Abstract
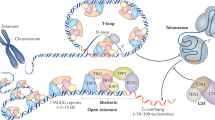
Telomeres arose from the need to stabilize natural chromosome ends, resulting in terminal chromatin structures with specific protective functions. Their constituent proteins also execute general functions within heterochromatin, mediating late replication and facilitating fork progression. Emerging insights into the mechanisms governing heterochromatin replication suggest telomeres and heterochromatin act in concert during development and aging. They also suggest a common evolutionary origin for these two chromosome regions that arose during eukaryogenesis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Giraud-Panis, M. J. et al. One identity or more for telomeres? Front. Oncol. 3, 48 (2013).
Heitz, E. Das Heterochromatin der Moose. Jarhb. Wiss. Bot. 69, 762–818 (1928).
Nishibuchi, G. & Déjardin, J. The molecular basis of the organization of repetitive DNA-containing constitutive heterochromatin in mammals. Chromosome Res. 25, 77–87 (2017).
Müller, H. J. The remaking of chromosomes. The Collecting Net 13, 182–198 (1938).
Gottschling, D. E., Aparicio, O. M., Billington, B. L. & Zakian, V. A. Position effect at S. cerevisiae telomeres: reversible repression of Pol II transcription. Cell 63, 751–762 (1990). This paper reveals that proximity to telomeres can cause repression of gene expression.
Gasser, S. M. & Cockell, M. M. The molecular biology of the SIR proteins. Gene 279, 1–16 (2001).
Maillet, L. et al. Evidence for silencing compartments within the yeast nucleus: a role for telomere proximity and Sir protein concentration in silencer-mediated repression. Genes Dev. 10, 1796–1811 (1996).
Palladino, F. et al. SIR3 and SIR4 proteins are required for the positioning and integrity of yeast telomeres. Cell 75, 543–555 (1993).
Gauchier, M. et al. SETDB1-dependent heterochromatin stimulates alternative lengthening of telomeres. Sci. Adv. 5, eaav3673 (2019).
Cubiles, M. D. et al. Epigenetic features of human telomeres. Nucleic Acids Res. 46, 2347–2355 (2018).
Jain, D., Hebden, A. K., Nakamura, T. M., Miller, K. M. & Cooper, J. P. HAATI survivors replace canonical telomeres with blocks of generic heterochromatin. Nature 467, 223–227 (2010). This paper provides evidence that fission yeast use heterochromatic blocks to replace telomeres and provide end protection in the absence of telomerase.
Fanti, L., Giovinazzo, G., Berloco, M. & Pimpinelli, S. The heterochromatin protein 1 prevents telomere fusions in Drosophila. Mol. Cell 2, 527–538 (1998).
Zhao, P. A., Rivera-Mulia, J. C. & Gilbert, D. M. DNA Replication. Advances in Experimental Medicine and Biology Vol. 1042 (Eds Masai, H. & Foiani, M.) Ch. 6 (Springer, 2017).
Cooper, J. P., Nimmo, E. R., Allshire, R. C. & Cech, T. R. Regulation of telomere length and function by a Myb-domain protein in fission yeast. Nature 385, 744–747 (1997).
Tazumi, A. et al. Telomere-binding protein Taz1 controls global replication timing through its localization near late replication origins in fission yeast. Genes Dev. 26, 2050–2062 (2012). This paper shows that Taz1 controls the replication timing of a subset of late origins by directly binding to telomeric repeats found proximal to origins.
Zofall, M., Smith, D. R., Mizuguchi, T., Dhakshnamoorthy, J. & Grewal, S. I. S. Taz1-shelterin promotes facultative heterochromatin assembly at chromosome-internal sites containing late replication origins. Mol. Cell 62, 862–874 (2016). This work shows that shelterin subunits assemble facultative heterochromatin at internal chromosomal sites that contain late-replicating origins.
Hardy, C. F., Balderes, D. & Shore, D. Dissection of a carboxy-terminal region of the yeast regulatory protein RAP1 with effects on both transcriptional activation and silencing. Mol. Cell. Biol. 12, 1209–1217 (1992).
Kedziora, S. et al. Rif1 acts through protein phosphatase 1 but independent of replication timing to suppress telomere extension in budding yeast. Nucleic Acids Res. 46, 3993–4003 (2018).
Miller, K. M., Ferreira, M. G. & Cooper, J. P. Taz1, Rap1 and Rif1 act both interdependently and independently to maintain telomeres. EMBO J. 24, 3128–3135 (2005).
Dan, J. et al. Rif1 maintains telomere length homeostasis of ESCs by mediating heterochromatin silencing. Dev. Cell 29, 7–19 (2014).
Hayano, M. et al. Rif1 is a global regulator of timing of replication origin firing in fission yeast. Genes Dev. 26, 137–150 (2012). This paper reveals that global replication timing in S. pombe is controlled by Rif1 in a Taz1-independent fashion.
Hiraga, S. et al. Rif1 controls DNA replication by directing protein phosphatase 1 to reverse Cdc7-mediated phosphorylation of the MCM complex. Genes Dev. 28, 372–383 (2014). This paper shows that the budding yeast protein Rif1 controls DNA replication genome-wide and that Rif1 exerts this control through PP1-mediated dephosphorylation of the MCM complex early in the cell cycle.
