Key Points
-
Short open reading frames (sORFs) of ∼100 codons in length are common and are distributed throughout the genome, but not all sORFs are biologically relevant.
-
sORFs are found on non-coding RNAs and within the 5′ leader and 3′ trailer regions of mRNAs. They can also overlap with the main protein-coding sequence of mRNAs.
-
The identification of sORFs that are translatable and that are likely to encode short peptides remains a major challenge. Three complementary approaches that are typically used to discover functional sORFs are bioinformatics, transcriptomics and proteomics.
-
Bioinformatic studies have identified a large pool of potentially translatable sORFs on the basis of sequence characteristics such as degree of conservation, coding potential and context of the initiation codon.
-
Global ribosome profiling has provided evidence of ribosome engagement at the start codon of many sORFs in various species, including yeast, insects, plants and mammals.
-
Proteomic studies using mass spectrometry on size-fractionated whole-cell lysates have identified several short peptides encoded by sORFs (sPEPs) in human tissues and cell lines.
-
Functional sPEPs have been identified in insects, plants and mammals, but only a small number of them have been fully characterized.
Abstract
Short open reading frames (sORFs) are a common feature of all genomes, but their coding potential has mostly been disregarded, partly because of the difficulty in determining whether these sequences are translated. Recent innovations in computing, proteomics and high-throughput analyses of translation start sites have begun to address this challenge and have identified hundreds of putative coding sORFs. The translation of some of these has been confirmed, although the contribution of their peptide products to cellular functions remains largely unknown. This Review examines this hitherto overlooked component of the proteome and considers potential roles for sORF-encoded peptides.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Change history
04 March 2014
In Table 2 (page 200) of the above article, the gene “RanGAP” was corrected to “SclA and SclB”, where Scl refers to the Sarcolamban gene in Drosophila melanogaster. The corresponding footnote was also corrected. The article has been corrected online. The editors apologize for this error.
References
Samayoa, J., Yildiz, F. H. & Karplus, K. Identification of prokaryotic small proteins using a comparative genomic approach. Bioinformatics 27, 1765–1771 (2011).
Hobbs, E. C., Fontaine, F., Yin, X. & Storz, G. An expanding universe of small proteins. Curr. Opin. Microbiol. 14, 167–173 (2011).
Law, G. L., Raney, A., Heusner, C. & Morris, D. R. Polyamine regulation of ribosome pausing at the upstream open reading frame of S-adenosylmethionine decarboxylase. J. Biol. Chem. 276, 38036–38043 (2001).
Lease, K. A. & Walker, J. C. The Arabidopsis unannotated secreted peptide database, a resource for plant peptidomics. Plant Physiol. 142, 831–838 (2006).
Hayden, C. & Bosco, G. Comparative genomic analysis of novel conserved peptide upstream open reading frames in Drosophila melanogaster and other dipteran species. BMC Genomics 9, 61 (2008).
Yang, X. et al. Discovery and annotation of small proteins using genomics, proteomics, and computational approaches. Genome Res. 21, 634–641 (2011).
Sleator, R. D. An overview of the current status of eukaryote gene prediction strategies. Gene 461, 1–4 (2010).
Brent, M. R. & Guigó, R. Recent advances in gene structure prediction. Curr. Opin. Struct. Biol. 14, 264–272 (2004).
Wang, J. et al. Vertebrate gene predictions and the problem of large genes. Nature Rev. Genet. 4, 741–749 (2003).
Hanada, K., Zhang, X., Borevitz, J. O., Li, W.-H. & Shiu, S.-H. A large number of novel coding small open reading frames in the intergenic regions of the Arabidopsis thaliana genome are transcribed and/or under purifying selection. Genome Res. 17, 632–640 (2007).
Cheng, H. et al. Small open reading frames: current prediction techniques and future prospect. Curr. Protein Pept. Sci. 12, 503–507 (2011).
