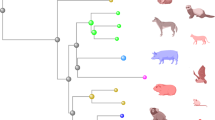
Key Points
-
Infection and immunity in humans occur in natural conditions, as opposed to experimental conditions in animal models. In our view, this accounts for most of the advantages and disadvantages of human studies, when compared with animal studies.
-
Immunocompetence can be defined as the immune status that results in self-healing infection, with or without symptoms, but of favourable outcome without any specific treatment. After exposure to the microbial world, human immunocompetence (to all microorganisms) is rare and immunodeficiency (to at least one microorganism) is, therefore, the rule rather than the exception.
-
All clinical and biological phenotypes in the course of an infectious disease result from the complex interaction between environmental (microbial and non-microbial) and host (genetic and non-genetic) factors. A forward-genetic dissection of immunity to infection is therefore possible.
-
The human model is an indispensable complement to animal studies, as it provides insight into the ecologically relevant and evolutionary selected functions of cells and molecules that are involved in immunity to infection. The forward-genetic dissection of immunity to infection in humans benefits from being an observational study of host–environment interaction in a natural ecosystem.
-
In some cases, 'wild-type' individuals are vulnerable to common infection, whereas mutants with relatively common Mendelian mutations are fully resistant and otherwise healthy. Examples include resistance to Plasmodium vivax associated with recessive mutations in Duffy antigen receptor for chemokines (DARC), resistance to HIV-1 with recessive mutations in CC-chemokine receptor 5 (CCR5) and resistance to noroviruses with recessive mutations in fucosyltransferase 2 (FUT2).
-
Studies of human genetics of infectious diseases have challenged several immunological theories and, to a lesser extent, data obtained in mice. Examples include the lack of viral diseases in children with mutations that impair antigen presentation to, or recognition by, cytotoxic CD8+ T cells, and the narrow spectrum of infections in children with mutations in the interleukin-12 (IL-12)–interferon-γ pathway or IL-1 receptor-associated kinase 4 (IRAK4).
-
The study of Mendelian 'holes' in immunity to infection has indicated the existence of 'pathogen-specific' genes in natural conditions of infection. Examples include the group of genetic defects of the terminal components of complement, associated with Neisseria infections, and epidermodysplasia verruciformis, associated with a selective susceptibility to human papillomaviruses.
-
There is a continuous spectrum between rare Mendelian vulnerability to poorly virulent microorganisms and common complex predisposition to virulent microorganisms. The elucidation of the genetic basis of common infectious diseases also sheds light on immunological mechanisms. Mendelian susceptibility to tuberculosis and complex susceptibility to leprosy have indicated the function of important immunological pathways.
Abstract
Tremendous progress has been achieved in developmental, cellular and molecular immunology in the past 20 years, largely due to studies using the mouse as a model system and the arrival of molecular genetics. Immunology is now faced with a difficult challenge. What are the functions of the individual cells and molecules in achieving immunity to infection? Renewed interest in animal models of disease has provided considerable insight in this area, but such models of infection suffer from the inherent limitation of being experimental. In humans, the complex host–environment interaction occurs in natural, as opposed to experimental, conditions. The human model is therefore an indispensable complement to animal models, as it allows an observational genetic dissection of immunity to infection.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Paul, W. E. Fundamental Imunology (Lippincott–Raven, Philadelphia, 1998).
Du Pasquier, L. The immune system of invertebrates and vertebrates. Comp. Biochem. Physiol. B Biochem. Mol. Biol. 129, 1–15 (2001).
Hoffmann, J. A. The immune response of Drosophila. Nature 426, 33–38 (2003).
Tzou, P., De Gregorio, E. & Lemaitre, B. How Drosophila combats microbial infection: a model to study innate immunity and host–pathogen interactions. Curr. Opin. Microbiol. 5, 102–110 (2002).
Fluhr, R. & Kaplan-Levy, R. N. Plant disease resistance: commonality and novelty in multicellular innate immunity. Curr. Top. Microbiol. Immunol. 270, 23–46 (2002).
Casanova, J. L., Schurr, E., Abel, L. & Skamene, E. Forward genetics of infectious diseases: immunological impact. Trends Immunol. 23, 469–472 (2002).
Buer, J. & Balling, R. Mice, microbes and models of infection. Nature Rev. Genet. 4, 195–205 (2003).
Kurz, C. L. & Ewbank, J. J. Caenorhabditis elegans: an emerging genetic model for the study of innate immunity. Nature Rev. Genet. 4, 380–390 (2003).
