Key Points
-
Genetic studies in IBS range from family and twin studies to candidate gene approaches and genome-wide association studies
-
Despite enlarged sample sizes, increased statistical power and meta-analyses, positive associations between gene variations and IBS subtypes are still scarce and many have not been reproduced
-
Epigenetic and pharmacogenetic approaches are in their infancy
-
A major pitfall in IBS research is the lack of large homogenized case–control cohorts recruited according to standardized and harmonized criteria
Abstract
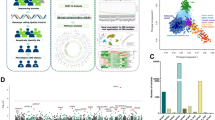
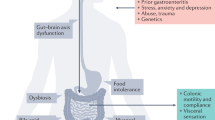
IBS is the most prevalent functional gastrointestinal disorder and phenotypically characterized by chronic abdominal discomfort, pain and altered defecation patterns. The pathophysiology of IBS is multifactorial, albeit with a substantial genetic component. To date, studies using various methodologies, ranging from family and twin studies to candidate gene approaches and genome-wide association studies, have identified several genetic variants in the context of IBS. Yet, despite enlarged sample sizes, increased statistical power and meta-analyses in the past 7 years, positive associations are still scarce and/or have not been reproduced. In addition, epigenetic and pharmacogenetic approaches remain in their infancy. A major hurdle is the lack of large homogenized case–control cohorts recruited according to standardized and harmonized criteria. The COST Action BM1106 GENIEUR (GENes in Irritable Bowel Syndrome Research Network EURope) has been established to address these obstacles. In this Review, the (epi)genetic working group of GENIEUR reports on the current state-of-the-art in the field, highlights fundamental flaws and pitfalls in current IBS (epi)genetic research and provides a vision on how to address and improve (epi)genetic approaches in this complex disorder in the future.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Drossman, D. A., Camilleri, M., Mayer, E. A. & Whitehead, W. E. AGA technical review on irritable bowel syndrome. Gastroenterology 123, 2108–2131 (2002).
Azpiroz, F. et al. Mechanisms of hypersensitivity in IBS and functional disorders. Neurogastroenterol. Motil. 19, 62–88 (2007).
Longstreth, G. F. et al. Functional bowel disorders. Gastroenterology 130, 1480–1491 (2006).
Spiller, R. C. et al. The Patient Health Questionnaire 12 Somatic Symptom scale as a predictor of symptom severity and consulting behaviour in patients with irritable bowel syndrome and symptomatic diverticular disease. Aliment. Pharmacol. Ther. 32, 811–820 (2010).
North, C. S. et al. The presentation of irritable bowel syndrome in the context of somatization disorder. Clin. Gastroenterol. Hepatol. 2, 787–795 (2004).
Frissora, C. L. & Koch, K. L. Symptom overlap and comorbidity of irritable bowel syndrome with other conditions. Curr. Gastroenterol. Rep. 7, 264–271 (2005).
Cole, J. A., Rothman, K. J., Cabral, H. J., Zhang, Y. & Farraye, F. A. Migraine, fibromyalgia, and depression among people with IBS: a prevalence study. BMC Gastroenterol. 6, 26 (2006).
Hillilä, M. T., Färkkilä, N. J. & Färkkilä, M. A. Societal costs for irritable bowel syndrome — a population based study. Scand. J. Gastroenterol. 45, 582–591 (2010).
Larauche, M., Mulak, A. & Tache, Y. Stress and visceral pain: from animal models to clinical therapies. Exp. Neurol. 233, 49–67 (2012).
Spiller, R. C. Role of infection in irritable bowel syndrome. J. Gastroenterol. 42 (Suppl. 17), 41–47 (2007).
Mayer, E. A. Gut feelings: the emerging biology of gut−brain communication. Nat. Rev. Neurosci. 12, 453–66 (2011).
Öhman, L., Törnblom, H. & Simren, M. Crosstalk at the mucosal border: importance of the gut microenvironment in IBS. Nat. Rev. Gastroenterol. Hepatol. 12, 36–49 (2015).