Basrai, M. A., Hieter, P. & Boeke, J. D. Small open reading frames: beautiful needles in the haystack. Genome Res. 7, 768–771 (1997).
Claverie, J.-M. Computational methods for the identification of genes in vertebrate genomic sequences. Hum. Mol. Genet. 6, 1735–1744 (1997).
Frith, M. C. et al. The abundance of short proteins in the mammalian proteome. PLoS Genet. 2, e52 (2006). This is the first study to examine the size and nature of the mammalian peptidome.
Hashimoto, Y., Kondo, T. & Kageyama, Y. Lilliputians get into the limelight: novel class of small peptide genes in morphogenesis. Dev. Growth Differ. 50, S269–S276 (2008).
Kastenmayer, J. P. et al. Functional genomics of genes with small open reading frames (sORFs) in S. cerevisiae. Genome Res. 16, 365–373 (2006).
Fälth, M. et al. SwePep, a database designed for endogenous peptides and mass spectrometry. Mol. Cell. Proteom. 5, 998–1005 (2006).
Slavoff, S. A. et al. Peptidomic discovery of short open reading frame-encoded peptides in human cells. Nature Chem. Biol. 9, 59–64 (2013). This work builds on previous studies to identify 90 human small proteins using mass spectrometry.
Fritsch, C. et al. Genome-wide search for novel human uORFs and N-terminal protein extensions using ribosomal footprinting. Genome Res. 22, 2208–2218 (2012).
Ingolia, N. T., Lareau, L. F. & Weissman, J. S. Ribosome profiling of mouse embryonic stem cells reveals the complexity and dynamics of mammalian proteomes. Cell 147, 789–802 (2011).
Lee, S. et al. Global mapping of translation initiation sites in mammalian cells at single-nucleotide resolution. Proc. Natl Acad. Sci. 109, E2424–E2432 (2012).
Takahashi, H., Takahashi, A., Naito, S. & Onouchi, H. BAIUCAS: a novel BLAST-based algorithm for the identification of upstream open reading frames with conserved amino acid sequences and its application to the Arabidopsis thaliana genome. Bioinformatics 28, 2231–2241 (2012).
Castellana, N. E. et al. Discovery and revision of Arabidopsis genes by proteogenomics. Proc. Natl Acad. Sci. 105, 21034–21038 (2008).
Vanderperre, B. et al. Direct detection of alternative open reading frames translation products in human significantly expands the proteome. PLoS ONE 8, e70698 (2013). This proteomic-based study has identified numerous short proteins in several human cell lines and tissues.
Menschaert, G. et al. Deep proteome coverage based on ribosome profiling aids mass spectrometry-based protein and peptide discovery and provides evidence of alternative translation products and near-cognate translation initiation events. Mol. Cell. Proteom. 12, 1780–1790 (2013). This study shows how ribosome profiling can aid short peptide discovery by mass spectrometry.
Hanada, K. et al. sORF finder: a program package to identify small open reading frames with high coding potential. Bioinformatics 26, 399–400 (2010).
Vanderperre, B., Lucier, J. F. & Roucou, X. HAltORF: a database of predicted out-of-frame alternative open reading frames in human. Database (Oxford) 2012, bas025 (2012).
Skarshewski, A. et al. uPEPperoni: an online tool for upstream open reading frame location and analysis of transcript conservation. BMC Bioinformatics http://dx.doi.org/10.1186/1471-2105-15-36 (2014).
Hurst, L. D. The Ka/Ks ratio: diagnosing the form of sequence evolution. Trends Genet. 18, 486–487 (2002).
Zhang, Z. & Dietrich, F. Identification and characterization of upstream open reading frames (uORF) in the 5′ untranslated regions (UTR) of genes in Saccharomyces cerevisiae. Curr. Genet. 48, 77–87 (2005).
Ladoukakis, E., Pereira, V., Magny, E., Eyre-Walker, A. & Couso, J. P. Hundreds of putatively functional small open reading frames in Drosophila. Genome Biol. 12, R118 (2011).
