Key Points
-
The rapid decrease in sequencing costs has led to the recent development of a plethora of epigenomic techniques that use short-read sequencing as a readout.
-
Enzymatic footprinting (using micrococcal nuclease (MNase), deoxyribonuclease (DNase), Tn5 transposase or methyltransferases) provides a means to assess the occupancy of both nucleosomes and non-histone proteins in a single experiment.
-
Enzymatic digestion of chromatin before or after chromatin immunoprecipitation (ChIP) greatly increases its resolution.
-
Mapping the last nucleotide added to a nascent RNA chain provides base-pair resolution maps of RNA polymerase occupancy.
Abstract
The widespread adoption of short-read DNA sequencing as a digital epigenomic readout platform has motivated the development of genome-wide tools that achieve base-pair resolution. New methods for footprinting and affinity purification of nucleosomes, RNA polymerases, chromatin remodellers and transcription factors have increased the resolution of epigenomic profiling by two orders of magnitude, leading to new insights into how the chromatin landscape affects gene regulation. These digital epigenomic tools have also been applied to directly profile both turnover kinetics and transcription in situ. In this Review, we describe how these new genome-wide tools allow interrogation of diverse aspects of the epigenome.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Jiang, C. & Pugh, B. F. Nucleosome positioning and gene regulation: advances through genomics. Nature Rev. Genet. 10, 161–172 (2009).
Talbert, P. B. & Henikoff, S. Histone variants — ancient wrap artists of the epigenome. Nature Rev. Mol. Cell Biol. 11, 264–275 (2010).
Zentner, G. E. & Henikoff, S. Regulation of nucleosome dynamics by histone modifications. Nature Struct. Mol. Biol. 20, 259–266 (2013).
Henikoff, S. Nucleosome destabilization in the epigenetic regulation of gene expression. Nature Rev. Genet. 9, 15–26 (2008).
Smith, Z. D. & Meissner, A. DNA methylation: roles in mammalian development. Nature Rev. Genet. 14, 204–220 (2013).
Zaret, K. S. & Carroll, J. S. Pioneer transcription factors: establishing competence for gene expression. Genes Dev. 25, 2227–2241 (2011).
Hargreaves, D. C. & Crabtree, G. R. ATP-dependent chromatin remodeling: genetics, genomics and mechanisms. Cell Res. 21, 396–420 (2011).
Flynn, R. A. & Chang, H. Y. Active chromatin and noncoding RNAs: an intimate relationship. Curr. Opin. Genet. Dev. 22, 172–178 (2012).
Stamatoyannopoulos, J. A. What does our genome encode? Genome Res. 22, 1602–1611 (2012).
Bernstein, B. E. et al. The NIH Roadmap Epigenomics Mapping Consortium. Nature Biotech. 28, 1045–1048 (2010).
Li, R. et al. De novo assembly of human genomes with massively parallel short read sequencing. Genome Res. 20, 265–272 (2010).
Barski, A. et al. High-resolution profiling of histone methylations in the human genome. Cell 129, 823–837 (2007).
Mikkelsen, T. S. et al. Genome-wide maps of chromatin state in pluripotent and lineage-committed cells. Nature 448, 553–560 (2007).
Johnson, D. S., Mortazavi, A., Myers, R. M. & Wold, B. Genome-wide mapping of in vivo protein–DNA interactions. Science 316, 1497–1502 (2007).
Krueger, F., Kreck, B., Franke, A. & Andrews, S. R. DNA methylome analysis using short bisulfite sequencing data. Nature Meth. 9, 145–151 (2012).
Brogaard, K., Xi, L., Wang, J.-P. & Widom, J. A map of nucleosome positions in yeast at base-pair resolution. Nature 486, 496–501 (2012).
Reeves, R. & Jones, A. Genomic transcriptional activity and the structure of chromatin. Nature 260, 495–500 (1976).
Weintraub, H. & Groudine, M. Chromosomal subunits in active genes have an altered conformation. Science 193, 848–856 (1976). References 17 and 18 describe the first uses of MNase and DNase, respectively, to map the accessibility of chromatin at specific genomic loci.