Casanova, J. L. & Abel, L. Genetic dissection of immunity to mycobacteria: the human model. Annu. Rev. Immunol. 20, 581–620 (2002).
Pasteur, L. Oeuvres complétes de Louis Pasteur, réunies par Pasteur Vallery-Radot (Masson et Cie, Paris, 1922–1939).
Dubos, R. J. Louis Pasteur, Free Lance of Science (Little Brown, Boston, 1950).
Abel, L. & Dessein, A. J. Genetic epidemiology of infectious diseases in humans: design of population-based studies. Emerg. Infect. Dis. 4, 593–603 (1998).
Kwiatkowski, D. Science, medicine, and the future: susceptibility to infection. BMJ 321, 1061–1065 (2000).
Burgner, D. & Levin, M. Genetic susceptibility to infectious diseases. Pediatr. Infect. Dis. J. 22, 1–6 (2003).
Bruton, O. C. Agammaglobulinemia. Pediatrics 9, 722–728 (1952). This is the first clinical description of a Mendelian primary immunodeficiency. Children with X-linked recessive Bruton's aggamaglobulinaemia lack peripheral B cells and mostly suffer from severe pyogenic bacterial infections.
Hitzig, W. H. The discovery of agammaglobulinaemia in 1952. Eur. J. Pediatr. 162, 289–304 (2003).
WHO. Primary immunodeficiency diseases: an update. Clin. Exp. Immunol. 132, 9–15 (2003).
Ochs, H., Smith C. I. E. & Puck, J. Primary Immunodeficiencies: a Molecular and Genetic Approach (Oxford Univ. Press, New York, 2003).
Fischer, A. Primary immunodeficiency diseases: an experimental model for molecular medicine. Lancet 357, 1863–1869 (2001).
Allison, A. C. Protection afforded by sickle cell trait against subtertian malarian infection. BMJ 1, 290–294 (1954). This is the first evidence for a non-Mendelian contribution of human genes to the development of an infectious disease. The haemoglobin S trait (sickle-cell disease, drapanocytosis) confers protection from Plasmodium falciparum malaria, probably accounting for the increase of this trait in endemic regions.
Weatherall, D. J. Common genetic disorders of the red cell and the 'malaria hypothesis'. Ann. Trop. Med. Parasitol. 81, 539–548 (1987).
Joliat, M. J. & Shultz, L. D. The molecular bases of spontaneous immunological mutations in the mouse and their homologous human diseases. Clin. Immunol. 101, 113–129 (2001).
Kaufmann, S. H. Bacterial and protozoal infections in genetically disrupted mice. Curr. Opin. Immunol. 6, 518–525 (1994).
van den Broek, M. F., Müller, U., Huang, S., Zinkernagel, R. M. & Aguet, M. Immune defence in mice lacking ype I and/or type II interferon receptors. Immunol. Rev. 148, 5–18 (1995).
Yap, G. S. & Sher, A. The use of germ line-mutated mice in understanding host-pathogen interactions. Cell Microbiol. 4, 627–634 (2002).
Dib, C. et al. A comprehensive genetic map of the human genome based on 5,264 microsatellites. Nature 380, 152–154 (1996).
Lander, E. S. et al. Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001).
Venter, J. C. et al. The sequence of the human genome. Science 291, 1304–1351 (2001).
Sachidanandam, R. et al. A map of human genome sequence variation containing 1.42 million single nucleotide polymorphisms. Nature 409, 928–933 (2001).
Nehls, M., Pfeifer, D., Schorpp, M., Hedrich, H. & Boehm, T. New member of the winged-helix protein family disrupted in mouse and rat nude mutations. Nature 372, 103–107 (1994).
Frank, J. et al. Exposing the human nude phenotype. Nature 398, 473–474 (1999).
Shiflett, S. L., Kaplan, J. & Ward, D. M. Chediak-Higashi syndrome: a rare disorder of lysosomes and lysosome related organelles. Pigment Cell Res. 15, 251–257 (2002).
Conley, M. E. & Howard, V. Clinical findings leading to the diagnosis of X-linked agammaglobulinemia. J. Pediatr. 141, 566–571 (2002).
Gambineri, E., Torgerson, T. R. & Ochs, H. D. Immune dysregulation, polyendocrinopathy, enteropathy, and X-linked inheritance (IPEX), a syndrome of systemic autoimmunity caused by mutations of FOXP3, a critical regulator of T-cell homeostasis. Curr. Opin. Rheumatol. 15, 430–435 (2003).
Lindenmann, J. Resistance of mice to mouse-adapted influenza A virus. Virology 16, 203–204 (1962).