Clamp, M. et al. Distinguishing protein-coding and noncoding genes in the human genome. Proc. Natl Acad. Sci. 104, 19428–19433 (2007).
Kozak, M. An analysis of 5′-noncoding sequences from 699 vertebrate messenger RNAs. Nucleic Acids Res. 15, 8125–8148 (1987).
Karlin, S., Campbell, A. M. & Mrázek, J. Comparative DNA analysis across diverse genomes. Annu. Rev. Genet. 32, 185–225 (1998).
Bateman, A. et al. The Pfam protein families database. Nucleic Acids Res. 32, D138–D141 (2004).
Hayden, C. & Jorgensen, R. Identification of novel conserved peptide uORF homology groups in Arabidopsis and rice reveals ancient eukaryotic origin of select groups and preferential association with transcription factor-encoding genes. BMC Biol. 5, 32 (2007).
Guillén, G. et al. Detailed analysis of putative genes encoding small proteins in legume genomes. Front. Plant Sci. 4, 208 (2013).
Castrignanò, T. et al. CSTminer: a web tool for the identification of coding and noncoding conserved sequence tags through cross-species genome comparison. Nucleic Acids Res. 32, W624–W627 (2004).
Badger, J. H. & Olsen, G. J. CRITICA: coding region identification tool invoking comparative analysis. Mol. Biol. Evol. 16, 512–524 (1999).
Kong, L. et al. CPC: assess the protein-coding potential of transcripts using sequence features and support vector machine. Nucleic Acids Res. 35, W345–W349 (2007).
Wang, Z., Gerstein, M. & Snyder, M. RNA-Seq: a revolutionary tool for transcriptomics. Nature Rev. Genet. 10, 57–63 (2009).
Raj, A. & van Oudenaarden, A. Nature, nurture, or chance: stochastic gene expression and its consequences. Cell 135, 216–226 (2008).
Ingolia, N. T., Ghaemmaghami, S., Newman, J. R. S. & Weissman, J. S. Genome-wide analysis in vivo of translation with nucleotide resolution using ribosome profiling. Science 324, 218–223 (2009).
Guttman, M., Russell, P., Ingolia, N. T., Weissman, J. S. & Lander, E. S. Ribosome profiling provides evidence that large noncoding RNAs do not encode proteins. Cell 154, 240–251 (2013).
Krug, K., Nahnsen, S. & Macek, B. Mass spectrometry at the interface of proteomics and genomics. Mol. BioSystems 7, 284–291 (2011).
Kapranov, P. et al. Large-scale transcriptional activity in chromosomes 21 and 22. Science 296, 916–919 (2002).
Okazaki, Y. et al. Analysis of the mouse transcriptome based on functional annotation of 60,770 full-length cDNAs. Nature 420, 563–573 (2002).
Birney, E. et al. Identification and analysis of functional elements in 1% of the human genome by the ENCODE pilot project. Nature 447, 799–816 (2007).
Clark, M. B. et al. The reality of pervasive transcription. PLoS Biol. 9, e1000625 (2011).
van Bakel, H., Nislow, C., Blencowe, B. J. & Hughes, T. R. Most “dark matter” transcripts are associated with known genes. PLoS Biol. 8, e1000371 (2010).
Struhl, K. Transcriptional noise and the fidelity of initiation by RNA polymerase II. Nature Struct. Mol. Biol. 14, 103–105 (2007).
Kellis, M., Patterson, N., Endrizzi, M., Birren, B. & Lander, E. S. Sequencing and comparison of yeast species to identify genes and regulatory elements. Nature 423, 241–254 (2003).
Kageyama, Y., Kondo, T. & Hashimoto, Y. Coding versus non-coding: translatability of short ORFs found in putative non-coding transcripts. Biochimie 93, 1981–1986 (2011).