Mattarocci, S. et al. Rif1 controls DNA replication timing in yeast through the PP1 phosphatase Glc7. Cell Rep. 7, 62–69 (2014). This work shows Rif1 is linked to the negative regulation of replication-origin firing through recruitment of the Glc7 phosphatase.
Dave, A., Cooley, C., Garg, M. & Bianchi, A. Protein phosphatase 1 recruitment by Rif1 regulates DNA replication origin firing by counteracting DDK activity. Cell Rep. 7, 53–61 (2014). This paper shows that Rif1 regulates replication timing in budding yeast through its interaction with PP1 phosphatases.
Seller, C. A. & O'Farrell, P. H. Rif1 prolongs the embryonic S phase at the Drosophila mid-blastula transition. PLoS Biol. 16, e2005687 (2018). This study demonstrates that Rif1 plays a direct role during development by delaying the replication of satellite sequences.
Cornacchia, D. et al. Mouse Rif1 is a key regulator of the replication-timing programme in mammalian cells. EMBO J. 31, 3678–3690 (2012).
Yamazaki, S. et al. Rif1 regulates the replication timing domains on the human genome. EMBO J. 31, 3667–3677 (2012).
Sukackaite, R. et al. Mouse Rif1 is a regulatory subunit of protein phosphatase 1 (PP1). Sci. Rep. 7, 2119 (2017).
Alver, R. C., Chadha, G. S., Gillespie, P. J. & Blow, J. J. Reversal of DDK-mediated MCM phosphorylation by Rif1-PP1 regulates replication initiation and replisome stability independently of ATR/Chk1. Cell Rep. 18, 2508–2520 (2017).
Foti, R. et al. Nuclear architecture organized by Rif1 underpins the replication-timing program. Mol. Cell 61, 260–273 (2016). This study identifies Rif1 as the molecular link between nuclear architecture and the establishment of replication timing in mammals.
Toteva, T. et al. Establishment of expression-state boundaries by Rif1 and Taz1 in fission yeast. Proc. Natl Acad. Sci. USA 114, 1093–1098 (2017).
Ogawa, S. et al. Shelterin promotes tethering of late replication origins to telomeres for replication-timing control. EMBO J. 37, e98997 (2018).
Hiraga, S. I. et al. Budding yeast Rif1 binds to replication origins and protects DNA at blocked replication forks. EMBO Rep. 20, e48152 (2019).
Moriyama, K., Yoshizawa-Sugata, N. & Masai, H. Oligomer formation and G-quadruplex binding by purified murine Rif1 protein, a key organizer of higher-order chromatin architecture. J. Biol. Chem. 293, 3607–3624 (2018).
Kanoh, Y. et al. Rif1 binds to G quadruplexes and suppresses replication over long distances. Nat. Struct. Mol. Biol. 22, 889–897 (2015).
Masai, H. et al. Rif1 promotes association of G-quadruplex (G4) by its specific G4 binding and oligomerization activities. Sci. Rep. 9, 8618 (2019).
Hasegawa, Y., Yamamoto, M., Miyamori, J. & Kanoh, J. Telomere DNA length-dependent regulation of DNA replication timing at internal late replication origins. Sci. Rep. 9, 9946 (2019).
Hafner, L. et al. Rif1 binding and control of chromosome-internal DNA replication origins is limited by telomere sequestration. Cell Rep. 23, 983–992 (2018). This work shows that global replication dynamics in budding yeast are controlled by telomeric sequestration of Rif1 by Rap1.
Mizuguchi, T. et al. Shelterin components mediate genome reorganization in response to replication stress. Proc. Natl Acad. Sci. USA 114, 5479–5484 (2017).
Mendez-Bermudez, A. et al. Genome-wide control of heterochromatin replication by the telomere capping protein TRF2. Mol. Cell 70, 449–461.e5 (2018). This paper provides evidence that TRF2 is involved in pericentromeric heterochromatin replication through the recruitment of the helicase RTEL1.
Postow, L. et al. Positive torsional strain causes the formation of a four-way junction at replication forks. J. Biol. Chem. 276, 2790–2796 (2001).
Rickman, K. & Smogorzewska, A. Advances in understanding DNA processing and protection at stalled replication forks. J. Cell Biol. 218, 1096–1107 (2019).
Miller, K. M., Rog, O. & Cooper, J. P. Semi-conservative DNA replication through telomeres requires Taz1. Nature 440, 824–828 (2006).
Zimmermann, M., Kibe, T., Kabir, S. & de Lange, T. TRF1 negotiates TTAGGG repeat-associated replication problems by recruiting the BLM helicase and the TPP1/POT1 repressor of ATR signaling. Genes Dev. 28, 2477–2491 (2014).
Ye, J. et al. TRF2 and Apollo cooperate with topoisomerase 2α to protect human telomeres from replicative damage. Cell 142, 230–242 (2010).
Li, F. et al. The BUB3-BUB1 complex promotes telomere DNA replication. Mol. Cell 70, 395–407.e4 (2018).