Horisberger, M. A., Stäheli, P. & Haller, O. Interferon induces a unique protein in mouse cells bearing a gene for resistance to influenza virus. Proc. Natl Acad. Sci. USA 80, 1910–1914 (1983).
Staeheli, P., Haller, O., Boll, W., Lindenmann, J. & Weissmann, C. Mx protein: constitutive expression in 3T3 cells transformed with cloned Mx cDNA confers selective resistance to influenza virus. Cell 44, 147–158 (1986).
Haller, O. & Kochs, G. Interferon-induced Mx proteins: dynamin-like GTPases with antiviral activity. Traffic 3, 710–717 (2002).
Gros, P., Skamene, E. & Forget, A. Genetic control of natural resistance to Mycobacterium bovis (BCG) in mice. J. Immunol. 127, 2417–2421 (1981).
Vidal, S., Malo, D., Vogan, K., Skamene, E. & Gros, P. Natural resistance to infection with intracellular parasites: isolation of a candidate for Bcg. Cell 73, 469–485 (1993).
Buu, N., Sanchez, F. & Schurr, E. The Bcg host-resistance gene. Clin. Infect. Dis. 31 (Suppl. 3), S81–S85 (2000).
Farrar, J. D. et al. Selective loss of type I interferon-induced STAT4 activation caused by a minisatellite insertion in mouse Stat2. Nature Immunol. 1, 65–69 (2000).
Wade, C. M. et al. The mosaic structure of variation in the laboratory mouse genome. Nature 420, 574–578 (2002).
Guenet, J. L. & Bonhomme, F. Wild mice: an ever-increasing contribution to a popular mammalian model. Trends Genet. 19, 24–31 (2003).
Appleby, M. W. & Ramsdell, F. A forward-genetic approach for analysis of the immune system. Nature Rev. Immunol. 3, 463–471 (2003).
Poltorak, A. et al. Defective LPS signaling in C3H/HeJ and C57BL/10ScCr mice: mutations in Tlr4 gene. Science 282, 2085–2088 (1998).
Qureshi, S. T. et al. Endotoxin-tolerant mice have mutations in Toll-like receptor 4 (Tlr4). J. Exp. Med. 189, 615–625 (1999).
Wright, E. K. et al. Naip5 affects host susceptibility to the intracellular pathogen Legionella pneumophila. Curr. Biol. 13, 27–36 (2003).
Diez, E. et al. Birc1e is the gene within the Lgn1 locus associated with resistance to Legionella pneumophila. Nature Genet. 33, 55–60 (2003).
Perelygin, A. A. et al. Positional cloning of the murine flavivirus resistance gene. Proc. Natl Acad. Sci. USA 99, 9322–9327 (2002).
Mashimo, T. et al. A nonsense mutation in the gene encoding 2′-5′-oligoadenylate synthetase/L1 isoform is associated with West Nile virus susceptibility in laboratory mice. Proc. Natl Acad. Sci. USA 99, 11311–11316 (2002).
Lee, S. H. et al. Susceptibility to mouse cytomegalovirus is associated with deletion of an activating natural killer cell receptor of the C-type lectin superfamily. Nature Genet. 28, 42–45 (2001).
Brown, M. G. et al. Vital involvement of a natural killer cell activation receptor in resistance to viral infection. Science 292, 934–937 (2001).
Daniels, K. A. et al. Murine cytomegalovirus is regulated by a discrete subset of natural killer cells reactive with monoclonal antibody to Ly49H. J. Exp. Med. 194, 29–44 (2001).
Holmes, K. V. & Dveksler, G. S. in Cellular Receptors for Animal Viruses (ed. Wimmer, E.) 403–443 (Cold Spring Harbor Laboratory Press, New York, 1994).
Taylor, G. M., Gao, Y. & Sanders, D. A. Fv-4: identification of the defect in Env and the mechanism of resistance to ecotropic murine leukemia virus. J. Virol. 75, 11244–11248 (2001).
Shaw, M. H. et al. A natural mutation in the Tyk2 pseudokinase domain underlies altered susceptibility of B10. Q/J mice to infection and autoimmunity. Proc. Natl Acad. Sci. USA 100, 11594–11599 (2003).
Lee, S. H. et al. Maneuvering for advantage: the genetics of mouse susceptibility to virus infection. Trends Genet. 19, 447–457 (2003).
Feigin, R. D. & Cherry, J. D. Textbook of Pediatric Infectious Diseases (W. B. Saunders Company, Philadelphia, USA, 1998).
