Abstract
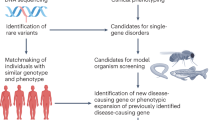
Model organisms have played a huge part in the history of studies of human genetic disease, both in identifying disease genes and characterizing their normal and abnormal functions. But is the importance of model organisms diminishing? The direct discovery of disease genes and variants in humans has been revolutionized, first by genome-wide association studies and now by whole-genome sequencing. Not only is it now much easier to directly identify potential disease genes in humans, but the genetic architecture that is being revealed in many cases is hard to replicate in model organisms. Furthermore, disease modelling can be done with increasing effectiveness using human cells. Where does this leave non-human models of disease?
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Aitman, T. J. et al. Progress and prospects in rat genetics: a community view. Nature Genet. 40, 516–522 (2008).
Shao, H. et al. Genetic architecture of complex traits: large phenotypic effects and pervasive epistasis. Proc. Natl Acad. Sci. USA 105, 19910–19914 (2008).
Rapp, J. P. Genetic analysis of inherited hypertension in the rat. Physiol. Rev. 80, 135–172 (2000).
Aitman, T. J. et al. Identification of Cd36 (Fat) as an insulin-resistance gene causing defective fatty acid and glucose metabolism in hypertensive rats. Nature Genet. 21, 76–83 (1999).
Hubner, N. et al. Integrated transcriptional profiling and linkage analysis for identification of genes underlying disease. Nature Genet. 37, 243–253 (2005).
Petretto, E. et al. Heritability and tissue specificity of expression quantitative trait loci. PLoS Genet. 2, e172 (2006).
Pravenec, M. et al. Identification of renal Cd36 as a determinant of blood pressure and risk for hypertension. Nature Genet. 40, 952–954 (2008).
Snyder, M. & Gallagher, J. E. Systems biology from a yeast omics perspective. FEBS Lett. 583, 3895–3899 (2009).
Chuang, H. Y., Hofree, M. & Ideker, T. A decade of systems biology. Ann. Rev. Cell Dev. Biol. 26, 721–744 (2010).
Engel, S. R. et al. Saccharomyces Genome Database provides mutant phenotype data. Nucleic Acids Res. 38, D433–D436 (2010).
Menne, T. F. et al. The Shwachman-Bodian-Diamond syndrome protein mediates translational activation of ribosomes in yeast. Nature Genet. 39, 486–495 (2007).
McGary, K. L. et al. Systematic discovery of nonobvious human disease models through orthologous phenotypes. Proc. Natl Acad. Sci. USA 107, 6544–6549 (2010).
Visscher, P. M, & Montgomery, G.W. Genome-wide association studies and human disease: from trickle to flood. JAMA 302, 2028–2029 (2009).
Paigen, K. One hundred years of mouse genetics: an intellectual history. I. The classical period (1902–1980). Genetics 163, 1–7 (2003).
Paigen, K. One hundred years of mouse genetics: an intellectual history. II. The molecular revolution (1981–2002). Genetics 163, 1227–1235 (2003).
Yuan, J., Shaham, S., Ledoux, S., Ellis, H. M. & Horvitz, H. R. The C. elegans cell death gene ced-3 encodes a protein similar to mammalian interleukin-1 β-converting enzyme. Cell 75, 641–652 (1993).
Savage, C. et al. Caenorhabditis elegans genes sma-2, sma-3, and sma-4 define a conserved family of transforming growth factor β pathway components. Proc. Natl Acad. Sci. USA 93, 790–794 (1996).
Lee, R. C., Feinbaum, R. L. & Ambros, V. The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 75, 843–854 (1993).
Reinhart, B. J. et al. The 21-nucleotide let-7 RNA regulates developmental timing in Caenorhabditis elegans. Nature 403, 901–906 (2000).
Fire, A. et al. Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 391, 806–811 (1998).
Chalfie, M., Tu, Y., Euskirchen, G., Ward, W. W. & Prasher, D. C. Green fluorescent protein as a marker for gene expression. Science 263, 802–805 (1994).
Abbott, A. Basel Declaration defends animal research. Nature 468, 742 (2010).
Flint, J. & Mackay, T. F. C. Genetic architecture of quantitative traits in mice, flies, and humans. Genome Res. 19, 723–733 (2009).
Mackay, T. F. C., Stone, E. A. & Ayroles, J. F. The genetics of quantitative traits: challenges and prospects. Nature Rev. Genet. 10, 565–577 (2009).
Manolio, T. A. et al. Finding the missing heritability of complex diseases. Nature 461, 747–753 (2009).
Ober, C., Loisel, D. A. & Gilad, Y. Sex-specific genetic architecture of human disease. Nature Rev. Genet. 9, 911–922 (2008).
Caspi, A. et al. Role of genotype in the cycle of violence in maltreated children. Science 297, 851–854 (2002).
Caspi, A. et al. Influence of life stress on depression: moderation by a polymorphism in the 5-HTT gene. Science 301, 386–389 (2003).
