communications (2014)
-
Design and synthesis of macrocyclic polyoxazoles
Chemical Research in Chinese Universities (2014)
-
Human single-stranded DNA binding proteins are essential for maintaining genomic stability
BMC Molecular Biology (2013)
-
The TPR-containing domain within Est1 homologs exhibits species-specific roles in telomerase interaction and telomere length homeostasis
BMC Molecular Biology (2011)