Gorbach, S. L., Barlett, J. G. & Blacklow, N. R. Infectious Diseases (W. B. Saunders Company, Philadelphia, USA, 1998).
Lecuit, M. et al. A transgenic model for listeriosis: role of internalin in crossing the intestinal barrier. Science 292, 1722–1725 (2001).
Favre, M., Ramoz, N. & Orth, G. Human papillomaviruses: general features. Clin. Dermatol. 15, 181–198 (1997).
Breitburd, F., Salmon, J. & Orth, G. The rabbit viral skin papillomas and carcinomas: a model for the immunogenetics of HPV-associated carcinogenesis. Clin. Dermatol. 15, 237–247 (1997).
Ramoz, N., Rueda, L. A., Bouadjar, B., Favre, M. & Orth, G. A susceptibility locus for epidermodysplasia verruciformis, an abnormal predisposition to infection with the oncogenic human papillomavirus type 5, maps to chromosome 17qter in a region containing a psoriasis locus. J. Invest. Dermatol. 112, 259–263 (1999).
Ramoz, N. et al. Mutations in two adjacent novel genes are associated with epidermodysplasia verruciformis. Nature Genet. 32, 579–581 (2002). This is the first identification of a Mendelian genetic aetiology for the syndrome of epidermodysplasia verruciformis, which is characterized by a specific susceptibility to oncogenic cutaneous papillomaviruses. The positional cloning approach allowed the discovery of two new genes located in tandem, which opens a new field in skin immunity.
Hernandez, P. A. et al. Mutations in the chemokine receptor gene CXCR4 are associated with WHIM syndrome, a combined immunodeficiency disease. Nature Genet. 34, 70–74 (2003). Reports the genetic aetiology of the WHIM (warts, hypogammaglobulinaemia, infections and myelokathexis) syndrome, characterized by severe infections caused by human papillomaviruses and other microorganisms.
Miller, L. H., Mason, S. J., Clyde, D. F. & McGinniss, M. H. The resistance factor to Plasmodium vivax in blacks. The Duffy-blood-group genotype, FyFy. N. Engl. J. Med. 295, 302–304 (1976).
Miller, L. H., Mason, S. J., Dvorak, J. A., McGinniss, M. H. & Rothman, I. K. Erythrocyte receptors for (Plasmodium knowlesi) malaria: Duffy blood group determinants. Science 189, 561–563 (1975). References 67 and 68 describe the first Mendelian resistance to common infectious diseases in humans. Individuals negative for the Duffy red-cell antigen were shown to be completely resistant to Plasmodium vivax , probably accounting for the increase of this genotype in endemic regions.
Miller, L. H., Aikawa, M., Johnson, J. G. & Shiroishi, T. Interaction between cytochalasin B-treated malarial parasites and erythrocytes. Attachment and junction formation. J. Exp. Med. 149, 172–184 (1979).
Tournamille, C., Colin, Y., Cartron, J. P. & Le Van Kim, C. Disruption of a GATA motif in the Duffy gene promoter abolishes erythroid gene expression in Duffy-negative individuals. Nature Genet. 10, 224–228 (1995). This paper elucidated the molecular basis of the lack of erythroid expression of Duffy in Plasmodium vivax -resistant individuals. Remarkably, a recessive regulatory single-nucleotide substitution accounts for the erythroid-specific lack of Duffy expression.
Nickel, R. G. et al. Determination of Duffy genotypes in three populations of African descent using PCR and sequence-specific oligonucleotides. Hum. Immunol. 60, 738–742 (1999).
Samson, M. et al. Resistance to HIV-1 infection in caucasian individuals bearing mutant alleles of the CCR5 chemokine receptor gene. Nature 382, 722–725 (1996).
Liu, R. et al. Homozygous defects in HIV-1 coreceptor accounts for resistance of some multiply-exposed individuals to HIV-1 infection. Cell 86, 367–377 (1996).
Dean, M. et al. Genetic restriction of HIV-1 infection and progression to AIDS by a deletion allele of the CKR5 structural gene. Science 273, 1856–1862 (1996). References 73 and 74 describe the first Mendelian resistance to HIV-1, caused by recessive mutations in CC-chemokine receptor 5 (I>CCR5).
Stephens, J. C. et al. Dating the origin of the CCR5-Δ32 AIDS-resistance allele by the coalescence of haplotypes. Am. J. Hum. Genet. 62, 1507–1515 (1998).