Ober, C. & Vercelli, D. Gene-environment interactions in human disease: nuisance or opportunity? Trends Genet. 27, 107–115 (2011).
Hill, W. G., Goddard, M. E. & Visscher, P. M. Data and theory point to mainly additive genetic variance for complex traits. PLoS Genet. 4, e1000008 (2008).
Shulman, J. M. & Feany, M. B. Genetic modifiers of tauopathy in Drosophila. Genetics 165, 1233–1242 (2003).
Lessing, D. & Bonini, N. M. Maintaining the brain: insight into human neurodegeneration from Drosophila melanogaster mutants. Nature Rev. Genet. 10, 359–370 (2009).
Rockman, M. V. Reverse engineering the genotype-phenotype map with natural genetic variation. Nature 456, 738–744 (2008).
Ayroles, J. F. et al. Systems genetics of complex traits in Drosophila melanogaster. Nature Genet. 41, 299–307 (2009).
Chen, Y. et al. Variations in DNA elucidate molecular networks that cause disease. Nature 452, 429–435 (2008).
Emilsson, V. et al. Genetics of gene expression and its effect on disease. Nature 452, 423–428 (2008).
Livet, J., et al. Transgenic strategies for combinatorial expression of fluorescent proteins in the nervous system. Nature 450, 56–62 (2007).
Kamei, M. et al. Endothelial tubes assemble from intracellular vacuoles in vivo. Nature 442, 453–456 (2006).
Nusslein-Volhard, C. & Wieschaus, E. Mutations affecting segment number and polarity in Drosophila. Nature 287, 795–801 (1980).
Chalfie, M., Horvitz, H. R. & Sulston, J. E. Mutations that lead to reiterations in the cell lineages of C. elegans. Cell 24, 59–69 (1981).
Driever, W. et al. A genetic screen for mutations affecting embryogenesis in zebrafish. Development 123, 37–46 (1996).
Haffter, P. et al. The identification of genes with unique and essential functions in the development of the zebrafish, Danio rerio. Development 123, 1–36 (1996).
Atanur, S. S. et al., The genome sequence of the spontaneously hypertensive rat: analysis and functional significance. Genome Res. 20, 791–803 (2010).
Johannesson, M. et al. A resource for the simultaneous high-resolution mapping of multiple quantitative trait loci in rats: the NIH heterogeneous stock. Genome Res. 19, 150–158 (2009).
Tong, C., Li, P., Wu, N. L., Yan, Y. & Ying, Q.L. Production of p53 gene knockout rats by homologous recombination in embryonic stem cells. Nature 467, 211–213 (2010).
Geurts, A. M. et al. Knockout rats via embryo microinjection of zinc-finger nucleases. Science 325, 433 (2009).
Jacob, H. J., Lazar, J., Dwinell, M. R., Moreno, C. & Geurts, A. M. Gene targeting in the rat: advances and opportunities. Trends Genet. 26, 510–518 (2010).
Heinig, M. et al. A trans-acting locus regulates an anti-viral expression network and type 1 diabetes risk. Nature 467, 460–464 (2010).
Fields, S. & Johnston, M. Cell biology. Whither model organism research? Science 307, 1885–1886 (2005).
Perocchi, F. et al. Assessing systems properties of yeast mitochondria through an interaction map of the organelle. PLoS Genet. 2, e170 (2006).
Hao, H. X. et al. SDH5, a gene required for flavination of succinate dehydrogenase, is mutated in paraganglioma. Science 325, 1139–1142 (2009).
Khurana, V. & Lindquist, S. Modelling neurodegeneration in Saccharomyces cerevisiae: why cook with baker's yeast? Nature Rev. Neurosci. 11, 436–449 (2010).
Marini, N. J., Thomas, P. D. & Rine, J. The use of orthologous sequences to predict the impact of amino acid substitutions on protein function. PLoS Genet. 6, e1000968 (2010).
Ehrenreich, I. M. et al. Dissection of genetically complex traits with extremely large pools of yeast segregants. Nature 464, 1039–1042 (2010).
Costanzo, M. et al. The genetic landscape of a cell. Science 327, 425–431 (2010).
Hartman, J. L, Garvik, B. & Hartwell, L. Principles for the buffering of genetic variation. Science 291, 1001–1004 (2001).
Dowell, R. D. et al. Genotype to phenotype: a complex problem. Science 328, 469 (2010).
Yang, H. et al. A customized and versatile high-density genotyping array for the mouse. Nature Methods 6, 663–666 (2009).
Austin, C. P. et al. The knockout mouse project. Nature Genet. 36, 921–924 (2004).
Boone, C., Bussey, H. & Andrews, B. J. Exploring genetic interactions and networks with yeast. Nature Rev. Genet. 8, 437–449 (2007).
Hunter, K. W. & Crawford, N. P. S. The future of mouse QTL mapping to diagnose disease in mice in the age of whole-genome association studies. Ann. Rev. Genet. 42, 131–141 (2008).