Sabo, P. J. et al. Genome-scale mapping of DNase I sensitivity in vivo using tiling DNA microarrays. Nature Meth. 3, 511–518 (2006).
Crawford, G. E. et al. DNase-chip: a high-resolution method to identify DNase I hypersensitive sites using tiled microarrays. Nature Meth. 3, 503–509 (2006).
Yuan, G.-C. et al. Genome-scale identification of nucleosome positions in S. cerevisiae. Science 309, 626–630 (2005).
Giresi, P. G., Kim, J., McDaniell, R. M., Iyer, V. R. & Lieb, J. D. FAIRE (formaldehyde-assisted isolation of regulatory elements) isolates active regulatory elements from human chromatin. Genome Res. 17, 877–885 (2007).
Auerbach, R. K. et al. Mapping accessible chromatin regions using Sono-seq. Proc. Natl Acad. Sci. USA 106, 14926–14931 (2009).
Sulkowski, E. & Laskowski, M. Mechanism of action of micrococcal nuclease on deoxyribonucleic acid. J. Biol. Chem. 237, 2620–2625 (1962).
Noll, M. Subunit structure of chromatin. Nature 251, 249–251 (1974).
Weber, C. M., Henikoff, J. G. & Henikoff, S. H2A.Z nucleosomes enriched over active genes are homotypic. Nature Struct. Mol. Biol. 17, 1500–1507 (2010).
Teves, S. S. & Henikoff, S. Heat shock reduces stalled RNA polymerase II and nucleosome turnover genome-wide. Genes Dev. 25, 2387–2397 (2011).
Kent, N. A., Adams, S., Moorhouse, A. & Paszkiewicz, K. Chromatin particle spectrum analysis: a method for comparative chromatin structure analysis using paired-end mode next-generation DNA sequencing. Nucleic Acids Res. 39, e26 (2011).
Henikoff, J. G., Belsky, J. A., Krassovsky, K., MacAlpine, D. M. & Henikoff, S. Epigenome characterization at single base-pair resolution. Proc. Natl Acad. Sci. USA 108, 18318–18323 (2011). References 28 and 29 show that mapping of a broad range of MNase-digested fragments gives precise information about positioning and occupancy of both nucleosomes and non-histone proteins in a single sample.
Krassovsky, K., Henikoff, J. G. & Henikoff, S. Tripartite organization of centromeric chromatin in budding yeast. Proc. Natl Acad. Sci. USA 109, 243–248 (2012).
Chung, H.-R. et al. The effect of micrococcal nuclease digestion on nucleosome positioning data. PLoS ONE 5, e15754 (2010).
Deniz, O. et al. Physical properties of naked DNA influence nucleosome positioning and correlate with transcription start and termination sites in yeast. BMC Genomics 12, 489 (2011).
Allan, J., Fraser, R. M., Owen-Hughes, T. & Keszenman-Pereyra, D. Micrococcal nuclease does not substantially bias nucleosome mapping. J. Mol. Biol. 417, 152–164 (2012).
Albert, I. et al. Translational and rotational settings of H2A.Z nucleosomes across the Saccharomyces cerevisiae genome. Nature 446, 572–576 (2007).
Hesselberth, J. R. et al. Global mapping of protein–DNA interactions in vivo by digital genomic footprinting. Nature Meth 6, 283–289 (2009).
Vierstra, J., Wang, H., John, S., Sandstrom, R. & Stamatoyannopoulos, J. A. Coupling transcription factor occupancy to nucleosome architecture with DNase-FLASH. Nature Meth 11, 66–72 (2014).
He, H. H. et al. Refined DNase-seq protocol and data analysis reveals intrinsic bias in transcription factor footprint identification. Nature Meth 11, 73–78 (2014).
Boyle, A. P. et al. High-resolution genome-wide in vivo footprinting of diverse transcription factors in human cells. Genome Res. 21, 456–464 (2011).