Fukudo, S. & Kanazawa, M. Gene, environment, and brain−gut interactions in irritable bowel syndrome. J. Gastroenterol. Hepatol. 26 (Suppl. 3), 110–115 (2011).
Buonavolonta, R. et al. Familial aggregation in children affected by functional gastrointestinal disorders. J. Pediatr. Gastroenterol. Nutr. 50, 500–505 (2010).
Saito, Y. A. et al. Irritable bowel syndrome aggregates strongly in families: a family-based case-control study. Neurogastroenterol. Motil. 20, 790–797 (2008).
Saito, Y. A. et al. Familial aggregation of irritable bowel syndrome: a family case-control study. Am. J. Gastroenterol. 105, 833–841 (2010).
Bellentani, S. et al. A simple score for the identification of patients at high risk of organic diseases of the colon in the family doctor consulting room. Fam. Pract. 7, 307–312 (1990).
Kalantar, J. S., Locke, G. R. 3rd, Zinsmeister, A. R., Beighley, C. M. & Talley, N. J. Familial aggregation of irritable bowel syndrome: a prospective study. Gut 52, 1703–1707 (2003).
Waehrens, R., Ohlsson, H., Sundquist, J., Sundquist, K. & Zoller, B. Risk of irritable bowel syndrome in first-degree, second-degree and third-degree relatives of affected individuals: a nationwide family study in Sweden. Gut 64, 215–221 (2015).
Whorwell, P. J., McCallum, M., Creed, F. H. & Roberts, C. T. Non-colonic features of irritable bowel syndrome. Gut 27, 37–40 (1986).
Saito, Y. A. The role of genetics in IBS. Gastroenterol. Clin. North Am. 40, 45–67 (2011).
Bengtson, M. B., Ronning, T., Vatn, M. H. & Harris, J. R. Irritable bowel syndrome in twins: genes and environment. Gut 55, 1754–1759 (2006).
Lembo, A., Zaman, M., Jones, M. & Talley, N. J. Influence of genetics on irritable bowel syndrome, gastro-oesophageal reflux and dyspepsia: a twin study. Aliment. Pharmacol. Ther. 25, 1343–1350 (2007).
Morris-Yates, A., Talley, N. J., Boyce, P. M., Nandurkar, S. & Andrews, G. Evidence of a genetic contribution to functional bowel disorder. Am. J. Gastroenterol. 93, 1311–1317 (1998).
Bengtson, M. B., Aamodt, G., Vatn, M. H. & Harris, J. R. Co-occurrence of IBS and symptoms of anxiety or depression, among Norwegian twins, is influenced by both heredity and intrauterine growth. BMC Gastroenterol. 15, 9 (2015).
McGowan, P. O. et al. Epigenetic regulation of the glucocorticoid receptor in human brain associates with childhood abuse. Nat. Neurosci. 12, 342–348 (2009).
Ke, X. et al. Intrauterine growth retardation affects expression and epigenetic characteristics of the rat hippocampal glucocorticoid receptor gene. Physiol. Genomics 42, 177–189 (2010).
Levy, R. L. et al. Irritable bowel syndrome in twins: heredity and social learning both contribute to etiology. Gastroenterology 121, 799–804 (2001).
Levy, R. L. et al. Increased somatic complaints and health-care utilization in children: effects of parent IBS status and parent response to gastrointestinal symptoms. Am. J. Gastroenterol. 99, 2442–2451 (2004).
Videlock, E. J. et al. Childhood trauma is associated with hypothalamic−pituitary−adrenal axis responsiveness in irritable bowel syndrome. Gastroenterology 137, 1954–1962 (2009).
Niesler, B. et al. 5-HTTLPR and STin2 polymorphisms in the serotonin transporter gene and irritable bowel syndrome: effect of bowel habit and sex. Eur. J. Gastroenterol. Hepatol. 22, 856–861 (2010).
Choi, Y. J. et al. Association between SLC6A4 serotonin transporter gene lainked polymorphic region and ADRA2A−1291C>G and irritable bowel syndrome in Korea. J. Neurogastroenterol. Motil. 20, 388–399 (2014).