Derrien, T. et al. The GENCODE v7 catalog of human long noncoding RNAs: analysis of their gene structure, evolution, and expression. Genome Res. 22, 1775–1789 (2012).
Iacono, M., Mignone, F. & Pesole, G. uAUG and uORFs in human and rodent 5′untranslated mRNAs. Gene 349, 97–105 (2005).
Crowe, M., Wang, X.-Q. & Rothnagel, J. A. Evidence for conservation and selection of upstream open reading frames suggests probable encoding of bioactive peptides. BMC Genomics 7, 16 (2006).
Neafsey, D. E. & Galagan, J. E. Dual modes of natural selection on upstream open reading frames. Mol. Biol. Evol. 24, 1744–1751 (2007).
Cvijovic, M., Dalevi, D., Bilsland, E., Kemp, G. & Sunnerhagen, P. Identification of putative regulatory upstream ORFs in the yeast genome using heuristics and evolutionary conservation. BMC Bioinformatics 8, 295 (2007).
Tran, M., Schultz, C. & Baumann, U. Conserved upstream open reading frames in higher plants. BMC Genomics 9, 361 (2008).
Vaughn, J. N., Ellingson, S. R., Mignone, F. & von Arnim, A. Known and novel post-transcriptional regulatory sequences are conserved across plant families. RNA 18, 368–384 (2012).
Wethmar, K., Smink, J. J. & Leutz, A. Upstream open reading frames: molecular switches in (patho)physiology. BioEssays 32, 885–893 (2010).
Pesole, G. et al. Analysis of oligonucleotide AUG start codon context in eukariotic mRNAs. Gene 261, 85–91 (2000).
Suzuki, Y. et al. Statistical analysis of the 5′ untranslated region of human mRNA using “oligo-capped” cDNA libraries. Genomics 64, 286–297 (2000).
Rogozin, I. B., Kochetov, A. V., Kondrashov, F. A., Koonin, E. V. & Milanesi, L. Presence of ATG triplets in 5′ untranslated regions of eukaryotic cDNAs correlates with a 'weak' context of the start codon. Bioinformatics 17, 890–900 (2001).
Yamashita, R., Suzuki, Y., Nakai, K. & Sugano, S. Small open reading frames in 5′ untranslated regions of mRNAs. C. R. Biol. 326, 987–991 (2003).
Chen, C. H., Liao, B. Y. & Chen, F. C. Exploring the selective constraint on the sizes of insertions and deletions in 5′ untranslated regions in mammals. BMC Evol. Biol. 11, 192 (2011).
Ribrioux, S., Brungger, A., Baumgarten, B., Seuwen, K. & John, M. R. Bioinformatics prediction of overlapping frameshifted translation products in mammalian transcripts. BMC Genomics 9, 122 (2008).
Michel, A. M. et al. Observation of dually decoded regions of the human genome using ribosome profiling data. Genome Res. 22, 2219–2229 (2012).
Chung, W.-Y., Wadhawan, S., Szklarczyk, R., Pond, S. K. & Nekrutenko, A. A. First look at ARFome: dual-coding genes in mammalian genomes. PLoS Comput. Biol. 3, e91 (2007).
Xu, H. et al. Length of the ORF, position of the first AUG and the Kozak motif are important factors in potential dual-coding transcripts. Cell Res. 20, 445–457 (2010).
Mercer, T. R. et al. Expression of distinct RNAs from 3′ untranslated regions. Nucleic Acids Res. 39, 2393–2403 (2011).
Chew, G.-L. et al. Ribosome profiling reveals resemblance between long non-coding RNAs and 5′ leaders of coding RNAs. Development 140, 2828–2834 (2013).
Bánfai, B. et al. Long noncoding RNAs are rarely translated in two human cell lines. Genome Res. 22, 1646–1657 (2012).