Neph, S. et al. An expansive human regulatory lexicon encoded in transcription factor footprints. Nature 489, 83–90 (2012).
Lazarides, E. & Lindberg, U. Actin is the naturally occurring inhibitor of deoxyribonuclease I. Proc. Natl Acad. Sci. USA 71, 4742–4746 (1974).
Grontved, L. et al. Rapid genome-scale mapping of chromatin accessibility in tissue. Epigenetics Chromatin 5, 10 (2012).
Adey, A. et al. Rapid, low-input, low-bias construction of shotgun fragment libraries by high-density in vitro transposition. Genome Biol. 11, R119 (2010).
Gangadharan, S., Mularoni, L., Fain-Thornton, J., Wheelan, S. J. & Craig, N. L. DNA transposon Hermes inserts into DNA in nucleosome-free regions in vivo. Proc. Natl Acad. Sci. 107, 21966–21972 (2010). This study describes a rapid, simple procedure for epigenomic analysis based on transposition of sequencing adapters into chromatin.
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nature Meth. 10, 1213–1218 (2013).
Henikoff, S. et al. The budding yeast Centromere DNA Element II wraps a stable Cse4 hemisome in either orientation in vivo. eLife 3, e01861 (2014).
Pan, C. Q., Landgraf, R. & Sigman, D. S. Drosophila engrailed-1, 10-phenanthroline chimeras as probes of homeodomain-DNA complexes. Protein Sci. 4, 2279–2288 (1995).
Pan, C. Q., Johnson, R. C. & Sigman, D. S. Identification of new fis binding sites by DNA scission with fis-1,10-phenanthroline-copper(I) chimeras. Biochemistry 35, 4326–4333 (1996).
Landgraf, R., Pan, C., Sutton, C., Pearson, L. & Sigman, D. S. Engineering of DNA binding proteins into site-specific cutters: reactivity of Trp repressor-1,10-phenanthroline chimeras. Protein Eng. 9, 603–610 (1996).
Izzo, A. et al. The genomic landscape of the somatic linker histone subtypes H1.1 to H1.5 in human cells. Cell Rep. 3, 2142–2154 (2013).
van Bemmel, J. G. et al. A network model of the molecular organization of chromatin in Drosophila. Mol. Cell 49, 759–771 (2013).
Jessen, W. J. et al. Mapping chromatin structure in vivo using DNA methyltransferases. Methods 33, 68–80 (2004).
Kelly, T. K. et al. Genome-wide mapping of nucleosome positioning and DNA methylation within individual DNA molecules. Genome Res. 22, 2497–2506 (2012).
Gerstein, M. B. et al. Integrative analysis of the Caenorhabditis elegans genome by the modENCODE project. Science 330, 1775–1787 (2010).
The modENCODE Consortium. Identification of functional elements and regulatory circuits by Drosophila modENCODE. Science 330, 1787–1797 (2010).
Wang, J. et al. Factorbook.org: a Wiki-based database for transcription factor-binding data generated by the ENCODE consortium. Nucleic Acids Res. 41, D171–D176 (2013).
Schmidt, D. et al. ChIP–seq: using high-throughput sequencing to discover protein–DNA interactions. Methods 48, 240–248 (2009).
Rhee, H. S. & Pugh, B. F. Comprehensive genome-wide protein–DNA interactions detected at single-nucleotide resolution. Cell 147, 1408–1419 (2011).
Nakahashi, H. et al. A genome-wide map of CTCF multivalency redefines the CTCF code. Cell Rep. 3, 1678–1689 (2013).
Yen, K., Vinayachandran, V., Batta, K., Koerber, R. T. & Pugh, B. F. Genome-wide nucleosome specificity and directionality of chromatin remodelers. Cell 149, 1461–1473 (2012).
Chen, J. et al. Single-molecule dynamics of enhanceosome assembly in embryonic stem cells. Cell 156, 1274–1285 (2014). This study suggests that TFs find their binding sites through trial-and-error sampling of degenerate motifs, which provides a potential explanation for the prevalence of low-occupancy ChIP–seq peaks with weak motifs.