Kim, H. J. et al. Association of distinct α2 adrenoceptor and serotonin transporter polymorphisms with constipation and somatic symptoms in functional gastrointestinal disorders. Gut 53, 829–837 (2004).
Van Kerkhoven, L. A., Laheij, R. J. & Jansen, J. B. Meta-analysis: a functional polymorphism in the gene encoding for activity of the serotonin transporter protein is not associated with the irritable bowel syndrome. Aliment. Pharmacol. Ther. 26, 979–986 (2007).
Yeo, A. et al. Association between a functional polymorphism in the serotonin transporter gene and diarrhoea predominant irritable bowel syndrome in women. Gut 53, 1452–1458 (2004).
Park, J. M. et al. Serotonin transporter gene polymorphism and irritable bowel syndrome. Neurogastroenterol. Motil. 18, 995–1000 (2006).
Kumar, S., Ranjan, P., Mittal, B. & Ghoshal, U. C. Serotonin transporter gene (SLC6A4) polymorphism in patients with irritable bowel syndrome and healthy controls. J. Gastrointestin. Liver Dis. 21, 31–38 (2012).
Pata, C. et al. Serotonin transporter gene polymorphism in irritable bowel syndrome. Am. J. Gastroenterol. 97, 1780–1784 (2002).
Saito, Y. A. et al. A genetic association study of 5-HTT LPR and GNβ3 C825T polymorphisms with irritable bowel syndrome. Neurogastroenterol. Motil. 19, 465–470 (2007).
Sikander, A. et al. Serotonin transporter promoter variant: analysis in Indian IBS patients and control population. J. Clin. Gastroenterol. 43, 957–961 (2009).
Lee, D. Y. et al. Serotonin transporter gene polymorphism in healthy adults and patients with irritable bowel syndrome. Kor. J. Gastroenterol. 43, 18–22 (in Korean) (2004).
Wang, B. M. et al. Serotonin transporter gene polymorphism in irritable bowel syndrome. Zhonghua Nei Ke Za Zhi 43, 439–441 (in Chinese) (2004).
Li, Y. et al. The association of serotonin transporter genetic polymorphisms and irritable bowel syndrome and its influence on tegaserod treatment in Chinese patients. Dig. Dis. Sci. 52, 2942–2949 (2007).
Yuan, J. et al. Association study of serotonin transporter SLC6A4 gene with Chinese Han irritable bowel syndrome. PLoS ONE 9, e84414 (2014).
Kohen, R. et al. The serotonin transporter polymorphism rs25531 is associated with irritable bowel syndrome. Dig. Dis. Sci. 54, 2663–2670 (2009).
Lesch, K. P. et al. Association of anxiety-related traits with a polymorphism in the serotonin transporter gene regulatory region. Science 274, 1527–1531 (1996).
Jarrett, M. E. et al. Relationship of SERT polymorphisms to depressive and anxiety symptoms in irritable bowel syndrome. Biol. Res. Nurs. 9, 161–169 (2007).
Blom, R. M. et al. Association between a serotonin transporter promoter polymorphism (5HTTLPR) and personality disorder traits in a community sample. J. Psychiatr. Res. 45, 1153–1159 (2011).
Camilleri, M. et al. Candidate genes and sensory functions in health and irritable bowel syndrome. Am. J. Physiol. Gastrointest. Liver Physiol. 295, G219–G225 (2008).
Farmer, A. D. et al. Psychophysiological responses to pain identify reproducible human clusters. Pain 154, 2266–2276 (2013).
Pezawas, L. et al. 5-HTTLPR polymorphism impacts human cingulate−amygdala interactions: a genetic susceptibility mechanism for depression. Nat. Neurosci. 8, 828–834 (2005).
Fukudo, S. et al. Impact of serotonin transporter gene polymorphism on brain activation by colorectal distention. NeuroImage 47, 946–951 (2009).
Spiller, R. C. Targeting the 5-HT3 receptor in the treatment of irritable bowel syndrome. Curr. Opin. Pharmacol. 11, 68–74 (2011).