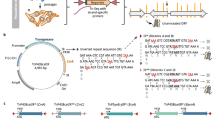
Hanada, K. et al. Small open reading frames associated with morphogenesis are hidden in plant genomes. Proc. Natl Acad. Sci. 110, 2395–2400 (2013). This is the first systematic characterization of short open reading frames using transgenic plants.
Oyama, M. et al. Analysis of small human proteins reveals the translation of upstream open reading frames of mRNAs. Genome Res. 14, 2048–2052 (2004). This is the first study to identify small proteins in human cells using mass spectrometry.
Oyama, M. et al. Diversity of translation start sites may define increased complexity of the human short ORFeome. Mol. Cell. Proteom. 6, 1000–1006 (2007).
Wang, R. F., Parkhurst, M. R., Kawakami, Y., Robbins, P. F. & Rosenberg, S. A. Utilization of an alternative open reading frame of a normal gene in generating a novel human cancer antigen. J. Exp. Med. 183, 1131–1140 (1996).
Ronsin, C. et al. A non-AUG-defined alternative open reading frame of the intestinal carboxyl esterase mRNA generates an epitope recognized by renal cell carcinoma-reactive tumor-infiltrating lymphocytes in situ. J. Immunol. 163, 483–490 (1999).
Frank, M. J. & Smith, L. G. A small, novel protein highly conserved in plants and animals promotes the polarized growth and division of maize leaf epidermal cells. Curr. Biol. 12, 849–853 (2002).
Rohrig, H., Schmidt, J., Miklashevichs, E., Schell, J. & John, M. Soybean ENDO40 encodes two peptides that bind sucrose synthase. Proc. Natl Acad. Sci. 99, 5 (2002).
Stuart, A. et al. The POLARIS gene of Arabidopsis encodes a predicted peptide required for correct root growth and leaf vascular patterning. Plant Cell 14, 16 (2002).
Narita, N. N. et al. Overexpression of a novel small peptide ROTUNDIFOLIA4 decreases cell proliferation and alters leaf shape in Arabidopsis thaliana. Plant J. 38, 699–713 (2004).
Abrar, Q. et al. HSPC300 and its role in neuronal connectivity. Neural Dev. 2, 18 (2007).
Colombani, J., Andersen, D. S. & Léopold, P. Secreted peptide Dilp8 coordinates Drosophila tissue growth with developmental timing. Science 336, 582–585 (2012).
Garelli, A., Gontijo, A. M., Miguela, V., Caparros, E. & Dominguez, M. Imaginal discs secrete insulin-like peptide 8 to mediate plasticity of growth and maturation. Science 336, 579–582 (2012).
Magny, E. G. et al. Conserved regulation of cardiac calcium uptake by peptides encoded in small open reading frames. Science 341, 1116–1120 (2013).
Galindo, M. I., Pueyo, J. I., Fouix, S., Bishop, S. A. & Couso, J. P. Peptides encoded by short ORFs control development and define a new eukaryotic gene family. PLoS Biol. 5, e106 (2007).
Kondo, T. et al. Small peptide regulators of actin-based cell morphogenesis encoded by a polycistronic mRNA. Nature Cell Biol. 9, 660–665 (2007).
Kondo, T. et al. Small peptides switch the transcriptional activity of Shavenbaby during Drosophila embryogenesis. Science 329, 336–339 (2010). This study identifies the molecular target of the small regulatory peptides encoded by a polycistronic mRNA that was previously thought to be a non-coding transcript.
Savard, J., Marques-Souza, H., Aranda, M. & Tautz, D. A segmentation gene in tribolium produces a polycistronic mRNA that codes for multiple conserved peptides. Cell 126, 559–569 (2006).
Li, B. et al. Ovol2, a mammalian homolog of Drosophila Ovo: gene structure, chromosomal mapping, and aberrant expression in blind-sterile mice. Genomics 80, 319–325 (2002).
Jorgensen, R. A. & Dorantes-Acosta, A. E. Conserved-peptide upstream open reading frames (CPuORFs) are associated with regulatory genes in angiosperms. Front. Plant Sci. 3, 191 (2012).