Guo, Y., Mahony, S. & Gifford, D. K. High resolution genome wide binding event finding and motif discovery reveals transcription factor spatial binding constraints. PLoS Comput. Biol. 8, e1002638 (2012).
Serandour, A., Brown, G., Cohen, J. & Carroll, J. Development of an Illumina-based ChIP-exonuclease method provides insight into FoxA1-DNA binding properties. Genome Biol. 14, R147 (2013).
Skene, P. J., Hernandez, A. E., Groudine, M. & Henikoff, S. The nucleosomal barrier to promoter escape by RNA polymerase II is overcome by the chromatin remodeler Chd1. eLife 3, e02042 (2014).
Zentner, G. E., Tsukiyama, T. & Henikoff, S. ISWI and CHD chromatin remodelers bind promoters but act in gene bodies. PLOS Genet. 9, e1003317 (2013).
Kasinathan, S., Orsi, G. A., Zentner, G. E., Ahmad, K. & Henikoff, S. High-resolution mapping of transcription factor binding sites on native chromatin. Nature Meth 11, 203–209 (2014).
Zentner, G. E. & Henikoff, S. Mot1 redistributes TBP from TATA-containing to TATA-less promoters. Mol. Cell. Biol. 33, 4996–5004 (2013).
Orsi, G. A. et al. High-resolution mapping defines the cooperative architecture of Polycomb response elements. Genome Res. 24, 809–820 (2014). References 64–67 show that native ChIP is applicable to a wide range of non-histone proteins.
Voss, T. C. & Hager, G. L. Dynamic regulation of transcriptional states by chromatin and transcription factors. Nature Rev. Genet. 15, 69–81 (2014).
Ahmad, K. & Henikoff, S. The histone variant H3.3 marks active chromatin by replication-independent nucleosome assembly. Mol. Cell 9, 1191–1200 (2002).
Mito, Y., Henikoff, J. G. & Henikoff, S. Genome-scale profiling of histone H3.3 replacement patterns. Nature Genet. 37, 1090–1097 (2005).
Mito, Y., Henikoff, J. G. & Henikoff, S. Histone replacement marks the boundaries of cis-regulatory domains. Science 315, 1408–1411 (2007).
Chow, C.-M. et al. Variant histone H3.3 marks promoters of transcriptionally active genes during mammalian cell division. EMBO Rep. 6, 354–360 (2005).
Jin, C. et al. H3.3/H2A. Z double variant-containing nucleosomes mark 'nucleosome-free regions' of active promoters and other regulatory regions. Nature Genet. 41, 941–945 (2009).
Ooi, S. L., Henikoff, J. G. & Henikoff, S. A native chromatin purification system for epigenomic profiling in Caenorhabditis elegans. Nucleic Acids Res. 38, e26 (2010).
Dion, M. F. et al. Dynamics of replication-independent histone turnover in budding yeast. Science 315, 1405–1408 (2007).
Jamai, A., Imoberdorf, R. M. & Strubin, M. Continuous histone H2B and transcription-dependent histone H3 exchange in yeast cells outside of replication. Mol. Cell 25, 345–355 (2007).
van Werven, F. J., van Teeffelen, H. A. A. M., Holstege, F. C. P. & Timmers, H. T. M. Distinct promoter dynamics of the basal transcription factor TBP across the yeast genome. Nature Struct. Mol. Biol. 16, 1043–1048 (2009).
Lickwar, C. R., Mueller, F., Hanlon, S. E., McNally, J. G. & Lieb, J. D. Genome-wide protein–DNA binding dynamics suggest a molecular clutch for transcription factor function. Nature 484, 251–255 (2012).
Kraushaar, D. et al. Genome-wide incorporation dynamics reveal distinct categories of turnover for the histone variant H3.3. Genome Biol. 14, R121 (2013).