Garsed, K. et al. A randomised trial of ondansetron for the treatment of irritable bowel syndrome with diarrhoea. Gut 63, 1617–1625 (2014).
Walstab, J., Rappold, G. & Niesler, B. 5-HT3 receptors: role in disease and target of drugs. Pharmacol. Ther. 128, 146–169 (2010).
Camilleri, M. et al. Efficacy and safety of alosetron in women with irritable bowel syndrome: a randomised, placebo-controlled trial. Lancet 355, 1035–1040 (2000).
Fukudo, S., Ida, M., Akiho, H., Nakashima, Y. & Matsueda, K. Effect of ramosetron on stool consistency in male patients with irritable bowel syndrome with diarrhea. Clin. Gastroenterol. Hepatol. 12, 953–959.e4 (2014).
Kapeller, J. et al. First evidence for an association of a functional variant in the microRNA-510 target site of the serotonin receptor-type 3E gene with diarrhea predominant irritable bowel syndrome. Hum. Mol. Genet. 17, 2967–2977 (2008).
Kapeller, J. et al. A coding variant in the serotonin receptor 3c subunit is associated with diarrhea predominant irritable bowel syndrome. Gastroenterology 136, A-155–A-156 (2009).
Zhang, Y., Huang, Y. & Bo, P. Association between diarrhea-predominant irritable bowel syndrome and HTR3A, HTR3E gene polymorphism in Yangzhou, Jiangsu province, China. Zhonghua Liu Xing Bing Xue Za Zhi 34, 721–724 (in Chinese) (2013).
Gu, Q. Y., Zhang, J., Feng, Y. C., Dai, G. R. & Du, W. P. Association of genetic polymorphisms in HTR3A and HTR3E with diarrhea predominant irritable bowel syndrome. Int. J. Clin. Exp. Med. 8, 4581–4585 (2015).
de Vries, D., ter Linde, J., van Herwaarden, M., Smout, A. & Samsom, M. Serotonin receptor 3a polymorphism C178t is associated with visceral hypersensitivity in GERD. Gastroenterology Abstr. 132, A276 (2007).
Mujakovic, S. et al. Serotonin receptor 3A polymorphism c.-42C > T is associated with severe dyspepsia. BMC Med. Genet. 12, 140 (2011).
Melke, J. et al. A polymorphism in the serotonin receptor 3A (HTR3A) gene and its association with harm avoidance in women. Arch. Gen. Psychiatry 60, 1017–1023 (2003).
Niesler, B. et al. Serotonin receptor gene HTR3A variants in schizophrenic and bipolar affective patients. Pharmacogenetics 11, 21–27 (2001).
Cloninger, C. R. A systematic method for clinical description and classification of personality variants. A Proposal. Arch. Gen. Psychiatry 44, 573–588 (1987).
Kilpatrick, L. A. et al. The HTR3A polymorphism c. -42C>T is associated with amygdala responsiveness in patients with irritable bowel syndrome. Gastroenterology 140, 1943–1951 (2011).
Yamada, K. et al. Distinguishable haplotype blocks in the HTR3A and HTR3B region in the Japanese reveal evidence of association of HTR3B with female major depression. Biol. Psychiatry 60, 192–201 (2006).
Hammer, C. et al. Functional variants of the serotonin receptor type 3A and B gene are associated with eating disorders. Pharmacogenet. Genomics 19, 790–799 (2009).
Hammer, C. et al. Replication of functional serotonin receptor type 3A and B variants in bipolar affective disorder: a European multicenter study. Transl. Psychiatry 2, e103 (2012).
Aibiki, L. et al. Impact of serotonin receptor-3 gene polymorphism on irritable bowel syndrome. Gastroenterology 132, A134–A135 (2007).
Ek, W. E. et al. Exploring the genetics of irritable bowel syndrome: a GWA study in the general population and replication in multinational case-control cohorts. Gut 64, 1774–1782 (2015).