Yang, H., Bell, T. A., Churchill, G. A. & Pardo-Manuel de Villena, F. On the subspecific origin of the laboratory mouse. Nature Genet. 39, 1100–1107 (2007).
Yang, H. et al. Subspecific origin and haplotye diversity in the laboratory mouse. Nature Genet. 43, 648–655 (2011).
Threadgill, D. W., Hunter, K. W. & Williams, R. W. Genetic dissection of complex and quantitative traits: from fantasy to reality via a community effort. Mamm. Genome 13, 175–178 (2002).
Churchill, G. A. et al. The Collaborative Cross, a community resource for the genetic analysis of complex traits. Nature Genet. 36, 1133–1137 (2004).
Aylor, D. L. et al. Genetic analysis of complex traits in the emerging collaborative cross. Genome Res. 15 Mar 2011 (doi:10.1101/gr.111310.110).
Rose, M. R., Mueller, L. D. & Burke, M. K. New experiments for an undivided genetics. Genetics 188, 1–10 (2011).
McClellan, J. & King M.-C. Genetic heterogeneity in human disease. Cell 141, 210–217 (2010).
Dickson, S. P., Wang, K., Krantz, I., Hakonarson, H. & Goldstein, D. B. Rare variants create synthetic genome-wide associations. PLoS Biol. 8, e1000294 (2010).
Milloz, J., Duveau, F., Nuez, I. & Felix, M. A. Intraspecific evolution of the intercellular signaling network underlying a robust developmental system. Genes Dev. 22, 3064–3075 (2008).
Li, Y. et al. Global genetic robustness of the alternative splicing machinery in Caenorhabditis elegans. Genetics 186, 405–410 (2010).
Chandler, C. H. Cryptic intraspecific variation in sex determination in Caenorhabditis elegans revealed by mutations. Heredity 105, 473–482 (2010).
Gerstein, M. B. et al. Integrative analysis of the Caenorhabditis elegans genome by the modENCODE project. Science 330, 1775–1787 (2010).
Roy, S. et al. Identification of functional elements and regulatory circuits by Drosophila modENCODE. Science 330, 1787–1797 (2010).
Fox Keller, E. A Feeling for the Organism: The Life and Work of Barbara McClintock (W. H. Freeman and Company, New York, 1983).
Threadgill, D. W., Pardo-Manuel de Villena, F., Miller, D. R., Miller, D. R. & Churchill, G. A. The collaborative cross: a recombinant inbred mouse population for the systems genetic era. ILAR J. 52, 24–31 (2011).
Teer, J. K. & Mullikin, J. C. Exome sequencing: the sweet spot before whole genomes. Hum. Mol. Genet. 19, R145–R151 (2010).
Nasevicius, A. & Ekker, S. C. Effective targeted gene 'knockdown' in zebrafish. Nature Genet. 26, 216–220 (2000).
Khanna, H. et al. A common allele in RPGRIP1L is a modifier of retinal degeneration in ciliopathies. Nature Genet. 41, 739–745 (2009).
Anderson, C. A., Soranzo, N., Zeggini, E. & Barrett, J. C. Synthetic associations are unlikely to account for many common disease genome-wide association signals. PLoS Biol. 9, e1000580 (2011).
Liston, A., Lesage, S., Gray, D. H., Boyd, R. L. & Goodnow, C. C. Genetic lesions in T-cell tolerance and thresholds for autoimmunity. Immunol. Rev. 204, 87–101 (2005).
Cheung, C. & Gonzalez, F. J. Humanized mouse lines and their application for prediction of human drug metabolism and toxicological risk assessment. J. Pharmacol. Exp. Ther. 327, 288–299 (2008).
Acknowledgements
T.J.A. acknowledges funding from the Medical Research Council (MRC) Clinical Sciences Centre, the Imperial British Heart Foundation Centre of Excellence, the European Union EURATRANS consortium and the Leducq Cardianet Transatlantic Network of Excellence. C.B. is grateful for comments provided by B. Andrews. G.A.C. and T.F.C.M. both acknowledge funding from the US National Institutes of Health (NIH), grant numbers GM076468 and GM45146, respectively.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Aitman, T., Boone, C., Churchill, G. et al. The future of model organisms in human disease research. Nat Rev Genet 12, 575–582 (2011). https://doi.org/10.1038/nrg3047
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrg3047
This article is cited by
-
Dissecting genetic architecture of startle response in Drosophila melanogaster using multi-omics information
Scientific Reports (2017)
-
Heterozygous connexin 50 mutation affects metabolic syndrome attributes in spontaneously hypertensive rat
Lipids in Health and Disease (2016)
-
A mammary adenocarcinoma murine model suitable for the study of cancer immunoediting
Journal of Biomedical Science (2014)
-
An SNP-based second-generation genetic map of Daphnia magna and its application to QTL analysis of phenotypic traits
BMC Genomics (2014)
-
Controlling complexity: the clinical relevance of mouse complex genetics
European Journal of Human Genetics (2013)