Lindesmith, L. et al. Human susceptibility and resistance to Norwalk virus infection. Nature Med. 9, 548–553 (2003). This is the third example of Mendelian resistance to common infections, as recessive mutations in fucosyltransferase 2 ( FUT2 ) are associated with resistance to gastroenteritis caused by noroviruses.
Marionneau, S. et al. Norwalk virus binds to histo-blood group antigens present on gastroduodenal epithelial cells of secretor individuals. Gastroenterology 122, 1967–1977 (2002).
Tsukada, S. et al. Deficient expression of a B cell cytoplasmic tyrosine kinase in human X-linked agammaglobulinemia. Cell 72, 279–290 (1993).
Zeidler, C. & Welte, K. Kostmann syndrome and severe congenital neutropenia. Semin. Hematol. 39, 82–88 (2002).
Kostmann, R. Infantile genetic agranulocytosis: a review with presentation of ten new cases. Acta Pediatr. Scand. 64, 362–368 (1975).
Kostmann, R. Infantile genetic agranulocytosis; agranulocytosis infantilis hereditaria.. Acta Paediatr. Scand. 45, 1–78 (1956). The first description of a Mendelian disorder of myeloid cells, with an autosomal recessive lack of polymorphonuclear leukocytes in children with severe infections.
Dale, D. C. et al. Mutations in the gene encoding neutrophil elastase in congenital and cyclic neutropenia. Blood 96, 2317–2322 (2000).
Person, R. E. et al. Mutations in proto-oncogene GFI1 cause human neutropenia and target ELA2. Nature Genet. 34, 308–312 (2003).
DiGeorge, A. M. A new concept of the cellular basis of immunity. J. Pediatr. 67, 907–908 (1965). Following Bruton's agammaglobulinaemia and Kostmann's agranulocytosis, the third description of a genetically determined complete lack of a leukocyte subset: patients with Di George syndrome are heterozygous for a dominant 22q11 deletion and lack peripheral T cells due to impaired thymus formation. This disorder is not strictly Mendelian, as the causal dominant gene(s) is not known and the penetrance of the T-cell anomaly is very low.
Perez, E. & Sullivan, K. E. Chromosome 22q11.2 deletion syndrome (DiGeorge and velocardiofacial syndromes). Curr. Opin. Pediatr. 14, 678–683 (2002).
Puel, A., Ziegler, S. F., Buckley, R. H. & Leonard, W. J. Defective IL7R expression in T−B+NK+ severe combined immunodeficiency. Nature Genet. 20, 394–397 (1998).
de la Salle, H. et al. Homozygous human TAP peptide transporter mutation in HLA class I deficiency. Science 265, 237–241 (1994). The first report of human inherited deficiency of transporter for antigen processing (TAP), a condition associated with the deficiency of HLA class I molecule expression and low blood counts of cytotoxic CD8+ T cells, but surprisingly not with viral diseases. This paper illustrates the paradoxical insights that can be provided by human genetic studies.
Gadola, S. D., Moins-Teisserenc, H. T., Trowsdale, J., Gross, W. L. & Cerundolo, V. TAP deficiency syndrome. Clin. Exp. Immunol. 121, 173–178 (2000).
Yabe, T. et al. A subject with a novel type I bare lymphocyte syndrome has tapasin deficiency due to deletion of 4 exons by Alu-mediated recombination. Blood 100, 1496–1498 (2002). The first report of inherited tapasin deficiency, also associated with HLA class I deficiency, but surprisingly not susceptibility to viral diseases.
de la Calle-Martin, O. et al. Familial CD8 deficiency due to a mutation in the CD8α gene. J. Clin. Invest. 108, 117–123 (2001). The first report of CD8 deficiency, associated with a lack of cytotoxic CD8+ T cells, but not with viral diseases.
Barry, M. & Bleackley, R. C. Cytotoxic T lymphocytes: all roads lead to death. Nature Rev. Immunol. 2, 401–409 (2002).
Li, Q. & Verma, I. M. NF-κB regulation in the immune system. Nature Rev. Immunol. 2, 725–734 (2002).
Puel, A. et al. Inherited disorders of NF-κB-mediated immunity in man. Curr. Opin. Immunol. (in the press).
Doffinger, R. et al. X-linked anhidrotic ectodermal dysplasia with immunodeficiency is caused by impaired NF-κB signaling. Nature Genet. 27, 277–285 (2001).
Courtois, G. et al. A hypermorphic mutation of IkBα is associated with autosomal dominant anhydrotic ectodermal dysplasia and T cell immunodeficiency. J. Clin. Invest. 112, 1108–1115 (2003).