Huang, C. et al. H3.3–H4 tetramer splitting events feature cell-type specific enhancers. PLoS Genet. 9, e1003558 (2013).
Deal, R. B., Henikoff, J. G. & Henikoff, S. Genome-wide kinetics of nucleosome turnover determined by metabolic labeling of histones. Science 328, 1161–1164 (2010).
Yang, F., Kemp, Christopher, J. & Henikoff, S. Doxorubicin enhances nucleosome turnover around promoters. Curr. Biol. 23, 782–787 (2013).
Gupta, P., Zlatanova, J. & Tomschik, M. Nucleosome assembly depends on the torsion in the DNA molecule: a magnetic tweezers study. Biophys. J. 97, 3150–3157 (2009).
Bermúdez, I., García-Martínez, J., Pérez-Ortín, J. E. & Roca, J. A method for genome-wide analysis of DNA helical tension by means of psoralen–DNA photobinding. Nucleic Acids Res. 38, e182 (2010).
Naughton, C. et al. Transcription forms and remodels supercoiling domains unfolding large-scale chromatin structures. Nature Struct. Mol. Biol. 20, 387–395 (2013).
Kouzine, F. et al. Transcription-dependent dynamic supercoiling is a short-range genomic force. Nature Struct. Mol. Biol. 20, 396–403 (2013).
Teves, S. S. & Henikoff, S. Transcription-generated torsional stress destabilizes nucleosomes. Nature Struct. Mol. Biol. 21, 88–94 (2014).
Poorey, K. et al. Measuring chromatin interaction dynamics on the second time scale at single-copy genes. Science 342, 369–372 (2013). This study indicates that a single long formaldehyde crosslinking time is unsuitable for inference of the relative occupancy or dynamics of a chromatin-binding factor.
Kulaeva, O. I., Hsieh, F.-K., Chang, H.-W., Luse, D. S. & Studitsky, V. M. Mechanism of transcription through a nucleosome by RNA polymerase II. Biochim. Biophys. Acta 1829, 76–83 (2013).
Adelman, K. & Lis, J. T. Promoter-proximal pausing of RNA polymerase II: emerging roles in metazoans. Nature Rev. Genet. 13, 720–731 (2012).
Hansen, K. D., Brenner, S. E. & Dudoit, S. Biases in Illumina transcriptome sequencing caused by random hexamer priming. Nucleic Acids Res. 38, e131 (2010).
Ståhlberg, A., Håkansson, J., Xian, X., Semb, H. & Kubista, M. Properties of the reverse transcription reaction in mRNA quantification. Clin. Chem. 50, 509–515 (2004).
Churchman, L. S. & Weissman, J. S. Nascent transcript sequencing visualizes transcription at nucleotide resolution. Nature 469, 368–373 (2011).
Gatehouse, J. & Thompson, A. in Plant Gene Transfer and Expression Protocols Vol. 49 Ch. 19, (ed. Jones, H.) 229–238 (Springer, 1995).
Core, L. J., Waterfall, J. J. & Lis, J. T. Nascent RNA. Sequencing reveals widespread pausing and divergent initiation at human promoters. Science 322, 1845–1848 (2008).
Kwak, H., Fuda, N. J., Core, L. J. & Lis, J. T. Precise maps of RNA polymerase reveal how promoters direct initiation and pausing. Science 339, 950–953 (2013).
Wuarin, J. & Schibler, U. Physical isolation of nascent RNA chains transcribed by RNA polymerase II: evidence for cotranscriptional splicing. Mol. Cell. Biol. 14, 7219–7225 (1994).
Weber, Christopher, M., Ramachandran, S. & Henikoff, S. Nucleosomes are context-specific, H2A.Z-modulated barriers to RNA polymerase. Mol. Cell 53, 819–830 (2014).
Macaulay, I. C. & Voet, T. Single cell genomics: advances and future perspectives. PLoS Genet. 10, e1004126 (2014).