Iidaka, T. et al. A variant C178T in the regulatory region of the serotonin receptor gene HTR3A modulates neural activation in the human amygdala. J. Neurosci. 25, 6460–6466 (2005).
Fukudo, S. et al. Impact of serotonin-3 receptor gene polymorphism on brain activation by rectal distention in human. Gastroenterology 136, A170 (2009).
Horjales-Araujo, E. et al. Polymorphism in serotonin receptor 3B is associated with pain catastrophizing. PLoS ONE 8, e78889 (2013).
Mulak, A. et al. Association of polymorphisms in 5-HT2A and 5-HT2C receptors genes with depressive and anxiety disorders in patients with irritable bowel syndrome. Gastroenterology 144, S725 (2013).
Jun, S., Kohen, R., Cain, K. C., Jarrett, M. E. & Heitkemper, M. M. Associations of tryptophan hydroxylase gene polymorphisms with irritable bowel syndrome. Neurogastroenterol. Motil. 23, 233–e116 (2011).
Jun, S. E., Kohen, R., Cain, K. C., Jarrett, M. E. & Heitkemper, M. M. TPH gene polymorphisms are associated with disease perception and quality of life in women with irritable bowel syndrome. Biol. Res. Nurs. 16, 95–104 (2014).
Pata, C. et al. Association of the -1438 G/A and 102 T/C polymorphism of the 5-Ht2A receptor gene with irritable bowel syndrome 5-Ht2A gene polymorphism in irritable bowel syndrome. J. Clin. Gastroenterol. 38, 561–566 (2004).
Beyder, A. et al. Loss-of-function of the voltage-gated sodium channel NaV1.5 (channelopathies) in patients with irritable bowel syndrome. Gastroenterology 146, 1659–1668 (2014).
Saito, Y. A. et al. Sodium channel mutation in irritable bowel syndrome: evidence for an ion channelopathy. Am. J. Physiol. Gastrointest. Liver Physiol. 296, G211–G218 (2009).
Wouters, M. M. et al. Genetic variants in CDC42 and NXPH1 as susceptibility factors for constipation and diarrhoea predominant irritable bowel syndrome. Gut 63, 1103–1111 (2014).
van den Oord, E. J. et al. Genomewide association analysis followed by a replication study implicates a novel candidate gene for neuroticism. Arch. Gen. Psychiatry 65, 1062–1071 (2008).
Sikander, A. et al. Association of alpha 2A adrenergic receptor gene (ADRA2A) polymorphism with irritable bowel syndrome, microscopic and ulcerative colitis. Clin. Chim. Acta 411, 59–63 (2010).
Camilleri, M. et al. Cannabinoid receptor 1 gene and irritable bowel syndrome: phenotype and quantitative traits. Am. J. Physiol. Gastrointest. Liver Physiol. 304, G553–G560 (2013).
Camilleri, M. et al. Genetic variation in endocannabinoid metabolism, gastrointestinal motility, and sensation. Am. J. Physiol. Gastrointest. Liver Physiol. 294, G13–G19 (2008).
Karling, P. et al. The relationship between the val158met catechol-O-methyltransferase (COMT) polymorphism and irritable bowel syndrome. PLoS ONE 6, e18035 (2011).
Jiang, Y., Nie, Y., Li, Y. & Zhang, L. Association of cannabinoid type 1 receptor and fatty acid amide hydrolase genetic polymorphisms in Chinese patients with irritable bowel syndrome. J. Gastroenterol. Hepatol. 29, 1186–1191 (2014).
Park, J. M. et al. Cannabinoid receptor 1 gene polymorphism and irritable bowel syndrome in the Korean population: a hypothesis-generating study. J. Clin. Gastroenterol. 45, 45–49 (2011).
Zhou, Q., Souba, W. W., Croce, C. M. & Verne, G. N. MicroRNA-29a regulates intestinal membrane permeability in patients with irritable bowel syndrome. Gut 59, 775–784 (2010).
Dunlop, S. P. et al. Abnormal intestinal permeability in subgroups of diarrhea-predominant irritable bowel syndromes. Am. J. Gastroenterol. 101, 1288–1294 (2006).