O'Neill, L. A. Signal transduction pathways activated by the IL-1 receptor/toll-like receptor superfamily. Curr. Top. Microbiol. Immunol. 270, 47–61 (2002).
Picard, C. et al. Pyogenic bacterial infections in humans with IRAK-4 deficiency. Science 299, 2076–2079 (2003). The first report of a human Mendelian defect that affects the common Toll-like receptor and interleukin-1 (IL-1)-receptor signalling (TIR) pathway. Surprisingly, patients with this condition suffer from a relatively narrow spectrum of infectious diseases, mostly limited to pyogenic bacteria.
Medvedev, A. E. et al. Distinct mutations in IRAK-4 confer hyporesponsiveness to lipopolysaccharide and interleukin-1 in a patient with recurrent bacterial infections. J. Exp. Med. 198, 521–531 (2003).
Janeway, C. A. Jr & Medzhitov, R. Innate immune recognition. Annu. Rev. Immunol. 20, 197–216 (2002).
Dinarello, C. A. The role of interleukin-1 in host responses to infectious diseases. Infect. Agents Dis. 1, 227–236 (1992).
Dinarello, C. A. Interleukin-1. Cytokine Growth Factor Rev. 8, 253–265 (1997).
Sieling, P. A. & Modlin, R. L. Toll-like receptors: mammalian 'taste receptors' for a smorgasbord of microbial invaders. Curr. Opin. Microbiol. 5, 70–75 (2002).
Dinarello, C. A. & Fantuzzi, G. Interleukin-18 and host defense against infection. J. Infect. Dis. 187, S370–S384 (2003).
Leddy, J. P., Frank, M. M., Gaither, T., Baum, J. & Klemperer, M. R. Hereditary deficiency of the sixth component of complement in man. I. Immunochemical, biologic, and family studies. J. Clin. Invest. 53, 544–553 (1974). The first description of a genetic defect of a component of the terminal phase of complement, associated with a selective susceptibility to Neisseria . It is also the first molecular elucidation of a Mendelian 'hole' in immunity to infection.
Wurzner, R. et al. Reference typing report for complement components C6, C7 and C9 including mutations leading to deficiencies. Exp. Clin. Immunogenet. 15, 268–285 (1998).
Würzner, R., Orren, A. & Lachmann, P. J. Inherited deficiencies of the terminal components of human complement. Immunodef. Rev. 3, 123–147 (1992).
Frank, M. M. Complement deficiencies. Pediatr. Clin. N. Am. 47, 1339–1354 (2000).
Newport, M. J. et al. A mutation in the interferon-γ-receptor gene and susceptibility to mycobacterial infection. N. Engl. J. Med. 335, 1941–1949 (1996).
Jouanguy, E. et al. Interferon-γ-receptor deficiency in an infant with fatal bacille Calmette-Guerin infection. N. Engl. J. Med. 335, 1956–1961 (1996). References 108 and 109 are the first descriptions of autosomal-recessive complete interferon-γ (IFN-γ)-receptor deficiency, in children with the syndrome of Mendelian susceptibility to mycobacterial diseases, which is characterized by a hole in immunity to infection and a narrow spectrum of infections, caused mostly by mycobacteria (and salmonella).
Levin, M. et al. Familial disseminated atypical mycobacterial infection in childhood: a human mycobacterial susceptibility gene? Lancet 345, 79–83 (1995).
Casanova, J. L. et al. Idiopathic disseminated bacillus Calmette-Guerin infection: a French national retrospective study. Pediatrics 98, 774–778 (1996).
Dorman, S. E. et al. Viral infections in interferon-γ receptor deficiency. J. Pediatr. 135, 640–643 (1999).
Roesler, J. et al. Recurrent mycobacterial and listeria infections in a child with interferon-γ receptor deficiency: mutational analysis and evaluation of therapeutic options. Exp. Haematol. 27, 1368–1374 (1999).
Dorman, S. E. & Holland, S. M. Interferon-γ and interleukin-12 pathway defects and human disease. Cytokine Growth Factor Rev. 11, 321–333 (2000).
Mosmann, T. R., Cherwinski, H., Bond, M. W., Giedlin, M. A. & Coffman, R. L. Two types of murine helper T cell clone. I. Definition according to profiles of lymphokine activities and secreted proteins. J. Immunol. 136, 2348–2357 (1986).
Döffinger, R. et al. Inheritable defects in IL-12- and IFNγ-mediated immunity and the TH1/TH2 paradigm in man. Allergy 54, 409–412 (1999).