Tomlinson, M. J., Tomlinson, S., Yang, X. B. & Kirkham, J. Cell separation: terminology and practical considerations. J. Tissue Eng. http://dx.doi.org/10.1177/2041731412472690 (2013).
Curran, S., McKay, J. A., McLeod, H. L. & Murray, G. I. Laser capture microscopy. Mol. Pathol. 53, 64–68 (2000).
Deal, R. B. & Henikoff, S. A simple method for gene expression and chromatin profiling of individual cell types within a tissue. Dev. Cell 18, 1030–1040 (2010).
Steiner, F. A., Talbert, P. B., Kasinathan, S., Deal, R. B. & Henikoff, S. Cell-type-specific nuclei purification from whole animals for genome-wide expression and chromatin profiling. Genome Res. 22, 766–777 (2012).
Henry, G. L., Davis, F. P., Picard, S. & Eddy, S. R. Cell type-specific genomics of Drosophila neurons. Nucleic Acids Res. 40, 9691–9704 (2012).
Schauer, T. et al. CAST-ChIP maps cell-type-specific chromatin states in the Drosophila central nervous system. Cell Rep. 5, 271–282 (2013).
Satterlee, J. S., Schubeler, D. & Ng, H.-H. Tackling the epigenome: challenges and opportunities for collaboration. Nature Biotech. 28, 1039–1044 (2010).
Lampa, S., Dahlo, M., Olason, P., Hagberg, J. & Spjuth, O. Lessons learned from implementing a national infrastructure in Sweden for storage and analysis of next-generation sequencing data. GigaScience 2, 9 (2013).
Kodama, Y., Shumway, M. & Leinonen, R. The sequence read archive: explosive growth of sequencing data. Nucleic Acids Res. 40, D54–D56 (2012).
Zhao, R., Bodnar, M. S. & Spector, D. L. Nuclear neighborhoods and gene expression. Curr. Opin. Genet. Dev. 19, 172–179 (2009).
Unnikrishnan, A., Gafken, P. R. & Tsukiyama, T. Dynamic changes in histone acetylation regulate origins of DNA replication. Nature Struct. Mol. Biol. 17, 430–437 (2010).
Kitamura, E., Blow, J. J. & Tanaka, T. U. Live-cell imaging reveals replication of individual replicons in eukaryotic replication factories. Cell 125, 1297–1308 (2006).
Nagano, T. et al. Single-cell Hi-C reveals cell-to-cell variability in chromosome structure. Nature 502, 59–64 (2013).
Ay, F., Bailey, T. L. & Noble, W. S. Statistical confidence estimation for Hi-C data reveals regulatory chromatin contacts. Genome Res. 24, 999–1011 (2014).
Mendenhall, E. M. et al. Locus-specific editing of histone modifications at endogenous enhancers. Nature Biotech. 31, 1133–1136 (2013).
Konermann, S. et al. Optical control of mammalian endogenous transcription and epigenetic states. Nature 500, 472–476 (2013).
Rusk, N. CRISPRs and epigenome editing. Nature Methods 11, 28 (2014).
Nishimura, K., Fukagawa, T., Takisawa, H., Kakimoto, T. & Kanemaki, M. An auxin-based degron system for the rapid depletion of proteins in nonplant cells. Nature Meth 6, 917–922 (2009).
Haruki, H., Nishikawa, J. & Laemmli, U. K. The anchor-away technique: rapid, conditional establishment of yeast mutant phenotypes. Mol. Cell 31, 925–932 (2008).
Landt, S. G. et al. ChIP–seq guidelines and practices of the ENCODE and modENCODE consortia. Genome Res. 22, 1813–1831 (2012).
Bailey, T. et al. Practical guidelines for the comprehensive analysis of ChIP–seq data. PLoS Comput. Biol. 9, e1003326 (2013).
Wade, J. T., Struhl, K., Busby, S. J. W. & Grainger, D. C. Genomic analysis of protein–DNA interactions in bacteria: insights into transcription and chromosome organization. Mol. Microbiol. 65, 21–26 (2007).
