Abstract
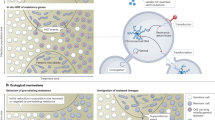
The evolution of antibiotic resistance can now be rapidly tracked with high-throughput technologies for bacterial genotyping and phenotyping. Combined with new approaches to evolve resistance in the laboratory and to characterize clinically evolved resistant pathogens, these methods are revealing the molecular basis and rate of evolution of antibiotic resistance under treatment regimens of single drugs or drug combinations. In this Progress article, we review these new tools for studying the evolution of antibiotic resistance and discuss how the genomic and evolutionary insights they provide could transform the diagnosis, treatment and predictability of antibiotic resistance in bacterial infections.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
World Health Organization. The evolving threat of antimicrobial resistance: options for action (World Health Organization, 2012).
Paterson, D. L. Resistance in Gram-negative bacteria: Enterobacteriaceae. Am. J. Med. 119, S20–S28; discussion S62–S70 (2006).
Didelot, X., Bowden, R., Wilson, D. J., Peto, T. E. & Crook, D. W. Transforming clinical microbiology with bacterial genome sequencing. Nature Rev. Genet. 13, 601–612 (2012).
Morar, M. & Wright, G. D. The genomic enzymology of antibiotic resistance. Annu. Rev. Genet. 44, 25–51 (2010).
Novais, A. et al. Evolutionary trajectories of β-lactamase CTX-M-1 cluster enzymes: predicting antibiotic resistance. PLoS Pathog. 6, e1000735 (2010).
Elena, S. F. & Lenski, R. E. Evolution experiments with microorganisms: the dynamics and genetic bases of adaptation. Nature Rev. Genet. 4, 457–469 (2003).
Toprak, E. et al. Evolutionary paths to antibiotic resistance under dynamically sustained drug selection. Nature Genet. 44, 101–105 (2012).
Lee, H. H., Molla, M. N., Cantor, C. R. & Collins, J. J. Bacterial charity work leads to population-wide resistance. Nature 467, 82–85 (2010).
Zhang, Q. et al. Acceleration of emergence of bacterial antibiotic resistance in connected microenvironments. Science 333, 1764–1767 (2011).
Torella, J. P., Chait, R. & Kishony, R. Optimal drug synergy in antimicrobial treatments. PLoS Comput. Biol. 6, e1000796 (2010).
Bonhoeffer, S., Lipsitch, M. & Levin, B. R. Evaluating treatment protocols to prevent antibiotic resistance. Proc. Natl Acad. Sci. USA 94, 12106–12111 (1997).
Michel, J. B., Yeh, P. J., Chait, R., Moellering, R. C. Jr & Kishony, R. Drug interactions modulate the potential for evolution of resistance. Proc. Natl Acad. Sci. USA 105, 14918–14923 (2008).
Hegreness, M., Shoresh, N., Damian, D., Hartl, D. & Kishony, R. Accelerated evolution of resistance in multidrug environments. Proc. Natl Acad. Sci. USA 105, 13977–13981 (2008).
Chait, R., Craney, A. & Kishony, R. Antibiotic interactions that select against resistance. Nature 446, 668–671 (2007).
Palmer, A. C., Angelino, E. & Kishony, R. Chemical decay of an antibiotic inverts selection for resistance. Nature Chem. Biol. 6, 105–107 (2010).
Yeh, P. J., Hegreness, M. J., Aiden, A. P. & Kishony, R. Drug interactions and the evolution of antibiotic resistance. Nature Rev. Microbiol. 7, 460–466 (2009).
Mwangi, M. M. et al. Tracking the in vivo evolution of multidrug resistance in Staphylococcus aureus by whole-genome sequencing. Proc. Natl Acad. Sci. USA 104, 9451–9456 (2007).
Lieberman, T. D. et al. Parallel bacterial evolution within multiple patients identifies candidate pathogenicity genes. Nature Genet. 43, 1275–1280 (2011).
Comas, I. et al. Whole-genome sequencing of rifampicin-resistant Mycobacterium tuberculosis strains identifies compensatory mutations in RNA polymerase genes. Nature Genet. 44, 106–110 (2012).
Harris, S. R. et al. Evolution of MRSA during hospital transmission and intercontinental spread. Science 327, 469–474 (2010).
Croucher, N. J. et al. Rapid pneumococcal evolution in response to clinical interventions. Science 331, 430–434 (2011).