Allen, J. E. & Maizels, R. M. TH1–TH2: reliable paradigm or dangerous dogma? Immunol. Today 18, 387–392 (1997).
Fieschi, C. & Casanova, J. L. The role of interleukin-12 in human infectious diseases: only a faint signature. Eur. J. Immunol. 33, 1461–1464 (2003).
Fieschi, C. et al. Low penetrance, broad resistance, and favorable outcome of interleukin-12 receptor β1 deficiency: medical and immunological implications. J. Exp. Med. 197, 527–535 (2003). The first description of a low clinical penetrance (for the case-definition phenotype) among genetic aetiologies of the syndrome of Mendelian susceptibility to mycobacterial diseases, raising the possibility that other, related phenotypes, such as tuberculosis, might be associated with IL-12-receptor deficiency.
Caragol, I. et al. Clinical tuberculosis in 2 of 3 siblings with interleukin-12 receptor β1 deficiency. Clin. Infect. Dis. 37, 302–306 (2003).
Altare, F. et al. Interleukin-12 receptor β1 deficiency in a patient with abdominal tuberculosis. J. Infect. Dis. 184, 231–236 (2001).
Jouanguy, E. et al. Partial interferon-γ receptor 1 deficiency in a child with tuberculoid bacillus Calmette–Guerin infection and a sibling with clinical tuberculosis. J. Clin. Invest. 100, 2658–2664 (1997). References 120–122 describe the first three families with a truly Mendelian tuberculosis: patients with a genetic defect of the IL-12–IFN-γ pathway developed clinical tuberculosis in the absence of any personal or even familial previous history of clinical disease caused by poorly virulent mycobacterial species.
Dupuis, S. et al. Human interferon-γ-mediated immunity is a genetically controlled continuous trait that determines the outcome of mycobacterial invasion. Immunol. Rev. 178, 129–137 (2000).
Abel, L. & Casanova, J. L. Genetic predisposition to clinical tuberculosis: bridging the gap between simple and complex inheritance. Am. J. Hum. Genet. 67, 274–277 (2000).
Kwiatkowski, D. Genetic susceptibility to malaria getting complex. Curr. Opin. Genet. Dev. 10, 320–324 (2000).
Abel, L. & Demenais, F. Detection of major genes for susceptibility to leprosy and its subtypes. Am. J. Hum. Genet. 42, 256–266 (1988).
Mira, M. T. et al. Chromosome 6q25 is linked to susceptibility to leprosy in a Vietnamese population. Nature Genet. 33, 412–415 (2003).
Siddiqui, M. R. et al. A major susceptibility locus for leprosy in India maps to chromosome 10p13. Nature Genet. 27, 439–441 (2001).
Marquet, S. et al. Genetic localization of a locus controlling the intensity of infection by Schistosoma mansoni on chromosome 5q31-q33. Nature Genet. 14, 181–184 (1996). The first identification of a major susceptibility region to a common infectious disease (schistosomiasis) by a genome-wide scan approach.
Dessein, A. J. et al. Severe hepatic fibrosis in Schistosoma mansoni infection is controlled by a major locus that is closely linked to the interferon-γ receptor gene. Am. J. Hum. Genet. 65, 709–721 (1999).
Plancoulaine, S., Gessain, A., van Beveren, M., Tortevoye, P. & Abel, L. Evidence for a recessive major gene predisposing to human herpesvirus 8 (HHV-8) infection in a population in which HHV-8 is endemic. J. Infect. Dis. 187, 1944–1950 (2003).
Plancoulaine, S. et al. Detection of a major gene predisposing to human T lymphotropic virus type I infection in children among an endemic population of African origin. J. Infect. Dis. 182, 405–412 (2000).
Beutler, B. & Rietschel, E. T. Innate immune sensing and its roots: the story of endotoxin. Nature Rev. Immunol. 3, 169–176 (2003).
Isaacs, A. & Lindenmann, J. Virus interference. I. The interferon. Proc. R. Soc. London B 147, 258–267 (1957).
Dupuis, S. et al. Impaired response to interferon-α/β and lethal viral disease in human STAT1 deficiency. Nature Genet. 33, 388–391 (2003).
Nathan, C. F., Murray, H. W., Wiebe, M. E. & Rubin, B. Y. Identification of interferon-γ as the lymphokine that activates human macrophage oxidative metabolism and antimicrobial activity. J. Exp. Med. 158, 670–689 (1983).