Martinez, C. et al. The jejunum of diarrhea-predominant irritable bowel syndrome shows molecular alterations in the tight junction signaling pathway that are associated with mucosal pathobiology and clinical manifestations. Am. J. Gastroenterol. 107, 736–746 (2012).
Martinez, C. et al. Diarrhoea-predominant irritable bowel syndrome: an organic disorder with structural abnormalities in the jejunal epithelial barrier. Gut 62, 1160–1168 (2013).
Villani, A. C. et al. Genetic risk factors for post-infectious irritable bowel syndrome following a waterborne outbreak of gastroenteritis. Gastroenterology 138, 1502–1513 (2010).
Bashashati, M. et al. Cytokine gene polymorphisms are associated with irritable bowel syndrome: a systematic review and meta-analysis. Neurogastroenterol. Motil. 24, e1102–e1566 (2012).
Czogalla, B. et al. A meta-analysis of immunogenetic Case−Control Association Studies in irritable bowel syndrome. Neurogastroenterol. Motil. 27, 717–727 (2015).
Lee, Y. J. & Park, K. S. Irritable bowel syndrome: emerging paradigm in pathophysiology. World J. Gastroenterol. 20, 2456–2469 (2014).
Zucchelli, M. et al. Association of TNFSF15 polymorphism with irritable bowel syndrome. Gut 60, 1671–1677 (2011).
Swan, C. et al. Identifying and testing candidate genetic polymorphisms in the irritable bowel syndrome (IBS): association with TNFSF15 and TNFα. Gut 62, 985–994 (2013).
Bamias, G. et al. Expression, localization, and functional activity of TL1A, a novel Th1-polarizing cytokine in inflammatory bowel disease. J. Immunol. 171, 4868–4874 (2003).
Kugathasan, S. & Cohen, S. Searching for new clues in inflammatory bowel disease: tell tales from pediatric IBD natural history studies. Gastroenterology 135, 1038–1041 (2008).
Bamias, G. et al. Upregulation and nuclear localization of TNF-like cytokine 1A (TL1A) and its receptors DR3 and DcR3 in psoriatic skin lesions. Exp. Dermatol. 20, 725–731 (2011).
Pappu, B. P. et al. TL1A−DR3 interaction regulates Th17 cell function and Th17-mediated autoimmune disease. J. Exp. Med. 205, 1049–1062 (2008).
Kamada, N. et al. TL1A produced by lamina propria macrophages induces Th1 and Th17 immune responses in cooperation with IL-23 in patients with Crohn's disease. Inflamm. Bowel Dis. 16, 568–575 (2010).
Meylan, F. et al. The TNF-family cytokine TL1A promotes allergic immunopathology through group 2 innate lymphoid cells. Mucosal Immunol. 7, 958–968 (2014).
van der Veek, P. P., van den Berg, M., de Kroon, Y. E., Verspaget, H. W. & Masclee, A. A. Role of tumor necrosis factor-α and interleukin-10 gene polymorphisms in irritable bowel syndrome. Am. J. Gastroenterol. 100, 2510–2516 (2005).
Gonsalkorale, W. M., Perrey, C., Pravica, V., Whorwell, P. J. & Hutchinson, I. V. Interleukin 10 genotypes in irritable bowel syndrome: evidence for an inflammatory component? Gut 52, 91–93 (2003).
Schmulson, M. et al. IL-10 and TNF-α polymorphisms in subjects with irritable bowel syndrome in Mexico. Rev. Esp. Enferm. Dig. 105, 392–399 (2013).
Barkhordari, E. et al. Proinflammatory cytokine gene polymorphisms in irritable bowel syndrome. J. Clin. Immunol. 30, 74–79 (2010).
Romero-Valdovinos, M. et al. Interleukin-8 and -10 gene polymorphisms in irritable bowel syndrome. Mol. Biol. Rep. 39, 8837–8843 (2012).