Simonet, T. et al. The human TTAGGG repeat factors 1 and 2 bind to a subset of interstitial telomeric sequences and satellite repeats. Cell Res. 21, 1028–1038 (2011).
Fouche, N. et al. The basic domain of TRF2 directs binding to DNA junctions irrespective of the presence of TTAGGG repeats. J. Biol. Chem. 281, 37486–37495 (2006).
Poulet, A. et al. TRF2 promotes, remodels and protects telomeric Holliday junctions. EMBO J. 28, 641–651 (2009).
Alabert, C. et al. Nascent chromatin capture proteomics determines chromatin dynamics during DNA replication and identifies unknown fork components. Nat. Cell Biol. 16, 281–291 (2014).
Garzón, J., Ursich, S., Lopes, M., Hiraga, S. I. & Donaldson, A. D. Human RIF1-protein phosphatase 1 prevents degradation and breakage of nascent DNA on replication stalling. Cell Rep. 27, 2558–2566.e4 (2019).
Mukherjee, C. et al. RIF1 promotes replication fork protection and efficient restart to maintain genome stability. Nat. Commun. 10, 3287 (2019).
Fouché, N., Özgür, S., Roy, D. & Griffith, J. D. Replication fork regression in repetitive DNAs. Nucleic Acids Res. 34, 6044–6050 (2006).
Verdun, R. E. & Karlseder, J. The DNA damage machinery and homologous recombination pathway act consecutively to protect human telomeres. Cell 127, 709–720 (2006).
Amiard, S. et al. A topological mechanism for TRF2-enhanced strand invasion. Nat. Struct. Mol. Biol. 14, 147–154 (2007).
Sarek, G., Vannier, J. B., Panier, S., Petrini, J. H. & Boulton, S. J. TRF2 recruits RTEL1 to telomeres in S phase to promote t-loop unwinding. Mol. Cell 57, 622–635 (2015).
Jaco, I. et al. Role of mammalian Rad54 in telomere length maintenance. Mol. Cell Biol. 23, 5572–5580 (2003).
Badie, S. et al. BRCA2 acts as a RAD51 loader to facilitate telomere replication and capping. Nat. Struct. Mol. Biol. 17, 1461–1469 (2010).
Sfeir, A. & de Lange, T. Removal of shelterin reveals the telomere end-protection problem. Science 336, 593–597 (2012).
Teixeira-Silva, A. et al. The end-joining factor Ku acts in the end-resection of double strand break-free arrested replication forks. Nat. Commun. 8, 1982 (2017).
Lam, Y. C. et al. SNMIB/Apollo protects leading-strand telomeres against NHEJ-mediated repair. EMBO J. 29, 2230–2241.
Mason, J. M. et al. The SNM1B/APOLLO DNA nuclease functions in resolution of replication stress and maintenance of common fragile site stability. Hum. Mol. Genet. 22, 4901–4913 (2013).
Aggarwal, M., Sommers, J. A., Morris, C. & Brosh, R. M. Jr. Delineation of WRN helicase function with EXO1 in the replicational stress response. DNA Repair (Amst.) 9, 765–776 (2010).
Guervilly, J. H. & Gaillard, P. H. SLX4: multitasking to maintain genome stability. Crit. Rev. Biochem. Mol. Biol. 53, 475–514 (2018).
Myung, K., Chen, C. & Kolodner, R. D. Multiple pathways cooperate in the suppression of genome instability in Saccharomyces cerevisiae. Nature 411, 1073–1076 (2001).
Margalef, P. et al. Stabilization of reversed replication forks by telomerase drives telomere catastrophe. Cell 172, 439–453.e14 (2018).
Sinha, A. K. et al. Broken replication forks trigger heritable DNA breaks in the terminus of a circular chromosome. PLoS Genet. 14, e1007256 (2018).
de Lange, T. A loopy view of telomere evolution. Front. Genet. 6, 321 (2015).
Lee, G. et al. Testing the retroelement invasion hypothesis for the emergence of the ancestral eukaryotic cell. Proc. Natl Acad. Sci. USA 115, 12465–12470 (2018).
Ye, J., Renault, V. M., Jamet, K. & Gilson, E. Transcriptional outcome of telomere signalling. Nat. Rev. Genet. 15, 491–503 (2014).
Acknowledgements
This work was supported by the cross-cutting Inserm program on aging (AGEMED), the National Reseach Agency (ANR) program TELOCHROM, the Sino-French Partenariat Hubert Curien (PHC) Cai Yuanpei program and the ‘Investments for the Future’ Labex SIGNALIFE (grant reference no. ANR-11-LABX-0028-01).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Beth Moorefield was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Mendez-Bermudez, A., Giraud-Panis, MJ., Ye, J. et al. Heterochromatin replication goes hand in hand with telomere protection. Nat Struct Mol Biol 27, 313–318 (2020). https://doi.org/10.1038/s41594-020-0400-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-020-0400-1
This article is cited by
-
Biomarkers of aging
Science China Life Sciences (2023)
-
The landscape of aging
Science China Life Sciences (2022)