Baldauf, S. L. Phylogeny for the faint of heart: a tutorial. Trends Genet. 19, 345–351 (2003).
Didelot, X. & Falush, D. Inference of bacterial microevolution using multilocus sequence data. Genetics 175, 1251–1266 (2007).
Cohen, T. & Murray, M. Modeling epidemics of multidrug-resistant M. tuberculosis of heterogeneous fitness. Nature Med. 10, 1117–1121 (2004).
Andersson, D. I. & Hughes, D. Antibiotic resistance and its cost: is it possible to reverse resistance? Nature Rev. Microbiol. 8, 260–271 (2010).
Hegreness, M., Shoresh, N., Hartl, D. & Kishony, R. An equivalence principle for the incorporation of favorable mutations in asexual populations. Science 311, 1615–1617 (2006).
Chubiz, L. M., Lee, M. C., Delaney, N. F. & Marx, C. J. FREQ-seq: a rapid, cost-effective, sequencing-based method to determine allele frequencies directly from mixed populations. PLoS ONE 7, e47959 (2012).
Levin-Reisman, I. et al. Automated imaging with ScanLag reveals previously undetectable bacterial growth phenotypes. Nature Methods 7, 737–739 (2010).
Goodarzi, H., Hottes, A. K. & Tavazoie, S. Global discovery of adaptive mutations. Nature Methods 6, 581–583 (2009).
Schenk, M. F., Szendro, I. G., Krug, J. & de Visser, J. A. Quantifying the adaptive potential of an antibiotic resistance enzyme. PLoS Genet. 8, e1002783 (2012).
Salverda, M. L. et al. Initial mutations direct alternative pathways of protein evolution. PLoS Genet. 7, e1001321 (2011).
van Opijnen, T., Bodi, K. L. & Camilli, A. Tn-seq: high-throughput parallel sequencing for fitness and genetic interaction studies in microorganisms. Nature Methods 6, 767–772 (2009).
Girgis, H. S., Hottes, A. K. & Tavazoie, S. Genetic architecture of intrinsic antibiotic susceptibility. PLoS ONE 4, e5629 (2009).
Poelwijk, F. J., Kiviet, D. J., Weinreich, D. M. & Tans, S. J. Empirical fitness landscapes reveal accessible evolutionary paths. Nature 445, 383–386 (2007).
Weinreich, D. M., Delaney, N. F., Depristo, M. A. & Hartl, D. L. Darwinian evolution can follow only very few mutational paths to fitter proteins. Science 312, 111–114 (2006).
Lozovsky, E. R. et al. Stepwise acquisition of pyrimethamine resistance in the malaria parasite. Proc. Natl Acad. Sci. USA 106, 12025–12030 (2009).
Tan, L., Serene, S., Chao, H. X. & Gore, J. Hidden randomness between fitness landscapes limits reverse evolution. Phys. Rev. Lett. 106, 198102 (2011).
Trindade, S. et al. Positive epistasis drives the acquisition of multidrug resistance. PLoS Genet. 5, e1000578 (2009).
Brown, K. M. et al. Compensatory mutations restore fitness during the evolution of dihydrofolate reductase. Mol. Biol. Evol. 27, 2682–2690 (2010).
Hall, A. R. & MacLean, R. C. Epistasis buffers the fitness effects of rifampicin- resistance mutations in Pseudomonas aeruginosa. Evolution 65, 2370–2379 (2011).
D'Costa, V. M., McGrann, K. M., Hughes, D. W. & Wright, G. D. Sampling the antibiotic resistome. Science 311, 374–377 (2006).
Sommer, M. O., Church, G. M. & Dantas, G. The human microbiome harbors a diverse reservoir of antibiotic resistance genes. Virulence 1, 299–303 (2010).
D'Costa, V. M. et al. Antibiotic resistance is ancient. Nature 477, 457–461 (2011).
Riesenfeld, C. S., Goodman, R. M. & Handelsman, J. Uncultured soil bacteria are a reservoir of new antibiotic resistance genes. Environ. Microbiol. 6, 981–989 (2004).
D'Costa, V. M. et al. Inactivation of the lipopeptide antibiotic daptomycin by hydrolytic mechanisms. Antimicrob. Agents Chemother. 56, 757–764 (2012).