Werner, M., Feller, A., Messenguy, F. & Piérard, A. The leader peptide of yeast gene CPA1 is essential for the translational repression of its expression. Cell 49, 805–813 (1987).
Gaba, A., Jacobson, A. & Sachs, M. S. Ribosome occupancy of the yeast CPA1 upstream open reading frame termination codon modulates nonsense-mediated mRNA decay. Mol. Cell 20, 449–460 (2005).
Rahmani, F. et al. Sucrose control of translation mediated by an upstream open reading frame-encoded peptide. Plant Physiol. 150, 1356–1367 (2009).
Hanfrey, C. et al. A dual upstream open reading frame-based autoregulatory circuit controlling polyamine-responsive translation. J. Biol. Chem. 280, 39229–39237 (2005).
Alatorre-Cobos, F. et al. Translational regulation of Arabidopsis XIPOTL1 is modulated by phosphocholine levels via the phylogenetically conserved upstream open reading frame 30. J. Exp. Bot. 63, 5203–5221 (2012).
Diba, F., Watson, C. S. & Gametchu, B. 5′UTR sequences of the glucocorticoid receptor 1A transcript encode a peptide associated with translational regulation of the glucocorticoid receptor. J. Cell. Biochem. 81, 149–161 (2001).
Pendleton, L. C., Goodwin, B. L., Solomonson, L. P. & Eichler, D. C. Regulation of endothelial argininosuccinate synthase expression and NO production by an upstream open reading frame. J. Biol. Chem. 280, 24252–24260 (2005).
Nguyen, H. L., Yang, X. & Omiecinski, C. J. Expression of a novel mRNA transcript for human microsomal epoxide hydrolase (EPHX1) is regulated by short open reading frames within its 5′-untranslated region. RNA 19, 752–766 (2013).
Akimoto, C. et al. Translational repression of the McKusick–Kaufman syndrome transcript by unique upstream open reading frames encoding mitochondrial proteins with alternative polyadenylation sites. Biochim. Biophys. Acta. 1830, 2728–2738 (2013).
Vanderperre, B. et al. An overlapping reading frame in the PRNP gene encodes a novel polypeptide distinct from the prion protein. FASEB J. 25, 2373–2386 (2011).
Bergeron, D. et al. An out-of-frame overlapping reading frame in the ataxin-1 coding sequence encodes a novel ataxin-1 interacting protein. J. Biol. Chem. 288, 21824–21835 (2013).
Joung, J. K. & Sander, J. D. TALENs: a widely applicable technology for targeted genome editing. Nature Rev. Mol. Cell Biol. 14, 49–55 (2013).
Mali, P., Esvelt, K. M. & Church, G. M. Cas9 as a versatile tool for engineering biology. Nature Methods 10, 957–963 (2013).
Staudt, A. C. & Wenkel, S. Regulation of protein function by 'microProteins'. EMBO Rep. 12, 35–42 (2011).
Calvo, S. E., Pagliarini, D. J. & Mootha, V. K. Upstream open reading frames cause widespread reduction of protein expression and are polymorphic among humans. Proc. Natl Acad. Sci. 106, 7507–7512 (2009).
Wen, Y. et al. Loss-of-function mutations of an inhibitory upstream ORF in the human hairless transcript cause Marie Unna hereditary hypotrichosis. Nature Genet. 41, 228–233 (2009). This study identified mutations in a highly conserved upstream open reading frame that are associated with genetic hair loss and suggests that an aberrant short peptide may result in disease.
Almansour, N. M., Pirogova, E., Coloe, P. J., Cosic, I. & Istivan, T. S. Investigation of cytotoxicity of negative control peptides versus bioactive peptides on skin cancer and normal cells: a comparative study. Future Med. Chem. 4, 1553–1565 (2012).