Biddie, S. C. et al. Transcription factor AP1 potentiates chromatin accessibility and glucocorticoid receptor binding. Mol. Cell 43, 145–155 (2011).
Biggin, Mark, D. Animal transcription networks as highly connected, quantitative continua. Dev. Cell 21, 611–626 (2011).
Fisher, W. W. et al. DNA regions bound at low occupancy by transcription factors do not drive patterned reporter gene expression in Drosophila. Proc. Natl Acad. Sci. USA 109, 21330–21335 (2012). This study shows that low-occupancy TF sites determined by ChIP–seq are often non-functional, which argues for cautious interpretation of such sites.
Marinov, G. K., Kundaje, A., Park, P. J. & Wold, B. J. Large-scale quality analysis of published ChIP–seq data. G3 4, 209–223 (2014). This analysis suggests that a substantial minority of published ChIP–seq data sets are of poor or intermediate quality.
Teytelman, L., Thurtle, D. M., Rine, J. & van Oudenaarden, A. Highly expressed loci are vulnerable to misleading ChIP localization of multiple unrelated proteins. Proc. Natl Acad. Sci. USA 110, 18602–18607 (2013).
Park, D., Lee, Y., Bhupindersingh, G. & Iyer, V. R. Widespread misinterpretable ChIP–seq bias in yeast. PLoS ONE 8, e83506 (2013). References 123 and 124 describe biases in X-ChIP–seq experiments that could lead to artefactual results.
Fan, X. & Struhl, K. Where does Mediator bind in vivo? PLoS ONE 4, e5029 (2009).
Jeronimo, C. & Robert, F. Kin28 regulates the transient association of Mediator with core promoters. Nature Struct. Mol. Biol. 21, 449–455 (2014).
Zhang, L., Zhang, K., Prändl, R. & Schöffl, F. Detecting DNA-binding of proteins in vivo by UV-crosslinking and immunoprecipitation. Biochem. Biophys. Res. Commun. 322, 705–711 (2004).
Vega, V. B., Cheung, E., Palanisamy, N. & Sung, W.-K. Inherent signals in sequencing-based chromatin-immunoprecipitation control libraries. PLoS ONE 4, e5241 (2009).
Teytelman, L. et al. Impact of chromatin structures on DNA processing for genomic analyses. PLoS ONE 4, e6700 (2009).
Acknowledgements
The authors thank S. Kasinathan and S. Ramachandran for critical reading of the manuscript and C. Weber for discussions. Work in the authors' laboratory is supported by the US National Institutes of Health grants 5U01 HG004274, U54 CA143862, and R01 ES020116 and by the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
Glossary
- Digital epigenomic analysis
-
The use of methods with sequencing-based readouts to interrogate the epigenome.
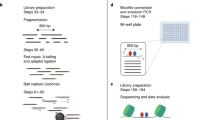
- Sequencing library
-
A collection of DNA fragments prepared for high-throughput sequencing by the addition of specific adapter sequences.
- Fragment midpoint-versus-length plots
-
(V-plots). Representations of paired-end sequencing data in which a point corresponding to the midpoint of a paired-end read is plotted in two-dimensional space. The x coordinate of the point represents the distance of the read midpoint from a defined genomic feature, and its y coordinate represents the length of the fragment from which it was derived.
- Affinity reagents
-
Antibodies or other molecules used to recover specific proteins from a complex mixture.
- Tagmentation
-
Simultaneous fragmentation and incorporation of sequencing adapters into chromatin using Tn5 transposase.
- Native ChIP
-
Chromatin immunoprecipitation (ChIP) using chromatin that has not been crosslinked with formaldehyde or any other crosslinking agent.
- Nuclear run-on
-
A technique in which transcription is reinitated in isolated nuclei to determine the rates at which genes are transcribed.
Rights and permissions
About this article
Cite this article
Zentner, G., Henikoff, S. High-resolution digital profiling of the epigenome. Nat Rev Genet 15, 814–827 (2014). https://doi.org/10.1038/nrg3798
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrg3798