Kobayashi, M. et al. Identification and purification of natural killer cell stimulatory factor (NKSF), a cytokine with multiple biologic effects on human lymphocytes. J. Exp. Med. 170, 827–845 (1989).
Trinchieri, G. Interleukin-12 and the regulation of innate resistance and adaptive immunity. Nature Rev. Immunol. 3, 133–146 (2003).
Sayos, J. et al. The X-linked lymphoproliferative-disease gene product SAP regulates signals induced through the co-receptor SLAM. Nature 395, 462–469 (1998).
Coffey, A. J. et al. Host response to EBV infection in X-linked lymphoproliferative disease results from mutations in an SH2-domain encoding gene. Nature Genet. 20, 129–135 (1998).
Nichols, K. E. et al. Inactivating mutations in an SH2 domain-encoding gene in X-linked lymphoproliferative syndrome. Proc. Natl Acad. Sci. USA 95, 13765–13770 (1998). References 139–141 were the first to identify the molecular basis of an X-linked-specific susceptibility to Epstein–Barr virus, another hole in immunity to infection.
Conley, M. E., Rohrer, J., Rapalus, L., Boylin, E. C. & Minegishi, Y. Defects in early B-cell development: comparing the consequences of abnormalities in pre-BCR signaling in the human and the mouse. Immunol. Rev. 178, 75–90 (2000).
Kung, C. et al. Mutations in the tyrosine phosphatase CD45 gene in a child with severe combined immunodeficiency disease. Nature Med. 6, 343–345 (2000).
Dadi, H. K., Simon, A. J. & Roifman, C. M. Effect of CD3δ deficiency on maturation of αβ and γδ T-cell lineages in severe combined immunodeficiency. N. Engl. J. Med. 349, 1821–1828 (2003).
Acknowledgements
We thank P. Kourilsky, P. Sansonetti and J.-C. Weill for critical reading; O. Haller, F. Novelli and R. Würzner for helpful discussions; and present and past members of the Laboratory of Human Genetics of Infectious Diseases for critical reading, helpful discussions and enthusiastic hard work. Our laboratory is supported by grants from the University of Paris René Descartes, INSERM, the European Union, the BNP-Paribas Foundation and the Schlumberger Foundation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Glossary
- SEGREGATION STUDIES
-
Statistical analyses of the familial distribution of a disease (or any phenotype) displaying some degree of intra-familial aggregation (clustering within families). The main goal of a segregation study is to test whether or not the observed familial distributions can be explained by the segregation of a major gene (that is, a gene with a marked effect on the heredity of the disease).
- MICROSATELLITE MARKERS
-
A microsatellite consists of a specific sequence of genomic DNA that contains mono-, di-, tri- or tetranucleotide repeats. Alleles at a specific location (locus) differ in the number of repeats. Microsatellites are inherited in a Mendelian manner, and are, therefore, used as multiallelic (highly polymorphic) genetic markers.
- SINGLE-NUCLEOTIDE POLYMORPHISMS
-
(SNPs). Positions in the genome at which at least two alternative nucleotides occur at appreciable frequencies (usually >1%) in the population. Similar to microsatellites, SNPs display Mendelian inheritance, and are used as genetic markers.
- GENOME-WIDE LINKAGE STUDY
-
Random search over the whole genome of chromosomal regions that are shared in excess by relatives showing a given phenotypic resemblance (for example, affected siblings). In practice, genome scans are done by genotyping 300–400 microsatellite markers that cover all chromosomes (average spacing ∼10 centimorgans) in a relevant sample (for example, 100–200 affected sibling pairs). These data are then analysed by an appropriate linkage method (see Box 1).
Rights and permissions
About this article
Cite this article
Casanova, JL., Abel, L. The human model: a genetic dissection of immunity to infection in natural conditions. Nat Rev Immunol 4, 55–66 (2004). https://doi.org/10.1038/nri1264
Issue Date:
DOI: https://doi.org/10.1038/nri1264
This article is cited by
-
Immunsystem und Allergien – eine unheilige Allianz
Der Internist (2022)
-
The network interplay of interferon and Toll-like receptor signaling pathways in the anti-Candida immune response
Scientific Reports (2021)
-
The human genetic determinism of life-threatening infectious diseases: genetic heterogeneity and physiological homogeneity?
Human Genetics (2020)
-
Inherited CARD9 Deficiency: Invasive Disease Caused by Ascomycete Fungi in Previously Healthy Children and Adults
Journal of Clinical Immunology (2018)
-
Natural variation of macrophage activation as disease-relevant phenotype predictive of inflammation and cancer survival
Nature Communications (2017)