Olivo-Diaz, A. et al. Findings related to IL-8 and IL-10 gene polymorphisms in a Mexican patient population with irritable bowel syndrome infected with Blastocystis. Parasitol. Res. 111, 487–491 (2012).
Qin, S. Y., Jiang, H. X., Lu, D. H. & Zhou, Y. Association of interleukin-10 polymorphisms with risk of irritable bowel syndrome: a meta-analysis. World J. Gastroenterol. 19, 9472–9480 (2013).
Santhosh, S. et al. Cytokine gene polymorphisms in irritable bowel syndrome in Indian population — a pilot case control study. Trop. Gastroenterol. 31, 30–33 (2010).
Shiotani, A. et al. S100A expression and interleukin-10 polymorphisms are associated with ulcerative colitis and diarrhea predominant irritable bowel syndrome. Dig. Dis. Sci. 58, 2314–2323 (2013).
Holliday, E. G. et al. Genome-wide association study identifies two novel genomic regions in irritable bowel syndrome. Am. J. Gastroenterol. 109, 770–772 (2014).
Saito, Y. A. et al. A candidate gene association study of functional 'psychiatric' polymorphisms in irritable bowel syndrome (IBS). Gastroenterology 138, S90 (2010).
Dinan, T. G., Cryan, J., Shanahan, F., Keeling, P. W. & Quigley, E. M. IBS: an epigenetic perspective. Nat. Rev. Gastroenterol. Hepatol. 7, 465–471 (2010).
Tran, L., Chaloner, A., Sawalha, A. H. & Greenwood Van-Meerveld, B. Importance of epigenetic mechanisms in visceral pain induced by chronic water avoidance stress. Psychoneuroendocrinology 38, 898–906 (2013).
van den Wijngaard, R. M. et al. Susceptibility to stress induced visceral hypersensitivity in maternally separated rats is transferred across generations. Neurogastroenterol. Motil. 25, e780–e790 (2013).
Zhou, Q. et al. MicroRNA 29 targets nuclear factor-kappaB-repressing factor and Claudin 1 to increase intestinal permeability. Gastroenterology 148, 158–169.e8 (2015).
Zhou, Q. et al. Decreased miR-199 augments visceral pain in patients with IBS through translational upregulation of TRPV1. Gut http://dx.doi.org/10.1136/gutjnl-2013-306464 (2015).
Fourie, N. H. et al. Elevated circulating miR-150 and miR-342-3p in patients with irritable bowel syndrome. Exp. Mol. Pathol. 96, 422–425 (2014).
Gheinani, A. H., Burkhard, F. C. & Monastyrskaya, K. Deciphering microRNA code in pain and inflammation: lessons from bladder pain syndrome. Cell. Mol. Life Sci. 70, 3773–3789 (2013).
Ehrenreich, H. & Nave, K. A. Phenotype-Based Genetic Association Studies (PGAS) — towards understanding the contribution of common genetic variants to schizophrenia subphenotypes. Genes (Basel) 5, 97–105 (2014).
Clarke, G. M. et al. Basic statistical analysis in genetic case-control studies. Nat. Protoc. 6, 121–133 (2011).
Balding, D. J. A tutorial on statistical methods for population association studies. Nat. Rev. Genet. 7, 781–791 (2006).
Laird, N. M. & Lange, C. The Fundamentals of Modern Statistical Genetics: Statistics for Biology and Health (Springer, 2010).
Bearcroft, C. P., Perrett, D. & Farthing, M. J. Postprandial plasma 5-hydroxytryptamine in diarrhoea predominant irritable bowel syndrome: a pilot study. Gut 42, 42–46 (1998).
Camilleri, M. et al. Serotonin-transporter polymorphism pharmacogenetics in diarrhea-predominant irritable bowel syndrome. Gastroenterology 123, 425–432 (2002).
Camilleri, M. et al. Pharmacogenetics of low dose clonidine in irritable bowel syndrome. Neurogastroenterol. Motil. 21, 399–410 (2009).