Chusri, S., Villanueva, I., Voravuthikunchai, S. P. & Davies, J. Enhancing antibiotic activity: a strategy to control Acinetobacter infections. J. Antimicrob. Chemother. 64, 1203–1211 (2009).
Lewis, K. Antibiotics: recover the lost art of drug discovery. Nature 485, 439–440 (2012).
Koser, C. U. et al. Rapid whole-genome sequencing for investigation of a neonatal MRSA outbreak. N. Engl. J. Med. 366, 2267–2275 (2012).
Snitkin, E. S. et al. Tracking a hospital outbreak of carbapenem-resistant Klebsiella pneumoniae with whole-genome sequencing. Sci. Transl. Med. 4, 148ra116 (2012).
Harris, S. R. et al. Whole-genome sequencing for analysis of an outbreak of meticillin-resistant Staphylococcus aureus: a descriptive study. Lancet Infect. Dis. 13, 130–136 (2012).
Acknowledgements
We thank T. Lieberman for discussions on phylogeny and comments on the manuscript. This work was supported in part by US National Institutes of Health grants R01GM081617 and US National Institute of General Medical Science Center grant P50GM068763, and the Novartis Institutes for BioMedical Research.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Glossary
- β-lactamase
-
An enzyme that can confer resistance to β-lactam antibiotics by catalysing their degradation.
- Commensal microbes
-
Microbes living on or in a host without causing disease, although they typically include opportunistic pathogens.
- Cross-resistance
-
The propensity of a genetic change that confers resistance to one drug also to affect resistance to a different drug (by either increasing or decreasing resistance).
- dN/dS
-
The ratio of mutation rates at nonsynonymous (N) and synonymous (S) sites. dN/dS is increased by selection for amino acid changes (a signature of adaptive selection) and decreased by selection against amino acid changes (purifying selection).
- Horizontal gene transfer
-
The acquisition of a gene by a means other than direct inheritance from a parent cell (vertical transfer). Common in many bacteria and archaea, mechanisms of horizontal gene transfer include transformation, conjugation and transduction.
- Maximum likelihood and Bayesian approaches
-
This definition applies to the context of phylogenetics. Phylogenetic trees can be constructed by maximum parsimony, maximum likelihood and Bayesian inference. Maximum parsimony methods select from all possible trees the one containing the fewest mutations. Trees chosen by maximum likelihood and other Bayesian methods may contain more mutations, as they weigh the relative probabilities of different mutations according to various models.
- Microfluidic device
-
Customized, microscopic chambers in which fluid flows can be precisely controlled. Applied to microbiology, these allow the study of bacterial behaviour in spatially and temporally controllable environments.
- Monotherapy
-
Chemical therapy by a single drug.
- Parallel evolution
-
When the same mutations (or a range of mutations in the same gene) repeatedly occur in independent lineages; this provides an indication that these mutations may have been fixed by positive selection rather than by chance.
- Proto-resistance genes
-
Evolutionary precursors to drug-resistance genes that do not yet contribute to drug resistance but may do so on mutation and selection by drug stress.
- Resistance cassettes
-
A genetic element containing one or more drug resistance genes, often carried in transposable elements or plasmids that facilitate horizontal gene transfer.
- Transposon mutagenesis
-
The insertion of transposons at random locations throughout a genome to generate a library of different gene disruptions. Transposons can be constructed with outward-facing promoters also to introduce gene overexpression into the library.
- Turbidostats
-
Devices that maintain constant cell density (turbidity) in a continuously growing microbial culture by routinely removing a small volume of culture and replacing it with fresh sterile media.
Rights and permissions
About this article
Cite this article
Palmer, A., Kishony, R. Understanding, predicting and manipulating the genotypic evolution of antibiotic resistance. Nat Rev Genet 14, 243–248 (2013). https://doi.org/10.1038/nrg3351
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrg3351
This article is cited by
-
Assessing the predictability of fungicide resistance evolution through in vitro selection
Journal of Plant Diseases and Protection (2024)
-
Green adeptness in synthesis of non-toxic copper and cobalt oxide nanocomposites with multifaceted bioactivities
Cancer Nanotechnology (2023)
-
Environmental modulation of global epistasis in a drug resistance fitness landscape
Nature Communications (2023)
-
Quantitative systems-based prediction of antimicrobial resistance evolution
npj Systems Biology and Applications (2023)
-
Translating eco-evolutionary biology into therapy to tackle antibiotic resistance
Nature Reviews Microbiology (2023)