Kozak, M. Point mutations define a sequence flanking the AUG initiator codon that modulates translation by eukaryotic ribosomes. Cell 44, 283–292 (1986).
Kozak, M. Effects of intercistronic length on the efficiency of reinitiation by eucaryotic ribosomes. Mol. Cell. Biol. 7, 3438–3445 (1987).
Morris, D. R. & Geballe, A. P. Upstream open reading frames as regulators of mRNA translation. Mol. Cell. Biol. 20, 8635–8642 (2000).
Ghilardi, N., Wiestner, A. & Skoda, R. C. Thrombopoietin production is inhibited by a translational mechanism. Blood 92, 4023–4030 (1998).
Calkhoven, C. F., Müller, C. & Leutz, A. Translational control of C/EBPα and C/EBPβ isoform expression. Genes Dev. 14, 1920–1932 (2000).
Hinnebusch, A. G. Translational regulation of yeast GCN4. J. Biol. Chem. 272, 21661–21664 (1997).
Child, S. J., Miller, M. K. & Geballe, A. P. Translational control by an upstream open reading frame in the HER-2/neu transcript. J. Biol. Chem. 274, 24335–24341 (1999).
Wang, X.-Q. & Rothnagel, J. A. Post-transcriptional regulation of the GLI1 oncogene by the expression of alternative 5′ untranslated regions. J. Biol. Chem. 276, 1311–1316 (2001).
Wang, X. Q. & Rothnagel, J. A. 5′-untranslated regions with multiple upstream AUG codons can support low-level translation via leaky scanning and reinitiation. Nucleic Acids Res. 32, 1382–1391 (2004).
Kozak, M. An analysis of vertebrate mRNA sequences: intimations of translational control. J. Cell Biol. 115, 887–903 (1991).
Hanyu-Nakamura, K., Sonobe-Nojima, H., Tanigawa, A. & Lasko, P. Drosophila Pgc protein inhibits P-TEFb recruitment to chromatin in primordial germ cells. Nature 451, 730–733 (2008).
Acknowledgements
This work was supported by a grant to J.A.R. from the Australian National Health and Medical Research Council (ID631551).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Glossary
- Short open reading frames
-
(sORFs). Open reading frames that are usually <100 codons in length but that can also be longer.
- Coding DNA sequence
-
(CDS). An open reading frame (ORF) that encodes a verified protein product.The CDS is typically the first ORF identified and characterized on an mRNA. It defines the end of the 5′ leader and the start of the 3′ trailer sequences.
- Ka/Ks test
-
A ratio that compares the number of nonsynonymous substitutions per nonsynonymous site with the number of synonymous substitutions per synonymous site.
- Transcription activator-like effector nucleases
-
(TALENs). Engineered enzymes that permit precise editing of genomes and that can be used to make specific sequence changes in model organisms such as Arabidopsis thaliana, zebrafish and mice.
- Microproteins
-
Negative regulators of multiprotein complexes. In this case, micro refers to the mechanism of action of these proteins rather than to their sizes.
Rights and permissions
About this article
Cite this article
Andrews, S., Rothnagel, J. Emerging evidence for functional peptides encoded by short open reading frames. Nat Rev Genet 15, 193–204 (2014). https://doi.org/10.1038/nrg3520
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrg3520
This article is cited by
-
A small protein coded within the mitochondrial canonical gene nd4 regulates mitochondrial bioenergetics
BMC Biology (2023)
-
Evidence of natural selection in the mitochondrial-derived peptides humanin and SHLP6
Scientific Reports (2023)
-
A novel tumor suppressor encoded by a 1p36.3 lncRNA functions as a phosphoinositide-binding protein repressing AKT phosphorylation/activation and promoting autophagy
Cell Death & Differentiation (2023)
-
Pseudogene MAPK6P4-encoded functional peptide promotes glioblastoma vasculogenic mimicry development
Communications Biology (2023)
-
Antioxidant peptides from plants: a review
Phytochemistry Reviews (2023)