Wong, B. S. et al. Pharmacogenetic trial of a cannabinoid agonist shows reduced fasting colonic motility in patients with nonconstipated irritable bowel syndrome. Gastroenterology 141, 1638–1647.e7 (2011).
Wong, B. S. et al. Randomized pharmacodynamic and pharmacogenetic trial of dronabinol effects on colon transit in irritable bowel syndrome-diarrhea. Neurogastroenterol. Motil. 24, 358–e169 (2012).
Rao, A. S. et al. Chenodeoxycholate in females with irritable bowel syndrome-constipation: a pharmacodynamic and pharmacogenetic analysis. Gastroenterology 139, 1549–1558.e1 (2010).
Wong, B. S. et al. Pharmacogenetics of the effects of colesevelam on colonic transit in irritable bowel syndrome with diarrhea. Dig. Dis. Sci. 57, 1222–1226 (2012).
Camilleri, M. et al. Effect of increased bile acid synthesis or fecal excretion in irritable bowel syndrome-diarrhea. Am. J. Gastroenterol. 109, 1621–1630 (2014).
Knowles, C. H., Lindberg, G., Panza, E. & De Giorgio, R. New perspectives in the diagnosis and management of enteric neuropathies. Nat. Rev. Gastroenterol. Hepatol. 10, 206–218 (2013).
Camilleri, M. Peripheral mechanisms in irritable bowel syndrome. N. Engl. J. Med. 367, 1626–1635 (2012).
Ellinghaus, D. et al. Association between variants of PRDM1 and NDP52 and Crohn's disease, based on exome sequencing and functional studies. Gastroenterology 145, 339–347 (2013).
Xu, S. et al. Exome sequencing identifies DLG1 as a novel gene for potential susceptibility to Crohn's disease in a Chinese family study. PLoS ONE 9, e99807 (2014).
Fiskerstrand, T. et al. Familial diarrhea syndrome caused by an activating GUCY2C mutation. N. Engl. J. Med. 366, 1586–1595 (2012).
Acknowledgements
This manuscript has resulted from the collaboration and network activities of the genetics/epigenetics Working Group (WG3) under the frame of the international network GENIEUR (GENes in Irritable Bowel Syndrome Research Network EURope), which is currently funded by the COST (COoperation in Science and Technology) programme (BM1106, www.GENIEUR.eu). We thank A. Farmer, R. Spiller and C. Fischer for fruitful discussion and comments on the manuscript.
Author information
Authors and Affiliations
Contributions
B.N. and M.G. developed the concept, designed, wrote, assembled input data, and edited the manuscript; B.N. and M.M.W. created and revised the figures; A.M., B.N. and L.K.P. summed up the genetics and epigenetics findings in Tables 1 and 2; M.M.W., L.K.P., M.B.B., E.F., G.N., C.A.D., A.M. and J.S. reviewed the literature, selected the data and wrote the manuscript. All authors discussed the results and implications and commented on the manuscript at all stages.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
Supplementary information
Supplementary Table 1
Summary of genetic association data in IBS. (DOC 180 kb)
Supplementary Table 2
Summary of epigenetic data in IBS. (DOC 47 kb)
Rights and permissions
About this article
Cite this article
Gazouli, M., Wouters, M., Kapur-Pojskić, L. et al. Lessons learned — resolving the enigma of genetic factors in IBS. Nat Rev Gastroenterol Hepatol 13, 77–87 (2016). https://doi.org/10.1038/nrgastro.2015.206
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrgastro.2015.206
This article is cited by
-
The serotonin receptor 3E variant is a risk factor for female IBS-D
Journal of Molecular Medicine (2022)
-
Disorders of the enteric nervous system — a holistic view
Nature Reviews Gastroenterology & Hepatology (2021)
-
The enteric nervous system in gastrointestinal disease etiology
Cellular and Molecular Life Sciences (2021)
-
New treatments and therapeutic targets for IBS and other functional bowel disorders
Nature Reviews Gastroenterology & Hepatology (2018)
-
A Review of Microbiota and Irritable Bowel Syndrome: Future in Therapies
Advances in Therapy (2018)